

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.10
(353 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
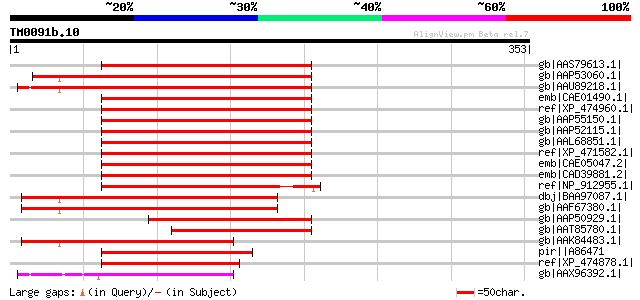
gb|AAS79613.1| putative copia-like polyprotein [Ipomoea trifida] 177 4e-43
gb|AAP53060.1| putative retroelement [Oryza sativa (japonica cul... 171 3e-41
gb|AAU89218.1| integrase core domain containing protein [Oryza s... 167 6e-40
emb|CAE01490.1| P0041A24.2 [Oryza sativa (japonica cultivar-grou... 164 3e-39
ref|XP_474960.1| OSJNBa0036B17.5 [Oryza sativa (japonica cultiva... 163 8e-39
gb|AAP55150.1| putative polyprotein [Oryza sativa (japonica cult... 162 1e-38
gb|AAP52115.1| putative copia-type pol polyprotein [Oryza sativa... 162 1e-38
gb|AAL68851.1| putative copia polyprotein [Sorghum bicolor] 162 2e-38
ref|XP_471582.1| OSJNBa0065J03.17 [Oryza sativa (japonica cultiv... 157 3e-37
emb|CAE05047.2| OSJNBa0049H08.8 [Oryza sativa (japonica cultivar... 157 5e-37
emb|CAD39881.2| OSJNBb0067G11.4 [Oryza sativa (japonica cultivar... 155 1e-36
ref|NP_912955.1| unnamed protein product [Oryza sativa (japonica... 154 3e-36
dbj|BAA97087.1| copia-type pol polyprotein-like [Arabidopsis tha... 142 1e-32
gb|AAF67380.1| Hypothetical protein T15F17.l [Arabidopsis thaliana] 140 4e-32
gb|AAP50929.1| putative polyprotein [Oryza sativa (japonica cult... 132 2e-29
gb|AAT85780.1| zinc knuckle domain containing protein [Oryza sat... 119 1e-25
gb|AAK84483.1| putative copia-like polyprotein [Lycopersicon esc... 116 9e-25
pir||A86471 F11O6.4 protein - Arabidopsis thaliana gi|12320956|g... 114 3e-24
ref|XP_474878.1| OSJNBa0021F22.5 [Oryza sativa (japonica cultiva... 101 3e-20
gb|AAX96392.1| retrotransposon protein, putative, Ty1-copia sub-... 89 2e-16
>gb|AAS79613.1| putative copia-like polyprotein [Ipomoea trifida]
Length = 1190
Score = 177 bits (449), Expect = 4e-43
Identities = 85/144 (59%), Positives = 114/144 (79%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L IE+L + +F++Q Y+ K+L+ FYMDKS PL T +V+RSL +KDPFRP E NE
Sbjct: 879 FCLGLQIEHLPNGIFIHQANYVNKLLEKFYMDKSHPLSTPMVVRSLEADKDPFRPKEDNE 938
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
++L PEVPYLSAI ALLYLA+ TR DI+F++NLLAR+S+SPT+RHW+G K +FRYL+G
Sbjct: 939 DILGPEVPYLSAIGALLYLANCTRPDIAFSVNLLARFSASPTRRHWSGVKHIFRYLQGTK 998
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ L+F + L+GY+DAGY+S
Sbjct: 999 DLGLYFENNQDITLMGYSDAGYMS 1022
>gb|AAP53060.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37532942|ref|NP_920773.1| putative retroelement
[Oryza sativa (japonica cultivar-group)]
gi|21671995|gb|AAM74357.1| Putative retroelement [Oryza
sativa (japonica cultivar-group)]
Length = 852
Score = 171 bits (433), Expect = 3e-41
Identities = 97/194 (50%), Positives = 128/194 (65%), Gaps = 4/194 (2%)
Query: 16 CIFMMRYEIVYVIINVY--DNNLLRL*KSSQKL*IC*RKNLR*KS*-E*NFCLALSIEYL 72
C+FM R E + II+VY D N++ + ++ + K + +CL L +E+
Sbjct: 498 CLFMKRSEHGFCIISVYVDDLNIIGTTQVIKEASSYLKTEFEMKELGKTTYCLGLQLEHT 557
