

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.9
(742 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
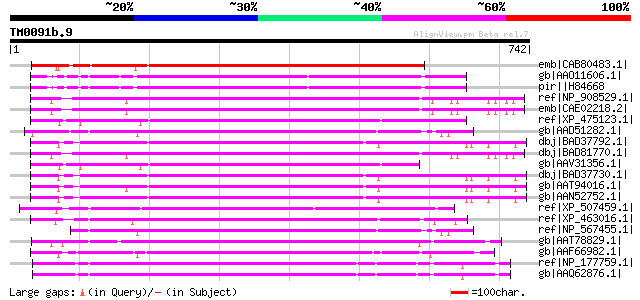
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB80483.1| hypothetical protein [Arabidopsis thaliana] gi|4... 472 e-131
gb|AAO11606.1| At2g27110/T20P8.16 [Arabidopsis thaliana] gi|2019... 380 e-104
pir||H84668 Mutator-like transposase [imported] - Arabidopsis th... 380 e-104
ref|NP_908529.1| P0702D12.15 [Oryza sativa (japonica cultivar-gr... 375 e-102
emb|CAE02218.2| OSJNBb0002N06.8 [Oryza sativa (japonica cultivar... 375 e-102
ref|XP_475123.1| putative Mutator-like transposase [Oryza sativa... 371 e-101
gb|AAD51282.1| far-red impaired response protein [Arabidopsis th... 368 e-100
dbj|BAD37792.1| far-red impaired response protein-like [Oryza sa... 365 2e-99
dbj|BAD81770.1| putative far-red impaired response protein [Oryz... 363 1e-98
gb|AAV31356.1| hypothetical protein [Oryza sativa (japonica cult... 362 2e-98
dbj|BAD37730.1| far-red impaired response protein-like [Oryza sa... 361 5e-98
gb|AAT94016.1| hypothetical protein [Oryza sativa (japonica cult... 358 3e-97
gb|AAN52752.1| Unknown protein [Oryza sativa (japonica cultivar-... 358 4e-97
ref|XP_507459.1| PREDICTED OJ1135_F06.10-1 gene product [Oryza s... 358 4e-97
ref|XP_463016.1| putative transposase [Oryza sativa (japonica cu... 357 1e-96
ref|NP_567455.1| far-red impaired response protein (FAR1) / far-... 353 9e-96
gb|AAT78829.1| putative FAR1 protein [Oryza sativa (japonica cul... 350 1e-94
gb|AAF66982.1| transposase [Zea mays] 350 1e-94
ref|NP_177759.1| far-red impaired responsive protein, putative [... 350 1e-94
gb|AAQ62876.1| At1g76320 [Arabidopsis thaliana] gi|62320160|dbj|... 349 2e-94
>emb|CAB80483.1| hypothetical protein [Arabidopsis thaliana]
gi|4467124|emb|CAB37558.1| hypothetical protein
[Arabidopsis thaliana] gi|22531237|gb|AAM97122.1|
unknown protein [Arabidopsis thaliana]
gi|15233732|ref|NP_195531.1| far-red impaired responsive
protein, putative [Arabidopsis thaliana]
gi|7485859|pir||T05645 hypothetical protein F20D10.300 -
Arabidopsis thaliana
Length = 788
Score = 472 bits (1215), Expect = e-131
Identities = 232/575 (40%), Positives = 358/575 (61%), Gaps = 20/575 (3%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR---AVL----VCCNEGQHKVKISRTE 84
G+EFES E + FYNS+A++ GF RV SS+ R A++ VC EG + RT+
Sbjct: 76 GLEFESEEAAKAFYNSYARRIGFSTRVSSSRRSRRDGAIIQRQFVCAKEGFRNMNEKRTK 135
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCH 144
+ + KR + R GC+ASL V + KW++ F DHNH +V P V +R H
Sbjct: 136 D-----REIKRPRTITRVGCKASLSVKMQDS-GKWLVSGFVKDHNHELVPPDQVHCLRSH 189
Query: 145 KKMSVAAKSLVEKFEEEGI-PTGKVAAIFNDGDST----FTNRDCWNYIRNVRRKNLDVG 199
+++S AK+L++ + G+ P ++A+ + FT DC NY+RN R+K+++ G
Sbjct: 190 RQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYMRNNRQKSIE-G 248
Query: 200 DAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKT 259
+ Q + DY ++ +N NFFY++Q ED + N+FW D ++ + + FGD +TFDTTY++
Sbjct: 249 EIQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFDTTYRS 308
Query: 260 NKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDL 319
N+Y +PFAPF G+N+H Q IL+GCA + +E+EASFVWLF T L AM P+SI TD D
Sbjct: 309 NRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSAHPPVSITTDHDA 368
Query: 320 AMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEE 379
++AA+ VFP +RHR C WHI+KK EKL+H++ K F+ D +C+ + S++ FE
Sbjct: 369 VIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHPSFESDFHKCVNLTESVEDFERC 428
Query: 380 WMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAST 439
W ++ +Y L+++EWLQ +Y R W+P+Y R TFFA M+ T RS+SIN++FD +++AST
Sbjct: 429 WFSLLDKYELRDHEWLQAIYSDRRQWVPVYLRDTFFADMSLTHRSDSINSYFDGYINAST 488
Query: 440 TLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL 499
L +F +EKA+ESRLE E K +Y++ + +L T S +E+ A+ +YTR +F +F+ EL
Sbjct: 489 NLSQFFKLYEKALESRLEKEVKADYDTMNSPPVLKTPSPMEKQASELYTRKLFMRFQEEL 548
Query: 500 EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
F K DG YQV+ +A V ++ A C +++EF GI+CRHI
Sbjct: 549 VGTLTFMASKADDDGDLVTYQVAKYGEAHKAHFVKFNVLEMRANCSCQMFEFSGIICRHI 608
Query: 560 LVIFQAKGVVQIPSHFIMERWTKDANRG-LEDTYN 593
L +F+ ++ +P ++I++RWT++A + D YN
Sbjct: 609 LAVFRVTNLLTLPPYYILKRWTRNAKSSVIFDDYN 643
>gb|AAO11606.1| At2g27110/T20P8.16 [Arabidopsis thaliana]
gi|20197414|gb|AAC77869.2| Mutator-like transposase
[Arabidopsis thaliana] gi|15982769|gb|AAL09732.1|
At2g27110/T20P8.16 [Arabidopsis thaliana]
gi|18401324|ref|NP_565636.1| far-red impaired responsive
protein, putative [Arabidopsis thaliana]
gi|30683396|ref|NP_850098.1| far-red impaired responsive
protein, putative [Arabidopsis thaliana]
Length = 851
Score = 380 bits (976), Expect = e-104
Identities = 210/622 (33%), Positives = 345/622 (54%), Gaps = 29/622 (4%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDST 90
+GMEF S ++ + FY+ ++++ GF +SK L+ +G V R S+
Sbjct: 51 VGMEFNSEKEAKSFYDEYSRQLGF-----TSK-----LLPRTDGSVSV---REFVCSSSS 97
Query: 91 NQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVA 150
++KR+ S C+A + + KW++ F +H H + S + +R + + +
Sbjct: 98 KRSKRRLS---ESCDAMVRIEL-QGHEKWVVTKFVKEHTHGLASSNMLHCLRPRRHFANS 153
Query: 151 AKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKR 210
KS + E +P+G + + D +S R N K DA + +Y KR
Sbjct: 154 EKSSYQ--EGVNVPSGMMY-VSMDANS----RGARNASMATNTKRTIGRDAHNLLEYFKR 206
Query: 211 KQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFV 270
Q EN FFYA+Q DED++M N+FW D+RSR++Y FGD +T DT Y+ N++ +PFAPF
Sbjct: 207 MQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVPFAPFT 266
Query: 271 GLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFP 330
G+N+H Q+IL+GCAL+ DES+ SF+WLFKT L AM + P+S++TDQD A++ A +VFP
Sbjct: 267 GVNHHGQAILFGCALILDESDTSFIWLFKTFLTAMRDQPPVSLVTDQDRAIQIAAGQVFP 326
Query: 331 ESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLK 390
+RH + W ++++ EKLAH+ F+ ++ CI + +I+ FE W ++ +Y+L
Sbjct: 327 GARHCINKWDVLREGQEKLAHVCLAYPSFQVELYNCINFTETIEEFESSWSSVIDKYDLG 386
Query: 391 ENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEK 450
+EWL LY R W+P+Y R +FFA + +Q +FFD +V+ TTL F +E+
Sbjct: 387 RHEWLNSLYNARAQWVPVYFRDSFFAAVFPSQGYS--GSFFDGYVNQQTTLPMFFRLYER 444
Query: 451 AVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKI 510
A+ES E E + + ++ + +L T S +E AA+++TR IF KF+ EL + T ++I
Sbjct: 445 AMESWFEMEIEADLDTVNTPPVLKTPSPMENQAANLFTRKIFGKFQEELVETFAHTANRI 504
Query: 511 RRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQ 570
DG ++V+N + + V A C +++E GILCRH+L +F ++
Sbjct: 505 EDDGTTSTFRVANFENDNKAYIVTFCYPEMRANCSCQMFEHSGILCRHVLTVFTVTNILT 564
Query: 571 IPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAY 630
+P H+I+ RWT++A +E D + E I R H+ REA A+ + EAY
Sbjct: 565 LPPHYILRRWTRNAKSMVE---LDEHVSENGHDSSIHRYNHLCREAIKYAEEGAITAEAY 621
Query: 631 NFIISEMKQTHKSAAAMKTSEG 652
N + ++++ K + ++ G
