

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079a.1
(139 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
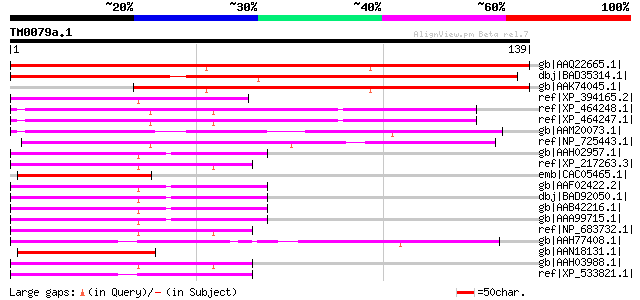
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ22665.1| At1g49590 [Arabidopsis thaliana] gi|15222782|ref|... 164 4e-40
dbj|BAD35314.1| formin-binding protein-related-like [Oryza sativ... 154 3e-37
gb|AAK74045.1| At1g49590/F14J22_9 [Arabidopsis thaliana] 114 5e-25
ref|XP_394165.2| PREDICTED: similar to RNA-binding protein 5 (RN... 52 3e-06
ref|XP_464248.1| putative RNA-binding protein 10 [Oryza sativa (... 50 2e-05
ref|XP_464247.1| putative RNA-binding protein 10 [Oryza sativa (... 50 2e-05
gb|AAM20073.1| unknown protein [Arabidopsis thaliana] gi|1797913... 49 3e-05
ref|NP_725443.1| CG8079-PB, isoform B [Drosophila melanogaster] ... 48 6e-05
gb|AAH02957.1| RBM5 protein [Homo sapiens] 47 1e-04
ref|XP_217263.3| PREDICTED: similar to RNA binding motif protein... 47 1e-04
emb|CAC05465.1| putative protein [Arabidopsis thaliana] 47 1e-04
gb|AAF02422.2| lung cancer tumor suppressor H37 [Homo sapiens] g... 47 1e-04
dbj|BAD92050.1| RNA binding motif protein 5 variant [Homo sapiens] 47 1e-04
gb|AAB42216.1| partial CDS, human putative tumor suppressor (U23... 47 1e-04
gb|AAA99715.1| putative tumor suppressor 47 1e-04
ref|NP_683732.1| RNA binding motif protein 5 [Mus musculus] gi|5... 47 1e-04
gb|AAH77408.1| Rbm5-prov protein [Xenopus laevis] 47 1e-04
gb|AAN18131.1| At5g09390/T5E8_190 [Arabidopsis thaliana] gi|2155... 47 1e-04
gb|AAH03988.1| Rbm5 protein [Mus musculus] 47 1e-04
ref|XP_533821.1| PREDICTED: similar to RNA-binding protein 5 (RN... 46 2e-04
>gb|AAQ22665.1| At1g49590 [Arabidopsis thaliana] gi|15222782|ref|NP_175382.1|
formin-binding protein-related [Arabidopsis thaliana]
Length = 242
Score = 164 bits (415), Expect = 4e-40
Identities = 87/142 (61%), Positives = 105/142 (73%), Gaps = 3/142 (2%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTK-KASSTSQS 59
W +S+SGYYY++TNG +YD +SGFYYSD+IG WVTQ+EAYA+ +S TK S
Sbjct: 100 WMLDSASGYYYNQTNGLHYDSQSGFYYSDSIGHWVTQDEAYAAVKTSSGTKVPLVKKPVS 159
Query: 60 NKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLG--KRKRPGEKPKVVSEEEKSALKA 117
+ G P G PGR+VT SLNP R VK A SS+ LG KRKR EKPK VS EEK+ALKA
Sbjct: 160 SSGAGPSVGKPPGRLVTASLNPKRAVKGAASSVDLGNNKRKRQDEKPKKVSAEEKAALKA 219
Query: 118 REAARKRVQEREKSLLGLYSKP 139
REAARKRV++REK LLGLY++P
Sbjct: 220 REAARKRVEDREKPLLGLYNRP 241
>dbj|BAD35314.1| formin-binding protein-related-like [Oryza sativa (japonica
cultivar-group)]
Length = 243
Score = 154 bits (390), Expect = 3e-37
Identities = 78/146 (53%), Positives = 101/146 (68%), Gaps = 14/146 (9%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSN 60
W+F+S+SGYYY K+ G Y+D SGFYYSD +GKWVTQEEAYA + T +A++ S+
Sbjct: 99 WEFDSTSGYYYDKSTGLYFDSNSGFYYSDGLGKWVTQEEAYAW----AKTSQANAGQSSS 154
Query: 61 KGNKP----------HNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEE 110
KP G PG VV LNP R VK APS++++ KRKR KPKV+S+E
Sbjct: 155 SQTKPTASVATVPTIKGGQAPGLVVKKPLNPMRTVKGAPSAIAVNKRKREDGKPKVISKE 214
Query: 111 EKSALKAREAARKRVQEREKSLLGLY 136
E++ALKAREAARKR+++REK L+GLY
Sbjct: 215 EEAALKAREAARKRMEDREKPLMGLY 240
>gb|AAK74045.1| At1g49590/F14J22_9 [Arabidopsis thaliana]
Length = 111
Score = 114 bits (285), Expect = 5e-25
Identities = 66/109 (60%), Positives = 77/109 (70%), Gaps = 3/109 (2%)
Query: 34 WVTQEEAYASPHFASTTK-KASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSL 92
WVTQ+EAYA+ +S TK S+ G P G PGR+VT SLNP R VK A SS+
