

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
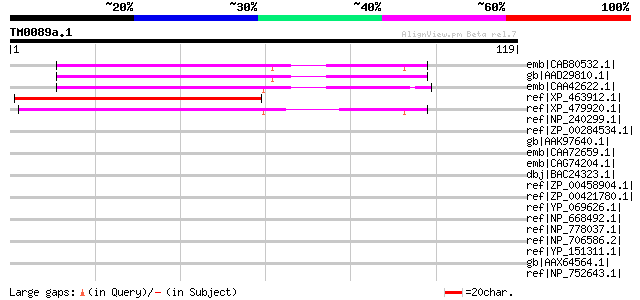
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB80532.1| glycine-rich protein 2 (GRP2) [Arabidopsis thali... 56 2e-07
gb|AAD29810.1| glycine-rich protein (AtGRP2) [Arabidopsis thalia... 54 7e-07
emb|CAA42622.1| nsGRP-2 [Nicotiana sylvestris] gi|121631|sp|P274... 53 2e-06
ref|XP_463912.1| putative Glycine-rich protein 2 [Oryza sativa (... 46 2e-04
ref|XP_479920.1| putative cold shock protein-1 [Oryza sativa (ja... 45 5e-04
ref|NP_240299.1| cold shock-like protein cspE [Buchnera aphidico... 44 0.001
ref|ZP_00284534.1| COG1278: Cold shock proteins [Burkholderia fu... 44 0.001
gb|AAK97640.1| cold-shock protein CspV [Vibrio cholerae] gi|9658... 44 0.001
emb|CAA72659.1| cold shock protein, CSPA [Vibrio cholerae] 44 0.001
emb|CAG74204.1| cold shock-like protein [Erwinia carotovora subs... 44 0.001
dbj|BAC24323.1| cspE [Wigglesworthia glossinidia endosymbiont of... 44 0.001
ref|ZP_00458904.1| Cold-shock protein, DNA-binding [Burkholderia... 43 0.002
ref|ZP_00421780.1| Cold-shock protein, DNA-binding [Burkholderia... 43 0.002
ref|YP_069626.1| putative cold shock protein [Yersinia pseudotub... 43 0.002
ref|NP_668492.1| cold shock protein [Yersinia pestis KIM] gi|454... 43 0.002
ref|NP_778037.1| cold shock protein CspA [Buchnera aphidicola st... 43 0.002
ref|NP_706586.2| cold shock protein [Shigella flexneri 2a str. 3... 42 0.003
ref|YP_151311.1| cold shock-like protein cspE [Salmonella enteri... 42 0.003
gb|AAX64564.1| RNA chaperone, negative regulator of cspA transcr... 42 0.003
ref|NP_752643.1| Cold shock-like protein cspE [Escherichia coli ... 42 0.003
>emb|CAB80532.1| glycine-rich protein 2 (GRP2) [Arabidopsis thaliana]
gi|4467155|emb|CAB37524.1| glycine-rich protein 2 (GRP2)
[Arabidopsis thaliana] gi|14532850|gb|AAK64107.1|
putative glycine-rich protein 2 [Arabidopsis thaliana]
gi|13430580|gb|AAK25912.1| putative glycine-rich protein
GRP2 [Arabidopsis thaliana] gi|22137152|gb|AAM91421.1|
AT4g38680/F20M13_240 [Arabidopsis thaliana]
gi|259445|gb|AAB24074.1| glycine-rich protein; atGRP
[Arabidopsis thaliana] gi|15234010|ref|NP_195580.1|
cold-shock DNA-binding family protein [Arabidopsis
thaliana] gi|14326487|gb|AAK60289.1|
AT4g38680/F20M13_240 [Arabidopsis thaliana]
gi|81624|pir||JQ1061 glycine-rich protein 2 -
Arabidopsis thaliana
Length = 203
Score = 56.2 bits (134), Expect = 2e-07
Identities = 40/98 (40%), Positives = 51/98 (51%), Gaps = 19/98 (19%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DDGG+DLF+ S I+S+GF LA E V+ + I I +
Sbjct: 15 VKWFDTQKGFGFITPDDGGDDLFVHQSSIRSEGFRSLAAEEAVEFEVEIDNNNRPKAIDV 74
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSS----GYGRGGG 98
+ PD AP+ G GG+ G GRGGG
Sbjct: 75 SG--------PDGAPVQGNSGGGSSGGRGGFGGGRGGG 104
>gb|AAD29810.1| glycine-rich protein (AtGRP2) [Arabidopsis thaliana]
gi|60543359|gb|AAX22277.1| At2g21060 [Arabidopsis