Sbjct: 622 NIALGQLREGGKKVSVVRKRIG 643
>pir||H84668 Mutator-like transposase [imported] - Arabidopsis thaliana
Length = 849
Score = 380 bits (976), Expect = e-104
Identities = 210/622 (33%), Positives = 345/622 (54%), Gaps = 29/622 (4%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDST 90
+GMEF S ++ + FY+ ++++ GF +SK L+ +G V R S+
Sbjct: 51 VGMEFNSEKEAKSFYDEYSRQLGF-----TSK-----LLPRTDGSVSV---REFVCSSSS 97
Query: 91 NQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVA 150
++KR+ S C+A + + KW++ F +H H + S + +R + + +
Sbjct: 98 KRSKRRLS---ESCDAMVRIEL-QGHEKWVVTKFVKEHTHGLASSNMLHCLRPRRHFANS 153
Query: 151 AKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKR 210
KS + E +P+G + + D +S R N K DA + +Y KR
Sbjct: 154 EKSSYQ--EGVNVPSGMMY-VSMDANS----RGARNASMATNTKRTIGRDAHNLLEYFKR 206
Query: 211 KQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFV 270
Q EN FFYA+Q DED++M N+FW D+RSR++Y FGD +T DT Y+ N++ +PFAPF
Sbjct: 207 MQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVPFAPFT 266
Query: 271 GLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFP 330
G+N+H Q+IL+GCAL+ DES+ SF+WLFKT L AM + P+S++TDQD A++ A +VFP
Sbjct: 267 GVNHHGQAILFGCALILDESDTSFIWLFKTFLTAMRDQPPVSLVTDQDRAIQIAAGQVFP 326
Query: 331 ESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLK 390
+RH + W ++++ EKLAH+ F+ ++ CI + +I+ FE W ++ +Y+L
Sbjct: 327 GARHCINKWDVLREGQEKLAHVCLAYPSFQVELYNCINFTETIEEFESSWSSVIDKYDLG 386
Query: 391 ENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEK 450
+EWL LY R W+P+Y R +FFA + +Q +FFD +V+ TTL F +E+
Sbjct: 387 RHEWLNSLYNARAQWVPVYFRDSFFAAVFPSQGYS--GSFFDGYVNQQTTLPMFFRLYER 444
Query: 451 AVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKI 510
A+ES E E + + ++ + +L T S +E AA+++TR IF KF+ EL + T ++I
Sbjct: 445 AMESWFEMEIEADLDTVNTPPVLKTPSPMENQAANLFTRKIFGKFQEELVETFAHTANRI 504
Query: 511 RRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQ 570
DG ++V+N + + V A C +++E GILCRH+L +F ++
Sbjct: 505 EDDGTTSTFRVANFENDNKAYIVTFCYPEMRANCSCQMFEHSGILCRHVLTVFTVTNILT 564
Query: 571 IPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAY 630
+P H+I+ RWT++A +E D + E I R H+ REA A+ + EAY
Sbjct: 565 LPPHYILRRWTRNAKSMVE---LDEHVSENGHDSSIHRYNHLCREAIKYAEEGAITAEAY 621
Query: 631 NFIISEMKQTHKSAAAMKTSEG 652
N + ++++ K + ++ G
Sbjct: 622 NIALGQLREGGKKVSVVRKRIG 643
>ref|NP_908529.1| P0702D12.15 [Oryza sativa (japonica cultivar-group)]
Length = 832
Score = 375 bits (964), Expect = e-102
Identities = 241/770 (31%), Positives = 380/770 (49%), Gaps = 85/770 (11%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV----RSSKPKRAVL-VCCNEGQHKVKISRTEE 85
+G F++ FYN +A GFG+R+ +++K +R + +CC+
Sbjct: 73 LGQRFKTERDAYNFYNVYAVSKGFGIRLDKDRKNTKNQRTMRQICCSH------------ 120
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSY-MRCH 144
K K ++R GC A + ++R S W + + HNH M V+ + H
Sbjct: 121 ---QGRNPKTKKPSVRIGCPAMMKINRSRAGSGWSVTKVVSAHNHPMKKSVGVTKNYQSH 177
Query: 145 KKMSVAAKSLVEKFEEEGIPT----GKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGD 200
++ + ++E+ + + G ++ + R + + R++ D
Sbjct: 178 NQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDRLAYAIRRDESSDD 237
Query: 201 AQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTN 260
Q D K Q + NFFY+IQ DE R+ N+FW A SRL+++ FGDVITFDTTYKTN
Sbjct: 238 MQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHFGDVITFDTTYKTN 297
Query: 261 KYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLA 320
KY+MPFAPFVG+NNH QS +GCALL++E+E SF WLF T + M GK PI I+TD +
Sbjct: 298 KYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGKVPIGILTDNCPS 357
Query: 321 MKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEW 380
M AA+ VFP + HR+C WH++KK E + +IY K+ FK+ + + + + + F W
Sbjct: 358 MAAAIRTVFPNTIHRVCKWHVLKKAKEFMGNIYSKRHTFKKAFHKVLTQTLTEEEFVAAW 417
Query: 381 MRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTT 440
+++ +YNL+++ +L+ ++ IR W +Y FFAGM TTQRSES N F FV S++
Sbjct: 418 HKLIRDYNLEKSVYLRHIWDIRRKWAFVYFSHRFFAGMTTTQRSESANHVFKMFVSPSSS 477
Query: 441 LQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL- 499
+ FV ++++ +L+ E E +++ + + T S +E HA+ +YTR +F F EL
Sbjct: 478 MNGFVKRYDRFFNEKLQKEDSEEFQTSNDKVEIKTRSPIEIHASQVYTRAVFQLFSEELT 537
Query: 500 EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
+ ++ + + V S R + V DL+ + C +++E GILC HI
Sbjct: 538 DSLSYMVKPGEDESTVQVVRMNSQESFLRKEYQVSCDLEREEFSCVCKMFEHKGILCSHI 597
Query: 560 LVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDT----LKILRRVHVQRE 615
L + G+ +IP +I++RWTKDA DT + G K D + R V + R+
Sbjct: 598 LRVLVQYGLSRIPERYILKRWTKDAR----DTIPPHLHGYKDDVDASQSRSYRHVMLNRK 653
Query: 616 ASFLADLAEESEEAY-------NFIISEMK------------QTHKSAAAMKTSEGVVLL 656
+A +A + + + N ++ +MK + K AA K S VL
Sbjct: 654 TVEVAKIANKDVQTFKMAMRVMNKLLEDMKNQLSLDDGDNSREAPKRAARHK-SRSTVLT 712
Query: 657 ESSEKNVNQVCSSEQVSEPPQLTIGN-------PHVSQTKGRKK-----DGGKMTQNSRF 704
E ++V + P + GN P +++GR K GG++ R
Sbjct: 713 EDGNEDVGGGDEDSEYDVPSEDEGGNLTADILPPLKKRSRGRPKVNRYKSGGEVASLKRR 772
Query: 705 KS-GLE----------VSLNKSVVK--------RKACHECGEHGHNSRTC 735
K G+E V V+ R+ CH C E GHNSRTC
Sbjct: 773 KEVGIEKKNTTNECESVEEENQVIDHEDEVPFPRRFCHTCHEAGHNSRTC 822
>emb|CAE02218.2| OSJNBb0002N06.8 [Oryza sativa (japonica cultivar-group)]
gi|50923349|ref|XP_472035.1| OSJNBb0002N06.8 [Oryza
sativa (japonica cultivar-group)]
Length = 885
Score = 375 bits (962), Expect = e-102
Identities = 240/770 (31%), Positives = 375/770 (48%), Gaps = 85/770 (11%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV-----RSSKPKRAVLVCCNEGQHKVKISRTEE 85
+G F++ FYN +A GFG+R+ + K + +CC+
Sbjct: 126 LGQRFKTERDAYNFYNVYAVSKGFGIRLDKDRMNTEKQRTMRQICCSH------------ 173
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSY-MRCH 144
K K ++R GC A + ++R S W + + HNH M V+ + H
Sbjct: 174 ---QGRNPKTKKPSVRIGCPAMMKINRSGAGSGWSVTKVVSTHNHPMKKSVGVTKNYQSH 230
Query: 145 KKMSVAAKSLVEKFEEEGIPT----GKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGD 200
++ + ++E+ + + G ++ + R + + R++ D
Sbjct: 231 NQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDRLAYAIRRDESSDD 290
Query: 201 AQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTN 260
Q D K Q + NFFY+IQ DE R+ N+FW A SRL+++ FGDVITFDTTYKTN
Sbjct: 291 MQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHFGDVITFDTTYKTN 350
Query: 261 KYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLA 320
KY+MPFAPFVG+NNH QS +GCALL++E+E SF WLF T + M GK PI I+TD +
Sbjct: 351 KYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGKVPIGILTDNCPS 410
Query: 321 MKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEW 380
M AA+ VFP + HR+C WH++KK E + +IY K+ FK+ + + + + + F W
Sbjct: 411 MAAAIRTVFPNTIHRVCKWHVLKKAKEFMGNIYSKRHTFKKAFHKVLTQTLTEEEFVAAW 470
Query: 381 MRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTT 440
+++ +YNL+++ +L+ ++ IR W +Y FFAGM TTQRSES N F FV S++
Sbjct: 471 HKLIRDYNLEKSVYLRHIWDIRRKWAFVYFSHRFFAGMTTTQRSESANHVFKMFVSPSSS 530