Sbjct: 2 WVTQDEAYAAVKTSSGTKVPLVKKPVSSSGAGPSVGKPPGRLVTASLNPKRAVKGAASSV 61
Query: 93 SLG--KRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSKP 139
LG KRKR EKPK VS EEK+ALKAREAARKRV++REK LLGLY++P
Sbjct: 62 DLGNNKRKRQDEKPKKVSAEEKAALKAREAARKRVEDREKPLLGLYNRP 110
>ref|XP_394165.2| PREDICTED: similar to RNA-binding protein 5 (RNA binding motif
protein 5) (Putative tumor suppressor LUCA15) (G15
protein) [Apis mellifera]
Length = 888
Score = 52.0 bits (123), Expect = 3e-06
Identities = 24/67 (35%), Positives = 37/67 (54%), Gaps = 3/67 (4%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY + G YYDP S +YY+ + W + +Y A+T +A+ TS
Sbjct: 538 YHYDESSGYYYDPSTGLYYDPNSQYYYNSHTQQFLYWDAESFSYQPAKTATTVAQATGTS 597
Query: 58 QSNKGNK 64
SN+ K
Sbjct: 598 TSNENKK 604
>ref|XP_464248.1| putative RNA-binding protein 10 [Oryza sativa (japonica
cultivar-group)] gi|49387753|dbj|BAD26241.1| putative
RNA-binding protein 10 [Oryza sativa (japonica
cultivar-group)]
Length = 889
Score = 49.7 bits (117), Expect = 2e-05
Identities = 38/135 (28%), Positives = 59/135 (43%), Gaps = 13/135 (9%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVT---QEEAYASPHFASTTKKA---- 53
WD SGYYY +GFYYD +G YY G W + Q + Y +++K A
Sbjct: 472 WD--EKSGYYYDSASGFYYDGNTGLYYDGNAGVWYSYDQQTQQYVPCSEQNSSKAAGDMA 529
Query: 54 ---SSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEE 110
+ TS+S+ N + P + + V+AA +S +L K+ E+ K +
Sbjct: 530 NTSTKTSESSGKNVVISAPAATIKQSEKTSLPEAVQAA-ASAALAAEKKEKERAKEIKLA 588
Query: 111 EKSALKAREAARKRV 125
K +L A + V
Sbjct: 589 SKGSLLANKKKMNNV 603
Score = 40.0 bits (92), Expect = 0.012
Identities = 27/121 (22%), Positives = 53/121 (43%), Gaps = 4/121 (3%)
Query: 7 SGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPH 66
SG+ + + +G+YYD SGFYY G + + A ++ + S + +
Sbjct: 468 SGFVWDEKSGYYYDSASGFYYDGNTGLYY---DGNAGVWYSYDQQTQQYVPCSEQNSSKA 524
Query: 67 NGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQ 126
G + TS + +NV + + ++ K+ P+ V +AL A + ++R +
Sbjct: 525 AGDMANTSTKTSESSGKNVVISAPAATI-KQSEKTSLPEAVQAAASAALAAEKKEKERAK 583
Query: 127 E 127
E
Sbjct: 584 E 584
>ref|XP_464247.1| putative RNA-binding protein 10 [Oryza sativa (japonica
cultivar-group)] gi|49387752|dbj|BAD26240.1| putative
RNA-binding protein 10 [Oryza sativa (japonica
cultivar-group)]
Length = 928
Score = 49.7 bits (117), Expect = 2e-05
Identities = 38/135 (28%), Positives = 59/135 (43%), Gaps = 13/135 (9%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVT---QEEAYASPHFASTTKKA---- 53
WD SGYYY +GFYYD +G YY G W + Q + Y +++K A
Sbjct: 472 WD--EKSGYYYDSASGFYYDGNTGLYYDGNAGVWYSYDQQTQQYVPCSEQNSSKAAGDMA 529
Query: 54 ---SSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEE 110
+ TS+S+ N + P + + V+AA +S +L K+ E+ K +
Sbjct: 530 NTSTKTSESSGKNVVISAPAATIKQSEKTSLPEAVQAA-ASAALAAEKKEKERAKEIKLA 588
Query: 111 EKSALKAREAARKRV 125
K +L A + V
Sbjct: 589 SKGSLLANKKKMNNV 603
Score = 40.0 bits (92), Expect = 0.012
Identities = 27/121 (22%), Positives = 53/121 (43%), Gaps = 4/121 (3%)
Query: 7 SGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPH 66
SG+ + + +G+YYD SGFYY G + + A ++ + S + +
Sbjct: 468 SGFVWDEKSGYYYDSASGFYYDGNTGLYY---DGNAGVWYSYDQQTQQYVPCSEQNSSKA 524
Query: 67 NGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQ 126
G + TS + +NV + + ++ K+ P+ V +AL A + ++R +
Sbjct: 525 AGDMANTSTKTSESSGKNVVISAPAATI-KQSEKTSLPEAVQAAASAALAAEKKEKERAK 583
Query: 127 E 127
E
Sbjct: 584 E 584
>gb|AAM20073.1| unknown protein [Arabidopsis thaliana] gi|17979131|gb|AAL49823.1|