Query: 73 KDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPYL 132
+ V ++Q YI K+L+ F M S P T +V+RSL V DPFRP E NEELL PE PYL
Sbjct: 558 PEGVLLHQYAYIQKILEKFNMKDSYPTRTPMVVRSLAVESDPFRPQENNEELLGPECPYL 617
Query: 133 SAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMS 191
SAI AL+YLA+ TR DI+FA+NLLAR+SS+PT+RHWNG KQ+FRYL+G D+ LFF
Sbjct: 618 SAIGALMYLANGTRPDIAFAVNLLARFSSAPTKRHWNGVKQIFRYLRGTQDLGLFFRKNQ 677
Query: 192 KLNLIGYADAGYLS 205
L + GYADAGYLS
Sbjct: 678 DLTMAGYADAGYLS 691
>gb|AAU89218.1| integrase core domain containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 1419
Score = 167 bits (422), Expect = 6e-40
Identities = 96/204 (47%), Positives = 134/204 (65%), Gaps = 5/204 (2%)
Query: 6 GRSQNDPTNSCIFMMRYEIVYVIINVY--DNNLLRL*KSSQKL*IC*RKNLR*KS*-E*N 62
G + ND C+F+ + E + II+VY D N++ + ++ + K +
Sbjct: 1056 GYANNDDC-LCVFIKKSENGFCIISVYVDDLNIIGTTRDIEEASTYLKAEFEMKDLGKTK 1114
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L VF++Q TY KVL+ F MDKS PL T +V+RSL V+KDPFRP E++E
Sbjct: 1115 FCLGLQLEHLPKGVFVHQSTYTKKVLEMFNMDKSHPLKTPMVVRSLEVDKDPFRPKEEDE 1174
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
+ L PEVPYLSAI L+YL + TR DI+FA+NLLARYS++PT+RHW G K + +YL+G
Sbjct: 1175 KPLGPEVPYLSAIGPLMYLTNCTRPDIAFAVNLLARYSAAPTRRHWVGVKTILKYLQGTR 1234
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ L+FP L + GY DAGY+S
Sbjct: 1235 DLGLWFPRKGDLTMEGYVDAGYMS 1258
>emb|CAE01490.1| P0041A24.2 [Oryza sativa (japonica cultivar-group)]
gi|50924536|ref|XP_472628.1| P0041A24.2 [Oryza sativa
(japonica cultivar-group)]
Length = 1376
Score = 164 bits (416), Expect = 3e-39
Identities = 81/144 (56%), Positives = 109/144 (75%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L + VFM+Q TY +VL F MDK PL T +V+RSL +KDPF+P E +E
Sbjct: 1072 FCLGLQLEHLPEGVFMHQSTYTKRVLGKFNMDKCHPLKTPMVVRSLEADKDPFKPKEDDE 1131
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
++L PEVPYL+AI AL+YLA+ TR DI+FA+NLLARYS++PT+RHW G K + RYL+G
Sbjct: 1132 KVLGPEVPYLNAIGALMYLANCTRPDIAFAVNLLARYSAAPTRRHWVGVKTILRYLRGTQ 1191
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ L+FP +++GY DAGY+S
Sbjct: 1192 DLGLWFPKNQDPSMVGYVDAGYMS 1215
>ref|XP_474960.1| OSJNBa0036B17.5 [Oryza sativa (japonica cultivar-group)]
gi|38344529|emb|CAD39862.2| OSJNBa0036B17.5 [Oryza
sativa (japonica cultivar-group)]
Length = 443
Score = 163 bits (412), Expect = 8e-39
Identities = 82/144 (56%), Positives = 104/144 (71%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L + ++Q Y K+L+ F MDKS P T +V+RSL+V KDPFRP E E
Sbjct: 136 FCLGLQLEHLPSGILVHQSAYAQKMLEKFNMDKSYPSKTPMVVRSLDVEKDPFRPKEDGE 195
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
++L P+ PYLSAI AL+YLA+ TR DI+F++NLLARYS++PT+RHW G K VFRYL G
Sbjct: 196 DVLGPDFPYLSAIGALMYLANSTRPDIAFSVNLLARYSAAPTKRHWTGVKNVFRYLNGTR 255
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ LFF LIGY DAGYLS
Sbjct: 256 DLGLFFKKNQGSTLIGYTDAGYLS 279