Query: 441 LQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL- 499
+ FV ++++ +L+ E E +++ + + T S +E HA+ +YTR +F F EL
Sbjct: 531 MNGFVKRYDRFFNEKLQKEDSEEFQTSNDKVEIKTRSPIEIHASQVYTRAVFQLFSEELT 590
Query: 500 EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
+ ++ + + V S R + V DL+ + C +++E GILC HI
Sbjct: 591 DSLSYMVKPGEDESTVQVVRMNSQESFLRKEYQVSCDLEREEFSCVCKMFEHKGILCSHI 650
Query: 560 LVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDT----LKILRRVHVQRE 615
L + G+ +IP +I++RWTKDA DT + G K D + R V + R+
Sbjct: 651 LRVLVQYGLSRIPERYILKRWTKDAR----DTIPPHLHGYKDDVDASQSRSYRHVMLNRK 706
Query: 616 ASFLADLAEESEEAY-------NFIISEMK------------QTHKSAAAMKTSEGVVLL 656
+A +A + + + N ++ +MK + K AA K S VL
Sbjct: 707 TVEVAKIANKDVQTFKMAMTVMNKLLEDMKNQLSLDDGDNSREAPKRAARHK-SRSTVLT 765
Query: 657 ESSEKNVNQVCSSEQVSEPPQLTIGN-------PHVSQTKGRKK-----DGGKMTQNSRF 704
E + V + P + GN P +++GR K GG++ R
Sbjct: 766 EDGNEEVGGGDEDSEYDVPSEDEGGNLTADILPPLKKKSRGRPKVNRYKSGGEVASLKRR 825
Query: 705 KS-GLE----------VSLNKSVVK--------RKACHECGEHGHNSRTC 735
K G+E V V+ R+ CH C E GHNSRTC
Sbjct: 826 KEVGIEKKNTTNECESVEEENQVIDHEDEVPFPRRFCHTCHEAGHNSRTC 875
>ref|XP_475123.1| putative Mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|46063440|gb|AAS79743.1| putative
Mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1510
Score = 371 bits (953), Expect = e-101
Identities = 221/639 (34%), Positives = 338/639 (52%), Gaps = 23/639 (3%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLV-----CCNEGQHKVKISRTEE 85
+GM F+++ V +FY S+A + GF VRV K + ++ C EG K R ++
Sbjct: 709 VGMIFDTLTDVEKFYKSYAHEAGFSVRVGQHKKQNEEILFKRYYCSREGYIK---ERVKD 765
Query: 86 IQDSTNQTKRKCSTI---RSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMR 142
+ D + + KRK + R GCEA ++V G+ K K+ I S +H+H VSP +R
Sbjct: 766 VSDESGKKKRKTPYMMETRCGCEAHIVVKLGSDK-KYRISSMIGEHSHGFVSPDKRHLLR 824
Query: 143 CHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCW-----NYIRNVRRKNLD 197
++ +S AKS + + I T + + D F N C NY ++R K D
Sbjct: 825 SNRTVSERAKSTLFSCHKASIGTSQAFRLLQVSDGGFQNVGCTLRNLQNYYHDLRCKIKD 884
Query: 198 VGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTY 257
DAQ +RK+ N FFY D+ R+V +FW DA R +Y +FGDV++ D+TY
Sbjct: 885 A-DAQMFVGQLERKKEVNPAFFYEFMVDKQGRLVRVFWADAICRKNYSVFGDVLSVDSTY 943
Query: 258 KTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQ 317
TN+Y+M F PF G+N+H QS+ G + L DE SFVWLF+T L+A GG P IITD+
Sbjct: 944 STNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLKATGGVAPRLIITDE 1003
Query: 318 DLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFE 377
D +MKAA+A++ P + HRLC+WHI++K PEK+ + F + +C+ GS FE
Sbjct: 1004 DASMKAAIAQILPNTVHRLCMWHIMEKVPEKVGPSIREDGEFWDRLHKCVWGSEDSDDFE 1063
Query: 378 EEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAG-MNTTQRSESINAFFDSFVD 436
EW IM +Y L NEW + IR+SWI Y AG ++TT RSES N+FF+ F+
Sbjct: 1064 SEWNSIMAKYGLIGNEWFSTKFDIRQSWILAYFMDIPLAGILSTTSRSESANSFFNRFIH 1123
Query: 437 ASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFK 496
T EF L+F+ A+E + + E K + S H + L T +E+ + IYT +F+KF+
Sbjct: 1124 RKLTFVEFWLRFDTALECQRQEELKADNISLHTNPKLMTPWDMEKQCSGIYTHEVFSKFQ 1183
Query: 497 GEL-EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGIL 555
+L + I + G + +S+ ++ V ++ + C +LYE GI
Sbjct: 1184 EQLIVARDHCIIQGISKSGDMKIVTISSLFEKER--VVQMNKSNMFGTCSCKLYESYGIP 1241
Query: 556 CRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKS-DTLKILRRVHVQR 614
CRHI+ + + + +IPS +IM+RW K R + N L EK+ D +++ R +
Sbjct: 1242 CRHIIQVLRGEKQNEIPSIYIMKRWEKICKREMFFDDEGNLLDEKAKDPMEVAMRKKISD 1301
Query: 615 EASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSEGV 653
+ L+DL E ++ I + +S K + V
Sbjct: 1302 SRNNLSDLVEPLQKIIPAAIVNKQDEFESFLGSKIPDEV 1340
>gb|AAD51282.1| far-red impaired response protein [Arabidopsis thaliana]
Length = 827
Score = 368 bits (944), Expect = e-100
Identities = 213/657 (32%), Positives = 350/657 (52%), Gaps = 31/657 (4%)
Query: 22 DIGGNSKLEI----GMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHK 77
D+G + L++ G++F++ E FY +AK GF +++S+ + +
Sbjct: 40 DVGFSGDLDLEPRNGIDFDTHEAAYIFYQEYAKSMGFTTSIKNSRRSKKTKDFIDAKFAC 99
Query: 78 VKISRTEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKS 137
+ T E +S+ + R+ + ++ C+AS+ V R KW+I F DHNH ++ P
Sbjct: 100 SRYGVTPE-SESSGSSSRRSTVKKTDCKASMHVKR-RPDGKWIIHEFVKDHNHELL-PAL 156
Query: 138 VSYMRCHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTN------RDCWNYIRNV 191
+ R + + +A K+ ++ T K+ + + N D + +
Sbjct: 157 AYHFRIQRNVKLAEKNNIDILHAVSERTKKMYVEMSRQSGGYKNIGSLLQTDVSSQVDKG 216
Query: 192 RRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVI 251
R L+ GD+Q + +Y KR + EN FFYAI +ED R+ NLFW DA+SR Y F DV+
Sbjct: 217 RYLALEEGDSQVLLEYFKRIKKENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFNDVV 276
Query: 252 TFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPI 311
+FDTTY +P A F+G+N+HSQ +L GCAL+ DES +FVWL KT L+AMGG+ P
Sbjct: 277 SFDTTYVKFNDKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIKTWLRAMGGRAPK 336
Query: 312 SIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSH 371
I+TDQD + +A++++ P +RH LWH+++K PE +H+ + F +CI S
Sbjct: 337 VILTDQDKFLMSAVSELLPNTRHCFALWHVLEKIPEYFSHVMKRHENFLLKFNKCIFRSW 396
Query: 372 SIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFF 431
+ F+ W +++ ++ L+ +EWL L++ R+ W+P + F AGM+T+QRSES+N+FF
Sbjct: 397 TDDEFDMRWWKMVSQFGLENDEWLLWLHEHRQKWVPTFMSDVFLAGMSTSQRSESVNSFF 456
Query: 432 DSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNI 491
D ++ TL+EF+ ++ +++R E E ++++ HK L + S E+ A+ YT I
Sbjct: 457 DKYIHKKITLKEFLRQYGVILQNRYEEESVADFDTCHKQPALKSPSPWEKQMATTYTHTI 516
Query: 492 FAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEF 551
F KF+ E+ + K + D ++V +C + D F V C ++E+
Sbjct: 517 FKKFQVEVLGVVACHPRKEKEDENMATFRVQDC-EKDDDFLVTWSKTKSELCCFCRMFEY 575
Query: 552 MGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVH 611
G LCRH L+I Q G IP +I++RWTKDA G+ GE +D + +
Sbjct: 576 KGFLCRHALMILQMCGFASIPPQYILKRWTKDAKSGVL-------AGEGADQI----QTR 624
Query: 612 VQRE---ASFLADLAEE---SEEAYNFIISEMKQTHKSAAAMKTSEGVVLLESSEKN 662
VQR S +L+EE SEE YN + + +T K+ M + + +S+ N
Sbjct: 625 VQRYNDLCSRATELSEEGCVSEENYNIALRTLVETLKNCVDMNNARNNITESNSQLN 681
>dbj|BAD37792.1| far-red impaired response protein-like [Oryza sativa (japonica
cultivar-group)] gi|51535335|dbj|BAD38595.1| far-red
impaired response protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 823
Score = 365 bits (938), Expect = 2e-99
Identities = 230/786 (29%), Positives = 380/786 (48%), Gaps = 95/786 (12%)
Query: 30 EIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVL-----VCCNEGQHKVKISRTE 84
+IG F FYN +A+ GFG+R ++ K + CC ++ R
Sbjct: 51 KIGQTFNEDSDGYAFYNLYARFTGFGIRRSKNRYKDGGVKSMQEFCC------IREGRDN 104
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSP-KSVSYMRC 143
+ R GC+A + ++R + KW + +F ++HNH M + + R
Sbjct: 105 SVTGPPT---------RIGCKAMVRLNRSSESQKWRVSAFVSEHNHEMKRDLQHTKHFRS 155