unknown protein [Arabidopsis thaliana]
Length = 1007
Score = 48.9 bits (115), Expect = 3e-05
Identities = 34/133 (25%), Positives = 56/133 (41%), Gaps = 21/133 (15%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSN 60
WD +SGYYY +G+YYD SG YY G W + ++ T + Q+N
Sbjct: 586 WD--EASGYYYDAASGYYYDGNSGLYYDSNSGLWYSYDQ--------QTQQYVPCPDQNN 635
Query: 61 KGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG-EKPKVVSEEEKSALKARE 119
+ N P + + K++ + + P EK + + ++A A
Sbjct: 636 ESKVTENQP----------DSAKKEKSSQQKVIISAATTPNVEKVLSLPDAVQAAAAAAI 685
Query: 120 AARKRVQEREKSL 132
A+ KR +ER K +
Sbjct: 686 ASEKREKERVKEI 698
>ref|NP_725443.1| CG8079-PB, isoform B [Drosophila melanogaster]
gi|21627145|gb|AAM68526.1| CG8079-PB, isoform B
[Drosophila melanogaster]
Length = 599
Score = 47.8 bits (112), Expect = 6e-05
Identities = 35/135 (25%), Positives = 61/135 (44%), Gaps = 13/135 (9%)
Query: 4 ESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVT-QEEAYASPHFASTTKKASSTSQSNKG 62
E+ + + Y T+G YYDPK+G+YY+ G + Y S A + + S +Q+N
Sbjct: 126 ENLNNFVYEHTSGMYYDPKTGYYYNAEYGLYYDGNNGCYYSYDHAKDSYEFHSQAQANDA 185
Query: 63 NKPHNGPLPGRV-------VTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSAL 115
KP + V V T + +KA K K EK K ++++KS
Sbjct: 186 AKPESEDEDLEVQFDELGGVITDHETLKKIKAEKQ-----KAKDQAEKSKRKAKKKKSKK 240
Query: 116 KAREAARKRVQEREK 130
+++ ++K + + K
Sbjct: 241 HSKKRSKKERRHKSK 255
>gb|AAH02957.1| RBM5 protein [Homo sapiens]
Length = 546
Score = 47.0 bits (110), Expect = 1e-04
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 4/72 (5%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y P S++ + S
Sbjct: 462 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYV-PAAESSSHQQSGLP 520
Query: 58 QSNKGNKPHNGP 69
+ +G + P
Sbjct: 521 PAKEGKEKKEKP 532
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 3/63 (4%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 459 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSHQQSG 518
Query: 63 NKP 65
P
Sbjct: 519 LPP 521
>ref|XP_217263.3| PREDICTED: similar to RNA binding motif protein 5 [Rattus
norvegicus]
Length = 815
Score = 47.0 bits (110), Expect = 1e-04
Identities = 24/71 (33%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKA---S 54
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y +S+ ++A S
Sbjct: 462 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSNQQAGLPS 521
Query: 55 STSQSNKGNKP 65
+ K KP
Sbjct: 522 TKEGKEKKEKP 532
Score = 34.7 bits (78), Expect = 0.51
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 3/60 (5%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 459 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSNQQAG 518
>emb|CAC05465.1| putative protein [Arabidopsis thaliana]
Length = 354
Score = 47.0 bits (110), Expect = 1e-04
Identities = 18/36 (50%), Positives = 23/36 (63%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQE 38
F+ SSGYYY + G+YYDP +G Y GKW +E
Sbjct: 303 FDESSGYYYSSSLGYYYDPNTGLYCYATTGKWYDEE 338
>gb|AAF02422.2| lung cancer tumor suppressor H37 [Homo sapiens]
gi|4140647|gb|AAD04159.1| RNA binding motif protein 5
[Homo sapiens] gi|13124794|sp|P52756|RBM5_HUMAN
RNA-binding protein 5 (RNA binding motif protein 5)
(Putative tumor suppressor LUCA15) (G15 protein)
gi|5032031|ref|NP_005769.1| RNA binding motif protein 5
[Homo sapiens]
Length = 815
Score = 47.0 bits (110), Expect = 1e-04
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 4/72 (5%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y P S++ + S
