

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.5
(140 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
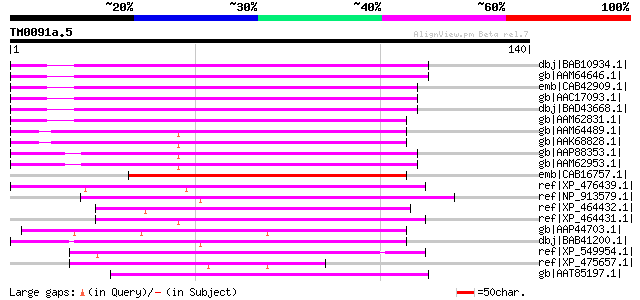
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB10934.1| unnamed protein product [Arabidopsis thaliana] g... 80 1e-14
gb|AAM64646.1| unknown [Arabidopsis thaliana] 79 2e-14
emb|CAB42909.1| putative protein [Arabidopsis thaliana] gi|28827... 73 1e-12
gb|AAC17093.1| unknown protein [Arabidopsis thaliana] gi|7486264... 72 3e-12
dbj|BAD43668.1| unknown protein [Arabidopsis thaliana] 72 3e-12
gb|AAM62831.1| unknown [Arabidopsis thaliana] gi|26451664|dbj|BA... 72 4e-12
gb|AAM64489.1| unknown [Arabidopsis thaliana] gi|23197634|gb|AAN... 64 6e-10
gb|AAK68828.1| Unknown protein [Arabidopsis thaliana] 62 3e-09
gb|AAP88353.1| At1g21010 [Arabidopsis thaliana] gi|18394938|ref|... 61 5e-09
gb|AAM62953.1| unknown [Arabidopsis thaliana] 61 5e-09
emb|CAB16757.1| putative protein [Arabidopsis thaliana] gi|72707... 60 1e-08
ref|XP_476439.1| hypothetical protein [Oryza sativa (japonica cu... 57 1e-07
ref|NP_913579.1| OSJNBa0086P08.11 [Oryza sativa (japonica cultiv... 56 2e-07
ref|XP_464432.1| unknown protein [Oryza sativa (japonica cultiva... 55 4e-07
ref|XP_464431.1| hypothetical protein [Oryza sativa (japonica cu... 54 8e-07
gb|AAP44703.1| hypothetical protein [Oryza sativa (japonica cult... 53 2e-06
dbj|BAB41200.1| TMV response-related gene product [Nicotiana tab... 53 2e-06
ref|XP_549954.1| unknown protein [Oryza sativa (japonica cultiva... 52 3e-06
ref|XP_475657.1| hypothetical protein [Oryza sativa (japonica cu... 50 9e-06
gb|AAT85197.1| hypothetical protein [Oryza sativa (japonica cult... 48 4e-05
>dbj|BAB10934.1| unnamed protein product [Arabidopsis thaliana]
gi|28972979|gb|AAO63814.1| unknown protein [Arabidopsis
thaliana] gi|28393574|gb|AAO42207.1| unknown protein
[Arabidopsis thaliana] gi|15240014|ref|NP_201459.1|
expressed protein [Arabidopsis thaliana]
Length = 156
Score = 80.1 bits (196), Expect = 1e-14
Identities = 40/113 (35%), Positives = 62/113 (54%), Gaps = 7/113 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG CAS + + ++ DG LQ+ PVK W +L +NP F+C+S+ M
Sbjct: 1 MGACASRESL-------RSDSAKLILLDGTLQEFSSPVKVWQILQKNPTSFVCNSDEMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSV 113
++ V NEEL+ +YF++P + PL E++ ALA+KA++AL S V
Sbjct: 54 DDAVSAVAGNEELRSGQLYFVLPLTWLNHPLRAEEMAALAVKASSALTKSGGV 106
>gb|AAM64646.1| unknown [Arabidopsis thaliana]
Length = 156
Score = 79.0 bits (193), Expect = 2e-14
Identities = 40/113 (35%), Positives = 62/113 (54%), Gaps = 7/113 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG CAS + + ++ DG LQ+ PVK W +L +NP F+C+S+ M
Sbjct: 1 MGACASRESL-------RSDSAKLILLDGTLQEFSSPVKVWQILQKNPTIFVCNSDEMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSV 113
++ V NEEL+ +YF++P + PL E++ ALA+KA++AL S V
Sbjct: 54 DDAVSAVAGNEELRPGQLYFVLPLTWLNHPLRAEEMAALAVKASSALTKSGGV 106