Query: 144 HKKMSVAAKSLVEKFEEEGI-PTGKVAAI--FNDGDST--FTNRDCWNYIRNVRRKNLDV 198
H + K +++ + G+ PT + + G S FT R +RR
Sbjct: 156 HNFIDEGTKRNIKEMVDNGMTPTAMYGLLSGMHGGPSLTPFTRRAVTRMAYAIRRDECS- 214
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
D Q ++ + Q + NFFY IQ D SR+ N+FW + SRLS+ FGD ITFDTTY+
Sbjct: 215 NDVQKTLNFFREMQCRSKNFFYTIQIDGASRIKNIFWCHSLSRLSFDHFGDAITFDTTYQ 274
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
TN+Y+MPF FVG+NNH Q+ ++GCALL++E+ +F WLF+T AM GK+P++I+TD
Sbjct: 275 TNRYNMPFGIFVGVNNHFQTAIFGCALLREETIEAFKWLFQTFTDAMHGKRPVAILTDNC 334
Query: 319 LAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEE 378
M+ A+ V+PE+ HR+C WH++K E L +IY K+S FK++ R + + FE+
Sbjct: 335 HQMEVAIKAVWPETIHRVCKWHVLKNAKENLGNIYSKRSSFKQEFHRVLNEPQTEAEFEK 394
Query: 379 EWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAS 438
W +M +YNL+ + +L+ ++ +++ W P Y R FFA M+TTQRSES+N +V S
Sbjct: 395 AWSDLMEQYNLESSVYLRRMWDMKKKWAPAYFREFFFARMSTTQRSESMNHVLKKYVKPS 454
Query: 439 TTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGE 498
++L F ++E R+EAE E +++ ++ T S +E+HA+ +YTR F++FK +
Sbjct: 455 SSLHGFAKRYENFYNDRIEAEDAEEHDTYNEKVSTLTSSPIEKHASRVYTRGAFSRFKEQ 514
Query: 499 LEKINRFTRHKIRRDGPKYVYQVSNCYD------ARDTFTVDIDLDSQIAKCGYELYEFM 552
+ F + ++V Q+ + D F V +DL Q CG +L+E +
Sbjct: 515 FKLSFSF---MVYHTSDQHVLQLVHIGDDTLQSWGSKEFKVQVDLTEQDLSCGCKLFEHL 571
Query: 553 GILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEK-SDTLKILRRVH 611
GI+C HI+ + G +IP +I++RWTKDA + ++ L +K + + + R
Sbjct: 572 GIICSHIIRVMVQYGFTEIPKKYILKRWTKDARDSIPKHLEESYLKDKEAASSRTYRNTL 631
Query: 612 VQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSE-------------------G 652
+ + A + L S E Y + + + M TS+ G
Sbjct: 632 LHKSALDMVRLGGTSSETYEKTVEVLTKLIGELQVMCTSQVVNNKEIHYGDRTIGKKPTG 691
Query: 653 VVLLES-----------------SEKNVNQVCSSEQVSEPPQLTIGN------PHVSQTK 689
V L +S E + Q S+ + S +T N P V +++
Sbjct: 692 VQLDDSVDSSDSEHGMSDEFCVADEDGIGQDVSAGEDSVDVDMTDVNEEDILPPEVRRSR 751
Query: 690 GRKKDGGKMTQNSRFKSGLEVSLNKSVVKRKA----------------CHECGEHGHNSR 733
GR + M++ + ++S K ++ C +CG HGH
Sbjct: 752 GRPRSTRLMSKGETSSKAKKKKASESTSKDESKNHAKGKKESTKQIRYCKQCGGHGHYKS 811
Query: 734 TCKKRN 739
TC +++
Sbjct: 812 TCGRKS 817
>dbj|BAD81770.1| putative far-red impaired response protein [Oryza sativa (japonica
cultivar-group)]
Length = 883
Score = 363 bits (932), Expect = 1e-98
Identities = 239/770 (31%), Positives = 374/770 (48%), Gaps = 85/770 (11%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRV-----RSSKPKRAVLVCCNEGQHKVKISRTEE 85
+G F++ FYN +A GFG+R+ + K + +CC+
Sbjct: 124 LGQRFKTERDAYNFYNVYAVSKGFGIRLDKDHMNTKKQRTMRQICCSH------------ 171
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSY-MRCH 144
K K ++R GC A + ++ S W + + HNH M V+ + H
Sbjct: 172 ---QGRNPKTKKPSVRIGCPAMMKINMSGAGSGWSVTKVVSAHNHPMKKSVGVTKNYQSH 228
Query: 145 KKMSVAAKSLVEKFEEEGIPT----GKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGD 200
++ + ++E+ + + G ++ + R + + R++ D
Sbjct: 229 NQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDRVAYAIRRDESSDD 288
Query: 201 AQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTN 260
Q D K Q + NFFY+IQ DE R+ N+FW A SRL+++ FGDVITFDTTYKTN
Sbjct: 289 MQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHFGDVITFDTTYKTN 348
Query: 261 KYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLA 320
KY+MPFAPFVG+NNH QS +GCALL++E+E SF WLF T + M GK PI I+TD +
Sbjct: 349 KYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGKVPIGILTDNCPS 408
Query: 321 MKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEW 380
M AA+ VFP + HR+C H++KK E + +IY K FK+ + + + + + F W
Sbjct: 409 MAAAIRTVFPNTIHRVCKSHVLKKAKEFMGNIYSKCHTFKKAFHKVLTQTLTEEEFVAAW 468
Query: 381 MRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTT 440
+++ +YNL+++ +L+ ++ IR W +Y FFAGM TTQRSES N F FV S++
Sbjct: 469 HKLIRDYNLEKSVYLRHIWDIRRKWAFVYFSHRFFAGMTTTQRSESANHVFKMFVSPSSS 528
Query: 441 LQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL- 499
+ FV ++++ +L+ E E +++ + + T S +E HA+ +YTR +F F EL
Sbjct: 529 MNGFVKRYDRFFNEKLQKEDSEEFQTSNDKVEIKTRSPIEIHASQVYTRAVFQLFSEELT 588
Query: 500 EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
+ I+ + + V S R + V DL+ + C +++E GILC HI
Sbjct: 589 DSISYMVKPGEDESTVQVVRMNSQESFLRKEYQVSCDLEREEFSCVCKMFEHKGILCSHI 648
Query: 560 LVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRR--VHV--QRE 615
L + G+ +IP +I++RWTKDA DT + G K D R +HV +R+
Sbjct: 649 LRVLVQYGLSRIPERYILKRWTKDAR----DTIPPHLHGYKDDVDASQSRSYIHVMLKRK 704
Query: 616 ASFLADLAEESEEAY-------NFIISEMK------------QTHKSAAAMKTSEGVVLL 656
+A +A + + + N ++ +MK + K AA K S VL
Sbjct: 705 TVEVAKIANKDVQTFKMAMTVMNKLLEDMKNQLSLDDGDNSREAPKRAARHK-SRSTVLT 763
Query: 657 ESSEKNVNQVCSSEQVSEPPQLTIGN-------PHVSQTKGRKK-----DGGKMTQNSRF 704
+ ++V + P + GN P +++GR K GG++ R
Sbjct: 764 KDGNEDVGGGDEDSEYDVPSEDEGGNLTADILPPLKKRSRGRPKVNRYKSGGEVASLKRR 823
Query: 705 KS-GLE----------VSLNKSVVK--------RKACHECGEHGHNSRTC 735
K G+E V V+ R+ CH C E GHNSRTC
Sbjct: 824 KEVGIEKKNTTNECESVEEENQVIDHDDEVPFPRRFCHTCHEAGHNSRTC 873
>gb|AAV31356.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 896
Score = 362 bits (930), Expect = 2e-98
Identities = 207/571 (36%), Positives = 311/571 (54%), Gaps = 22/571 (3%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLV-----CCNEGQHKVKISRTEE 85
+GM F+++ V +FY S+A + GF VRV K + ++ C EG K R ++
Sbjct: 100 VGMIFDTLTDVEKFYKSYAHEAGFSVRVGQHKKQNEEILFKRYYCSREGYIK---ERVKD 156
Query: 86 IQDSTNQTKRKCSTI---RSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMR 142
+ D + + KRK + R GCEA ++V G+ K K+ I S +H+H VSP +R
Sbjct: 157 VSDESGKKKRKTPYMMETRCGCEAHIVVKLGSDK-KYRISSMIGEHSHGFVSPDKRHLLR 215
Query: 143 CHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCW-----NYIRNVRRKNLD 197
++ +S AKS + + I T + + D F N C NY ++R K D
Sbjct: 216 SNRTVSERAKSTLFSCHKASIGTSQAFRLLQVSDGGFQNVGCTLRNLQNYYHDLRCKIKD 275
Query: 198 VGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTY 257
DAQ +RK+ N FFY D+ R+V +FW DA R +Y +FGDV++ D+TY
Sbjct: 276 A-DAQMFVGQLERKKEVNPAFFYEFMVDKQGRLVRVFWADAICRKNYSVFGDVLSVDSTY 334
Query: 258 KTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQ 317
TN+Y+M F PF G+N+H QS+ G + L DE SFVWLF+T L+A GG P IITD+
Sbjct: 335 STNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLKATGGVAPRLIITDE 394
Query: 318 DLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFE 377
D +MKAA+A++ P + HRLC+WHI++K PEK+ + F + +C+ GS FE
Sbjct: 395 DASMKAAIAQILPNTVHRLCMWHIMEKVPEKVGPSIREDGEFWDRLHKCVWGSEDSDDFE 454
Query: 378 EEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAG-MNTTQRSESINAFFDSFVD 436
EW IM +Y L NEW + IR+SWI Y AG ++TT RSES N+FF+ F+
Sbjct: 455 SEWNSIMAKYGLIGNEWFSTKFDIRQSWILAYFMDIPLAGILSTTSRSESANSFFNRFIH 514
Query: 437 ASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFK 496
T EF L+F+ A+E + + E K + S H + L T +E+ + IYT +F+KF+
Sbjct: 515 RKLTFVEFWLRFDTALECQRQEELKADNISLHTNPKLMTPWDMEKQCSGIYTHEVFSKFQ 574