thaliana] gi|16323178|gb|AAL15323.1| At2g21060/F26H11.18
[Arabidopsis thaliana] gi|17366505|sp|Q38896|GRP2B_ARATH
Glycine-rich protein 2b (AtGRP2b)
gi|15226451|ref|NP_179702.1| cold-shock DNA-binding
family protein / glycine-rich protein (GRP2)
[Arabidopsis thaliana] gi|1063684|gb|AAA91165.1| AtGRP2b
Length = 201
Score = 54.3 bits (129), Expect = 7e-07
Identities = 37/94 (39%), Positives = 48/94 (50%), Gaps = 15/94 (15%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DGG+DLF+ S I+S+GF LA E V+ + + I +
Sbjct: 19 VKWFDTQKGFGFITPSDGGDDLFVHQSSIRSEGFRSLAAEESVEFDVEVDNSGRPKAIEV 78
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGG 98
+ PD AP+ G GG S G G GG
Sbjct: 79 SG--------PDGAPVQGNSGGGGSSGGRGGFGG 104
>emb|CAA42622.1| nsGRP-2 [Nicotiana sylvestris] gi|121631|sp|P27484|GRP2_NICSY
Glycine-rich protein 2
Length = 214
Score = 52.8 bits (125), Expect = 2e-06
Identities = 39/95 (41%), Positives = 48/95 (50%), Gaps = 16/95 (16%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL-------TILIPM 64
+K F DQKGF IT DDGG DLF+ S I+S+GF LAE E V+ + T + +
Sbjct: 13 VKWFSDQKGFGFITPDDGGEDLFVHQSGIRSEGFRSLAEGETVEFEVESGGDGRTKAVDV 72
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGGY 99
PD A + G GG G G GGGY
Sbjct: 73 TG--------PDGAAVQGGRGGGGGGGGRG-GGGY 98
>ref|XP_463912.1| putative Glycine-rich protein 2 [Oryza sativa (japonica
cultivar-group)] gi|51963818|ref|XP_506692.1| PREDICTED
OSJNBb0088N06.21 gene product [Oryza sativa (japonica
cultivar-group)] gi|41052743|dbj|BAD07599.1| putative
Glycine-rich protein 2 [Oryza sativa (japonica
cultivar-group)] gi|41052630|dbj|BAD08139.1| putative
Glycine-rich protein 2 [Oryza sativa (japonica
cultivar-group)]
Length = 241
Score = 45.8 bits (107), Expect = 2e-04
Identities = 25/58 (43%), Positives = 35/58 (60%)
Query: 2 MMSVSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
M + + R +K F+D KGF I+ DDG DLF+ S IK+ GF LAE E V+ ++
Sbjct: 1 MAAAARHRGTVKWFNDTKGFGFISPDDGSEDLFVHQSSIKADGFRSLAEGEQVEFAIS 58
>ref|XP_479920.1| putative cold shock protein-1 [Oryza sativa (japonica
cultivar-group)] gi|51964662|ref|XP_507115.1| PREDICTED
P0582D05.112 gene product [Oryza sativa (japonica
cultivar-group)] gi|29467522|dbj|BAC66711.1| putative
cold shock protein-1 [Oryza sativa (japonica
cultivar-group)]
Length = 197
Score = 44.7 bits (104), Expect = 5e-04
Identities = 36/108 (33%), Positives = 49/108 (45%), Gaps = 24/108 (22%)
Query: 3 MSVSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL---- 58
M+ V+ +K FD KGF IT DDGG DLF+ S +KS G+ L + + V+ +
Sbjct: 1 MASERVKGTVKWFDATKGFGFITPDDGGEDLFVHQSSLKSDGYRSLNDGDVVEFSVGSGN 60
Query: 59 ---TILIPMAALRR*RD*RPD*API*GTCCGGNDSS-----GYGRGGG 98
T + + AP G GG+ S GYG GGG
Sbjct: 61 DGRTKAVDVT------------APGGGALTGGSRPSGGGDRGYGGGGG 96
>ref|NP_240299.1| cold shock-like protein cspE [Buchnera aphidicola str. APS
(Acyrthosiphon pisum)] gi|21672738|ref|NP_660805.1|
cold shock like protein CspE [Buchnera aphidicola str.