Sbjct: 462 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYV-PAAESSSHQQSGLP 520
Query: 58 QSNKGNKPHNGP 69
+ +G + P
Sbjct: 521 PAKEGKEKKEKP 532
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 3/63 (4%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 459 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSHQQSG 518
Query: 63 NKP 65
P
Sbjct: 519 LPP 521
>dbj|BAD92050.1| RNA binding motif protein 5 variant [Homo sapiens]
Length = 505
Score = 47.0 bits (110), Expect = 1e-04
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 4/72 (5%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y P S++ + S
Sbjct: 152 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYV-PAAESSSHQQSGLP 210
Query: 58 QSNKGNKPHNGP 69
+ +G + P
Sbjct: 211 PAKEGKEKKEKP 222
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 3/63 (4%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 149 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSHQQSG 208
Query: 63 NKP 65
P
Sbjct: 209 LPP 211
>gb|AAB42216.1| partial CDS, human putative tumor suppressor (U23946) [Homo
sapiens]
Length = 698
Score = 47.0 bits (110), Expect = 1e-04
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 4/72 (5%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y P S++ + S
Sbjct: 345 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYV-PAAESSSHQQSGLP 403
Query: 58 QSNKGNKPHNGP 69
+ +G + P
Sbjct: 404 PAKEGKEKKEKP 415
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 3/63 (4%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 342 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSHQQSG 401
Query: 63 NKP 65
P
Sbjct: 402 LPP 404
>gb|AAA99715.1| putative tumor suppressor
Length = 815
Score = 47.0 bits (110), Expect = 1e-04
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 4/72 (5%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKASSTS 57
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y P S++ + S
Sbjct: 462 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYV-PAAESSSHQQSGLP 520
Query: 58 QSNKGNKPHNGP 69
+ +G + P
Sbjct: 521 PAKEGKEKKEKP 532
Score = 36.2 bits (82), Expect = 0.17
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 3/63 (4%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A S+S G
Sbjct: 459 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAESSSHQQSG 518
Query: 63 NKP 65
P
Sbjct: 519 LPP 521
>ref|NP_683732.1| RNA binding motif protein 5 [Mus musculus]
gi|55777254|gb|AAH43058.1| RNA binding motif protein 5
[Mus musculus] gi|54611282|gb|AAH31899.1| RNA binding
motif protein 5 [Mus musculus]
gi|15528488|emb|CAC69136.1| RNA binding motif protein 5
[Mus musculus] gi|54611437|gb|AAH23854.1| RNA binding
motif protein 5 [Mus musculus]
gi|32451706|gb|AAH54729.1| RNA binding motif protein 5
[Mus musculus]
Length = 815
Score = 46.6 bits (109), Expect = 1e-04
Identities = 24/71 (33%), Positives = 36/71 (49%), Gaps = 6/71 (8%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKA---S 54
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y AS+ ++ S
Sbjct: 462 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAEASSNQQTGLPS 521
Query: 55 STSQSNKGNKP 65
+ K KP
Sbjct: 522 TKEGKEKKEKP 532
Score = 33.5 bits (75), Expect = 1.1
Identities = 18/60 (30%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A ++S G
Sbjct: 459 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAEASSNQQTG 518
>gb|AAH77408.1| Rbm5-prov protein [Xenopus laevis]
Length = 749
Score = 46.6 bits (109), Expect = 1e-04
