

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079c.14
(171 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
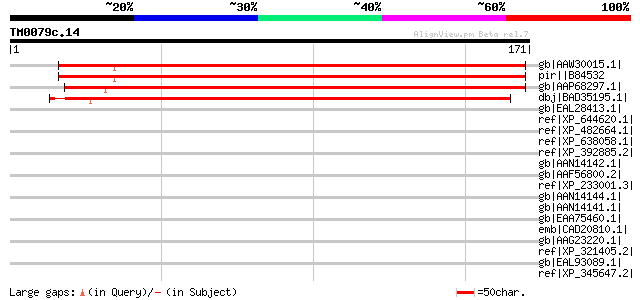
Score E
Sequences producing significant alignments: (bits) Value
gb|AAW30015.1| At2g15695 [Arabidopsis thaliana] gi|54606848|gb|A... 166 2e-40
pir||B84532 hypothetical protein At2g15690 [imported] - Arabidop... 166 2e-40
gb|AAP68297.1| At5g44250 [Arabidopsis thaliana] gi|21555160|gb|A... 155 5e-37
dbj|BAD35195.1| unknown protein [Oryza sativa (japonica cultivar... 148 5e-35
gb|EAL28413.1| GA18044-PA [Drosophila pseudoobscura] 36 0.37
ref|XP_644620.1| hypothetical protein DDB0203357 [Dictyostelium ... 36 0.48
ref|XP_482664.1| hypothetical protein [Oryza sativa (japonica cu... 35 1.1
ref|XP_638058.1| hypothetical protein DDB0186498 [Dictyostelium ... 35 1.1
ref|XP_392885.2| PREDICTED: similar to ENSANGP00000006433 [Apis ... 35 1.1
gb|AAN14142.1| CG9983-PC, isoform C [Drosophila melanogaster] gi... 33 2.4
gb|AAF56800.2| CG9983-PA, isoform A [Drosophila melanogaster] gi... 33 2.4
ref|XP_233001.3| PREDICTED: similar to Exosome component 3 [Ratt... 33 2.4
gb|AAN14144.1| CG9983-PF, isoform F [Drosophila melanogaster] gi... 33 2.4
gb|AAN14141.1| CG9983-PE, isoform E [Drosophila melanogaster] gi... 33 2.4
gb|EAA75460.1| hypothetical protein FG05224.1 [Gibberella zeae P... 33 4.1
emb|CAD20810.1| Keratin 13 [Oncorhynchus mykiss] 32 5.4
gb|AAG23220.1| glycine-rich RNA-binding protein [Sorghum bicolor] 32 5.4
ref|XP_321405.2| ENSANGP00000008532 [Anopheles gambiae str. PEST... 32 9.1
gb|EAL93089.1| cytochrome P450, putative [Aspergillus fumigatus ... 32 9.1
ref|XP_345647.2| PREDICTED: similar to growth differentiation fa... 32 9.1
>gb|AAW30015.1| At2g15695 [Arabidopsis thaliana] gi|54606848|gb|AAV34772.1|
At2g15695 [Arabidopsis thaliana]
gi|20197706|gb|AAM15216.1| unknown protein [Arabidopsis
thaliana] gi|18397898|ref|NP_565378.1| expressed protein
[Arabidopsis thaliana]
Length = 420
Score = 166 bits (420), Expect = 2e-40
Identities = 80/165 (48%), Positives = 109/165 (65%), Gaps = 11/165 (6%)
Query: 17 VRGGNIYWGRKHATDFR-----------GIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLV 65
+ GG +YWG+K + G+VVIF W S+ + L +FV+LYSSLGWNSLV
Sbjct: 2 IGGGRVYWGKKLDKEMEDAAVVDGGGSNGVVVIFVWSSINENQLMNFVDLYSSLGWNSLV 61
Query: 66 CYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRC 125
C A +L+A E + LAF ++ EL+EELK++ CPV+F AFS KAC+YKVLQ+I C
Sbjct: 62 CRADFLTAVYPEMALSLAFHLLSELVEELKSRPCPVIFLAFSGAPKACMYKVLQVIMDDC 121
Query: 126 ETPHCLHNYHLLRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
E + L+R C+SGH+YDSGPLD TSD +FALHP++ ++
Sbjct: 122 EAQIHPDDSQLVRTCLSGHVYDSGPLDFTSDLNVKFALHPTIRRM 166
>pir||B84532 hypothetical protein At2g15690 [imported] - Arabidopsis thaliana
Length = 989
Score = 166 bits (420), Expect = 2e-40
Identities = 80/165 (48%), Positives = 109/165 (65%), Gaps = 11/165 (6%)
Query: 17 VRGGNIYWGRKHATDFR-----------GIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLV 65
+ GG +YWG+K + G+VVIF W S+ + L +FV+LYSSLGWNSLV
Sbjct: 2 IGGGRVYWGKKLDKEMEDAAVVDGGGSNGVVVIFVWSSINENQLMNFVDLYSSLGWNSLV 61
Query: 66 CYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRC 125
C A +L+A E + LAF ++ EL+EELK++ CPV+F AFS KAC+YKVLQ+I C
Sbjct: 62 CRADFLTAVYPEMALSLAFHLLSELVEELKSRPCPVIFLAFSGAPKACMYKVLQVIMDDC 121
Query: 126 ETPHCLHNYHLLRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
E + L+R C+SGH+YDSGPLD TSD +FALHP++ ++
Sbjct: 122 EAQIHPDDSQLVRTCLSGHVYDSGPLDFTSDLNVKFALHPTIRRM 166
>gb|AAP68297.1| At5g44250 [Arabidopsis thaliana] gi|21555160|gb|AAM63792.1| unknown
[Arabidopsis thaliana] gi|9759526|dbj|BAB10992.1|
unnamed protein product [Arabidopsis thaliana]
gi|15241450|ref|NP_199238.1| expressed protein
[Arabidopsis thaliana] gi|15450862|gb|AAK96702.1|
Unknown protein [Arabidopsis thaliana]
Length = 403
Score = 155 bits (391), Expect = 5e-37
Identities = 76/154 (49%), Positives = 106/154 (68%), Gaps = 2/154 (1%)
Query: 19 GGNIYWGRKHAT--DFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAHYLSAFRD 76
GGN YW +K+ + IVV+FAW+S + L++ V+LYSSL W+SLVC++ +L+ F
Sbjct: 5 GGNYYWRKKNNNGGESEAIVVVFAWMSSEERNLKNHVDLYSSLLWDSLVCHSQFLNMFLP 64
Query: 77 ESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPHCLHNYHL 136
+ LA VV EL++ELK K P+VFA+FS G AC+YKVLQ+++G CET + L
Sbjct: 65 DKAADLASNVVSELVKELKAKPVPLVFASFSGGPNACMYKVLQILEGTCETGLNPDDCRL 124
Query: 137 LRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
+RNC+SG IYDS P+D TSD G R A+HP+ K+
Sbjct: 125 VRNCISGFIYDSCPVDFTSDLGARLAVHPTTLKM 158
>dbj|BAD35195.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 405
Score = 148 bits (374), Expect = 5e-35
Identities = 72/156 (46%), Positives = 95/156 (60%), Gaps = 7/156 (4%)
Query: 14 GCAVRGGNIYWG----RKHATDFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAH 69
GC GG YW A RG+VV+F WV + L+ FV LY+SLGW LVC+
Sbjct: 4 GC---GGRFYWAPAPPSPSAAGARGVVVVFGWVWSDEAQLRPFVELYASLGWRCLVCHPD 60
Query: 70 YLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPH 129
++ + E LA V+ EL++E K K P VFA+FS GSK C+YKV+QL+DG CE
Sbjct: 61 LVALYLSEKAASLASGVISELVKEFKVKPLPTVFASFSGGSKGCMYKVIQLLDGNCEGDA 120
Query: 130 CLHNYHLLRNCVSGHIYDSGPLDVTSDFGFRFALHP 165
+ +Y L+RNC+ G IYDSGP+D SD G +F +P
Sbjct: 121 TMKDYRLVRNCICGQIYDSGPVDFFSDVGTQFLQNP 156
>gb|EAL28413.1| GA18044-PA [Drosophila pseudoobscura]
Length = 772
Score = 36.2 bits (82), Expect = 0.37
Identities = 43/155 (27%), Positives = 59/155 (37%), Gaps = 22/155 (14%)
Query: 2 RGAESEGGGGGGGCAVRGGNI---YWGRK---------HATDFRGIVVIFAWV-SVPQTL 48
RGA S GGGGGGG V G +I W RK IV IF+W+ S
Sbjct: 364 RGASSGGGGGGGGGTVPGKSIGPPPWNRKGPFRCGPYFDRLPDEAIVRIFSWLDSCELCT 423
Query: 49 LQDFVNLYSSLGWNSLV--CYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAF 106
+ + + W ++ C + T+ + F +L + +CP V
Sbjct: 424 VARVCRRFEQVAWRPVLWKCITLRGEHLNGDKTLKMIF---RQLCGQSCNGACPEVERVM 480
Query: 107 SAGSKACLYKVLQLIDGRCETPHCLHNYHLLRNCV 141
A K LQL+ RC P H L+ CV
Sbjct: 481 LADGCRISDKGLQLLTRRC--PELTHLQ--LQTCV 511
>ref|XP_644620.1| hypothetical protein DDB0203357 [Dictyostelium discoideum]
gi|66822551|ref|XP_644630.1| hypothetical protein
DDB0217163 [Dictyostelium discoideum]
gi|60472754|gb|EAL70704.1| hypothetical protein
DDB0217163 [Dictyostelium discoideum]
gi|60472736|gb|EAL70686.1| hypothetical protein
DDB0203357 [Dictyostelium discoideum]
Length = 354
Score = 35.8 bits (81), Expect = 0.48
Identities = 26/126 (20%), Positives = 56/126 (43%), Gaps = 8/126 (6%)
Query: 35 IVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEEL 94
+ +I W+ L + LY G+ ++ YL F + LA+ ++ L+ E
Sbjct: 91 MAIIVGWIKSNPKHLNKYSKLYLDNGFVTISFSPSYLCHFFPKKMKELAYNFLEFLVSEN 150