Sg (Schizaphis graminum)] gi|21623383|gb|AAM68016.1|
cold shock like protein CspE [Buchnera aphidicola str.
Sg (Schizaphis graminum)] gi|10039151|dbj|BAB13185.1|
cold shock-like protein cspE [Buchnera aphidicola str.
APS (Acyrthosiphon pisum)]
gi|54036912|sp|P63237|CSPE_BUCAI Cold shock-like
protein cspE (CSP-E) gi|25296105|pir||A84987 cold
shock-like protein cspE [imported] - Buchnera sp.
(strain APS) gi|54036913|sp|P63238|CSPE_BUCAP Cold
shock-like protein cspE (CSP-E)
Length = 69
Score = 43.9 bits (102), Expect = 0.001
Identities = 22/55 (40%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQSVEFEIT 55
>ref|ZP_00284534.1| COG1278: Cold shock proteins [Burkholderia fungorum LB400]
Length = 67
Score = 43.9 bits (102), Expect = 0.001
Identities = 23/43 (53%), Positives = 29/43 (66%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F+D KGF IT DDGG DLF S+I+S+GF L E + V
Sbjct: 6 VKWFNDAKGFGFITPDDGGEDLFAHFSEIRSEGFKSLQENQKV 48
>gb|AAK97640.1| cold-shock protein CspV [Vibrio cholerae]
gi|9658370|gb|AAF96830.1| cold shock domain family
protein [Vibrio cholerae O1 biovar eltor str. N16961]
gi|15601687|ref|NP_233318.1| cold shock domain family
protein [Vibrio cholerae O1 biovar eltor str. N16961]
gi|20138009|sp|Q9KL16|CSPV_VIBCH Cold shock protein
cspV
Length = 70
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/47 (42%), Positives = 33/47 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL 58
+K F++ KGF +T D+GGND+F+ + I+S+GF LAE + V ++
Sbjct: 9 VKWFNETKGFGFLTQDNGGNDVFVHFNSIQSEGFKTLAEGQRVSFIV 55
>emb|CAA72659.1| cold shock protein, CSPA [Vibrio cholerae]
Length = 70
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/47 (42%), Positives = 33/47 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL 58
+K F++ KGF +T D+GGND+F+ + I+S+GF LAE + V ++
Sbjct: 9 VKWFNETKGFGFLTQDNGGNDVFVHLNSIQSEGFKTLAEGQRVSFIV 55
>emb|CAG74204.1| cold shock-like protein [Erwinia carotovora subsp. atroseptica
SCRI1043] gi|50120233|ref|YP_049400.1| cold shock-like
protein [Erwinia carotovora subsp. atroseptica
SCRI1043]
Length = 69
Score = 43.5 bits (101), Expect = 0.001
Identities = 22/55 (40%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 MSKIKGSVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQRVEFEIT 55
>dbj|BAC24323.1| cspE [Wigglesworthia glossinidia endosymbiont of Glossina
brevipalpis] gi|32490926|ref|NP_871180.1| hypothetical
protein WGLp177 [Wigglesworthia glossinidia
endosymbiont of Glossina brevipalpis]
Length = 69
Score = 43.5 bits (101), Expect = 0.001
Identities = 22/55 (40%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQRVEFEIT 55
>ref|ZP_00458904.1| Cold-shock protein, DNA-binding [Burkholderia cenocepacia HI2424]
gi|67104843|gb|EAM21961.1| Cold-shock protein,
DNA-binding [Burkholderia cenocepacia HI2424]
Length = 170
Score = 43.1 bits (100), Expect = 0.002
Identities = 24/50 (48%), Positives = 31/50 (62%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+SM +K F+D KGF IT D GG+DLF S+I+ GF LAE + V
Sbjct: 102 ISMDTGTVKWFNDNKGFGFITPDSGGDDLFAHFSEIRGDGFKTLAEGQKV 151
>ref|ZP_00421780.1| Cold-shock protein, DNA-binding [Burkholderia vietnamiensis G4]
