

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.21
(1215 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
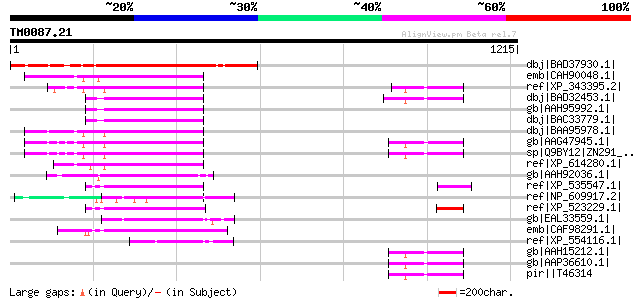
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAD37930.1| zinc finger protein-like [Oryza sativa (japonica... 610 e-173
emb|CAH90048.1| hypothetical protein [Pongo pygmaeus] 143 4e-32
ref|XP_343395.2| PREDICTED: zinc finger protein 291 [Rattus norv... 142 5e-32
dbj|BAD32453.1| mKIAA1454 protein [Mus musculus] 142 5e-32
gb|AAH95992.1| Zfp291 protein [Mus musculus] 142 5e-32
dbj|BAC33779.1| unnamed protein product [Mus musculus] 142 5e-32
dbj|BAA95978.1| KIAA1454 protein [Homo sapiens] 142 5e-32
gb|AAG47945.1| MSTP063 [Homo sapiens] 140 2e-31
sp|Q9BY12|ZN291_HUMAN Zinc finger protein 291 gi|13539684|gb|AAK... 140 2e-31
ref|XP_614280.1| PREDICTED: similar to KIAA1454 protein, partial... 140 3e-31
gb|AAH92036.1| Unknown (protein for MGC:85007) [Xenopus laevis] 140 3e-31
ref|XP_535547.1| PREDICTED: similar to zinc finger protein 291 [... 139 6e-31
ref|NP_609917.2| CG18397-PA [Drosophila melanogaster] gi|2294679... 132 7e-29
ref|XP_523229.1| PREDICTED: similar to zinc finger protein 291 [... 127 3e-27
gb|EAL33559.1| GA14918-PA [Drosophila pseudoobscura] 124 1e-26
emb|CAF98291.1| unnamed protein product [Tetraodon nigroviridis] 123 3e-26
ref|XP_554116.1| ENSANGP00000026011 [Anopheles gambiae str. PEST... 105 9e-21
gb|AAH15212.1| ZNF291 protein [Homo sapiens] gi|30582363|gb|AAP3... 102 1e-19
gb|AAP36610.1| Homo sapiens zinc finger protein 291 [synthetic c... 102 1e-19
pir||T46314 hypothetical protein DKFZp434M1511.1 - human (fragme... 102 1e-19
>dbj|BAD37930.1| zinc finger protein-like [Oryza sativa (japonica cultivar-group)]
gi|51535126|dbj|BAD37789.1| zinc finger protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 996
Score = 610 bits (1574), Expect = e-173
Identities = 361/594 (60%), Positives = 435/594 (72%), Gaps = 29/594 (4%)
Query: 1 MTTSPHRADILSSSLEAFRKIQQERASLKLSNNTEHAMSKCLTSESVGNMNGTHNAKNSR 60
MT+SPHR +ILSSSLEAF++IQ E A + TE S S G ++G+ + +
Sbjct: 417 MTSSPHRQEILSSSLEAFQRIQLELARKQAGITTES-----FASSSSGEVSGSSSKLTTA 471
Query: 61 TKSRKHIGSSDANQGNLNVKKHNIEGTKPCDAIRVQKGYIPPESLLTSEVDLAKLPSLEN 120
+ + I +Q L+ + I G + ++ G PP+++ +S K SLE
Sbjct: 472 SATVGSISLKVESQVKLSDTEKKIAGERQSKDT-IKSGRSPPQNMPSSSAKSRK-GSLEP 529
Query: 121 SSALATAKVKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDNLRRQMQLPEKDKEKRSTS 180
S + + KDK E +K RSTD +R EK EK++ +
Sbjct: 530 ISEVEKHNFR-------------KDKELPENKFDKL-RSTDTAKRTTVHTEK--EKQNAA 573
Query: 181 LGKSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTS 240
KS++AWKEKRNWEDIL SP R SSR+S+SP +GRK ER R LHDKLMSPEKKK++
Sbjct: 574 PRKSLDAWKEKRNWEDILKSPVR-SSRVSHSPGVGRKVPERARVLHDKLMSPEKKKRSAL 632
Query: 241 DLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRS 300
D+KKEAEEKHARALRIRS+LE+ERVQ+LQRTS+KLNRV+EW AVR KLRE M ARHQRS
Sbjct: 633 DMKKEAEEKHARALRIRSQLESERVQRLQRTSEKLNRVNEWQAVRSSKLREVMNARHQRS 692
Query: 301 ESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKS 360
ESRHEA+LAQVAKRAGDES+KVNEVRFITSLNEENKK +LRQKLHGSE+RRAEKL VIK+
Sbjct: 693 ESRHEAYLAQVAKRAGDESTKVNEVRFITSLNEENKKFLLRQKLHGSEMRRAEKLLVIKT 752
Query: 361 KQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQL 420
KQKED+AREEAVLERRK++EAEK+QRLAE+QRKKEEA IRREEERKASSAAREARA EQ
Sbjct: 753 KQKEDIAREEAVLERRKILEAEKMQRLAEIQRKKEEAIIRREEERKASSAAREARAAEQQ 812
Query: 421 RRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERANL--RDQSSPLLRRSLN 478
RRKE RAKAQQEEAELLAQKLAE+L ESEQRRK YLEQIRERA++ RDQ SP RR +
Sbjct: 813 RRKEIRAKAQQEEAELLAQKLAEKLRESEQRRKYYLEQIRERASMDFRDQPSPFQRRFPS 872
Query: 479 KDGQGRSTPNNSSDDSQTNIASGLGSSLGIGSITLQHSMKRRIKRIRQKLMALKYEFLEP 538
KD Q RS+ NS +DSQ I+S + G+ S MKRRIK+IRQ+LMALK++F+E
Sbjct: 873 KDNQNRSSSANSGEDSQI-ISSANAAESGVKSFN-STQMKRRIKKIRQRLMALKHDFVE- 929
Query: 539 PLGGESAGIGYRVAVGAARAKVGRWLQELQRLRQARKEGATSIGLIISEMIKYL 592
PL GE+ GI +R A+G A+AK+ RWLQ+LQRLRQARKEGA SIGLI+S+M K L
Sbjct: 930 PLIGENTGIVHRSALGTAKAKLSRWLQDLQRLRQARKEGAASIGLIVSDMTKVL 983
>emb|CAH90048.1| hypothetical protein [Pongo pygmaeus]
Length = 732
Score = 143 bits (360), Expect = 4e-32
Identities = 120/449 (26%), Positives = 226/449 (49%), Gaps = 29/449 (6%)
Query: 36 HAMSKCLTSESVGNMNGTHNAKNSRTKSRKHIGSSDANQGNLNVKKHNIEGTKPCDAIRV 95
H+ L ++ ++ + KNS + ++ +S+ + +++ + ++ ++V
Sbjct: 153 HSCDHPLAEKTQFTVSTLDDVKNSGSIQDNYVRTSEISAVHIDTECVSVMLQADAPPLQV 212
Query: 96 QKGYIPPESL-LTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEVINE 154
+ P E + +E+D + + ++ +++ L+ A+V AD+L ++A I E
Sbjct: 213 NEEKFPAEKARIENEMDPSDISNV-SAANLSMAEVLAKKEELADRLEKANEEAIASAIAE 271
Query: 155 KNPRSTDNLRRQMQLPEKDK--------EKRSTSLGK-SMNAWKEKRNWEDILSS-PFRV 204
+ + L R+++ E + S S+G S++ +W D+L+ R
Sbjct: 272 E-----EQLTREIEAEENNDINIETDNDSDFSASMGSGSVSFCGMSMDWNDVLADCEARE 326
Query: 205 SSRMSYS------PSLGRKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRS 258
S R + S R + +H+KL SP +K+ T ++ KK+ EEK +A ++R
Sbjct: 327 SWRQNTSWGDIVEEEPARPPGHGIH-MHEKLSSPSRKR-TIAESKKKHEEKQMKAQQLRE 384
Query: 259 ELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDE 318
+L E+ KLQ+ ++ V +W + R M + +E + E L + K+A +E
