

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.11
(146 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
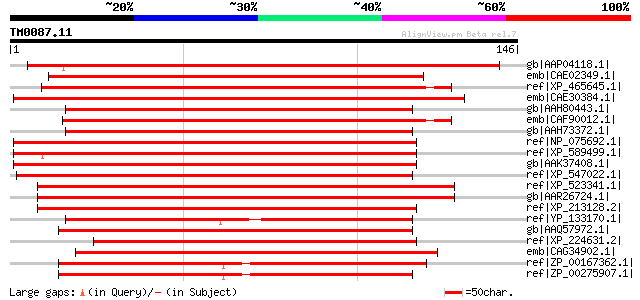
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP04118.1| unknown protein [Arabidopsis thaliana] gi|9279721... 164 4e-40
emb|CAE02349.1| OSJNBb0072M01.10 [Oryza sativa (japonica cultiva... 159 2e-38
ref|XP_465645.1| XTP3-transactivated protein A-like [Oryza sativ... 154 6e-37
emb|CAE30384.1| novel protein [Danio rerio] gi|37589724|gb|AAH59... 135 2e-31
gb|AAH80443.1| MGC89294 protein [Xenopus tropicalis] gi|56118426... 130 9e-30
emb|CAF90012.1| unnamed protein product [Tetraodon nigroviridis] 129 1e-29
gb|AAH73372.1| MGC80796 protein [Xenopus laevis] 126 1e-28
ref|NP_075692.1| hypothetical protein LOC66422 [Mus musculus] gi... 118 3e-26
ref|XP_589499.1| PREDICTED: similar to selenophosphate synthetas... 117 4e-26
gb|AAK37408.1| RS21-C6 protein [Rattus norvegicus] gi|13752373|g... 117 4e-26
ref|XP_547022.1| PREDICTED: hypothetical protein XP_547022 [Cani... 113 8e-25
ref|XP_523341.1| PREDICTED: similar to XTP3-transactivated prote... 112 1e-24
gb|AAR26724.1| XTP3-transactivated protein A [Homo sapiens] gi|1... 112 2e-24
ref|XP_213128.2| PREDICTED: similar to RS21-C6 protein [Rattus n... 108 3e-23
ref|YP_133170.1| hypothetical protein PBPRB1504 [Photobacterium ... 107 8e-23
gb|AAQ57972.1| conserved hypothetical protein [Chromobacterium v... 106 1e-22
ref|XP_224631.2| PREDICTED: similar to RS21-C6 protein [Rattus n... 105 2e-22
emb|CAG34902.1| hypothetical protein [Desulfotalea psychrophila ... 104 5e-22
ref|ZP_00167362.1| COG1694: Predicted pyrophosphatase [Ralstonia... 103 1e-21
ref|ZP_00275907.1| COG1694: Predicted pyrophosphatase [Ralstonia... 97 8e-20
>gb|AAP04118.1| unknown protein [Arabidopsis thaliana] gi|9279721|dbj|BAB01311.1|
unnamed protein product [Arabidopsis thaliana]
gi|28393817|gb|AAO42317.1| unknown protein [Arabidopsis
thaliana] gi|30687841|ref|NP_189167.2| expressed protein
[Arabidopsis thaliana]
Length = 141
Score = 164 bits (415), Expect = 4e-40
Identities = 82/139 (58%), Positives = 106/139 (75%), Gaps = 3/139 (2%)
Query: 6 SNGFDGARD---VSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGE 62
+ G +G D VSLQ LSK++ +FA+ R W++YHSPRNLLLA+VGEVGELSEIFQWKGE
Sbjct: 2 NKGEEGGEDKEVVSLQTLSKKMDDFAKARDWEKYHSPRNLLLAMVGEVGELSEIFQWKGE 61
Query: 63 VARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAIN 122
VARG P+W ++K HL EELSDVLLYLVRL+D CG+DLG+AAL KI NA KYPV +
Sbjct: 62 VARGCPDWKEEEKVHLGEELSDVLLYLVRLSDACGVDLGKAALRKIELNAIKYPVPKKTD 121
Query: 123 NGNSKTGEKTTAVVKIDNQ 141
+ GE +++ +K++ +
Sbjct: 122 DHCVGDGEDSSSKIKLNEE 140