>gb|AAP55150.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37537122|ref|NP_922863.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|18266644|gb|AAL67590.1| putative polyprotein [Oryza
sativa]
Length = 419
Score = 162 bits (411), Expect = 1e-38
Identities = 83/144 (57%), Positives = 106/144 (72%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L + + ++Q TY KVL+ F M K+ PL T +V+RSL+ +KD FRP + E
Sbjct: 111 FCLVLQVEHLPNGILVHQSTYTKKVLERFNMSKAHPLKTPMVVRSLDKDKDLFRPRSETE 170
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
E L PE+PYLSAI AL+YLA+ TR DI+F +NLLARYS SPT+RHW G K V RYL+G
Sbjct: 171 EELGPEIPYLSAIGALMYLANCTRPDIAFVVNLLARYSVSPTRRHWVGVKTVLRYLQGTQ 230
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ LFFP L ++GYADAGYLS
Sbjct: 231 DLGLFFPRQRDLTMVGYADAGYLS 254
>gb|AAP52115.1| putative copia-type pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37531052|ref|NP_919828.1| putative
copia-type pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|15209145|gb|AAK91878.1| Putative
copia-type pol polyprotein [Oryza sativa]
Length = 1225
Score = 162 bits (411), Expect = 1e-38
Identities = 80/144 (55%), Positives = 109/144 (75%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +++L + VFM+Q TY +VL F MDK PL T +V+RSL +KDPF+P E +E
Sbjct: 921 FCLGLQLKHLPEGVFMHQSTYTKRVLGKFNMDKCHPLKTPMVVRSLEADKDPFKPKEDDE 980
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
E+L PEVPYLSAI AL+YLA+ TR +I+FA+NLLARYS++PT+RHW G K + +YL+G
Sbjct: 981 EVLGPEVPYLSAIGALMYLANCTRPNIAFAVNLLARYSATPTRRHWVGVKTILKYLRGTQ 1040
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ L+FP +++GY DAGY+S
Sbjct: 1041 DLGLWFPKNQDPSMVGYVDAGYMS 1064
>gb|AAL68851.1| putative copia polyprotein [Sorghum bicolor]
Length = 1082
Score = 162 bits (409), Expect = 2e-38
Identities = 78/144 (54%), Positives = 106/144 (73%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L +F++Q Y+ KVL+ F MDK+ P T +V+RSL +N DPFRP ++ E
Sbjct: 825 FCLGLQLEHLSTGIFVHQSAYVQKVLERFNMDKAYPSKTPMVVRSLGLNTDPFRPQDEGE 884
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
E+L P VPYLSA+ AL+YLA+ TR DI+FA+NLLAR+S++PT+RHW G KQ+ RY+ G
Sbjct: 885 EILGPHVPYLSAVGALMYLANSTRPDIAFAVNLLARHSAAPTKRHWTGVKQILRYVNGTR 944
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ LFF + +IGY DAGY S
Sbjct: 945 DLGLFFQRGTNPEMIGYTDAGYQS 968
>ref|XP_471582.1| OSJNBa0065J03.17 [Oryza sativa (japonica cultivar-group)]
gi|38568047|emb|CAD40421.3| OSJNBa0065J03.17 [Oryza
sativa (japonica cultivar-group)]
Length = 864
Score = 157 bits (398), Expect = 3e-37
Identities = 79/144 (54%), Positives = 101/144 (69%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L + ++Q Y K+L+ F MDKS P +V+RSL+V KDPFRP E E
Sbjct: 557 FCLGLQLEHLPSGILVHQSAYTQKILEKFNMDKSYPSKIPMVVRSLDVEKDPFRPKEDGE 616
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
++L P+ PYLSAI AL+YLA+ TR DI+F++NLLARYS++PT+ HW G K VFRYL G
Sbjct: 617 DVLGPDFPYLSAIGALMYLANSTRPDIAFSVNLLARYSAAPTKHHWTGVKNVFRYLNGTR 676