Query: 497 GEL-EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGIL 555
+L + I + G + +S+ ++ V ++ + C +LYE GI
Sbjct: 575 EQLIVARDHCIIQGISKSGDMKIVTISSLFEKER--VVQMNKSNMFGTCSCKLYESYGIP 632
Query: 556 CRHILVIFQAKGVVQIPSHFIMERWTKDANR 586
CRHI+ + + + +IPS +IM+RW K R
Sbjct: 633 CRHIIQVLRGEKQNEIPSIYIMKRWEKICKR 663
>dbj|BAD37730.1| far-red impaired response protein-like [Oryza sativa (japonica
cultivar-group)] gi|51535313|dbj|BAD38573.1| far-red
impaired response protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 845
Score = 361 bits (926), Expect = 5e-98
Identities = 228/786 (29%), Positives = 379/786 (48%), Gaps = 95/786 (12%)
Query: 30 EIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVL-----VCCNEGQHKVKISRTE 84
+IG F FYN +++ GFG+R ++ K + CC ++ R
Sbjct: 51 KIGQTFNEDSDGYAFYNLYSRFTGFGIRRSKNRYKDGGVKSMQEFCC------IREGRDN 104
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSP-KSVSYMRC 143
+ R GC+A + ++R + KW + +F ++HNH M + + R
Sbjct: 105 SVTGPPT---------RIGCKAMVRLNRSSESQKWRVSAFVSEHNHEMKRDLQHTKHFRS 155
Query: 144 HKKMSVAAKSLVEKFEEEGI-PTGKVAAI--FNDGDST--FTNRDCWNYIRNVRRKNLDV 198
H + K +++ + G+ PT + + G S FT R +RR
Sbjct: 156 HNFIDEGTKRNIKEMVDNGMTPTAMYGLLSGMHGGPSLTPFTRRAVTRMAYAIRRDECS- 214
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
D Q ++ + Q + NFFY IQ D SR+ N+FW + SRLS+ FGD ITFDTTY+
Sbjct: 215 NDVQKTLNFFREMQCRSKNFFYTIQIDGASRIKNIFWCHSLSRLSFDHFGDAITFDTTYQ 274
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
TN+Y+MPF FVG+NNH Q+ ++GCALL++E+ +F WLF+T AM GK+P +I+TD
Sbjct: 275 TNRYNMPFGIFVGVNNHFQTAIFGCALLREETIEAFKWLFQTFTDAMHGKRPAAILTDNC 334
Query: 319 LAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEE 378
M+ A+ V+PE+ HR+C WH++K E L +IY K+S FK++ R + + FE+
Sbjct: 335 HQMEVAIKAVWPETIHRVCKWHVLKNAKENLGNIYSKRSSFKQEFHRVLNEPQTEAEFEK 394
Query: 379 EWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAS 438
W +M +YNL+ + +L+ ++ +++ W P Y R F+A M+TTQRSES+N +V S
Sbjct: 395 AWSDLMEQYNLESSVYLRRMWDMKKKWAPDYFREFFYARMSTTQRSESMNHVLKKYVKPS 454
Query: 439 TTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGE 498
++L F ++E R+EAE E +++ ++ T S +E+HA+ +YTR F++FK +
Sbjct: 455 SSLHGFAKRYENFYNDRIEAEDAEEHDTYNEKVSTLTSSPIEKHASRVYTRGAFSRFKEQ 514
Query: 499 LEKINRFTRHKIRRDGPKYVYQVSNCYD------ARDTFTVDIDLDSQIAKCGYELYEFM 552
+ F + ++V Q+ + D F V +DL Q CG +L+E +
Sbjct: 515 FKLSFSF---MVYHTSDQHVLQLVHIGDDTLQSWGSKEFKVQVDLTEQDLSCGCKLFEHL 571
Query: 553 GILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEK-SDTLKILRRVH 611
GI+C HI+ + G +IP +I++RWTKDA + ++ L +K + + + R
Sbjct: 572 GIICSHIIRVMVQYGFTEIPKKYILKRWTKDARDSIPKHLEESYLKDKEAASSRTYRNTL 631
Query: 612 VQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSE-------------------G 652
+ + A + L S E Y + + + M TS+ G
Sbjct: 632 LHKSALDMVRLGGTSSETYEKTVEVLTKLIGELQVMCTSQVVNNKEIHCGDRTIGKKPTG 691
Query: 653 VVLLES-----------------SEKNVNQVCSSEQVSEPPQLTIGN------PHVSQTK 689
V L +S E + Q S+ + S +T N P V +++
Sbjct: 692 VQLDDSVDSSDSEHGMSDEFCVADEDGIGQDVSAGEDSVDVDMTDVNEEDILPPEVRRSR 751
Query: 690 GRKKDGGKMTQNSRFKSGLEVSLNKSVVKRKA----------------CHECGEHGHNSR 733
GR + M++ + ++S K ++ C +CG HGH
Sbjct: 752 GRPRSTRLMSKGETSSKAKKKKASESTSKDESKNHAKGKKESTKQIRYCKQCGGHGHYKS 811
Query: 734 TCKKRN 739
TC +++
Sbjct: 812 TCGRKS 817
>gb|AAT94016.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|51038153|gb|AAT93956.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 909
Score = 358 bits (920), Expect = 3e-97
Identities = 229/786 (29%), Positives = 379/786 (48%), Gaps = 95/786 (12%)
Query: 30 EIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVL-----VCCNEGQHKVKISRTE 84
+IG F FYN +A+ GFG+R ++ K + CC ++ R
Sbjct: 137 KIGQTFNEDSHGYAFYNLYARFTGFGIRRSKNRYKDGGVKSMQEFCC------IREGRDN 190
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSP-KSVSYMRC 143
+ R G +A + ++R + KW + +F ++HNH M + + R
Sbjct: 191 SVTGPPT---------RIGYKAMVRLNRSSESQKWRVSAFVSEHNHEMKRDLQHTKHFRS 241
Query: 144 HKKMSVAAKSLVEKFEEEGI-PT---GKVAAIFNDGDST-FTNRDCWNYIRNVRRKNLDV 198
H + K +++ + G+ PT G ++ + T FT R +RR
Sbjct: 242 HNFIDEGTKRNIKEMVDNGMTPTTMYGLLSGMHGGPSLTPFTRRAVTRMAYAIRRDECS- 300
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
D Q ++ + Q + NFFY IQ D SR+ N+FW + SRLS+ FGD ITFDTTY+
Sbjct: 301 NDVQKTLNFFREMQCRSKNFFYTIQIDGASRIKNIFWCHSLSRLSFDHFGDAITFDTTYQ 360
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
TN+Y+MPF FVG+NNH Q+ ++GCALL++E+ +F WLF+T AM GK+P +I+TD
Sbjct: 361 TNRYNMPFGIFVGVNNHFQTAIFGCALLREETIEAFKWLFQTFTDAMHGKRPAAILTDNC 420
Query: 319 LAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEE 378
M+ A+ V+PE+ HR+C WH++K E L +IY K+S FK++ R + + FE+
Sbjct: 421 HQMEVAIKAVWPETIHRVCKWHVLKNAKENLGNIYSKRSSFKQEFHRVLNEPQTEAEFEK 480
Query: 379 EWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAS 438
W +M +YNL+ + +L+ ++ +++ W P Y R FFA M+TTQRSES+N +V S
Sbjct: 481 AWSDLMEQYNLESSVYLRRMWDMKKKWAPAYFREFFFARMSTTQRSESMNHVLKKYVKPS 540
Query: 439 TTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGE 498
++L F ++E R+EAE E +++ ++ T S +E+HA+ +YTR F++FK +
Sbjct: 541 SSLHGFAKRYENFYNDRIEAEDAEEHDTYNEKVSTLTSSPIEKHASRVYTRGEFSRFKEQ 600
Query: 499 LEKINRFTRHKIRRDGPKYVYQVSNCYD------ARDTFTVDIDLDSQIAKCGYELYEFM 552
+ F + ++V Q+ + D F V +DL Q CG +L+E +
Sbjct: 601 FKLSFSF---MVYHTSDQHVLQLVHIGDDTLQSWGSKEFKVQVDLTEQDLSCGCKLFEHL 657
Query: 553 GILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEK-SDTLKILRRVH 611
GI+C HI+ + G +IP +I++RWTKDA + ++ L +K + + + R
Sbjct: 658 GIICSHIIRVMVQYGFTEIPKKYILKRWTKDARDSIPKHLEESYLKDKEAASSRTYRNTL 717
Query: 612 VQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSE-------------------G 652
+ + A + L S E Y + + + M TS+ G
Sbjct: 718 LHKSALDMVRLGGTSSETYEKTVEVLTKLIGELQVMCTSQVVNNKEIHCGDRTIGKKPTG 777
Query: 653 VVLLES-----------------SEKNVNQVCSSEQVSEPPQLTIGN------PHVSQTK 689
V L +S E + Q S+ + S +T N P V +++
Sbjct: 778 VQLDDSVDSSDSEHGMSDEFCVADEDGIGQDVSAGEDSVDVDMTDVNEEDILPPEVRRSR 837
Query: 690 GRKKDGGKMTQNSRFKSGLEVSLNKSVVKRKA----------------CHECGEHGHNSR 733
GR + M++ + ++S K ++ C +CG HGH
Sbjct: 838 GRPRSTRLMSKGETSSKAKKKKASESTSKDESKNHAKGKKESTKQIRYCKQCGGHGHYKS 897
Query: 734 TCKKRN 739
TC +++
Sbjct: 898 TCGRKS 903
>gb|AAN52752.1| Unknown protein [Oryza sativa (japonica cultivar-group)]
gi|34902304|ref|NP_912498.1| Unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 909
Score = 358 bits (919), Expect = 4e-97
Identities = 227/786 (28%), Positives = 377/786 (47%), Gaps = 95/786 (12%)
Query: 30 EIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVL-----VCCNEGQHKVKISRTE 84
+IG F FYN +A+ GFG+R ++ K + CC ++ R
Sbjct: 137 KIGQTFNEDSDGYAFYNLYARFTGFGIRRSKNRYKDGGVKSMQEFCC------IREGRDN 190
Query: 85 EIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSP-KSVSYMRC 143
+ R GC+A + ++R + KW + +F ++HNH M + + R
Sbjct: 191 SVTGPPT---------RIGCKAMVRLNRSSESQKWRVSAFVSEHNHEMKRDLQHTKHFRS 241
Query: 144 HKKMSVAAKSLVEKFEEEGI-PTGKVAAI--FNDGDST--FTNRDCWNYIRNVRRKNLDV 198