Sbjct: 385 KLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEFKREVQLQAIVKKAQEE 444
Query: 319 SSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKL 378
+KVNE+ FI +L +NK+ + KL E R E + + +Q+E AR+EAV ER++
Sbjct: 445 EAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQEEKQARDEAVQERKRA 504
Query: 379 IEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER----AKAQQEEA 434
+EAE+ R+ E+ K++E + R E++R+ ARE A E+ R +EER AQQE
Sbjct: 505 LEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERARDREERLAALTAAQQEAM 564
Query: 435 ELLAQKLAERLNESEQRRKIYLEQIRERA 463
E L +K+ + +ES +R +EQ +E+A
Sbjct: 565 EELQKKIQLKHDESIRRHMEQIEQRKEKA 593
>ref|XP_343395.2| PREDICTED: zinc finger protein 291 [Rattus norvegicus]
Length = 1401
Score = 142 bits (359), Expect = 5e-32
Identities = 118/397 (29%), Positives = 199/397 (49%), Gaps = 36/397 (9%)
Query: 92 AIRVQKGYIPPESL-----LTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDK 146
A+ K +P E++ T D++ + + + S A AK K L AD+L ++
Sbjct: 385 ALPTDKDRVPAETVGMVTETTDPSDISNVSAADWSMAEVLAK-KEEL---ADRLEKANEE 440
Query: 147 APTEVINEKNPRSTDNLRRQMQLPEKDK--------EKRSTSLGK-SMNAWKEKRNWEDI 197
A I E+ + L R+++ E + S S+G S++ +W D+
Sbjct: 441 AIASAIAEE-----EQLTREIEAEENNDINIETDNDSDFSASMGSGSLSFCGMSLDWNDV 495
Query: 198 LSSPFRVSSRMSYSPSLGRKSAERVRTL-------HDKLMSPEKKKKTTSDLKKEAEEKH 250
L+ + + S G E L H+KL SP +K+ T ++ KK+ EEKH
Sbjct: 496 LAD-YEARESWRQNTSWGDIVEEEPARLPGHGIHMHEKLSSPSRKR-TIAESKKKYEEKH 553
Query: 251 ARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQ 310
+A ++R +L E+ KLQ+ ++ V +W + R M + +E + E L
Sbjct: 554 MKAQQLREKLREEKSLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEFKREVQLQA 613
Query: 311 VAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREE 370
+ K+A +E +KVNE+ FI +L +NK+ + KL E R E + + +Q+E AR+E
Sbjct: 614 IVKKAQEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQEEKQARDE 673
Query: 371 AVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER---- 426
AV ER++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R +EER
Sbjct: 674 AVQERKRALEAERQARVEELLTKRKEQEARIEQQRQEKEKAREDAARERARDREERLAAL 733
Query: 427 AKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
AQQE E L +K+ + +ES +R +EQ +E+A
Sbjct: 734 TAAQQEAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 770
Score = 103 bits (256), Expect = 5e-20
Identities = 59/184 (32%), Positives = 100/184 (54%), Gaps = 17/184 (9%)
Query: 915 LLSALSETGLVSLPSLLTAVLLQAN-NRSSSEP----------VATGVLKVLNNVALLDL 963
L +AL T L + +L +L + +S+ P VA L+ N+ A+LDL
Sbjct: 1173 LTAALQATDLAGVLHMLYCILFHGTISDASTSPKESYTQNTIQVAIQSLRFFNSFAVLDL 1232
Query: 964 VFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGHFALFHPGNQ 1023
Q ++ L + H+ S +L HC+ SL+ E + +G+F + HP NQ
Sbjct: 1233 SAFQSIVGAEGLSLAFRHMASSLLGHCSQV------SCESLLHEVIVCVGYFTVNHPDNQ 1286
Query: 1024 AVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQQELSVDMLL 1083
+++ G+ PT+L K+C LPF +FSDP L+ +L +L+AAC+ QNK +++QE+S +L
Sbjct: 1287 VIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACHNNHQNKLILEQEMSCVLLA 1346
Query: 1084 SLLR 1087
+ ++
Sbjct: 1347 TFIQ 1350
>dbj|BAD32453.1| mKIAA1454 protein [Mus musculus]
Length = 1393
Score = 142 bits (359), Expect = 5e-32
Identities = 92/286 (32%), Positives = 157/286 (54%), Gaps = 20/286 (6%)
Query: 183 KSMNAWKEKRNWEDILSS-PFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSD 241
++ +W++ +W DI+ P R+ + +H+KL SP +K+ T ++
Sbjct: 492 EARESWRQNTSWGDIVEEEPARLPGHGIH--------------MHEKLSSPSRKR-TIAE 536
Query: 242 LKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSE 301
KK+ EEKH +A ++R +L E+ KLQ+ ++ V +W + R M + +E
Sbjct: 537 SKKKYEEKHMKAQQLREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAE 596
Query: 302 SRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSK 361
+ E L + K+A +E +KVNE+ FI +L +NK+ + KL E R E + + +
Sbjct: 597 FKREVQLQAIVKKAQEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRR 656
Query: 362 QKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLR 421
Q+E AR+EAV ER++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R
Sbjct: 657 QEEKQARDEAVQERKRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERAR 716
Query: 422 RKEER----AKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
+EER AQQE E L +K+ + +ES +R +EQ +E+A
Sbjct: 717 DREERLAALTAAQQEAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 762
Score = 101 bits (252), Expect = 1e-19
Identities = 60/202 (29%), Positives = 104/202 (50%), Gaps = 17/202 (8%)
Query: 897 ITTQKNEKSSNLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN-NRSSSEP--------- 946
+T + + N Q L +AL T L + +L +L + + + P
Sbjct: 1147 VTGRSSSIFDNNHQDPTGLTAALQATDLAGVLHMLYCILFHGTISDAGTSPKDSYTQNTI 1206
Query: 947 -VATGVLKVLNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLM 1005
VA L+ N+ A+LDL Q ++ L + H+ S +L HC+ SL+
Sbjct: 1207 QVAIQSLRFFNSFAILDLSAFQSVVGAEGLSLAFRHMASSLLGHCSQV------SCESLL 1260
Query: 1006 LESLSLLGHFALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYG 1065
E + +G+F + HP NQ +++ G+ PT+L K+C LPF +FSDP L+ +L +L+ AC+
Sbjct: 1261 HEVIVCVGYFTVNHPDNQVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLITACHN 1320
Query: 1066 CEQNKFVVQQELSVDMLLSLLR 1087
QNK +++QE+S +L + ++
Sbjct: 1321 NHQNKLILEQEMSCVLLATFIQ 1342