gi|67535017|gb|EAM31755.1| Cold-shock protein,
DNA-binding [Burkholderia vietnamiensis G4]
gi|46312719|ref|ZP_00213313.1| COG1278: Cold shock
proteins [Burkholderia cepacia R18194]
Length = 67
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/43 (51%), Positives = 30/43 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F+D KGF IT D+GG+DLF S+I++ GF LAE + V
Sbjct: 6 VKWFNDSKGFGFITPDNGGDDLFAHFSEIRADGFKTLAENQKV 48
>ref|YP_069626.1| putative cold shock protein [Yersinia pseudotuberculosis IP
32953] gi|51588717|emb|CAH20328.1| putative cold shock
protein [Yersinia pseudotuberculosis IP 32953]
gi|15980582|emb|CAC92838.1| putative cold shock protein
[Yersinia pestis CO92] gi|16122808|ref|NP_406121.1|
putative cold shock protein [Yersinia pestis CO92]
gi|25296124|pir||AG0316 probable cold shock protein
cspE [imported] - Yersinia pestis (strain CO92)
Length = 69
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/55 (40%), Positives = 34/55 (61%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I S GF LAE + V+ +T
Sbjct: 1 MSKIKGSVKWFNESKGFGFITPEDGSKDVFVHFSAIASNGFKTLAEGQRVEFEIT 55
>ref|NP_668492.1| cold shock protein [Yersinia pestis KIM]
gi|45440950|ref|NP_992489.1| putative cold shock
protein [Yersinia pestis biovar Medievalis str. 91001]
gi|45435809|gb|AAS61366.1| putative cold shock protein
[Yersinia pestis biovar Medievalis str. 91001]
gi|21957922|gb|AAM84743.1| cold shock protein [Yersinia
pestis KIM]
Length = 84
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/55 (40%), Positives = 34/55 (61%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I S GF LAE + V+ +T
Sbjct: 16 MSKIKGSVKWFNESKGFGFITPEDGSKDVFVHFSAIASNGFKTLAEGQRVEFEIT 70
>ref|NP_778037.1| cold shock protein CspA [Buchnera aphidicola str. Bp (Baizongia
pistaciae)] gi|27904309|gb|AAO27142.1| cold shock
protein CspA [Buchnera aphidicola str. Bp (Baizongia
pistaciae)] gi|38372204|sp|Q89A90|CSPE_BUCBP Cold
shock-like protein cspE (CSP-E)
Length = 69
Score = 42.7 bits (99), Expect = 0.002
Identities = 21/55 (38%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I+S GF L+E + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLSEGQSVEFEIT 55
>ref|NP_706586.2| cold shock protein [Shigella flexneri 2a str. 301]
gi|56383262|gb|AAN42293.2| cold shock protein [Shigella
flexneri 2a str. 301] gi|30062187|ref|NP_836358.1| cold
shock protein [Shigella flexneri 2a str. 2457T]
gi|30040432|gb|AAP16164.1| cold shock protein [Shigella
flexneri 2a str. 2457T] gi|26986712|gb|AAN86720.1| CspE
[Escherichia coli] gi|1786841|gb|AAC73724.1| cold shock
protein; RNA chaperone, transcription antiterminator,
affects expression of rpoS and uspA [Escherichia coli
K12] gi|1651256|dbj|BAA35266.1| CspE protein
[Escherichia coli K12] gi|16128606|ref|NP_415156.1| RNA
chaperone, transcription antiterminator, affects
expression of rpoS and uspA [Escherichia coli K12]
gi|12513525|gb|AAG54958.1| cold shock protein
[Escherichia coli O157:H7 EDL933]
gi|3851642|gb|AAC72388.1| unknown [Vibrio cholerae]
gi|544110|sp|P36997|CSPE_ECOLI Cold shock-like protein