H + K +++ + G+ PT + + G S FT R +RR
Sbjct: 242 HNFIDEGTKRNIKEMVDNGMTPTAMYGLLSGMHGGPSLTPFTRRAVTRMAYAIRRDECS- 300
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYK 258
D Q ++ + Q + NFFY IQ D SR+ N+FW + SRLS+ FGD ITFDTTY+
Sbjct: 301 NDVQKTLNFFREMQCRSKNFFYTIQIDGASRIKNIFWCHSLSRLSFDHFGDAITFDTTYQ 360
Query: 259 TNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQD 318
TN+Y+MPF FV +NNH Q+ ++GCALL++E+ +F WLF+T AM G +P +I+TD
Sbjct: 361 TNRYNMPFGIFVDVNNHFQTAIFGCALLREETIEAFKWLFQTFTDAMHGNRPAAILTDNC 420
Query: 319 LAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEE 378
M+ A+ V+PE+ HR+C WH++K E L +IY K+S FK++ R + + F++
Sbjct: 421 HQMEVAIKAVWPETIHRVCKWHVLKNAKENLGNIYSKRSSFKQEFHRVLNEPQTEAEFDK 480
Query: 379 EWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDAS 438
W +M +YNL+ + +L+ ++ +++ W P Y R FFA M+TTQRSES+N +V S
Sbjct: 481 AWSDLMEQYNLESSVYLRRMWDMKKKWAPAYFREFFFARMSTTQRSESMNHVLKKYVKPS 540
Query: 439 TTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGE 498
++L F ++E R+EAE E +++ ++ T S +E+HA+ +YTR F++FK +
Sbjct: 541 SSLHGFAKRYENFYNDRIEAEDAEEHDTYNEKVSTLTSSPIEKHASRVYTRGAFSRFKEQ 600
Query: 499 LEKINRFTRHKIRRDGPKYVYQVSNCYD------ARDTFTVDIDLDSQIAKCGYELYEFM 552
+ F + ++V Q+ + D F V +DL Q CG +L+E +
Sbjct: 601 FKLSFSF---MVYHTSDQHVLQLVHIGDDTLQSWGSKEFKVQVDLTEQDLSCGCKLFEHL 657
Query: 553 GILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEK-SDTLKILRRVH 611
GI+C HI+ + G +IP +I++RWTKDA + ++ L +K + + + R
Sbjct: 658 GIICSHIIRVMVQYGFTEIPKKYILKRWTKDARDSIPKHLEESYLKDKEAASSRTYRNTL 717
Query: 612 VQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSE-------------------G 652
+ + A + L S E Y + + + M TS+ G
Sbjct: 718 LHKSALDMVRLGGTSSETYEKTVEVLTKLIGELQVMCTSQVVNNKEIHCGDRTIGKKPTG 777
Query: 653 VVLLES-----------------SEKNVNQVCSSEQVSEPPQLTIGN------PHVSQTK 689
V L +S E + Q S+ + S +T N P V +++
Sbjct: 778 VQLDDSVDSSDSEPGMSDEFCVADEDGIGQDVSAGEDSVDVDMTDVNEEDILPPEVRRSR 837
Query: 690 GRKKDGGKMTQNSRFKSGLEVSLNKSVVKRKA----------------CHECGEHGHNSR 733
GR + M++ + ++S K ++ C +CG HGH
Sbjct: 838 GRPRSTRLMSKGETSSKAKKKKASESTSKDESKNHAKGKKESTKQIRYCKQCGGHGHYKS 897
Query: 734 TCKKRN 739
TC +++
Sbjct: 898 TCGRKS 903
>ref|XP_507459.1| PREDICTED OJ1135_F06.10-1 gene product [Oryza sativa (japonica
cultivar-group)] gi|51963940|ref|XP_506753.1| PREDICTED
OJ1135_F06.10-1 gene product [Oryza sativa (japonica
cultivar-group)]
Length = 721
Score = 358 bits (919), Expect = 4e-97
Identities = 207/632 (32%), Positives = 329/632 (51%), Gaps = 30/632 (4%)
Query: 15 TSILDTSDIGG--NSKLEIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKR------A 66
T L D GG N ++GM F S + +FYNS+A+ GF +R + +
Sbjct: 58 TPQLGDHDQGGLPNCVPKVGMTFNSENEAYDFYNSYARNVGFSIRKNHANTRADGSLCSK 117
Query: 67 VLVCCNEGQHKVKISRTEEIQDSTNQTKRKCSTIRSGCEASL--IVSRGTTKSKWMIKSF 124
+C NEGQ + +T ++K S+ RS C+A L +SR + W ++
Sbjct: 118 YFLCSNEGQ---------PVASTTQPGRKKRSSTRSDCKARLQFYISR---EGIWTVQKV 165
Query: 125 NNDHNHVMVSPKSVSYMRCHKKMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDC 184
DH H +VSP +R ++++ + + +V + +EGI + +F N
Sbjct: 166 ELDHKHYLVSPDKSHMLRSQRRLTPSYQQIVNEMRQEGITAADMQRVFRQWSRGAENVHL 225
Query: 185 WNYIRNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSY 244
++ +RK L AQ + +Y K KQ EN +FFYA+Q ++D R+ N FW D ++ + Y
Sbjct: 226 LK--KDTQRKYLQPSYAQKLLEYLKNKQTENPSFFYAVQLNDDGRIANFFWTDCQAIVDY 283
Query: 245 QLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQA 304
FGDV++FDTT++TNK+ MPFAPFVG N+H Q IL+G +L+ DES SF WLF+T L A
Sbjct: 284 ACFGDVVSFDTTFETNKFEMPFAPFVGTNHHKQPILFGASLIYDESSESFNWLFQTFLTA 343
Query: 305 MGGKKPISIITDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMK 364
M GK+P +I TD + ++ VFP S HRLCL HI + L H+ F D K
Sbjct: 344 MSGKQPATIFTDPSAEIIKSVRLVFPNSSHRLCLRHICHNAVKHLNHVICNHPEFLSDFK 403
Query: 365 RCIRGSHSIQSFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRS 424
R I S+ F+ +W ++ YNL N+W+ LY +RE W +Y+R +F+A M T Q +
Sbjct: 404 RRIYEERSVTFFDLKWKELVNAYNLDGNDWMNNLYAMREKWAAVYSRDSFYADMMTIQNA 463
Query: 425 ESINAFFDSFVDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRI-LSTGSKLEEHA 483
E + +F L EF+ ++EK + S + E + +Y SR S + + + A
Sbjct: 464 EGTSDALKNF-RRKLCLPEFLEEYEKCITSFRQNELEADYNSRQTSPVPYVPDLPMLKTA 522
Query: 484 ASIYTRNIFAKFKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAK 543
A YTRN+++ F+ E +K+ + + +D Y+++ + + V D+ ++
Sbjct: 523 AESYTRNLYSHFEEEFKKLFTLSCSLLSQDRTISTYKLTPLNSEEEAYVVFNSEDTTVS- 581
Query: 544 CGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDT 603
C +YE G+LC+H L + + PSH++ +RWTK A GL N++ G S
Sbjct: 582 CSCRMYECTGMLCKHALRVLNYSNIFTFPSHYVYKRWTKYAKAGLFGCRNNSQSGNGSSM 641
Query: 604 LKILRRVHVQREASFLADLAEESEEAYNFIIS 635
L+ R + ++ +A SE+A F+ S
Sbjct: 642 LRCAR---ISQKMHSVALRYSMSEKALQFLES 670
>ref|XP_463016.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|38093745|gb|AAR10861.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
Length = 813
Score = 357 bits (915), Expect = 1e-96
Identities = 217/651 (33%), Positives = 337/651 (51%), Gaps = 49/651 (7%)
Query: 30 EIGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPK------RAVLVCCNEGQHKVKISRT 83
++GM FES +K E YN++A K GF +R + K + + +VC +GQ
Sbjct: 86 QVGMSFESKDKAYEMYNTYAGKVGFSIRKSNVKRRSNGTIYQKHMVCNKQGQQ------- 138
Query: 84 EEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRC 143
+T T R+ C+A + S K WM++ +HNH +VSP +R
Sbjct: 139 --------ETSSSLDTTRTCCKARVQFSV-CRKEIWMVQKVVLEHNHDLVSPNKSHKLRS 189
Query: 144 HKKMSVAAKSLVEKFEEEGIPTGKVAAIFND-----GDSTFTNRDCWNYIRNVRRKNLDV 198
+++ A + L+ + E G+ +V + F+ DC N I R+K L+
Sbjct: 190 QRRVIEADRQLIGQIREAGMKPAQVYGFMKEWYGGADKVPFSKMDCNNEIGRERKKYLES 249
Query: 199 GDAQAVFDYCKRKQVENSNFFYAIQCD-EDSRMVNLFWVDARSRLSYQLFGDVITFDTTY 257
D Q + +Y + KQ+E+ FFYAIQ D ED R+ N FW D +S + Y FGD ++FDTT+
Sbjct: 250 NDTQTLLEYLRNKQLEDPTFFYAIQVDKEDGRIANFFWADGQSIMDYACFGDFVSFDTTF 309
Query: 258 KTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQ 317
TNK MPFAP +G N+H Q+I++G ALL +++ SFVWLF+T L AM GK P +I TDQ
Sbjct: 310 DTNKCEMPFAPLLGTNHHKQTIIFGAALLFNQTIESFVWLFETFLTAMSGKHPSTIFTDQ 369
Query: 318 DLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQ-SIFKRDMKRCIRGSHSIQSF 376
D AM AA+A VF + HRLCLWHI + L+H+ HK + F D KRC+ S F
Sbjct: 370 DAAMAAAIAFVFRNTSHRLCLWHIYLNGGKNLSHVIHKHPNKFLADFKRCVYEDRSEYYF 429
Query: 377 EEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVD 436
+ W ++ EYNL++N+W+ LYK+RE W ++ R++F A + +TQRSE +N +
Sbjct: 430 NKMWHELLSEYNLEDNKWISNLYKLREKWAIVF-RNSFTADITSTQRSEGMNNVYKKRFR 488
Query: 437 ASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGS-KLEEHAASIYTRNIFAKF 495
L E +++ +K + E E +++SRH + + + + + AA YTR+++++
Sbjct: 489 RKLGLSELLVECDKVSATLRENELDADFKSRHSNPVTYIPNLPMLKIAAESYTRSMYSEL 548
Query: 496 KGELEKINRFTRHKIRRDGPKYVYQVSNC-YDARDTFTVDIDLDSQIAKCGYELYEFMGI 554
+ E ++ + ++ +G + V YD TV + C YE +G+
Sbjct: 549 EDEFKQQFTLSCKLLKTEGATLTFVVMPMEYD--HEATVVFNPTEMTITCSCRKYECIGL 606