>emb|CAE02349.1| OSJNBb0072M01.10 [Oryza sativa (japonica cultivar-group)]
gi|38345698|emb|CAE01918.2| OSJNBb0070J16.14 [Oryza
sativa (japonica cultivar-group)]
gi|50926452|ref|XP_473173.1| OSJNBb0070J16.14 [Oryza
sativa (japonica cultivar-group)]
Length = 137
Score = 159 bits (401), Expect = 2e-38
Identities = 74/108 (68%), Positives = 90/108 (82%)
Query: 12 ARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWS 71
A DVSL++LSK+L +FA+ R W+ YH+PRNLLLA++ EVGELSE+F WKGEVA+GLP W
Sbjct: 27 ASDVSLKELSKKLDDFAKERDWEMYHAPRNLLLAMIAEVGELSELFMWKGEVAKGLPGWK 86
Query: 72 CDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTS 119
+KEHL EELSDVLLYL+RL+D+CG+DLG AA KIVKNA KYP S
Sbjct: 87 ESEKEHLGEELSDVLLYLIRLSDMCGVDLGDAATRKIVKNAVKYPAPS 134
>ref|XP_465645.1| XTP3-transactivated protein A-like [Oryza sativa (japonica
cultivar-group)] gi|32352194|dbj|BAC78590.1|
hypothetical protein [Oryza sativa (japonica
cultivar-group)] gi|47848239|dbj|BAD22064.1|
XTP3-transactivated protein A-like [Oryza sativa
(japonica cultivar-group)] gi|47848144|dbj|BAD21925.1|
XTP3-transactivated protein A-like [Oryza sativa
(japonica cultivar-group)]
Length = 172
Score = 154 bits (388), Expect = 6e-37
Identities = 75/118 (63%), Positives = 93/118 (78%), Gaps = 2/118 (1%)
Query: 10 DGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPN 69
+ A V L++L +R+A+FA R W+Q+HSPRNLLLALVGEVGELSEIFQWKGEV +GLP
Sbjct: 17 EAAATVGLEELRRRMADFARERDWEQFHSPRNLLLALVGEVGELSEIFQWKGEVPKGLPG 76
Query: 70 WSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAINNGNSK 127
W +K HL EEL+DVLLYLVRL+D+CG+DLG AAL K+ NA+KYP + G+SK
Sbjct: 77 WDEAEKVHLGEELADVLLYLVRLSDMCGVDLGSAALRKLEINARKYPASQC--KGSSK 132
>emb|CAE30384.1| novel protein [Danio rerio] gi|37589724|gb|AAH59602.1| Hypothetical
protein MGC73273 [Danio rerio]
gi|41152132|ref|NP_957065.1| hypothetical protein
LOC393744 [Danio rerio]
Length = 163
Score = 135 bits (340), Expect = 2e-31
Identities = 64/130 (49%), Positives = 89/130 (68%)
Query: 2 ESSKSNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKG 61
E ++ F + +L+D+ + AEF + R W+Q+H PRNLLLALVGEVGE+SE+FQW+G
Sbjct: 33 EHEHADRFTFTSEPTLEDIRRMQAEFTDERNWNQFHQPRNLLLALVGEVGEVSELFQWRG 92
Query: 62 EVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAI 121
EVA GLP+W+ ++EHL +ELSDVL+YLV LA+ C +DL +A L K+ N KYP +
Sbjct: 93 EVAEGLPDWTEPEREHLAQELSDVLIYLVELAEKCHVDLPRAVLRKMALNRLKYPASKVH 152
Query: 122 NNGNSKTGEK 131
+ T K
Sbjct: 153 GSAKKYTEYK 162
>gb|AAH80443.1| MGC89294 protein [Xenopus tropicalis]
gi|56118426|ref|NP_001007937.1| MGC89294 protein
[Xenopus tropicalis]
Length = 123
Score = 130 bits (326), Expect = 9e-30
Identities = 60/100 (60%), Positives = 78/100 (78%)