Identities = 36/132 (27%), Positives = 63/132 (47%), Gaps = 14/132 (10%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSN 60
+ ++ SSGYYY G YYDP S +YY+ +TQ+ Y + A T QS
Sbjct: 475 YQYDESSGYYYDPQTGLYYDPNSQYYYNS-----LTQQYLYWDGEKQTYLPAADGTGQS- 528
Query: 61 KGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEK-PKVVSEEEKSALKARE 119
G +P NG PG + + K P S + + + E+ K +++++++ + +
Sbjct: 529 -GAQP-NGANPG-----TSKEGKEKKEKPKSKTAQQIAKDMERWAKSLNKQKENFKNSFQ 581
Query: 120 AARKRVQEREKS 131
R +ER++S
Sbjct: 582 PLSSRDEERKES 593
>gb|AAN18131.1| At5g09390/T5E8_190 [Arabidopsis thaliana]
gi|21553355|gb|AAM62448.1| unknown [Arabidopsis
thaliana] gi|21703117|gb|AAM74500.1| AT5g09390/T5E8_190
[Arabidopsis thaliana] gi|18415962|ref|NP_568211.1|
CD2-binding protein-related [Arabidopsis thaliana]
Length = 351
Score = 46.6 bits (109), Expect = 1e-04
Identities = 18/37 (48%), Positives = 23/37 (61%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEE 39
F+ SSGYYY + G+YYDP +G Y GKW +E
Sbjct: 298 FDESSGYYYSSSLGYYYDPNTGLYCYATTGKWYRYDE 334
>gb|AAH03988.1| Rbm5 protein [Mus musculus]
Length = 520
Score = 46.6 bits (109), Expect = 1e-04
Identities = 24/71 (33%), Positives = 36/71 (49%), Gaps = 6/71 (8%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGK---WVTQEEAYASPHFASTTKKA---S 54
+ ++ SSGYYY T G YYDP S +YY+ + W ++E Y AS+ ++ S
Sbjct: 167 YQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAEASSNQQTGLPS 226
Query: 55 STSQSNKGNKP 65
+ K KP
Sbjct: 227 TKEGKEKKEKP 237
Score = 33.5 bits (75), Expect = 1.1
Identities = 18/60 (30%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Query: 6 SSGYYYHKTNGFYYDPKSGFYY---SDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKG 62
+S Y Y +++G+YYDP +G YY S +TQ+ Y + A ++S G
Sbjct: 164 TSTYQYDESSGYYYDPTTGLYYDPNSQYYYNSLTQQYLYWDGEKETYVPAAEASSNQQTG 223
>ref|XP_533821.1| PREDICTED: similar to RNA-binding protein 5 (RNA binding motif
protein 5) (Putative tumor suppressor LUCA15) (G15
protein) [Canis familiaris]
Length = 1253
Score = 46.2 bits (108), Expect = 2e-04
Identities = 23/65 (35%), Positives = 32/65 (48%), Gaps = 5/65 (7%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSN 60
+ ++ SSGYYY T G YYDP S +YY+ +TQ+ Y + A S+S
Sbjct: 900 YQYDESSGYYYDPTTGLYYDPNSQYYYNS-----LTQQYLYWDGEKETYVPAAESSSHQQ 954
Query: 61 KGNKP 65
G P
Sbjct: 955 TGLPP 959
Score = 36.2 bits (82), Expect = 0.17
Identities = 22/65 (33%), Positives = 28/65 (42%), Gaps = 10/65 (15%)
Query: 5 SSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAY-----ASPHFASTTKKASSTSQS 59
SS Y Y G+YYDP +G YY TQ+E Y SP +K SS+
Sbjct: 125 SSDCYIYDSATGYYYDPLAGTYYDPN-----TQQEVYVPQDPGSPEEEEIKEKKSSSQGK 179
Query: 60 NKGNK 64
+ K
Sbjct: 180 SSSKK 184
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.306 0.125 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 249,207,635
Number of Sequences: 2540612
Number of extensions: 9833383
Number of successful extensions: 45413
Number of sequences better than 10.0: 407
Number of HSP's better than 10.0 without gapping: 164
Number of HSP's successfully gapped in prelim test: 244
Number of HSP's that attempted gapping in prelim test: 44716
Number of HSP's gapped (non-prelim): 780
length of query: 139
length of database: 863,360,394
effective HSP length: 115
effective length of query: 24
effective length of database: 571,190,014
effective search space: 13708560336
effective search space used: 13708560336
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0079a.1