Query: 555 LCRHILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKI------LR 608
LC+H L +F V +PSH+I+ RWTK A G + G + +TLK
Sbjct: 607 LCKHALRVFNMNKVFTLPSHYILNRWTKYAKSG----FYIQKQGSEKETLKAHAARISRH 662
Query: 609 RVHVQREASFLADLAEESEEAYNFIISE-----MKQTHKSAAAMKTSEGVV 654
V+ + S +L ++ E+A N + E K KS S G V
Sbjct: 663 ATSVELKCSVSKELLDDLEQAINKLDLEADNSLSKMQEKSCEVPLNSNGCV 713
>ref|NP_567455.1| far-red impaired response protein (FAR1) / far-red impaired
responsive protein (FAR1) [Arabidopsis thaliana]
Length = 768
Score = 353 bits (907), Expect = 9e-96
Identities = 198/587 (33%), Positives = 319/587 (53%), Gaps = 26/587 (4%)
Query: 88 DSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKM 147
+S+ + R+ + ++ C+AS+ V R KW+I F DHNH ++ P + R + +
Sbjct: 50 ESSGSSSRRSTVKKTDCKASMHVKR-RPDGKWIIHEFVKDHNHELL-PALAYHFRIQRNV 107
Query: 148 SVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTN------RDCWNYIRNVRRKNLDVGDA 201
+A K+ ++ T K+ + + N D + + R L+ GD+
Sbjct: 108 KLAEKNNIDILHAVSERTKKMYVEMSRQSGGYKNIGSLLQTDVSSQVDKGRYLALEEGDS 167
Query: 202 QAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNK 261
Q + +Y KR + EN FFYAI +ED R+ NLFW DA+SR Y F DV++FDTTY
Sbjct: 168 QVLLEYFKRIKKENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFNDVVSFDTTYVKFN 227
Query: 262 YSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAM 321
+P A F+G+N+HSQ +L GCAL+ DES +FVWL KT L+AMGG+ P I+TDQD +
Sbjct: 228 DKLPLALFIGVNHHSQPMLLGCALVADESMETFVWLIKTWLRAMGGRAPKVILTDQDKFL 287
Query: 322 KAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWM 381
+A++++ P +RH LWH+++K PE +H+ + F +CI S + F+ W
Sbjct: 288 MSAVSELLPNTRHCFALWHVLEKIPEYFSHVMKRHENFLLKFNKCIFRSWTDDEFDMRWW 347
Query: 382 RIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTL 441
+++ ++ L+ +EWL L++ R+ W+P + F AGM+T+QRSES+N+FFD ++ TL
Sbjct: 348 KMVSQFGLENDEWLLWLHEHRQKWVPTFMSDVFLAGMSTSQRSESVNSFFDKYIHKKITL 407
Query: 442 QEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEK 501
+EF+ ++ +++R E E ++++ HK L + S E+ A+ YT IF KF+ E+
Sbjct: 408 KEFLRQYGVILQNRYEEESVADFDTCHKQPALKSPSPWEKQMATTYTHTIFKKFQVEVLG 467
Query: 502 INRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILV 561
+ K + D ++V +C + D F V C ++E+ G LCRH L+
Sbjct: 468 VVACHPRKEKEDENMATFRVQDC-EKDDDFLVTWSKTKSELCCFCRMFEYKGFLCRHALM 526
Query: 562 IFQAKGVVQIPSHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQRE---ASF 618
I Q G IP +I++RWTKDA G+ GE +D + + VQR S
Sbjct: 527 ILQMCGFASIPPQYILKRWTKDAKSGVL-------AGEGADQI----QTRVQRYNDLCSR 575
Query: 619 LADLAEE---SEEAYNFIISEMKQTHKSAAAMKTSEGVVLLESSEKN 662
+L+EE SEE YN + + +T K+ M + + +S+ N
Sbjct: 576 ATELSEEGCVSEENYNIALRTLVETLKNCVDMNNARNNITESNSQLN 622
>gb|AAT78829.1| putative FAR1 protein [Oryza sativa (japonica cultivar-group)]
Length = 1066
Score = 350 bits (898), Expect = 1e-94
Identities = 218/706 (30%), Positives = 351/706 (48%), Gaps = 50/706 (7%)
Query: 32 GMEFESIEKVREFYNSFAKKNGFGVRV-------RSSKPKRAVLVCCNEG------QHKV 78
GMEF+ E FYN +A + GF R+ R VC EG +++V
Sbjct: 71 GMEFDDEEDAWTFYNVYAHRVGFSTRISVMHRSRRDGSIMSRQFVCAKEGFRTYRGKNEV 130
Query: 79 KISRTEEIQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSV 138
+DS + +R + R GC+A + V + +W + HNH +V P
Sbjct: 131 SPVAAGSGEDS-GRGRRTRAVTRVGCKAMIRVKK-QDNGRWAVTKLETAHNHPLVPPNQA 188
Query: 139 SYMRCHKKMSVAAKSLVEKFEEEGIPT--GKVAAIFNDGDSTFTNRDCWNYIRNVRRKNL 196
+R HK +S K + GIP G + AI + ++ V
Sbjct: 189 HCLRPHKPLSECGKQ-----RQFGIPRNGGMLLAIEPPPPPISPPMPQTSVLQVVPHYTR 243
Query: 197 D-VGD-AQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFD 254
D +GD A+ + DY KR Q E+ FFYA+Q + + N+FW DAR+R++Y+ FGD + D
Sbjct: 244 DGIGDHARVILDYVKRMQAEDPAFFYAMQFIDGHPVGNIFWADARARMAYKHFGDAVFLD 303
Query: 255 TTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISII 314
K NKY +P F G+N+H Q +L+GCA++ D SEASFVWLF+TLL AM G P S+
Sbjct: 304 DYCKRNKYQLPLVAFTGVNHHCQPVLFGCAIIGDNSEASFVWLFETLLLAMSGHHPDSLT 363
Query: 315 TDQDLAMKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQ 374
T+ D A++ A KV P +RHR C WHI+ + +KL+H+ + ++ CI +I
Sbjct: 364 TEHDSAIQLAALKVLPRTRHRFCRWHILNETHDKLSHLSDEFPSLHEELVNCINMPETID 423
Query: 375 SFEEEWMRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSF 434
FE + ++ + +EWL +Y R+ W+P+Y R TFF ++ + S ++FFD +
Sbjct: 424 EFEVNFKALISKVGPGNSEWLYSVYNCRQHWVPVYLRDTFFGDESSKEECASRSSFFDGY 483
Query: 435 VDASTTLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAK 494
+ A T Q F+ ++EKA++ E E KE +E+++ + T S +E+ A +YTR++F K
Sbjct: 484 ISAKTDPQSFIQQYEKALDCCYEKEVKEEFETKYSLPEIKTSSPIEKQGADLYTRSMFLK 543
Query: 495 FKGELEKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGI 554
F+ EL + + ++ DG +Y+V+ + V+ + A C +++E +GI
Sbjct: 544 FQQELVDASVSSLEVMKEDGKSRIYKVTKSAGSEKPHMVEFNSFGSSATCSCQMFEHLGI 603
Query: 555 LCRHILVIFQAKGVVQIPSHFIMERWTKDA---------------NRGLEDTYNDNDLGE 599
+CRHIL +F +GV +PS +I++RWTK A N E+ + + GE
Sbjct: 604 VCRHILTVFGTQGVSSLPSQYIVKRWTKYAMERSPDKKIDEVSKVNEPKEEQKSGAEDGE 663
Query: 600 KSDTLKILRRVHVQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSEGVVLLES- 658
+S T R + REA A+ S E Y + +++ K G V +
Sbjct: 664 QSQT---WRYNSLCREALRYAEEGASSVEVYIVAMQALQEAANKVNMAKRGIGQVAPNAP 720
Query: 659 -SEKNVNQVCSSEQVSEPPQLTIGNPHVSQTKGRKKD-GGKMTQNS 702
+ + +E P+++ +Q K RK++ K T+NS
Sbjct: 721 LAVMPIAAQLPAEGFRNVPEISF-----NQRKKRKRNSNNKTTENS 761
>gb|AAF66982.1| transposase [Zea mays]
Length = 709
Score = 350 bits (897), Expect = 1e-94
Identities = 227/682 (33%), Positives = 352/682 (51%), Gaps = 43/682 (6%)
Query: 31 IGMEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVL-----VCCNEGQHKVKISRTEE 85
IGM F+S++++ FY ++A + GF VR+ + K V+ VC EG T
Sbjct: 27 IGMSFDSLDELEGFYKTYAHECGFSVRIGAQGKKNDVVEHKRFVCSREGF-------TRR 79
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHK 145
++ NQ K R GC A + V G K ++ I SF +HNH +VSP + ++R ++
Sbjct: 80 CAEAKNQKKH--FETRCGCNARVYVRLGQDK-RYYIASFVEEHNHGLVSPDKIPFLRSNR 136
Query: 146 KMSVAAKSLVEKFEEEGIPTGKVAAIFNDGDSTFTN-----RDCWNYIRNVRRKNLDVGD 200
+ AK+ + + I T + + D F N RD NY R +R K + D
Sbjct: 137 TICQRAKTTLFTCHKASIGTSQAYRLLQVSDG-FDNIGCMKRDLQNYYRGLREK-IKNAD 194
Query: 201 AQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTN 260
AQ +RK+ NS FFY DE ++V + W DA R SY FGD+++ D TY TN
Sbjct: 195 AQLFVAQMERKKEANSAFFYDFAVDEHGKLVYICWADATCRKSYTHFGDLVSVDATYSTN 254
Query: 261 KYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLA 320
+Y+M FAPF G+N+H Q + +G A L +E S+ WLF+T L AMGGK P IITD+D +
Sbjct: 255 QYNMRFAPFTGVNHHMQRVFFGAAFLANEKIESYEWLFRTFLVAMGGKAPRLIITDEDAS 314
Query: 321 MKAALAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEW 380
+K+A+ P++ HRLC+WHI++K EK+ H F + C+ GS + + FE W
Sbjct: 315 IKSAIRTTLPDTIHRLCMWHIMEKVSEKVGHPTSHDKEFWDALNTCVWGSETPEEFEMRW 374
Query: 381 MRIMVEYNLKENEWLQGLYKIRESWIPIYNRSTFFAG-MNTTQRSESINAFFDSFVDAST 439
+M Y L+ NEWL YKIRESWIP + T AG + TT RSES N+FF+ F+
Sbjct: 375 NALMDAYGLESNEWLANRYKIRESWIPAFFMDTPLAGVLRTTSRSESANSFFNRFIHRKL 434