Query: 17 LQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKE 76
++D+ + A+F R W+Q+H PRNLLLALVGEVGE++E+FQWKGEVA GLP W+ +E
Sbjct: 1 MEDIRRLQAQFTAERDWNQFHQPRNLLLALVGEVGEVAELFQWKGEVAEGLPGWTPSQRE 60
Query: 77 HLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
L ELSDVL+YL+ LA+ C +DL QAALAK+ NA+KYP
Sbjct: 61 ALSHELSDVLIYLLELAEKCHVDLPQAALAKMELNAKKYP 100
>emb|CAF90012.1| unnamed protein product [Tetraodon nigroviridis]
Length = 117
Score = 129 bits (324), Expect = 1e-29
Identities = 59/112 (52%), Positives = 84/112 (74%), Gaps = 2/112 (1%)
Query: 16 SLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDK 75
+++D+ + AEF + R W+Q+H PRNLLLA+VGEVGE++E+FQW+G+ A GLP WS D+
Sbjct: 4 TIEDIRRMQAEFTDERDWNQFHQPRNLLLAMVGEVGEVAELFQWRGDAAEGLPGWSETDR 63
Query: 76 EHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAINNGNSK 127
E+L ELSDVL+YLV LA+ C +DL QA L K+ N +KYP + +G++K
Sbjct: 64 ENLAHELSDVLIYLVELAEKCHVDLPQAVLRKMALNRRKYPASKV--HGSAK 113
>gb|AAH73372.1| MGC80796 protein [Xenopus laevis]
Length = 129
Score = 126 bits (317), Expect = 1e-28
Identities = 57/100 (57%), Positives = 77/100 (77%)
Query: 17 LQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKE 76
++D+ + ++F R W+Q+H PRNLLLALVGEVGE++E+FQWKGEVA GLP+W+ +E
Sbjct: 1 MEDIRRLQSQFTAERDWNQFHQPRNLLLALVGEVGEVAELFQWKGEVAEGLPDWTPSQRE 60
Query: 77 HLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
L ELSDVL+YL+ LA+ C +DL QA L K+ NA+KYP
Sbjct: 61 ALSHELSDVLIYLLELAEKCHVDLPQAVLTKLQLNAKKYP 100
>ref|NP_075692.1| hypothetical protein LOC66422 [Mus musculus]
gi|13435502|gb|AAH04623.1| RIKEN cDNA 2410015N17 [Mus
musculus] gi|6539656|gb|AAF15970.1| RS21-C6 [Mus
musculus] gi|12846168|dbj|BAB27056.1| unnamed protein
product [Mus musculus] gi|12846006|dbj|BAB26992.1|
unnamed protein product [Mus musculus]
gi|12834434|dbj|BAB22909.1| unnamed protein product [Mus
musculus]
Length = 170
Score = 118 bits (295), Expect = 3e-26
Identities = 58/116 (50%), Positives = 81/116 (69%)
Query: 2 ESSKSNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKG 61
+S+ + F + + +L+D+ + AEFA R W+Q+H PRNLLLALVGEVGEL+E+FQWK
Sbjct: 16 DSAAARPFRFSPEPTLEDIRRLHAEFAAERDWEQFHQPRNLLLALVGEVGELAELFQWKS 75
Query: 62 EVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPV 117
+ G W ++ L+EELSDVL+YLV LA C +DL QA ++K+ N Q+YPV
Sbjct: 76 DTEPGPQAWPPKERAALQEELSDVLIYLVALAARCHVDLPQAVISKMDTNRQRYPV 131
>ref|XP_589499.1| PREDICTED: similar to selenophosphate synthetase 2 [Bos taurus]
Length = 619
Score = 117 bits (294), Expect = 4e-26
Identities = 61/117 (52%), Positives = 82/117 (69%), Gaps = 1/117 (0%)
Query: 2 ESSKSNG-FDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWK 60
ES+ + G F + + +L+D+ + AEFA R W+Q+H PRNLLLALVGEVGEL+E+FQWK