>emb|CAB42909.1| putative protein [Arabidopsis thaliana] gi|28827350|gb|AAO50519.1|
unknown protein [Arabidopsis thaliana]
gi|28393455|gb|AAO42149.1| unknown protein [Arabidopsis
thaliana] gi|15230307|ref|NP_190649.1| expressed protein
[Arabidopsis thaliana] gi|7485709|pir||T08401
hypothetical protein F18B3.80 - Arabidopsis thaliana
Length = 152
Score = 73.2 bits (178), Expect = 1e-12
Identities = 38/110 (34%), Positives = 61/110 (54%), Gaps = 7/110 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG CAS + T ++ DG LQ+ PVK W +L +NP F+C+S+ M
Sbjct: 1 MGACASRESR-------RTETAKLILPDGTLQEFSTPVKVWQILQKNPTSFVCNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS 110
+ V +E+L+ +YF++P + PL +++ ALA+KA++ALA S
Sbjct: 54 DDAVLAVPGSEDLRPGELYFVLPLTWLNHPLRADEMAALAVKASSALAKS 103
>gb|AAC17093.1| unknown protein [Arabidopsis thaliana] gi|7486264|pir||T02422
hypothetical protein At2g23690 [imported] - Arabidopsis
thaliana gi|15224053|ref|NP_179949.1| expressed protein
[Arabidopsis thaliana]
Length = 278
Score = 72.0 bits (175), Expect = 3e-12
Identities = 37/110 (33%), Positives = 62/110 (55%), Gaps = 7/110 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MGIC+S + +T ++ DG++ + PVK +VL +NP F+C+S+ M
Sbjct: 1 MGICSSYEST-------QVATAKLILHDGRMMEFTSPVKVGYVLQKNPMCFICNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS 110
+ ++ + +EE QL +YF +P S L E++ ALA+KA++AL S
Sbjct: 54 DNVVSAISADEEFQLGQLYFALPLSSLHHSLKAEEMAALAVKASSALMRS 103
>dbj|BAD43668.1| unknown protein [Arabidopsis thaliana]
Length = 163
Score = 72.0 bits (175), Expect = 3e-12
Identities = 37/110 (33%), Positives = 62/110 (55%), Gaps = 7/110 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MGIC+S + +T ++ DG++ + PVK +VL +NP F+C+S+ M
Sbjct: 1 MGICSSYEST-------QVATAKLILHDGRMMEFTSPVKVGYVLQKNPMCFICNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS 110
+ ++ + +EE QL +YF +P S L E++ ALA+KA++AL S
Sbjct: 54 DNVVSAISADEEFQLGQLYFALPLSSLHHSLKAEEMAALAVKASSALMRS 103
>gb|AAM62831.1| unknown [Arabidopsis thaliana] gi|26451664|dbj|BAC42928.1| unknown
protein [Arabidopsis thaliana]
gi|28973267|gb|AAO63958.1| unknown protein [Arabidopsis
thaliana] gi|18420002|ref|NP_568020.1| expressed protein
[Arabidopsis thaliana]
Length = 168
Score = 71.6 bits (174), Expect = 4e-12
Identities = 38/107 (35%), Positives = 61/107 (56%), Gaps = 7/107 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MGIC+S++ +T ++ DG++ + PVK +VL + P F+C+S+ M
Sbjct: 1 MGICSSSEST-------QVATAKLILQDGRMMEFANPVKVGYVLLKYPMCFICNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
+ + +EELQL IYF +P R PL E++ ALA+KA++AL
Sbjct: 54 DDAVAAISADEELQLGQIYFALPLCWLRQPLKAEEMAALAVKASSAL 100
>gb|AAM64489.1| unknown [Arabidopsis thaliana] gi|23197634|gb|AAN15344.1| Unknown
protein [Arabidopsis thaliana]
gi|18411155|ref|NP_565136.1| expressed protein
[Arabidopsis thaliana] gi|12323985|gb|AAG51956.1|
unknown protein; 83277-83927 [Arabidopsis thaliana]
gi|25406456|pir||B96794 unknown protein F14G6.20
[imported] - Arabidopsis thaliana