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ LFF LIGY DA YLS
Sbjct: 677 DLGLFFKKNQDSTLIGYTDASYLS 700
>emb|CAE05047.2| OSJNBa0049H08.8 [Oryza sativa (japonica cultivar-group)]
gi|50923521|ref|XP_472121.1| OSJNBa0049H08.8 [Oryza
sativa (japonica cultivar-group)]
Length = 810
Score = 157 bits (397), Expect = 5e-37
Identities = 79/144 (54%), Positives = 102/144 (69%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +++L + ++Q Y K+L+ F MDKS P T +V+RSL+V KDPFRP E E
Sbjct: 660 FCLGLQLDHLPSRILVHQSAYTQKILEKFNMDKSYPSKTPMVVRSLDVEKDPFRPKEYGE 719
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
++L P+ PYLSAI AL+YLA+ TR DI+F++NLLARYS++PT+RHW K VFRYL G
Sbjct: 720 DVLGPDFPYLSAIGALMYLANSTRPDIAFSVNLLARYSAAPTKRHWTEVKNVFRYLNGTR 779
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ LFF LIGY DA YLS
Sbjct: 780 DLGLFFKKNQDSTLIGYTDARYLS 803
>emb|CAD39881.2| OSJNBb0067G11.4 [Oryza sativa (japonica cultivar-group)]
gi|38344721|emb|CAE05263.2| OSJNBb0115I09.25 [Oryza
sativa (japonica cultivar-group)]
gi|50922245|ref|XP_471483.1| OSJNBb0115I09.25 [Oryza
sativa (japonica cultivar-group)]
Length = 986
Score = 155 bits (393), Expect = 1e-36
Identities = 78/144 (54%), Positives = 107/144 (74%), Gaps = 1/144 (0%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L +E+L + VFM+Q TY +VL F M+K PL T +V++SL +KDPF+P E +E
Sbjct: 836 FCLDLQLEHLPEGVFMHQSTYTKRVLGKFNMNKCHPLKTPMVVQSLEADKDPFKPKEDDE 895
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
E+L PEVPYLSAI AL+YLA+ TR DI FA+NLLARYS++PT+RHW G K + YL+G
Sbjct: 896 EVLGPEVPYLSAIGALMYLANCTRPDIVFAVNLLARYSATPTRRHWVGVKTILTYLRGTQ 955
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS 205
D+ L+FP +++GY +AG++S
Sbjct: 956 DLGLWFPKNQDPSMVGYVNAGHMS 979
>ref|NP_912955.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 1321
Score = 154 bits (390), Expect = 3e-36
Identities = 82/153 (53%), Positives = 107/153 (69%), Gaps = 12/153 (7%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
+CL L +E+ + V ++Q YI K+L+ F + S P T +V+RSL V DPFRP E N+
Sbjct: 1078 YCLGLQLEHTHEGVLLHQSAYIQKILEKFNIKDSYPTRTPMVVRSLAVESDPFRPQENNQ 1137
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
ELL PE PYLSAI AL+YLA+ TR DI+FA+NLLAR+SS+PT+RHWNG K++FRYL+G
Sbjct: 1138 ELLGPECPYLSAIGALMYLANGTRPDIAFAVNLLARFSSAPTKRHWNGVKKIFRYLRGTQ 1197
Query: 182 DMSLFFPFMSKLNLIGYADAGYLS---KYLKQV 211
D+ L + GYADAGYLS K L Q+
Sbjct: 1198 DLD--------LTMTGYADAGYLSDPHKALSQI 1222
>dbj|BAA97087.1| copia-type pol polyprotein-like [Arabidopsis thaliana]
Length = 1123
Score = 142 bits (359), Expect = 1e-32
Identities = 81/179 (45%), Positives = 116/179 (64%), Gaps = 5/179 (2%)
Query: 9 QNDPTNSCIFMMRYEIV-YVIINVY--DNNLLRL*KSSQKL*IC*RKNLR*KS*-E*NFC 64
+NDP + CIF+ ++ +VII VY D N+L + +K K + FC
Sbjct: 787 KNDPISPCIFIKKFASKGFVIIAVYVDDLNILGTSGEIAQTVEYLKKEFEMKDLGKTKFC 846
Query: 65 LALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEEL 124
L L +EY+ + ++Q+ Y VLK F MDK+ PL + + +RSL ++ DPF P + +EE+