Sbjct: 464 ESTTATGPFSFSSEPTLEDIRRLHAEFAAERDWEQFHQPRNLLLALVGEVGELAELFQWK 523
Query: 61 GEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPV 117
+ G WS ++ L+EELSDVL+YLV LA C +DL QA L K+ N ++YPV
Sbjct: 524 PDEEPGPQAWSPRERAALQEELSDVLIYLVALAARCRVDLPQAVLCKMDTNRRRYPV 580
>gb|AAK37408.1| RS21-C6 protein [Rattus norvegicus] gi|13752373|gb|AAK38638.1|
RS21-C6-like protein [Rattus norvegicus]
gi|20302079|ref|NP_620247.1| RS21-C6 protein [Rattus
norvegicus]
Length = 170
Score = 117 bits (294), Expect = 4e-26
Identities = 57/116 (49%), Positives = 81/116 (69%)
Query: 2 ESSKSNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKG 61
+S+ + F + + +L+D+ + AEFA R W+Q+H PRNLLLALVGEVGEL+E+FQWK
Sbjct: 16 DSAAAGPFSFSPEPTLEDIRRLHAEFAAERDWEQFHQPRNLLLALVGEVGELAELFQWKS 75
Query: 62 EVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPV 117
+ G W ++ L+EELSDVL+YLV LA C +DL +A ++K+ N Q+YPV
Sbjct: 76 DAEPGPQAWQPKERAALQEELSDVLIYLVALAARCHVDLPRAVISKMDTNRQRYPV 131
>ref|XP_547022.1| PREDICTED: hypothetical protein XP_547022 [Canis familiaris]
Length = 170
Score = 113 bits (283), Expect = 8e-25
Identities = 57/114 (50%), Positives = 80/114 (70%)
Query: 3 SSKSNGFDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGE 62
++ ++ F + + +L+D+ + AEFA R W Q+H PRNLLLALVGEVGEL+E+FQWK +
Sbjct: 17 TAAASPFSFSPEPTLEDIRRLHAEFAAERDWGQFHQPRNLLLALVGEVGELAELFQWKPD 76
Query: 63 VARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
G WS ++ L+EELSDVL+YLV LA C +DL QA L+K+ N ++YP
Sbjct: 77 EEPGPQAWSPRERAALQEELSDVLIYLVALAARCHVDLPQAVLSKMDLNRRRYP 130
>ref|XP_523341.1| PREDICTED: similar to XTP3-transactivated protein A [Pan
troglodytes]
Length = 170
Score = 112 bits (281), Expect = 1e-24
Identities = 58/120 (48%), Positives = 80/120 (66%)
Query: 9 FDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLP 68
F + + +L+D+ + AEFA R W+Q+H PRNLLLALVGEVGEL+E+FQWK + G
Sbjct: 23 FSFSSEPTLEDIRRLHAEFAAERDWEQFHQPRNLLLALVGEVGELAELFQWKTDGEPGPQ 82
Query: 69 NWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAINNGNSKT 128
WS ++ L+EELSDVL+YLV LA C +DL A L+K+ N ++YP A ++ T
Sbjct: 83 GWSPRERAALQEELSDVLIYLVALAARCRVDLPLAVLSKMDINRRRYPAHLARSSSRKYT 142
>gb|AAR26724.1| XTP3-transactivated protein A [Homo sapiens]
gi|13129100|ref|NP_077001.1| hypothetical protein
LOC79077 [Homo sapiens] gi|10437250|dbj|BAB15025.1|
unnamed protein product [Homo sapiens]
gi|12654993|gb|AAH01344.1| XTP3-transactivated protein A
[Homo sapiens] gi|13897521|gb|AAK48422.1| RS21C6 [Homo
sapiens] gi|13182763|gb|AAK14927.1| CDA03 [Homo sapiens]
gi|48146787|emb|CAG33616.1| XTP3TPA [Homo sapiens]
Length = 170
Score = 112 bits (280), Expect = 2e-24
Identities = 58/120 (48%), Positives = 80/120 (66%)