Length = 216
Score = 64.3 bits (155), Expect = 6e-10
Identities = 40/117 (34%), Positives = 62/117 (52%), Gaps = 13/117 (11%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVL----------SQNPNH 50
MG+C S + +T IV +G L++ PV A VL S + ++
Sbjct: 1 MGLCVSVN---RNEYVSSSTTAKIVTINGDLREYDVPVLASQVLESESTSSSSSSSSSSY 57
Query: 51 FLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
FLC+S+S+Y + + +E LQ N IYF++P SK + LS D+ ALA+KA+ A+
Sbjct: 58 FLCNSDSLYYDDFIPAIESDEILQANQIYFVLPISKRQYRLSASDMAALAVKASVAI 114
>gb|AAK68828.1| Unknown protein [Arabidopsis thaliana]
Length = 216
Score = 62.0 bits (149), Expect = 3e-09
Identities = 39/117 (33%), Positives = 61/117 (51%), Gaps = 13/117 (11%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVL----------SQNPNH 50
MG+C S + +T IV +G L++ PV A VL S + ++
Sbjct: 1 MGLCVSVN---RNEYVSSSTTAKIVTINGDLREYDVPVLASQVLESESTSSSSSSSSSSY 57
Query: 51 FLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
FLC+S+S+Y + + +E LQ N IYF++P SK + LS D+ ALA+ A+ A+
Sbjct: 58 FLCNSDSLYYDDFIPAIESDEILQANQIYFVLPISKRQYRLSASDMAALAVXASVAI 114
>gb|AAP88353.1| At1g21010 [Arabidopsis thaliana] gi|18394938|ref|NP_564129.1|
expressed protein [Arabidopsis thaliana]
Length = 210
Score = 61.2 bits (147), Expect = 5e-09
Identities = 40/123 (32%), Positives = 63/123 (50%), Gaps = 17/123 (13%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVL-------------SQN 47
MGIC S + + TV IV +G L++ PV A VL S+
Sbjct: 1 MGICVSFRREDSNSS----PTVKIVTVNGDLREYNVPVIASQVLEAESAAAYSSSSSSRP 56
Query: 48 PNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
++F+C S+S+Y + + E LQ + IYF++P SK + L+ D+ ALA+KA+ A+
Sbjct: 57 SSYFICDSDSLYYDDFIPAIKSEEPLQADQIYFVLPISKRQSRLTASDMAALAVKASVAI 116
Query: 108 AHS 110
+S
Sbjct: 117 QNS 119
>gb|AAM62953.1| unknown [Arabidopsis thaliana]
Length = 210
Score = 61.2 bits (147), Expect = 5e-09
Identities = 40/123 (32%), Positives = 63/123 (50%), Gaps = 17/123 (13%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVL-------------SQN 47
MGIC S + + TV IV +G L++ PV A VL S+
Sbjct: 1 MGICVSFRREDSNSS----PTVKIVTVNGDLREYNVPVIASQVLEAESAAAYSSSSSSRP 56
Query: 48 PNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
++F+C S+S+Y + + E LQ + IYF++P SK + L+ D+ ALA+KA+ A+
Sbjct: 57 SSYFICDSDSLYYDDFIPAIKSEEPLQADQIYFVLPISKRQSRLTASDMAALAVKASVAI 116
Query: 108 AHS 110
+S
Sbjct: 117 QNS 119
>emb|CAB16757.1| putative protein [Arabidopsis thaliana] gi|7270707|emb|CAB80390.1|
putative protein [Arabidopsis thaliana]
gi|25371565|pir||A85440 hypothetical protein AT4g37240
[imported] - Arabidopsis thaliana
Length = 144
Score = 59.7 bits (143), Expect = 1e-08
Identities = 30/75 (40%), Positives = 45/75 (60%)
Query: 33 QLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLS 92
+ PVK +VL + P F+C+S+ M + + +EELQL IYF +P R PL
Sbjct: 2 EFANPVKVGYVLLKYPMCFICNSDDMDFDDAVAAISADEELQLGQIYFALPLCWLRQPLK 61
Query: 93 LEDLCALAIKANAAL 107
E++ ALA+KA++AL
Sbjct: 62 AEEMAALAVKASSAL 76