Sbjct: 847 LGLQLEYIDKGILVHQKAYTETVLKRFNMDKAHPLTSPMQVRSLGLDSDPFGPKKDDEEI 906
Query: 125 LDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALD 182
L PE+PYLSAI AL+YL+SHTR DI FA+NLL+R+SS PT+RHW G K + RYL+G +D
Sbjct: 907 LGPEMPYLSAIGALMYLSSHTRPDICFAVNLLSRFSSCPTKRHWEGIKHLLRYLQGTID 965
>gb|AAF67380.1| Hypothetical protein T15F17.l [Arabidopsis thaliana]
Length = 1141
Score = 140 bits (354), Expect = 4e-32
Identities = 80/179 (44%), Positives = 115/179 (63%), Gaps = 5/179 (2%)
Query: 9 QNDPTNSCIFMMRYEIV-YVIINVY--DNNLLRL*KSSQKL*IC*RKNLR*KS*-E*NFC 64
+NDP + CIF+ ++ +VII VY D N+L + +K K + FC
Sbjct: 787 KNDPISPCIFIKKFASKGFVIIAVYVDDLNILGTSGEIAQTVEYLKKEFEMKDLGKTKFC 846
Query: 65 LALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEEL 124
L L +EY+ + ++QR Y +LK F MDK+ PL + + +RSL ++ DPF P + +EE+
Sbjct: 847 LGLQLEYVDKGILVHQRAYTETILKRFNMDKAHPLTSPMQVRSLGLDSDPFGPKKDDEEI 906
Query: 125 LDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALD 182
L PE+PYLSAI A +YL+SHTR DI FA+NLL+R+SS PT+RHW G K + RYL+G +D
Sbjct: 907 LGPEMPYLSAIGAWMYLSSHTRPDICFAVNLLSRFSSCPTKRHWEGIKHLLRYLQGTID 965
>gb|AAP50929.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50921131|ref|XP_470926.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 639
Score = 132 bits (331), Expect = 2e-29
Identities = 67/112 (59%), Positives = 82/112 (72%), Gaps = 1/112 (0%)
Query: 95 KSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPYLSAIKALLYLASHTRLDISFAIN 154
KS P T +V+RSL+V KDPFRP E E++L P+ PYLSAI AL+Y A+ TR DI+F++
Sbjct: 445 KSYPSKTPMVVRSLDVEKDPFRPKEDGEDVLGPDFPYLSAIGALMYFANSTRPDIAFSVK 504
Query: 155 LLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKLNLIGYADAGYLS 205
LLARYS++PT+RHW G K VFRYL G D+ LFF LIGY DAGYLS
Sbjct: 505 LLARYSAAPTKRHWTGVKNVFRYLNGTRDLGLFFKKNQDSTLIGYTDAGYLS 556
>gb|AAT85780.1| zinc knuckle domain containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 1329
Score = 119 bits (298), Expect = 1e-25
Identities = 57/96 (59%), Positives = 76/96 (78%), Gaps = 1/96 (1%)
Query: 111 NKDPFRP*EKNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG 170
+KDPF+P + +EE+L PEVPYLSAI AL+YLA TR DI+FA+NLLARYS++PT+RHW G
Sbjct: 1071 DKDPFKPKKDDEEMLGPEVPYLSAIGALMYLAYCTRPDIAFALNLLARYSATPTRRHWVG 1130
Query: 171 -KQVFRYLKGALDMSLFFPFMSKLNLIGYADAGYLS 205
K + RYL+G D+ L+FP +++GY DAGY+S
Sbjct: 1131 VKTILRYLRGTQDLGLWFPKNQDPSMVGYVDAGYMS 1166
>gb|AAK84483.1| putative copia-like polyprotein [Lycopersicon esculentum]
Length = 779
Score = 116 bits (291), Expect = 9e-25
Identities = 69/147 (46%), Positives = 91/147 (60%), Gaps = 3/147 (2%)
Query: 9 QNDPTNSCIFMMRYEIVYVIINVY--DNNLLRL*KSSQKL*IC*RKNLR*KS*-E*NFCL 65
+ND CIF+ R +VII VY D N++ K + C ++ K + FCL
Sbjct: 547 KNDSICPCIFIKRLGSEFVIIAVYVDDLNIIGTRKELLEAVECLKREFEMKDLGKTKFCL 606
Query: 66 ALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEELL 125
L IE L + + ++Q TY K+LK FYMD S PL T +V+RSL++N DPFRP E +EELL
Sbjct: 607 GLQIENLSNGILVHQSTYTEKILKRFYMDNSHPLSTPMVVRSLDINTDPFRPQENDEELL 666