>gb|AAH95992.1| Zfp291 protein [Mus musculus]
Length = 863
Score = 142 bits (359), Expect = 5e-32
Identities = 92/286 (32%), Positives = 157/286 (54%), Gaps = 20/286 (6%)
Query: 183 KSMNAWKEKRNWEDILSS-PFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSD 241
++ +W++ +W DI+ P R+ + +H+KL SP +K+ T ++
Sbjct: 490 EARESWRQNTSWGDIVEEEPARLPGHGIH--------------MHEKLSSPSRKR-TIAE 534
Query: 242 LKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSE 301
KK+ EEKH +A ++R +L E+ KLQ+ ++ V +W + R M + +E
Sbjct: 535 SKKKYEEKHMKAQQLREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAE 594
Query: 302 SRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSK 361
+ E L + K+A +E +KVNE+ FI +L +NK+ + KL E R E + + +
Sbjct: 595 FKREVQLQAIVKKAQEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRR 654
Query: 362 QKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLR 421
Q+E AR+EAV ER++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R
Sbjct: 655 QEEKQARDEAVQERKRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERAR 714
Query: 422 RKEER----AKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
+EER AQQE E L +K+ + +ES +R +EQ +E+A
Sbjct: 715 DREERLAALTAAQQEAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 760
>dbj|BAC33779.1| unnamed protein product [Mus musculus]
Length = 864
Score = 142 bits (359), Expect = 5e-32
Identities = 92/286 (32%), Positives = 157/286 (54%), Gaps = 20/286 (6%)
Query: 183 KSMNAWKEKRNWEDILSS-PFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSD 241
++ +W++ +W DI+ P R+ + +H+KL SP +K+ T ++
Sbjct: 497 EARESWRQNTSWGDIVEEEPARLPGHGIH--------------MHEKLSSPSRKR-TIAE 541
Query: 242 LKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSE 301
KK+ EEKH +A ++R +L E+ KLQ+ ++ V +W + R M + +E
Sbjct: 542 SKKKYEEKHMKAQQLREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAE 601
Query: 302 SRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSK 361
+ E L + K+A +E +KVNE+ FI +L +NK+ + KL E R E + + +
Sbjct: 602 FKREVQLQAIVKKAQEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRR 661
Query: 362 QKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLR 421
Q+E AR+EAV ER++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R
Sbjct: 662 QEEKQARDEAVQERKRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERAR 721
Query: 422 RKEER----AKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
+EER AQQE E L +K+ + +ES +R +EQ +E+A
Sbjct: 722 DREERLAALTAAQQEAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 767
>dbj|BAA95978.1| KIAA1454 protein [Homo sapiens]
Length = 1265
Score = 142 bits (359), Expect = 5e-32
Identities = 121/452 (26%), Positives = 227/452 (49%), Gaps = 35/452 (7%)
Query: 36 HAMSKCLTSESVGNMNGTHNAKNSRTKSRKHIGSSDANQGNLNVKKHNI---EGTKPCDA 92
H+ L ++ ++ + KNS + ++ +S+ + +++ + ++ GT P
Sbjct: 349 HSCDHPLAEKTQFTVSTLDDVKNSGSIRDNYVRTSEISAVHIDTECVSVMLQAGTPP--- 405
Query: 93 IRVQKGYIPPESL-LTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEV 151
++V + P E + +E+D + + ++ +++ L+ A+V AD+L ++A
Sbjct: 406 LQVNEEKFPAEKARIENEMDPSDISNV-SAANLSMAEVLAKKEELADRLEKANEEAIASA 464
Query: 152 INEKNPRSTDNLRRQMQLPEKDK--------EKRSTSLGK-SMNAWKEKRNWEDILSSPF 202
I E+ + L R+++ E + S S+G S++ +W D+L+ +
Sbjct: 465 IAEE-----EQLTREIEAEENNDINIETDNDSDFSASMGSGSVSFCGMSMDWNDVLAD-Y 518
Query: 203 RVSSRMSYSPSLGRKSAERVRT-------LHDKLMSPEKKKKTTSDLKKEAEEKHARALR 255
+ S G E +H+KL SP +K+ T ++ KK+ EEK +A +
Sbjct: 519 EARESWRQNTSWGDIVEEEPARPPGHGIHMHEKLSSPSRKR-TIAESKKKHEEKQMKAQQ 577
Query: 256 IRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRA 315
+R +L E+ KLQ+ ++ V +W + R M + +E + E L + K+A
Sbjct: 578 LREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEFKREVQLQAIVKKA 637
Query: 316 GDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLER 375
+E +KVNE+ FI +L +NK+ + KL E R E + + +Q+E AR+EAV ER
Sbjct: 638 QEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQEEKQARDEAVQER 697
Query: 376 RKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER----AKAQQ 431
++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R +EER AQQ
Sbjct: 698 KRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERARDREERLAALTAAQQ 757
Query: 432 EEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
E E L +K+ + +ES +R +EQ +E+A
Sbjct: 758 EAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 789
>gb|AAG47945.1| MSTP063 [Homo sapiens]
Length = 1153
Score = 140 bits (354), Expect = 2e-31
Identities = 125/452 (27%), Positives = 224/452 (48%), Gaps = 41/452 (9%)
Query: 36 HAMSKCLTSESVGNMNGTHNAKNSRTKSRKHIGSSDANQGNLNVKKHNI---EGTKPCDA 92
H+ L ++ ++ + KNS + ++ +S+ + +++ + ++ GT P
Sbjct: 87 HSCDHPLAEKTQFTVSTLDDVKNSGSIRDNYVRTSEISAVHIDTECVSVMLQAGTPP--- 143
Query: 93 IRVQKGYIPPESL-LTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEV 151
++V + P E + +E+D + + NS A AK K L AD+L ++A
Sbjct: 144 LQVNEEKFPAEKARIENEMDPS---DISNSMAEVLAK-KEEL---ADRLEKANEEAIASA 196
Query: 152 INEKNPRSTDNLRRQMQLPEKDK--------EKRSTSLGK-SMNAWKEKRNWEDILSSPF 202
I E+ + L R+++ E + S S+G S++ +W D+L+ +
Sbjct: 197 IAEE-----EQLTREIEAEENNDINIETDNDSDFSASMGSGSVSFCGMSMDWNDVLAD-Y 250
Query: 203 RVSSRMSYSPSLGRKSAERVRT-------LHDKLMSPEKKKKTTSDLKKEAEEKHARALR 255
+ S G E +H+KL SP +K+ T ++ KK+ EEK +A +