>ref|XP_476439.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|34394505|dbj|BAC83793.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 205
Score = 57.0 bits (136), Expect = 1e-07
Identities = 40/124 (32%), Positives = 59/124 (47%), Gaps = 12/124 (9%)
Query: 1 MGICASTQYAGKGGNHGMQ-STVNIVHFDGKLQQLKEPVKAWHVLSQ-----------NP 48
MG+C S+ A GM ST ++ G+L++ P A VL
Sbjct: 1 MGLCLSSGVAAAMAEAGMAASTAMVLLPTGELREYPRPATAARVLEDVAAAEGEEEDVGR 60
Query: 49 NHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALA 108
FLC ++ M P+ V EL+ IYF++P R + E++ ALA+KA+AALA
Sbjct: 61 RFFLCDADKMGFEGPVAAVAAAAELRPGQIYFVLPGEVRRRGMRREEVAALAVKASAALA 120
Query: 109 HSSS 112
+SS
Sbjct: 121 AASS 124
>ref|NP_913579.1| OSJNBa0086P08.11 [Oryza sativa (japonica cultivar-group)]
Length = 145
Score = 55.8 bits (133), Expect = 2e-07
Identities = 32/102 (31%), Positives = 53/102 (51%), Gaps = 1/102 (0%)
Query: 20 STVNIVHFDGKLQQLKEPVKAWHVLSQNPNH-FLCSSESMYVGSPMTPVVPNEELQLNHI 78
+T +V D + Q P S + + F+C S+ + +P + ++ LQL +
Sbjct: 38 TTAKVVFQDSSMAQFVAPTSMGRASSASASTCFVCCSDELRFDAPARAMAVHDALQLGQL 97
Query: 79 YFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSV 120
YF++P S PLS +D+ ALA+KA AAL S++ SS+
Sbjct: 98 YFVLPVSALHRPLSDQDMAALAVKAIAALGASATAASMDSSI 139
>ref|XP_464432.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|46389831|dbj|BAD15394.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 192
Score = 55.1 bits (131), Expect = 4e-07
Identities = 29/88 (32%), Positives = 50/88 (55%), Gaps = 3/88 (3%)
Query: 24 IVHFDGKLQQLK---EPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYF 80
++ DG L++L P A + + + F+C+S+++Y + P E L+ IYF
Sbjct: 22 VIAADGSLKELHAAASPAVADVLRGEGESFFVCNSDTLYFNEQPPAMAPGEALRPGQIYF 81
Query: 81 LVPRSKSRLPLSLEDLCALAIKANAALA 108
++P + PLS D+ ALA++A+AALA
Sbjct: 82 VLPAAMLGQPLSTADMAALAVRASAALA 109
>ref|XP_464431.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|46389830|dbj|BAD15393.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 200
Score = 53.9 bits (128), Expect = 8e-07
Identities = 29/92 (31%), Positives = 52/92 (56%), Gaps = 3/92 (3%)
Query: 24 IVHFDGKLQQLKEPVKAWHVL---SQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYF 80
++ DG L++L VL + + F+C+S+++Y + P E L+ IYF
Sbjct: 26 VITADGSLKELAVSSAVADVLRGEGEGRSFFVCNSDALYFNEQPPALAPGEALRPGQIYF 85
Query: 81 LVPRSKSRLPLSLEDLCALAIKANAALAHSSS 112
++P + PLS D+ ALA++A+AALA +++
Sbjct: 86 VLPAAMLGQPLSTADMAALAVRASAALAAAAA 117
>gb|AAP44703.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50918533|ref|XP_469663.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|40539051|gb|AAR87308.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 170
Score = 52.8 bits (125), Expect = 2e-06
Identities = 36/114 (31%), Positives = 59/114 (51%), Gaps = 10/114 (8%)
Query: 4 CASTQYAGKGGNH--GMQSTVNIVHFDGKLQQL------KEPVKAWHVLSQNPNHFLCSS 55
C +T A G G + +V DG L++ + VKA + FLCS+
Sbjct: 7 CEATAVAAVNGRSASGEATAARVVLADGALRRFPGGTRASQAVKAAGGGGGGSSWFLCSA 66
Query: 56 ESMYVGSPMTPVV--PNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