cspE (CSP-E) gi|13360120|dbj|BAB34085.1| cold shock
protein [Escherichia coli O157:H7]
gi|15829916|ref|NP_308689.1| cold shock protein
[Escherichia coli O157:H7] gi|833769|gb|AAA67556.1|
gicA gene product gi|15800338|ref|NP_286350.1| cold
shock protein [Escherichia coli O157:H7 EDL933]
gi|471099|dbj|BAA05856.1| CspE (MsmC) [Escherichia
coli]
Length = 69
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/55 (38%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 55
>ref|YP_151311.1| cold shock-like protein cspE [Salmonella enterica subsp. enterica
serovar Paratyphi A str. ATCC 9150]
gi|29142638|ref|NP_805980.1| cold shock-like protein
cspE [Salmonella enterica subsp. enterica serovar Typhi
Ty2] gi|56128493|gb|AAV77999.1| cold shock-like protein
cspE [Salmonella enterica subsp. enterica serovar
Paratyphi A str. ATCC 9150] gi|16501881|emb|CAD05106.1|
cold shock-like protein cspE [Salmonella enterica
subsp. enterica serovar Typhi]
gi|16419141|gb|AAL19580.1| RNA chaperone, negative
regulator of cspA transcription [Salmonella typhimurium
LT2] gi|29138269|gb|AAO69840.1| cold shock-like protein
cspE [Salmonella enterica subsp. enterica serovar Typhi
Ty2] gi|16759589|ref|NP_455206.1| cold shock-like
protein cspE [Salmonella enterica subsp. enterica
serovar Typhi str. CT18] gi|16764006|ref|NP_459621.1|
negative regulator [Salmonella typhimurium LT2]
gi|25296131|pir||AH0579 cold shock-like protein cspE
[imported] - Salmonella enterica subsp. enterica
serovar Typhi (strain CT18)
Length = 69
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/55 (38%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 55
>gb|AAX64564.1| RNA chaperone, negative regulator of cspA transcription
[Salmonella enterica subsp. enterica serovar
Choleraesuis str. SC-B67] gi|62179228|ref|YP_215645.1|
RNA chaperone, negative regulator of cspA transcription
[Salmonella enterica subsp. enterica serovar
Choleraesuis str. SC-B67]
Length = 79
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/55 (38%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 11 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 65
>ref|NP_752643.1| Cold shock-like protein cspE [Escherichia coli CFT073]
gi|26107003|gb|AAN79187.1| Cold shock-like protein cspE
[Escherichia coli CFT073]
Length = 97
Score = 42.4 bits (98), Expect = 0.003
Identities = 21/55 (38%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 29 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 83
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 186,539,123
Number of Sequences: 2540612
Number of extensions: 6299414
Number of successful extensions: 40115
Number of sequences better than 10.0: 302
Number of HSP's better than 10.0 without gapping: 254
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 39757
Number of HSP's gapped (non-prelim): 357
length of query: 119
length of database: 863,360,394
effective HSP length: 95
effective length of query: 24
effective length of database: 622,002,254
effective search space: 14928054096
effective search space used: 14928054096
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0089a.1