Sbjct: 251 EARESWRQNTSWGDIVEEEPARPPGHGIHMHEKLSSPSRKR-TIAESKKKHEEKQMKAQQ 309
Query: 256 IRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRA 315
+R +L E+ KLQ+ ++ V +W + R M + +E + E L + K+A
Sbjct: 310 LREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEFKREVQLQAIVKKA 369
Query: 316 GDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLER 375
+E +KVNE+ FI +L +NK+ + KL E R E + + +Q+E AR+EAV ER
Sbjct: 370 QEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQEEKQARDEAVQER 429
Query: 376 RKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER----AKAQQ 431
++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R +EER AQQ
Sbjct: 430 KRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERARDREERLAALTAAQQ 489
Query: 432 EEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
E E L +K+ + +ES +R +EQ +E+A
Sbjct: 490 EAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 521
Score = 102 bits (254), Expect = 8e-20
Identities = 62/193 (32%), Positives = 101/193 (52%), Gaps = 18/193 (9%)
Query: 907 NLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN--NRSSSEP----------VATGVLKV 954
N Q L +AL T L + +L VL + S++ P VA L+
Sbjct: 916 NNRQDPTGLTAALQATDLAGVLHMLYCVLFHGTILDPSTASPKENYTQNTIQVAIQSLRF 975
Query: 955 LNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGH 1014
N+ A L L Q ++ L + H+ S +L HC+ + SL+ E + +G+
Sbjct: 976 FNSFAALHLPAFQSIVGAEGLSLAFRHMASSLLGHCSQVF------CESLLHEVIVCVGY 1029
Query: 1015 FALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQ 1074
F + HP NQ +++ G+ PT+L K+C LPF +FSDP L+ +L +L+AACY QNK +++
Sbjct: 1030 FTVNHPDNQVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACYNNHQNKIILE 1089
Query: 1075 QELSVDMLLSLLR 1087
QE+S +L + ++
Sbjct: 1090 QEMSCVLLATFIQ 1102
>sp|Q9BY12|ZN291_HUMAN Zinc finger protein 291 gi|13539684|gb|AAK29205.1| zinc finger
protein 291 [Homo sapiens] gi|16507198|ref|NP_065894.1|
zinc finger protein 291 [Homo sapiens]
Length = 1399
Score = 140 bits (354), Expect = 2e-31
Identities = 125/452 (27%), Positives = 224/452 (48%), Gaps = 41/452 (9%)
Query: 36 HAMSKCLTSESVGNMNGTHNAKNSRTKSRKHIGSSDANQGNLNVKKHNI---EGTKPCDA 92
H+ L ++ ++ + KNS + ++ +S+ + +++ + ++ GT P
Sbjct: 333 HSCDHPLAEKTQFTVSTLDDVKNSGSIRDNYVRTSEISAVHIDTECVSVMLQAGTPP--- 389
Query: 93 IRVQKGYIPPESL-LTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEV 151
++V + P E + +E+D + + NS A AK K L AD+L ++A
Sbjct: 390 LQVNEEKFPAEKARIENEMDPS---DISNSMAEVLAK-KEEL---ADRLEKANEEAIASA 442
Query: 152 INEKNPRSTDNLRRQMQLPEKDK--------EKRSTSLGK-SMNAWKEKRNWEDILSSPF 202
I E+ + L R+++ E + S S+G S++ +W D+L+ +
Sbjct: 443 IAEE-----EQLTREIEAEENNDINIETDNDSDFSASMGSGSVSFCGMSMDWNDVLAD-Y 496
Query: 203 RVSSRMSYSPSLGRKSAERVRT-------LHDKLMSPEKKKKTTSDLKKEAEEKHARALR 255
+ S G E +H+KL SP +K+ T ++ KK+ EEK +A +
Sbjct: 497 EARESWRQNTSWGDIVEEEPARPPGHGIHMHEKLSSPSRKR-TIAESKKKHEEKQMKAQQ 555
Query: 256 IRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRA 315
+R +L E+ KLQ+ ++ V +W + R M + +E + E L + K+A
Sbjct: 556 LREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEFKREVQLQAIVKKA 615
Query: 316 GDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLER 375
+E +KVNE+ FI +L +NK+ + KL E R E + + +Q+E AR+EAV ER
Sbjct: 616 QEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQEEKQARDEAVQER 675
Query: 376 RKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER----AKAQQ 431
++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R +EER AQQ
Sbjct: 676 KRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERARDREERLAALTAAQQ 735
Query: 432 EEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
E E L +K+ + +ES +R +EQ +E+A
Sbjct: 736 EAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 767
Score = 102 bits (253), Expect = 1e-19
Identities = 62/193 (32%), Positives = 100/193 (51%), Gaps = 18/193 (9%)
Query: 907 NLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN--NRSSSEP----------VATGVLKV 954
N Q L +AL T L + +L VL + S++ P VA L+
Sbjct: 1162 NNRQDPTGLTAALQATDLAGVLHMLYCVLFHGTILDPSTASPKENYTQNTIQVAIQSLRF 1221
Query: 955 LNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGH 1014
N+ A L L Q ++ L + H+ S +L HC+ SL+ E + +G+
Sbjct: 1222 FNSFAALHLPAFQSIVGAEGLSLAFRHMASSLLGHCSQV------SCESLLHEVIVCVGY 1275
Query: 1015 FALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQ 1074
F + HP NQ +++ G+ PT+L K+C LPF +FSDP L+ +L +L+AACY QNK +++
Sbjct: 1276 FTVNHPDNQVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACYNNHQNKIILE 1335
Query: 1075 QELSVDMLLSLLR 1087
QE+S +L + ++
Sbjct: 1336 QEMSCVLLATFIQ 1348
>ref|XP_614280.1| PREDICTED: similar to KIAA1454 protein, partial [Bos taurus]
Length = 497
Score = 140 bits (352), Expect = 3e-31
Identities = 108/376 (28%), Positives = 190/376 (49%), Gaps = 26/376 (6%)
Query: 106 LTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDNLRR 165
+ SE+D + + ++ +++ L+ A+V AD+L ++A I E+ + L R
Sbjct: 62 IESEMDPSDISNV-SAANLSMAEVLAKKEELADRLEKANEEAIASAIAEE-----EQLTR 115
Query: 166 QMQLPEKDKEKRSTSLGKSMNAWKEK-------RNWEDILSSPFRVSSRMSYSPSLGRKS 218
+++ E + T +A NW D+L+ + + S G
Sbjct: 116 EIEAEENNDINLETDNDSDFSASMGSGSVSFCGMNWNDVLAD-YEARESWRQNTSWGDIV 174
Query: 219 AERVRT-------LHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRT 271