Query: 126 DPEVPYLSAIKALLYLASHTRLDISFA 152
E PYLSAI AL+YL ++TR DI FA
Sbjct: 667 GDETPYLSAIGALMYLVNNTRPDICFA 693
>pir||A86471 F11O6.4 protein - Arabidopsis thaliana gi|12320956|gb|AAG50601.1|
hypothetical protein [Arabidopsis thaliana]
gi|9989047|gb|AAG10810.1| Hypothetical protein
[Arabidopsis thaliana]
Length = 484
Score = 114 bits (286), Expect = 3e-24
Identities = 58/103 (56%), Positives = 74/103 (71%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L IE+ +D +F++Q Y K+LK F MDK+ PL T +V RSLNV DPFRP E NE
Sbjct: 288 FCLGLQIEHFQDGIFVHQSNYTKKILKRFNMDKANPLSTPMVNRSLNVENDPFRPCEDNE 347
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQ 165
+ L P+VPY+SAI L+YLA+ TR DI+FA NLLAR ++ Q
Sbjct: 348 DFLGPKVPYMSAIGGLMYLANCTRPDIAFATNLLARSMTNHIQ 390
>ref|XP_474878.1| OSJNBa0021F22.5 [Oryza sativa (japonica cultivar-group)]
gi|38346472|emb|CAE03711.2| OSJNBa0021F22.5 [Oryza
sativa (japonica cultivar-group)]
Length = 911
Score = 101 bits (252), Expect = 3e-20
Identities = 51/94 (54%), Positives = 69/94 (73%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL+L +E+L + ++Q Y K+L+ F MDKS P T +V+RSL+V KD FRP E E
Sbjct: 779 FCLSLQLEHLPFGILVHQNAYTQKILEKFNMDKSYPSKTPMVVRSLDVEKDLFRPKEDGE 838
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLL 156
++L P+ PYLSAI AL+YLA+ TR DISF++NLL
Sbjct: 839 DVLGPDFPYLSAIGALMYLANSTRPDISFSVNLL 872
>gb|AAX96392.1| retrotransposon protein, putative, Ty1-copia sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 460
Score = 89.0 bits (219), Expect = 2e-16
Identities = 62/152 (40%), Positives = 89/152 (57%), Gaps = 8/152 (5%)
Query: 6 GRSQNDPTNSCIFMMRYEIVYVIINVYDNNLLRL*KSSQKL*IC*RKNLR*KS*-----E 60
G S ND CIF+ R + II+VY ++L + ++Q + R NL+ + +
Sbjct: 129 GNSNNDDC-PCIFIKRSSTGFCIISVYVDDL-NIIGNTQDINEA-RHNLKAEFEMKDLGK 185
Query: 61 *NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EK 120
FCL L IE+L + Q Y+ K+L+ F M+K+ P T +V+RSL KD FRP E+
Sbjct: 186 TRFCLGLQIEHLHSGFLVRQSAYVQKLLEKFNMNKAYPNKTPIVVRSLCKEKDIFRPREE 245
Query: 121 NEELLDPEVPYLSAIKALLYLASHTRLDISFA 152
EE+L PE PYLS I AL+Y A+ TR +I+FA
Sbjct: 246 GEEILGPEYPYLSLIGALMYFANSTRPNIAFA 277
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.357 0.160 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 496,920,864
Number of Sequences: 2540612
Number of extensions: 18024656
Number of successful extensions: 74831
Number of sequences better than 10.0: 595
Number of HSP's better than 10.0 without gapping: 270
Number of HSP's successfully gapped in prelim test: 325
Number of HSP's that attempted gapping in prelim test: 73952
Number of HSP's gapped (non-prelim): 705
length of query: 353
length of database: 863,360,394
effective HSP length: 129
effective length of query: 224
effective length of database: 535,621,446
effective search space: 119979203904
effective search space used: 119979203904
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0091b.10