+ + +G+ + V +EELQ +YF++P + R PL E++ ALA++A+AAL
Sbjct: 67 DGLELGAAVAAVGGGDDEELQPGQLYFVLPAAMRRRPLQAEEMAALAVRASAAL 120
>dbj|BAB41200.1| TMV response-related gene product [Nicotiana tabacum]
Length = 205
Score = 52.8 bits (125), Expect = 2e-06
Identities = 33/110 (30%), Positives = 57/110 (51%), Gaps = 4/110 (3%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNH---FLCSSES 57
MG C S+ + S+ ++ +G+L+Q P+ VL + F+C+S+
Sbjct: 1 MGACLSSSSVIDAKDQ-KPSSAYVISTNGELRQYTVPINVSQVLQSEMSSEASFICNSDR 59
Query: 58 MYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
+Y + + +LQ IYF++P SK + LS ++ ALA+KA+AAL
Sbjct: 60 LYFDDFIPRLDSEYQLQPGQIYFVLPTSKLQYRLSASEMAALAVKASAAL 109
>ref|XP_549954.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|53791247|dbj|BAD52452.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 201
Score = 52.0 bits (123), Expect = 3e-06
Identities = 34/97 (35%), Positives = 51/97 (52%), Gaps = 2/97 (2%)
Query: 17 GMQSTV-NIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQL 75
G ST I+H DG + +L PV+A ++ P F+C S + VG + V +E L+
Sbjct: 16 GSSSTACKIIHVDGTVTRLARPVRASELMVDYPGQFVCDSGRLAVGCRVPGVAADELLEP 75
Query: 76 NHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSS 112
YFL+P L+ E++ AL+ +AA A SSS
Sbjct: 76 RRAYFLLPMDMLYSVLTDEEMAALS-SFHAATAASSS 111
>ref|XP_475657.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50080294|gb|AAT69629.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 170
Score = 50.4 bits (119), Expect = 9e-06
Identities = 27/74 (36%), Positives = 41/74 (54%), Gaps = 5/74 (6%)
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFL---CSSESMYVGSPMTPVV--PNE 71
G + IVH +G++++ PV A +L+ NPNH L C S+ V T ++ P+
Sbjct: 16 GALDLIRIVHLNGRVEEYGRPVAAGEILAANPNHVLSKPCCSQGGAVAVRRTILIVSPDS 75
Query: 72 ELQLNHIYFLVPRS 85
EL+ IYFL+P S
Sbjct: 76 ELERGEIYFLIPAS 89
>gb|AAT85197.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 167
Score = 48.1 bits (113), Expect = 4e-05
Identities = 26/86 (30%), Positives = 42/86 (48%)
Query: 28 DGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKS 87
DG L+Q+ P A ++ P HFL + + VG+ + + +EEL+L +Y P +
Sbjct: 17 DGGLRQVALPATAAELMMDAPGHFLADARAARVGARLAALSADEELELGAVYATFPMKRL 76
Query: 88 RLPLSLEDLCALAIKANAALAHSSSV 113
PL+ D+ LA A S+ V
Sbjct: 77 GTPLAPADMARLAAVATREARRSAKV 102
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.130 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 246,321,045
Number of Sequences: 2540612
Number of extensions: 9381272
Number of successful extensions: 20367
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 48
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 20302
Number of HSP's gapped (non-prelim): 85
length of query: 140
length of database: 863,360,394
effective HSP length: 116
effective length of query: 24
effective length of database: 568,649,402
effective search space: 13647585648
effective search space used: 13647585648
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0091a.5