E +H+KL SP +K+ T ++ KK+ EEK +A ++R +L ++ KLQ+
Sbjct: 175 EEEPARPPGHGIHMHEKLSSPSRKR-TIAESKKKHEEKQMKAQQLREKLREQKTLKLQKL 233
Query: 272 SQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSL 331
++ V +W + R M + +E + E L + K+A +E +KVNE+ FI +L
Sbjct: 234 LEREKEVRKWKEGLLDQRRRMMEEKLLHAEFKREVQLQAIVKKAQEEEAKVNEIAFINTL 293
Query: 332 NEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQ 391
+NK+ + KL E R E + + +Q+E AR+EAV ER++ +EAE+ R+ E+
Sbjct: 294 EAQNKRHDVLSKLKEYEQRLNELQEERQRRQEEKQARDEAVQERKRALEAERQARVEELL 353
Query: 392 RKKEEAQIRREEERKASSAAREARAIEQLRRKEER----AKAQQEEAELLAQKLAERLNE 447
K++E + R E++R+ ARE A E+ R +EER AQQE E L +K+ + +E
Sbjct: 354 MKRKEQEARIEQQRQEKEKAREDAARERARDREERLAALTAAQQEAMEELQKKIQLKHDE 413
Query: 448 SEQRRKIYLEQIRERA 463
S +R +EQ +E+A
Sbjct: 414 SIRRHMEQIEQRKEKA 429
>gb|AAH92036.1| Unknown (protein for MGC:85007) [Xenopus laevis]
Length = 683
Score = 140 bits (352), Expect = 3e-31
Identities = 125/414 (30%), Positives = 201/414 (48%), Gaps = 33/414 (7%)
Query: 89 PCDAIRVQKGYIPPESLLTSEVDLAKLPSLENSSALATAKVKTHLGSGADKLLSLKDKAP 148
PC + + G P L S V KL + A+V AD+L ++A
Sbjct: 258 PCITDKARGGRKPGSDLDLSNVSAGKL---------SMAEVLAKKEELADRLEKANEEAI 308
Query: 149 TEVINEKNPRSTDNLRRQMQLPEKDKEKR-STSLGKSMNAWKE-KRNWEDILSS-PFRVS 205
I E+ + + + E D E S S+G + + +W ++L+ R S
Sbjct: 309 ASAIAEEEQLTREIEAEENNDIEIDNESDFSASIGSASLLYNGMSMDWNEVLADYEARES 368
Query: 206 SRMSYS--------PSLGRKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIR 257
SR + S PS R + +H+KL SP +K+ T ++ KK+ EEK +A ++R
Sbjct: 369 SRQNTSWGDMVEEEPS--RPPGHGIH-MHEKLSSPSRKR-TIAESKKKHEEKQLKAQQLR 424
Query: 258 SELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGD 317
+L E+ KLQ+ ++ V +W + R+ M + +E + E L + K+A
Sbjct: 425 EKLREEKAVKLQKLLEREKDVRKWKEGLFEQKRKMMDEKLLHAEFKREVQLQAIVKKAQV 484
Query: 318 ESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRK 377
E +KVNE+ FI +L +NK+ + KL E R E + + +Q+E AR+EAV ER++
Sbjct: 485 EEAKVNEIAFINTLEAQNKRHDVLAKLKEYEQRLNELQEERQQRQEEKQARDEAVQERKR 544
Query: 378 LIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER----AKAQQEE 433
+EAE+ R+ E+ K++E + R E +R+ ARE A E+ R +EER AQQE
Sbjct: 545 ALEAERQARVEELLMKRKEQEARIEMQRQEKERAREDAARERARDREERLAALTAAQQEA 604
Query: 434 AELLAQKLAERLNESEQRRKIYLEQIRERANLRDQSSPLLRRSLNKDGQGRSTP 487
E L +K+ + +ES +R +LEQI +R + S R N D + TP
Sbjct: 605 MEELQKKIQLKHDESVRR---HLEQIEQRKEKAAELSS--GRHANTDDAPKLTP 653
>ref|XP_535547.1| PREDICTED: similar to zinc finger protein 291 [Canis familiaris]
Length = 3564
Score = 139 bits (350), Expect = 6e-31
Identities = 92/285 (32%), Positives = 154/285 (53%), Gaps = 18/285 (6%)
Query: 183 KSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSDL 242
++ W++ +W DI V + P G +H+KL SP +K+ T ++
Sbjct: 474 EARETWRQNTSWGDI------VEEEPARPPGHGIH-------MHEKLSSPSRKR-TIAES 519
Query: 243 KKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSES 302
KK+ EEK +A ++R +L E+ KLQ+ ++ V +W + R M + +E
Sbjct: 520 KKKHEEKQMKAQQLREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEF 579
Query: 303 RHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQ 362
+ E L + K+A +E +KVNE+ FI +L +NK+ + KL E R E + + +Q
Sbjct: 580 KREVQLQAIVKKAQEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQ 639
Query: 363 KEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRR 422
+E AR+EAV ER++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R
Sbjct: 640 EEKQARDEAVQERKRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERARD 699
Query: 423 KEER----AKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
+EER AQQE E L +K+ + +ES +R +EQ +E+A
Sbjct: 700 REERLAALTAAQQEAMEELQKKIQLKHDESIRRHMEQIEQRKEKA 744
Score = 59.7 bits (143), Expect = 6e-07
Identities = 29/83 (34%), Positives = 50/83 (59%)
Query: 1025 VLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQQELSVDMLLS 1084
V++ G+ PT+L K+C LPF +FS+P L +L +L+AAC+ N+ +++QE+S +L +
Sbjct: 3329 VVQSGRHPTVLQKLCQLPFQYFSEPRLTKVLFPSLLAACFNNRHNRTILEQEMSCVLLAT 3388
Query: 1085 LLRSCKSAAPATQLITLDNSATD 1107
++ QL T + A D
Sbjct: 3389 FVQDVILPGRLGQLQTPASLAAD 3411
>ref|NP_609917.2| CG18397-PA [Drosophila melanogaster] gi|22946790|gb|AAF53722.3|
CG18397-PA [Drosophila melanogaster]
gi|61675681|gb|AAX51656.1| LD01527p [Drosophila
melanogaster]
Length = 1732
Score = 132 bits (332), Expect = 7e-29
Identities = 102/327 (31%), Positives = 172/327 (52%), Gaps = 16/327 (4%)
Query: 221 RVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSE 280
R + LH KL SP +++ LKK + K ARA + R+ L+ E+ KLQ Q +RV +
Sbjct: 989 RAQELHQKLSSPSRRRSLQETLKKY-QAKQARAQQKRNLLQQEKAAKLQ---QLFSRVED 1044
Query: 281 WHAVRHLKL---REGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKK 337
A ++ + R+ M R QR+ E +L Q+ ++A DE K+ E+ FI ++ +NK+
Sbjct: 1045 VKAAKNQIIEDKRQKMQGRLQRAAENREQYLKQIIEKAHDEEKKLKEINFIKNIEAQNKR 1104
Query: 338 LILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEA 397
L L + +E R + Q + + +E LA+E AV RR+ +E E+L +L +M + E
Sbjct: 1105 LDLLESSKETEGRLQDLEQERQKRVEEKLAKEAAVERRRQALEKERLLKLEKMNETRLEK 1164
Query: 398 QIRREEERKASSAAREARAIEQLRRKEERAKA----QQEEAELLAQKLAERLNESEQRRK 453
+ R + ++ R+A A E+ R +EER A QQ+ E L +K+ ++ ES +R +
Sbjct: 1165 EQRIGKMQEQKEKQRQALAREKARDREERLLALQVQQQQTTEELQRKILQKQMESARRHE 1224