Query: 95 KTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPHCLHNYHLLRNCVSGHIYDSGPLDVT 154
+ + P++F FS G+ ++ +L++ + + L + G I+DS P ++
Sbjct: 151 EKIARPIIFQVFS-GNMVFQSEIFKLLNEEIK-------FKKLIPFIKGQIFDSCPSKIS 202
Query: 155 SDFGFR 160
+ F+
Sbjct: 203 EEQAFQ 208
>ref|XP_482664.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|42409449|dbj|BAD09806.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|42408340|dbj|BAD09493.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 330
Score = 34.7 bits (78), Expect = 1.1
Identities = 13/21 (61%), Positives = 15/21 (70%)
Query: 4 AESEGGGGGGGCAVRGGNIYW 24
A S+GGGGGGGC GG Y+
Sbjct: 47 ASSDGGGGGGGCRSNGGRFYF 67
>ref|XP_638058.1| hypothetical protein DDB0186498 [Dictyostelium discoideum]
gi|60466509|gb|EAL64561.1| hypothetical protein
DDB0186498 [Dictyostelium discoideum]
Length = 305
Score = 34.7 bits (78), Expect = 1.1
Identities = 27/120 (22%), Positives = 52/120 (42%), Gaps = 9/120 (7%)
Query: 33 RGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAHYLSAFRDESTVPLAFCVVDELIE 92
R I ++ W+ Q LL ++NL++S G+N+ A Y A ++ ++
Sbjct: 60 RPISIVLGWMGSTQKLLLKYINLWTSRGFNTFSYRADYFETLAILGLRLKAMHMLKQIST 119
Query: 93 ELKTK-SC-PVVFAAFSAGSKACLYKVLQLIDGRCETPHCLHNYHLLRNCVSGHIYDSGP 150
LK + +C ++F FS G + +++ + E Y + + G + DS P
Sbjct: 120 YLKERPNCDTIIFHIFSNGGGFLYWALIEFMLANDE-------YKFIHPMIKGVVMDSLP 172
>ref|XP_392885.2| PREDICTED: similar to ENSANGP00000006433 [Apis mellifera]
Length = 1685
Score = 34.7 bits (78), Expect = 1.1
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1137 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1190
>gb|AAN14142.1| CG9983-PC, isoform C [Drosophila melanogaster]
gi|7301687|gb|AAF56801.1| CG9983-PB, isoform B
[Drosophila melanogaster] gi|24650836|ref|NP_733251.1|
CG9983-PC, isoform C [Drosophila melanogaster]
gi|17738267|ref|NP_524543.1| CG9983-PB, isoform B
[Drosophila melanogaster] gi|133253|sp|P07909|ROA1_DROME
Heterogeneous nuclear ribonucleoprotein A1 (hnRNP core
protein A1-A) (PEN repeat clone P9)
gi|908757|gb|AAA70426.1| unknown protein
gi|157654|gb|AAA28624.1| nulcear ribonucleoprotein
Length = 365
Score = 33.5 bits (75), Expect = 2.4
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 257 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 285
>gb|AAF56800.2| CG9983-PA, isoform A [Drosophila melanogaster]
gi|24650831|ref|NP_733249.1| CG9983-PA, isoform A
[Drosophila melanogaster] gi|157652|gb|AAA28622.1|
nuclear ribonucleoprotein
Length = 364
Score = 33.5 bits (75), Expect = 2.4
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 256 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 284
>ref|XP_233001.3| PREDICTED: similar to Exosome component 3 [Rattus norvegicus]
Length = 287
Score = 33.5 bits (75), Expect = 2.4
Identities = 13/24 (54%), Positives = 15/24 (62%)
Query: 1 MRGAESEGGGGGGGCAVRGGNIYW 24
+R E GGGGGGG GG +YW
Sbjct: 90 LRHKEPSGGGGGGGGGGGGGGVYW 113
>gb|AAN14144.1| CG9983-PF, isoform F [Drosophila melanogaster]
gi|23172512|gb|AAN14143.1| CG9983-PD, isoform D
[Drosophila melanogaster] gi|24650840|ref|NP_733253.1|
CG9983-PF, isoform F [Drosophila melanogaster]
gi|24650838|ref|NP_733252.1| CG9983-PD, isoform D
[Drosophila melanogaster] gi|16769554|gb|AAL28996.1|
LD38464p [Drosophila melanogaster]
gi|157651|gb|AAA28621.1| nuclear ribonucleoprotein
Length = 361