Query: 440 TLQEFVLKFEKAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGEL 499
EF L+F+ A+E + E K ++ S H + +L T +E+ A+ +YT +F F+ E+
Sbjct: 435 CFVEFWLRFDTALERQRHEELKADHISIHSTPVLRTPWVVEKQASILYTHKVFKIFQEEV 494
Query: 500 --EKINRFTRHKIRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCR 557
+ + ++D K+V V + RD V + +C +L+E +GI C
Sbjct: 495 IAARDHCSVLGTTQQDAVKFV--VVSDGSMRDR-VVQWCTSNIFGRCSCKLFEKIGIPCC 551
Query: 558 HILVIFQAKGVVQIPSHFIMERWTKDANRGLEDTYND--NDLGEK----SDTLKILRRVH 611
HI++ + + + ++PS +I++RW R E Y+D N L EK ++ K +
Sbjct: 552 HIILAMRGEKLYELPSSYILKRWETRCKR--ECVYDDDGNLLEEKPEDANEAEKRKKITV 609
Query: 612 VQREASFLADLAEESEEAYNFIISEMKQTHKSAAAMKTSEGVVLLESSEKNVNQVCSSEQ 671
V+ + A+ S EA +F++S + +S + +S E E +
Sbjct: 610 VRNKIEEAIQRAKSSNEAMDFLVSSVLNIGESLGHIVSSMVQPTQEEYENFIG------- 662
Query: 672 VSEPPQLTIGNPHVSQTKGRKK 693
P + I P+ ++KGR K
Sbjct: 663 CKIPADIQIHPPNDVRSKGRSK 684
>ref|NP_177759.1| far-red impaired responsive protein, putative [Arabidopsis
thaliana] gi|6573712|gb|AAF17632.1| T23E18.25
[Arabidopsis thaliana]
Length = 732
Score = 350 bits (897), Expect = 1e-94
Identities = 212/694 (30%), Positives = 356/694 (50%), Gaps = 28/694 (4%)
Query: 33 MEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDSTNQ 92
MEFE+ E FY +AK GFG SS+ RA + ++ ++ D+ N
Sbjct: 1 MEFETHEDAYLFYKDYAKSVGFGTAKLSSRRSRASKEFIDAKFSCIRYGSKQQSDDAINP 60
Query: 93 TKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVAAK 152
++ + GC+AS+ V R KW + SF +HNH ++ P+ Y R H+ +
Sbjct: 61 R----ASPKIGCKASMHVKR-RPDGKWYVYSFVKEHNHDLL-PEQAHYFRSHRNTELVKS 114
Query: 153 SLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQ 212
+ ++ P + D F + N RR LD GDA+ + ++ R Q
Sbjct: 115 NDSRLRRKKNTPLTDCKHLSAYHDLDFIDGYMRNQHDKGRRLVLDTGDAEILLEFLMRMQ 174
Query: 213 VENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGL 272
EN FF+A+ ED + N+FWVDA+ Y+ F DV++F+T+Y +KY +P FVG+
Sbjct: 175 EENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVLFVGV 234
Query: 273 NNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFPES 332
N+H Q +L GC LL D++ ++VWL ++ L AMGG+KP ++TDQ+ A+KAA+A V PE+
Sbjct: 235 NHHVQPVLLGCGLLADDTVYTYVWLMQSWLVAMGGQKPKVMLTDQNNAIKAAIAAVLPET 294
Query: 333 RHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLKEN 392
RH CLWH++ + P L + Q F + + +CI S S + F+ W++++ +++L++
Sbjct: 295 RHCYCLWHVLDQLPRNLDYWSMWQDTFMKKLFKCIYRSWSEEEFDRRWLKLIDKFHLRDV 354
Query: 393 EWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEKAV 452
W++ LY+ R+ W P + R FAG++ RSES+N+ FD +V T+L+EF+ + +
Sbjct: 355 PWMRSLYEERKFWAPTFMRGITFAGLSMRCRSESVNSLFDRYVHPETSLKEFLEGYGLML 414
Query: 453 ESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIRR 512
E R E E K ++++ H++ L + S E+ +Y+ IF +F +LE + H +
Sbjct: 415 EDRYEEEAKADFDAWHEAPELKSPSPFEKQMLLVYSHEIFRRF--QLEVLGAAACHLTKE 472
Query: 513 DGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIP 572
Y V + +D + VD D C +E+ G LCRH +V+ Q GV IP
Sbjct: 473 SEEGTTYSVKD-FDDEQKYLVDWDEFKSDIYCSCRSFEYKGYLCRHAIVVLQMSGVFTIP 531
Query: 573 SHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNF 632
+++++RWT +A R + +L + + I R + R A L + S+E+Y+
Sbjct: 532 INYVLQRWT-NAARNRHQISRNLELVQSN----IRRFNDLCRRAIILGEEGSLSQESYDI 586
Query: 633 IISEMKQTHKSAAA--------MKTSEGVVLLESSEKNVNQVCS-SEQVSEPPQLTIGN- 682
+ MK+ K A + E + + NQ S S Q+ P + GN
Sbjct: 587 AMFAMKEAFKQCAVTINTIKHPARCEEAAIQAGDPVQEENQYGSTSTQIGPEPNIHAGNV 646
Query: 683 PHVSQTKGRKKDGGKMTQNSRFKSGLEVSLNKSV 716
P ++T+ K+ + N+ K V+ +++V
Sbjct: 647 PWQAETRREKRS----SLNNTSKKAKHVAQSETV 676
>gb|AAQ62876.1| At1g76320 [Arabidopsis thaliana] gi|62320160|dbj|BAD94369.1|
putative phytochrome A signaling protein [Arabidopsis
thaliana]
Length = 732
Score = 349 bits (896), Expect = 2e-94
Identities = 212/694 (30%), Positives = 356/694 (50%), Gaps = 28/694 (4%)
Query: 33 MEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDSTNQ 92
MEFE+ E FY +AK GFG SS+ RA + ++ ++ D+ N
Sbjct: 1 MEFETHEDAYLFYKDYAKSVGFGTAKLSSRRSRASKEFIDAKFSCIRYGSKQQSDDAINP 60
Query: 93 TKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVAAK 152
++ + GC+AS+ V R KW + SF +HNH ++ P+ Y R H+ +
Sbjct: 61 R----ASPKIGCKASMHVKR-RPDGKWYVYSFVKEHNHDLL-PEQAHYFRSHRNTELVKS 114
Query: 153 SLVEKFEEEGIPTGKVAAIFNDGDSTFTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQ 212
+ ++ P + D F + N RR LD GDA+ + ++ R Q
Sbjct: 115 NDSRLRRKKNTPLTDCKHLSAYHDLDFIDGYMRNQHDKGRRLVLDTGDAEILLEFLMRMQ 174
Query: 213 VENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGL 272
EN FF+A+ ED + N+FWVDA+ Y+ F DV++F+T+Y +KY +P FVG+
Sbjct: 175 EENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVLFVGV 234
Query: 273 NNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAALAKVFPES 332
N+H Q +L GC LL D++ ++VWL ++ L AMGG+KP ++TDQ+ A+KAA+A V PE+
Sbjct: 235 NHHVQPVLLGCGLLADDTVYTYVWLMQSWLVAMGGQKPKVMLTDQNNAIKAAIAAVLPET 294
Query: 333 RHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIMVEYNLKEN 392
RH CLWH++ + P L + Q F + + +CI S S + F+ W++++ +++L++
Sbjct: 295 RHCYCLWHVLDQLPRNLDYWSMWQDTFMKKLFKCIYRSWSEEEFDRRWLKLIDKFHLRDV 354
Query: 393 EWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFEKAV 452
W++ LY+ R+ W P + R FAG++ RSES+N+ FD +V T+L+EF+ + +
Sbjct: 355 PWMRSLYEERKFWAPTFMRGITFAGLSMRCRSESVNSLFDRYVHPETSLKEFLEGYGLML 414
Query: 453 ESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHKIRR 512
E R E E K ++++ H++ L + S E+ +Y+ IF +F +LE + H +
Sbjct: 415 EDRYEEEAKADFDAWHEAPELKSPSPFEKQMLLVYSHEIFRRF--QLEVLGAAACHLTKE 472
Query: 513 DGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHILVIFQAKGVVQIP 572
Y V + +D + VD D C +E+ G LCRH +V+ Q GV IP
Sbjct: 473 SEEGTTYSVKD-FDDEQKYLVDWDEFKSDIYCSCRSFEYKGYLCRHAIVVLQMSGVFTIP 531
Query: 573 SHFIMERWTKDANRGLEDTYNDNDLGEKSDTLKILRRVHVQREASFLADLAEESEEAYNF 632
+++++RWT +A R + +L + + I R + R A L + S+E+Y+
Sbjct: 532 INYVLQRWT-NAARNRHQISRNLELVQSN----IRRFNDLCRRAIILGEEGSLSQESYDV 586
Query: 633 IISEMKQTHKSAAA--------MKTSEGVVLLESSEKNVNQVCS-SEQVSEPPQLTIGN- 682
+ MK+ K A + E + + NQ S S Q+ P + GN
Sbjct: 587 AMFAMKEAFKQCAVTINTIKHPARCEEAAIQAGDPVQEENQYGSTSTQIGPEPNIHAGNV 646
Query: 683 PHVSQTKGRKKDGGKMTQNSRFKSGLEVSLNKSV 716
P ++T+ K+ + N+ K V+ +++V
Sbjct: 647 PWQAETRREKRS----SLNNTSKKAKHVAQSETV 676
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,183,706,463
Number of Sequences: 2540612
Number of extensions: 47838354
Number of successful extensions: 143990
Number of sequences better than 10.0: 464
Number of HSP's better than 10.0 without gapping: 250
Number of HSP's successfully gapped in prelim test: 214
Number of HSP's that attempted gapping in prelim test: 142998
Number of HSP's gapped (non-prelim): 716
length of query: 742
length of database: 863,360,394
effective HSP length: 136
effective length of query: 606
effective length of database: 517,837,162
effective search space: 313809320172
effective search space used: 313809320172
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0091b.9