Query: 454 IYLEQIRERANLRDQSSPLLRRSLNKDGQGRSTP-NNSSDDSQTNIASGLGSSLGI--GS 510
+E IR+RA + + P + + Q S N + S TN L SSL G+
Sbjct: 1225 ENIEHIRQRA--LELTIPTRQADEGRGDQDVSEDILNGNATSTTNEDCDLSSSLSEVGGN 1282
Query: 511 ITLQHSMKRRIKRIRQKLMALKYEFLE 537
S K+++K+++Q++ E+LE
Sbjct: 1283 NAHTRSYKKKMKKLKQRMNQCAAEYLE 1309
Score = 58.5 bits (140), Expect = 1e-06
Identities = 102/534 (19%), Positives = 194/534 (36%), Gaps = 103/534 (19%)
Query: 11 LSSSLEAFRKIQQERASLKLSNNTEHAMSKCLTSESVGNMNGTHNAKNSRTKSRKHIGSS 70
+SS L + I ++LK SN E + N T A+N ++ ++ +
Sbjct: 573 MSSRLSSREDITSSTSTLKASN------------EQLSNSRNTLKAENQHSEPKQIQPPT 620
Query: 71 DANQGNLNVKKHNIEGTKPCDAIRVQKGY--IPPESLLTSEVDLAKLPSLENSSALATAK 128
DA+ G L VK + GY +P +LL + + +
Sbjct: 621 DADDGWLTVKNRRRTSMHWANRFNQPTGYASLPTLALLNEQQKEQEHKEKQKGEDDGKVI 680
Query: 129 VKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDNLRRQMQLPEKDKEKRSTSLGKSMNAW 188
VKT +S K KAP EV K S R +++ + +
Sbjct: 681 VKT---------ISAKTKAPIEVAKAKAKTSIVITRPEIKNAKAKVNSFPVQKSNTNQVK 731
Query: 189 KEKRNWEDILSSPFRVSS-----RMSYSPSLGRKS----------AERVRTLHDKLMSPE 233
K ++ + ++P ++S S+S +GR S ++ +LH + M E
Sbjct: 732 KPEKQEKSDTTAPAAIASSSNTRNSSHSICVGRNSIIKRQKSDLTGLKMTSLHKEYMRSE 791
Query: 234 KKKKTTSDLKKEAEEKHARA-------LRIRSELETERVQKLQRTSQKLNRVSE-WHAVR 285
K K++ ++H + + +E ++ K+Q + + E + ++
Sbjct: 792 KNALRKLQQKEQGNQQHNSSSSSAETVVESCNEDHSKIDIKIQTNCEFSKTIGELYESIA 851
Query: 286 HLKLREGMYARH----------------QRSESRHEAFLAQVAKRAGDESSKVNEVR--- 326
H KL G + E R+E L +V + + ++ +
Sbjct: 852 HCKLPSGSLKTNASTLSACDENEEQNTDDNEEERNERILGEVQESLERQIRELEQTEIDV 911
Query: 327 ---------------------------------FITSLNEENKKLILRQKLHGSEL---R 350
F+T ++EN L LR + S++
Sbjct: 912 DTETDETDCEVQLEEQDDGVDGLEMGSGDDSAVFVTMSDDENASLELRYQALLSDMSWNE 971
Query: 351 RAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSA 410
RAE L +++ R + + +KL + + L E +K + Q R +++R
Sbjct: 972 RAEALATLQAYVARHPGRAQEL--HQKLSSPSRRRSLQETLKKYQAKQARAQQKRNLLQQ 1029
Query: 411 AREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERAN 464
+ A+ + R E+ A+ + E QK+ RL + + R+ YL+QI E+A+
Sbjct: 1030 EKAAKLQQLFSRVEDVKAAKNQIIEDKRQKMQGRLQRAAENREQYLKQIIEKAH 1083
>ref|XP_523229.1| PREDICTED: similar to zinc finger protein 291 [Pan troglodytes]
Length = 1410
Score = 127 bits (318), Expect = 3e-27
Identities = 89/292 (30%), Positives = 154/292 (52%), Gaps = 20/292 (6%)
Query: 183 KSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSDL 242
++ +W++ +W DI V + P G +H+KL SP +K+ T ++
Sbjct: 541 EARESWRQNTSWGDI------VEEEPARPPGHGIH-------MHEKLSSPSRKR-TIAES 586
Query: 243 KKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSEWHAVRHLKLREGMYARHQRSES 302
KK+ EEK +A ++R +L E+ KLQ+ ++ V +W + R M + +E
Sbjct: 587 KKKHEEKQMKAQQLREKLREEKTLKLQKLLEREKDVRKWKEELLDQRRRMMEEKLLHAEF 646
Query: 303 RHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQ 362
+ E L + K+A +E +KVNE+ FI +L +NK+ + KL E R E + + +Q
Sbjct: 647 KREVQLQAIVKKAQEEEAKVNEIAFINTLEAQNKRHDVLSKLKEYEQRLNELQEERQRRQ 706
Query: 363 KEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRR 422
+E AR+EAV ER++ +EAE+ R+ E+ K++E + R E++R+ ARE A E+ R+
Sbjct: 707 EEKQARDEAVQERKRALEAERQARVEELLMKRKEQEARIEQQRQEKEKAREDAARERARK 766
Query: 423 KE--ERAKAQQEEAELLAQKL--AERLNESE--QRRKIYLEQIRERANLRDQ 468
E E ++ +++ + AE L + E QR K ++I+ R N R Q
Sbjct: 767 FENWEHLSLKKYIIDIVVESTAPAEALKDGEERQRNKKKAKKIKARMNFRLQ 818
Score = 68.9 bits (167), Expect = 1e-09
Identities = 28/65 (43%), Positives = 47/65 (72%)
Query: 1023 QAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQQELSVDML 1082
Q +++ G+ PT+L K+C LPF +FSDP L+ +L +L+AACY QNK +++QE+S +L
Sbjct: 1295 QVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACYNNHQNKIILEQEMSCVLL 1354
Query: 1083 LSLLR 1087
+ ++
Sbjct: 1355 ATFIQ 1359
Score = 41.2 bits (95), Expect = 0.22
Identities = 36/132 (27%), Positives = 56/132 (42%), Gaps = 18/132 (13%)
Query: 907 NLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN--NRSSSEP----------VATGVLKV 954
N Q L +AL T L + +L VL + S++ P VA L+
Sbjct: 1108 NNRQDPTGLTAALQATDLAGVLHMLYCVLFHGTILDPSTASPKENYTQNTIQVAIQSLRF 1167
Query: 955 LNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGH 1014
N+ A L L Q ++ L + H+ S +L HC+ SL+ E + +G+
Sbjct: 1168 FNSFAALHLPAFQSIVGAEGLSLAFRHMASSLLGHCSQV------SCESLLHEVIVCVGY 1221
Query: 1015 FALFHPGNQAVL 1026
F + HP NQ V+
Sbjct: 1222 FTVNHPDNQYVI 1233
>gb|EAL33559.1| GA14918-PA [Drosophila pseudoobscura]
Length = 1706
Score = 124 bits (312), Expect = 1e-26
Identities = 100/332 (30%), Positives = 170/332 (51%), Gaps = 29/332 (8%)
Query: 221 RVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQKLNRVSE 280
R + LH KL SP +++ LKK + K ARA + R L+ E+ KLQ Q +RV +
Sbjct: 973 RAQELHQKLSSPSRRRSLQETLKKY-QAKQARAHQKRVVLQQEKAAKLQ---QLFDRVED 1028
Query: 281 WHAVRHLKL---REGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKK 337
A + + R M R QR+ E +L Q+ +A DE K+ E+ FI ++ +NK+
Sbjct: 1029 VKAAKKQLIEDKRMKMQGRLQRAAENRENYLKQIIVKAHDEDKKLKEINFIKNMEAQNKR 1088
Query: 338 LILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEA 397
L L + E R E Q + + E LA+E AV RR+ +E E+L +L +M + E
Sbjct: 1089 LDLIESSKEIEGRLQELEQERQKRVDEKLAKEAAVERRRQALEKERLLKLEKMNETRLEK 1148