Score = 33.5 bits (75), Expect = 2.4
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 253 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 281
>gb|AAN14141.1| CG9983-PE, isoform E [Drosophila melanogaster]
gi|24650833|ref|NP_733250.1| CG9983-PE, isoform E
[Drosophila melanogaster] gi|157653|gb|AAA28623.1|
nuclear ribonucleoprotein
Length = 360
Score = 33.5 bits (75), Expect = 2.4
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 252 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 280
>gb|EAA75460.1| hypothetical protein FG05224.1 [Gibberella zeae PH-1]
gi|46121691|ref|XP_385400.1| hypothetical protein
FG05224.1 [Gibberella zeae PH-1]
Length = 657
Score = 32.7 bits (73), Expect = 4.1
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 4/35 (11%)
Query: 2 RGAESEGGGGGGGCA----VRGGNIYWGRKHATDF 32
+G + EGGGGGGG + + GG IY+G A +F
Sbjct: 11 QGEQREGGGGGGGFSFNKLILGGAIYFGLNAAMNF 45
>emb|CAD20810.1| Keratin 13 [Oncorhynchus mykiss]
Length = 492
Score = 32.3 bits (72), Expect = 5.4
Identities = 13/20 (65%), Positives = 16/20 (80%)
Query: 6 SEGGGGGGGCAVRGGNIYWG 25
S GGGGGGG A+R G++Y G
Sbjct: 41 SMGGGGGGGGAMRSGSVYGG 60
>gb|AAG23220.1| glycine-rich RNA-binding protein [Sorghum bicolor]
Length = 170
Score = 32.3 bits (72), Expect = 5.4
Identities = 15/31 (48%), Positives = 18/31 (57%)
Query: 4 AESEGGGGGGGCAVRGGNIYWGRKHATDFRG 34
A+S GGGGGGG GG Y GR+ + G
Sbjct: 84 AQSRGGGGGGGGYGGGGGGYGGRREGGGYGG 114
>ref|XP_321405.2| ENSANGP00000008532 [Anopheles gambiae str. PEST]
gi|55233667|gb|EAA00899.2| ENSANGP00000008532
[Anopheles gambiae str. PEST]
Length = 929
Score = 31.6 bits (70), Expect = 9.1
Identities = 24/84 (28%), Positives = 39/84 (45%), Gaps = 2/84 (2%)
Query: 1 MRGAESEGGGGGGGCAVRGGNIYWGRKHATDFRGI-VVIFAWVSVPQTLLQDFVNLYSSL 59
M G +S GG GGG + G +W KH +R + +V+F +S+P V+ S++
Sbjct: 1 MTGYDSLGGNGGGPSSTGGTVCHW-LKHMKLYRVVLIVVFCLLSLPFVFYNLLVHEESNV 59
Query: 60 GWNSLVCYAHYLSAFRDESTVPLA 83
+ + L F D S + A
Sbjct: 60 QHSDIHRTRSQLFTFEDVSPLKAA 83
>gb|EAL93089.1| cytochrome P450, putative [Aspergillus fumigatus Af293]
Length = 525
Score = 31.6 bits (70), Expect = 9.1
Identities = 19/64 (29%), Positives = 31/64 (47%)
Query: 71 LSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPHC 130
L A RDES P A + E + K+ F AFSAG+++C+ + + ++
Sbjct: 429 LVAHRDESVFPQADQYIPERWLGEEGKALQPYFVAFSAGARSCIGRNISYLEQTKAIATL 488
Query: 131 LHNY 134
+H Y
Sbjct: 489 VHRY 492
>ref|XP_345647.2| PREDICTED: similar to growth differentiation factor 7 [Rattus
norvegicus]
Length = 544
Score = 31.6 bits (70), Expect = 9.1
Identities = 15/27 (55%), Positives = 16/27 (58%)
Query: 2 RGAESEGGGGGGGCAVRGGNIYWGRKH 28
RGA+ GGGGGGG GG GR H
Sbjct: 409 RGAQGSGGGGGGGGGGGGGGGGAGRGH 435
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.141 0.461
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 321,786,635
Number of Sequences: 2540612
Number of extensions: 13560332
Number of successful extensions: 93647
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 93565
Number of HSP's gapped (non-prelim): 39
length of query: 171
length of database: 863,360,394
effective HSP length: 119
effective length of query: 52
effective length of database: 561,027,566
effective search space: 29173433432
effective search space used: 29173433432
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0079c.14