Query: 9 FDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLP 68
F + + +L+D+ + AEFA R W+Q+H PRNLLLALVGEVGEL+E+FQWK + G
Sbjct: 23 FSFSPEPTLEDIRRLHAEFAAERDWEQFHQPRNLLLALVGEVGELAELFQWKTDGEPGPQ 82
Query: 69 NWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAINNGNSKT 128
WS ++ L+EELSDVL+YLV LA C +DL A L+K+ N ++YP A ++ T
Sbjct: 83 GWSPRERAALQEELSDVLIYLVALAARCRVDLPLAVLSKMDINRRRYPAHLARSSSRKYT 142
>ref|XP_213128.2| PREDICTED: similar to RS21-C6 protein [Rattus norvegicus]
Length = 170
Score = 108 bits (269), Expect = 3e-23
Identities = 54/109 (49%), Positives = 75/109 (68%)
Query: 9 FDGARDVSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLP 68
F + + +L+D+ + AEFA R W+Q+H PRNLLLALVGEVG+L+E+FQ K E
Sbjct: 23 FSFSPEPTLEDIRRLHAEFAAKRDWEQFHQPRNLLLALVGEVGDLAELFQXKSEAEPDPQ 82
Query: 69 NWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPV 117
W ++ L+EELSDVL+YLV LA C +DL +A ++K+ N Q+YPV
Sbjct: 83 AWQPKERAALQEELSDVLIYLVALAARCHVDLPRAVISKMDTNRQRYPV 131
>ref|YP_133170.1| hypothetical protein PBPRB1504 [Photobacterium profundum SS9]
gi|46916605|emb|CAG23370.1| hypothetical protein
[Photobacterium profundum SS9]
Length = 116
Score = 107 bits (266), Expect = 8e-23
Identities = 51/101 (50%), Positives = 75/101 (73%), Gaps = 4/101 (3%)
Query: 17 LQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQW-KGEVARGLPNWSCDDK 75
++ L + L EFA+ R W+Q+H+P+NL++AL GE+GEL+EIFQW E + LP + +
Sbjct: 5 IKQLQRTLTEFAQERDWEQFHTPKNLVMALNGEIGELTEIFQWLTPEQSLSLPE---NKQ 61
Query: 76 EHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
EHLEEEL+DV++YL+RLAD C +D+ +A K++KN KYP
Sbjct: 62 EHLEEELADVMMYLLRLADKCEVDIIEACHKKLIKNKAKYP 102
>gb|AAQ57972.1| conserved hypothetical protein [Chromobacterium violaceum ATCC
12472] gi|34495748|ref|NP_899963.1| hypothetical protein
CV0293 [Chromobacterium violaceum ATCC 12472]
Length = 110
Score = 106 bits (264), Expect = 1e-22
Identities = 55/102 (53%), Positives = 70/102 (67%)
Query: 15 VSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDD 74
+ + L+ L FA+ RGW+++H+PRNLLLALVGEVGEL+EIFQW + A
Sbjct: 7 IDTRGLTAALQRFADERGWNRHHTPRNLLLALVGEVGELAEIFQWLDDDAAARLREDPAQ 66
Query: 75 KEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
HL+EE++DVLLYL RLA V G+DL A K+VKNA KYP
Sbjct: 67 FTHLQEEIADVLLYLTRLAMVTGVDLDAAVRDKMVKNAIKYP 108
>ref|XP_224631.2| PREDICTED: similar to RS21-C6 protein [Rattus norvegicus]
Length = 289
Score = 105 bits (262), Expect = 2e-22
Identities = 50/93 (53%), Positives = 67/93 (71%)
Query: 25 AEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSD 84
AEFA R W+Q+H PRNLLL LVGEVG+L+E+FQWK + G W ++ L++ELSD
Sbjct: 158 AEFAAERDWEQFHQPRNLLLTLVGEVGKLAELFQWKSDEEPGPQAWQPKERAALQKELSD 217
Query: 85 VLLYLVRLADVCGLDLGQAALAKIVKNAQKYPV 117