Query: 398 QIRREEERKASSAAREARAIEQLRRKEERAKA----QQEEAELLAQKLAERLNESEQRRK 453
+ R + ++ R+A A E+ R +EER A QQ+ E L +K+ ++ ES +R +
Sbjct: 1149 EQRIGKMQEQKERQRQALAREKARDREERLLALQVQQQQTTEELQRKIQQKQVESARRHE 1208
Query: 454 IYLEQIRERANLRDQSSPLLRRSLNKDGQG--------RSTPNNSSDDSQTNIASGLGSS 505
+E IR+RA ++ + P + +DG+G + +N+ D ++ S +G +
Sbjct: 1209 ENIEHIRQRA--QELTMPTRQ---PEDGRGDHEETDEALNHTSNTEDGDMSSTRSDVGGT 1263
Query: 506 LGIGSITLQHSMKRRIKRIRQKLMALKYEFLE 537
G S K+++K+++Q++ E+LE
Sbjct: 1264 GG-----HSRSYKKKMKKLKQRMSQNAAEYLE 1290
Score = 48.5 bits (114), Expect = 0.001
Identities = 107/548 (19%), Positives = 204/548 (36%), Gaps = 93/548 (16%)
Query: 20 KIQQERASLKLSNNTEH--AMSKCLTSESVGNMNGTHNAKNSRTKSRKHI---------- 67
K +R+++ L N SK + E +G+ T A N + S +
Sbjct: 550 KALPKRSTITLVANQRRNAGSSKVSSREDIGSSTSTLKASNEQLSSSRSTLKMEDRHSEP 609
Query: 68 ----GSSDANQGNLNVKKHNIEGTKPCDAIRVQKGY--IPPESLLTSEV----DLAKLPS 117
+DAN G L VK + GY +P +LL + D+ K P
Sbjct: 610 KQLQQKADANDGWLTVKNRRRSSMHWSNRFNQPTGYASLPTLALLNEQQREKEDMEKKP- 668
Query: 118 LENSSALATAKVKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDNLRRQMQLPEKDKEKR 177
+ ++ ATA + G GA S+ T KN ++ N ++ P + EK
Sbjct: 669 IPQTAKTATATTEASKGKGAAAKTSMTAAGGTRP-EIKNAKAKVNSFPPLKAPARKPEKP 727
Query: 178 STSLGKSMNAWKEKRNWEDILSSPFRVSSRMSYSPSLGRKSAE----RVRTLHDKLMSPE 233
K+ +A + + ++ V++R S ++ R+ ++ ++ +LH + M E
Sbjct: 728 EKHQDKAASACGARSSLP--ATATVYVAARTS---TIKRQKSDLTGLKMTSLHKEYMRSE 782
Query: 234 KKKKTTSDLKKEAEEKHARALRIRSELETERVQ------KLQRTSQKLNRVSE-WHAVRH 286
K +KE ++ + + + +E + K+Q + + E + ++
Sbjct: 783 KNALRKQQ-QKEHDQDDGSSTSVDTVVELSNDEHHKIDIKIQTNCEFSKTIGELYESIAR 841
Query: 287 LKLREG------------------------------MYARHQRSESRHEAFLAQVAKRAG 316
KL G + Q S R L Q
Sbjct: 842 CKLPNGFLKPEANTLSACDENDEQNTDDNEEERDERILVEEQESLERQIRELEQTEIDVD 901
Query: 317 DESSKVN-EVR----------------FITSLNEENKKLILRQKLHGSEL---RRAEKLQ 356
E+ + + EV+ F+T ++EN L LR + S++ RAE L
Sbjct: 902 TETDETDCEVQLDLEQEDLMGTDDAAVFVTMSDDENASLELRYQALLSDMSWNERAEALA 961
Query: 357 VIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARA 416
+++ R + + +KL + + L E +K + Q R ++R + A+
Sbjct: 962 TLQAYVARHPGRAQEL--HQKLSSPSRRRSLQETLKKYQAKQARAHQKRVVLQQEKAAKL 1019
Query: 417 IEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERANLRDQSSPLLRRS 476
+ R E+ A+++ E K+ RL + + R+ YL+QI +A+ D+ +
Sbjct: 1020 QQLFDRVEDVKAAKKQLIEDKRMKMQGRLQRAAENRENYLKQIIVKAHDEDKKLKEINFI 1079
Query: 477 LNKDGQGR 484
N + Q +
Sbjct: 1080 KNMEAQNK 1087
>emb|CAF98291.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1214
Score = 123 bits (309), Expect = 3e-26
Identities = 112/436 (25%), Positives = 204/436 (46%), Gaps = 43/436 (9%)
Query: 114 KLPSLENSSALATAKVKTHLGSGADKLLSLKDKAPTEVINEKNPRSTDNLRRQMQLPEKD 173
KL S+ + + L+T + + + ++L +KA E I + L R++Q D
Sbjct: 329 KLESVLDPTELSTPQSMAEVLAKKEELADRLEKANEEAIASAIAEE-EQLTREIQAENND 387
Query: 174 KEKRST--------SLGKSMN------------AWKEKRNWEDILSSPFRVSSRMSYSPS 213
E + S G S++ +W++ +W D+ V S P
Sbjct: 388 LETDNDFSSAIGNGSCGASVDWSDMLADYDAHESWRQNTSWGDM------VEEEPSRPPG 441
Query: 214 LGRKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIRSELETERVQKLQRTSQ 273
G +H+KL SP +K+ T ++ KK+ EEK +A ++R +L E+ KLQ+ +
Sbjct: 442 HGIH-------MHEKLSSPSRKR-TIAESKKKHEEKQLKAQQLRDKLREEKTLKLQKLLE 493
Query: 274 KLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNE 333
+ V +W + R M + +E + E L + K+A +E +KVNE+ FI +L
Sbjct: 494 REKEVRKWKEELLEQRRRMMDEKLLHAEFKRELQLQAIVKKAQEEEAKVNEIAFINTLEA 553
Query: 334 ENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRK 393
+NK+ KL+ E R E + + +Q+E AR+EAV ER++++EAE+ R+ E+ +
Sbjct: 554 QNKRHDALAKLNEYEQRLNELQEDRQRRQEEKQARDEAVQERKRVLEAERQARVEELLMR 613
Query: 394 KEEAQIRREEERKASSAAREARAIEQLRRKEE---RAKAQQEEAELLAQKLAERLNES-- 448
++E + R E++R+ ARE A E+ R + A + ++ L++ +A +ES
Sbjct: 614 RKEQEARIEQQRQEKEKAREDAARERARSPQRCTCLATPRAKDINRLSETVAASRDESCP 673
Query: 449 -EQRRKIYLEQIRERANLRDQSSPLLRRSLNKDGQGRSTPNNSSDDSQTNIASGLGSSLG 507
++ K Y + +A + D + L KD + S N S L +LG
Sbjct: 674 MKKWAKEYETSVETKAQVPDSPYKAKLQRLVKD-LLKHLQGQDSGQWANNKVSSLDRTLG 732
Query: 508 -IGSITLQHSMKRRIK 522
+ I + + ++ +K
Sbjct: 733 ELSRILEKQATEKEVK 748
Score = 38.5 bits (88), Expect = 1.4
Identities = 35/139 (25%), Positives = 60/139 (42%), Gaps = 24/139 (17%)
Query: 896 NITTQKNEKSSNLAQPVVFLLSALSETGLVSLPSLLTAVLLQANNRSSSEP--------- 946
NI K++ S+ L+ + L T LV + L +LL + P
Sbjct: 926 NIFDNKHQDSTGLS-------ALLQSTDLVGVVHTLYCILLHGFLPEMASPRQELYDPGV 978
Query: 947 --VATGVLKVLNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSL 1004
VA ++ LN+ ALLDL Q +L L + H++S +L +C+ +
Sbjct: 979 IRVALQGIRFLNSFALLDLSAFQCVLGAEGLSLAFRHIISSLLWYCSQH------SSEEV 1032
Query: 1005 MLESLSLLGHFALFHPGNQ 1023
+ E + +G+F + +P NQ
Sbjct: 1033 LHEVIICVGYFTVNNPDNQ 1051
>ref|XP_554116.1| ENSANGP00000026011 [Anopheles gambiae str. PEST]
gi|55236418|gb|EAL39299.1| ENSANGP00000026011 [Anopheles
gambiae str. PEST]
Length = 281
Score = 105 bits (262), Expect = 9e-21
Identities = 78/254 (30%), Positives = 132/254 (51%), Gaps = 16/254 (6%)
Query: 288 KLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGS 347
K R M R QR+ +L K+A DE K+ E+ FI +L +NK+L +
Sbjct: 17 KKRLRMEERLQRAAENRSQYLKDKIKKAHDEEEKLREIAFIKNLEAQNKRLDFLESCKEQ 76
Query: 348 ELRRAEKLQVIKSKQKEDLAREEAVLERRKL-IEAEKLQRLAEMQRKKEEAQIRREEERK 406
E+ R + L+ + K+ E+ A +EA +ERR+ IE E+ ++L +M + E + R + ++
Sbjct: 77 EV-RLQDLETERQKRAEEKAAKEAAVERRRQEIEQERQKKLQQMSENRREREKRVGKMQE 135
Query: 407 ASSAAREARAIEQLRRKEERAKAQQEE----AELLAQKLAERLNESEQRRKIYLEQIRER 462
R+A A E+ R +EER A Q++ AE L +K+ ++ ES +R + +EQIR+R
Sbjct: 136 QKERQRQALAREKARDREERLLALQQQKLATAEELQRKIIQKQQESARRHEENIEQIRQR 195
Query: 463 ANLRDQSSPLLRRSLNKDGQGRSTPNNSSDDSQTNIASGLGSSLGIGSITLQHSMKRRIK 522
A + + + RS ++ N D + + S SS G G+ + K+R+K
Sbjct: 196 A---VEMATIPSRSADE-------LRNDEQDEENSSVSERESSHGGGASMVPKINKKRLK 245
Query: 523 RIRQKLMALKYEFL 536
++RQKL E+L
Sbjct: 246 KLRQKLSTAAEEYL 259
Score = 43.1 bits (100), Expect = 0.057
Identities = 62/241 (25%), Positives = 103/241 (42%), Gaps = 46/241 (19%)
Query: 217 KSAERVRTLHDKLMSPEKKKKTTSDLK----KEAEEKHARALRI---RSELETERVQKLQ 269
K+ E D L S ++++ DL+ K AEEK A+ + R E+E ER +KLQ
Sbjct: 58 KNLEAQNKRLDFLESCKEQEVRLQDLETERQKRAEEKAAKEAAVERRRQEIEQERQKKLQ 117
Query: 270 RTSQKLNRVSEWHAVRHLKLREGMYARHQRSESRHEAFLAQVAKRAGDESSKVNEVRFIT 329
+ S+ NR + RE + Q + R LA+ R
Sbjct: 118 QMSE--NR----------REREKRVGKMQEQKERQRQALAREKAR--------------- 150
Query: 330 SLNEENKKLILRQKLHGSELRRAEKLQ-VIKSKQKEDLAREEAVLE--RRKLIEAEKL-Q 385
+ E + L L+Q+ +L AE+LQ I KQ+E R E +E R++ +E +
Sbjct: 151 --DREERLLALQQQ----KLATAEELQRKIIQKQQESARRHEENIEQIRQRAVEMATIPS 204
Query: 386 RLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERL 445
R A+ R E+ + + SS A + ++ +K R K +++ A++ L
Sbjct: 205 RSADELRNDEQDEENSSVSERESSHGGGASMVPKINKK--RLKKLRQKLSTAAEEYLSEL 262
Query: 446 N 446
N
Sbjct: 263 N 263
>gb|AAH15212.1| ZNF291 protein [Homo sapiens] gi|30582363|gb|AAP35408.1| zinc finger
protein 291 [Homo sapiens]
Length = 292
Score = 102 bits (253), Expect = 1e-19
Identities = 62/193 (32%), Positives = 100/193 (51%), Gaps = 18/193 (9%)
Query: 907 NLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN--NRSSSEP----------VATGVLKV 954
N Q L +AL T L + +L VL + S++ P VA L+
Sbjct: 55 NNRQDPTGLTAALQATDLAGVLHMLYCVLFHGTILDPSTASPKENYTQNTIQVAIQSLRF 114
Query: 955 LNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGH 1014
N+ A L L Q ++ L + H+ S +L HC+ SL+ E + +G+
Sbjct: 115 FNSFAALHLPAFQSIVGAEGLSLAFRHMASSLLGHCSQV------SCESLLHEVIVCVGY 168
Query: 1015 FALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQ 1074
F + HP NQ +++ G+ PT+L K+C LPF +FSDP L+ +L +L+AACY QNK +++
Sbjct: 169 FTVNHPDNQVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACYNNHQNKIILE 228
Query: 1075 QELSVDMLLSLLR 1087
QE+S +L + ++
Sbjct: 229 QEMSCVLLATFIQ 241
>gb|AAP36610.1| Homo sapiens zinc finger protein 291 [synthetic construct]
Length = 293
Score = 102 bits (253), Expect = 1e-19
Identities = 62/193 (32%), Positives = 100/193 (51%), Gaps = 18/193 (9%)
Query: 907 NLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN--NRSSSEP----------VATGVLKV 954
N Q L +AL T L + +L VL + S++ P VA L+
Sbjct: 55 NNRQDPTGLTAALQATDLAGVLHMLYCVLFHGTILDPSTASPKENYTQNTIQVAIQSLRF 114
Query: 955 LNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGH 1014
N+ A L L Q ++ L + H+ S +L HC+ SL+ E + +G+
Sbjct: 115 FNSFAALHLPAFQSIVGAEGLSLAFRHMASSLLGHCSQV------SCESLLHEVIVCVGY 168
Query: 1015 FALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQ 1074
F + HP NQ +++ G+ PT+L K+C LPF +FSDP L+ +L +L+AACY QNK +++
Sbjct: 169 FTVNHPDNQVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACYNNHQNKIILE 228
Query: 1075 QELSVDMLLSLLR 1087
QE+S +L + ++
Sbjct: 229 QEMSCVLLATFIQ 241
>pir||T46314 hypothetical protein DKFZp434M1511.1 - human (fragment)
gi|6808363|emb|CAB70841.1| hypothetical protein [Homo
sapiens]
Length = 266
Score = 102 bits (253), Expect = 1e-19
Identities = 62/193 (32%), Positives = 100/193 (51%), Gaps = 18/193 (9%)
Query: 907 NLAQPVVFLLSALSETGLVSLPSLLTAVLLQAN--NRSSSEP----------VATGVLKV 954
N Q L +AL T L + +L VL + S++ P VA L+
Sbjct: 29 NNRQDPTGLTAALQATDLAGVLHMLYCVLFHGTILDPSTASPKENYTQNTIQVAIQSLRF 88
Query: 955 LNNVALLDLVFLQRMLARPDLKMEIFHLMSFMLSHCASRWKTPNDQVGSLMLESLSLLGH 1014
N+ A L L Q ++ L + H+ S +L HC+ SL+ E + +G+
Sbjct: 89 FNSFAALHLPAFQSIVGAEGLSLAFRHMASSLLGHCSQV------SCESLLHEVIVCVGY 142
Query: 1015 FALFHPGNQAVLRWGKSPTILHKVCDLPFIFFSDPELMPILAGTLVAACYGCEQNKFVVQ 1074
F + HP NQ +++ G+ PT+L K+C LPF +FSDP L+ +L +L+AACY QNK +++
Sbjct: 143 FTVNHPDNQVIVQSGRHPTVLQKLCQLPFQYFSDPRLIKVLFPSLIAACYNNHQNKIILE 202
Query: 1075 QELSVDMLLSLLR 1087
QE+S +L + ++
Sbjct: 203 QEMSCVLLATFIQ 215
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.129 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,857,825,924
Number of Sequences: 2540612
Number of extensions: 76436393
Number of successful extensions: 453561
Number of sequences better than 10.0: 13808
Number of HSP's better than 10.0 without gapping: 2022
Number of HSP's successfully gapped in prelim test: 12502
Number of HSP's that attempted gapping in prelim test: 312823
Number of HSP's gapped (non-prelim): 58226
length of query: 1215
length of database: 863,360,394
effective HSP length: 140
effective length of query: 1075
effective length of database: 507,674,714
effective search space: 545750317550
effective search space used: 545750317550
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0087.21