VL+YLV LA C +DL +A ++K+ N Q+YPV
Sbjct: 218 VLIYLVALAARCHVDLPRAVISKMDTNRQRYPV 250
>emb|CAG34902.1| hypothetical protein [Desulfotalea psychrophila LSv54]
gi|51244025|ref|YP_063909.1| hypothetical protein DP0173
[Desulfotalea psychrophila LSv54]
Length = 122
Score = 104 bits (259), Expect = 5e-22
Identities = 50/104 (48%), Positives = 70/104 (67%)
Query: 20 LSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLE 79
+ ++L FA+ R WDQ+HSP+NL +AL GE GEL EIFQW E N DK+
Sbjct: 9 IQEKLRVFAQERNWDQFHSPKNLAMALAGEAGELLEIFQWLTEDESKKENIKAKDKQLAA 68
Query: 80 EELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSAINN 123
EEL+D+ LYL+R+AD ++L +A AK+ KNA+KYP++ A +N
Sbjct: 69 EELADIQLYLLRIADKLDINLEEATFAKLEKNAEKYPISLAKDN 112
>ref|ZP_00167362.1| COG1694: Predicted pyrophosphatase [Ralstonia eutropha JMP134]
Length = 110
Score = 103 bits (256), Expect = 1e-21
Identities = 51/107 (47%), Positives = 73/107 (67%), Gaps = 3/107 (2%)
Query: 15 VSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWK-GEVARGLPNWSCD 73
+ L+DL + +F E RGW +YHSP+NL +AL EV EL EIFQW+ E +RG+ + D
Sbjct: 4 IDLKDLQQAAIDFGEARGWGKYHSPKNLAMALSVEVAELVEIFQWQTEEQSRGI--MATD 61
Query: 74 DKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYPVTSA 120
++EH+E+EL+D+ +YL +L G+DL A AK+ NA+KYPV A
Sbjct: 62 EREHVEQELADITIYLTQLVTALGVDLDAAVRAKMEMNAKKYPVKKA 108
>ref|ZP_00275907.1| COG1694: Predicted pyrophosphatase [Ralstonia metallidurans CH34]
Length = 107
Score = 97.1 bits (240), Expect = 8e-20
Identities = 48/103 (46%), Positives = 69/103 (66%), Gaps = 3/103 (2%)
Query: 15 VSLQDLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQWK-GEVARGLPNWSCD 73
+ + +L + A F E RGW +YHSP+NL +AL EV EL EIFQW+ E +RG+ S D
Sbjct: 4 IDISNLQQAAAAFGEARGWGKYHSPKNLAMALSVEVAELVEIFQWQTEEESRGI--MSTD 61
Query: 74 DKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNAQKYP 116
++ H+E+EL+D+ +YL +L G+DL A AK+ NA+KYP
Sbjct: 62 ERAHVEQELADITIYLTQLVTALGVDLDAAVQAKMEMNARKYP 104
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.132 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 223,710,918
Number of Sequences: 2540612
Number of extensions: 8127899
Number of successful extensions: 16319
Number of sequences better than 10.0: 162
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 71
Number of HSP's that attempted gapping in prelim test: 16172
Number of HSP's gapped (non-prelim): 163
length of query: 146
length of database: 863,360,394
effective HSP length: 122
effective length of query: 24
effective length of database: 553,405,730
effective search space: 13281737520
effective search space used: 13281737520
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0087.11


