

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.3
(267 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
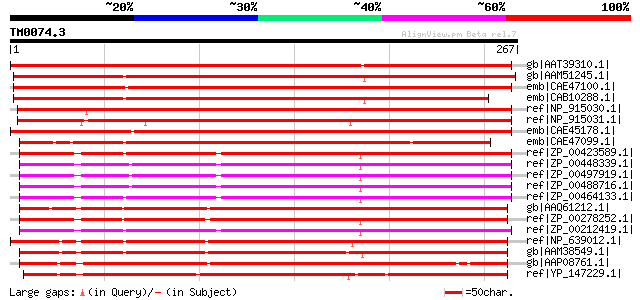
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT39310.1| putative dioxygenase [Solanum demissum] 387 e-106
gb|AAM51245.1| unknown protein [Arabidopsis thaliana] gi|1529302... 376 e-103
emb|CAE47100.1| 4,5-DOPA dioxygenase extradiol [Beta vulgaris] 367 e-100
emb|CAB10288.1| hypothetical protein [Arabidopsis thaliana] gi|7... 357 2e-97
ref|NP_915030.1| P0471B04.17 [Oryza sativa (japonica cultivar-gr... 353 2e-96
ref|NP_915031.1| P0471B04.18 [Oryza sativa (japonica cultivar-gr... 335 1e-90
emb|CAE45178.1| 4,5-DOPA dioxygenase extradiol [Portulaca grandi... 327 3e-88
emb|CAE47099.1| 4,5 dioxygenase extradiol [Physcomitrella patens] 287 2e-76
ref|ZP_00423589.1| Catalytic LigB subunit of aromatic ring-openi... 218 1e-55
ref|ZP_00448339.1| COG3384: Uncharacterized conserved protein [B... 217 2e-55
ref|ZP_00497919.1| COG3384: Uncharacterized conserved protein [B... 217 3e-55
ref|ZP_00488716.1| COG3384: Uncharacterized conserved protein [B... 216 4e-55
ref|ZP_00464133.1| Catalytic LigB subunit of aromatic ring-openi... 214 3e-54
gb|AAQ61212.1| conserved hypothetical protein [Chromobacterium v... 212 8e-54
ref|ZP_00278252.1| COG3384: Uncharacterized conserved protein [B... 211 1e-53
ref|ZP_00212419.1| COG3384: Uncharacterized conserved protein [B... 211 2e-53
ref|NP_639012.1| hypothetical protein XCC3666 [Xanthomonas campe... 203 5e-51
gb|AAM38549.1| conserved hypothetical protein [Xanthomonas axono... 202 1e-50
gb|AAP08761.1| hypothetical protein [Bacillus cereus ATCC 14579]... 200 4e-50
ref|YP_147229.1| hypothetical protein GK1376 [Geobacillus kausto... 200 4e-50
>gb|AAT39310.1| putative dioxygenase [Solanum demissum]
Length = 306
Score = 387 bits (995), Expect = e-106
Identities = 186/265 (70%), Positives = 219/265 (82%), Gaps = 2/265 (0%)
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKE-VFPPRPTSILVISGHWDTAVPTVNV 59
+ +K+TF+ISHGSPTLSIDESL AR FL+S+K+ + +P SILVIS HW+T+ PTVN
Sbjct: 38 LPVKETFFISHGSPTLSIDESLPARNFLKSFKERFMINQKPNSILVISAHWETSEPTVNS 97
Query: 60 VDSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAW 119
+ DTI+DFYGFPK MYQLKYPAPG+P LAKRVK++L GF V EDK RGLDHGAW
Sbjct: 98 IRGRQDTIHDFYGFPKSMYQLKYPAPGSPELAKRVKDVLMASGFPIVHEDKNRGLDHGAW 157
Query: 120 VPLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALER 179
VPL+LMYPEADIPVCQLSVQ N DGT+HYN+GKALA LKDEGVLI+GSGSA HNLRAL
Sbjct: 158 VPLMLMYPEADIPVCQLSVQPNRDGTYHYNLGKALASLKDEGVLIIGSGSATHNLRALGP 217
Query: 180 HATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGE 239
V++ WA+EFDNWLK+ALL GR++DVN+Y+ KAPHAK AHPWP+H YPLHVA+GAAGE
Sbjct: 218 SKNVSS-WALEFDNWLKDALLSGRHQDVNNYDMKAPHAKVAHPWPEHIYPLHVALGAAGE 276
Query: 240 NSKAKLIHSSIDLGSLSYASYQFTS 264
+LIH S DLG+LSYASY+F S
Sbjct: 277 GVNGELIHHSWDLGALSYASYRFPS 301
>gb|AAM51245.1| unknown protein [Arabidopsis thaliana] gi|15293029|gb|AAK93625.1|
unknown protein [Arabidopsis thaliana]
gi|26452573|dbj|BAC43371.1| unknown protein [Arabidopsis
thaliana] gi|42558925|sp|Q949R4|DIOXL_ARATH 4,5-DOPA
dioxygenase extradiol-like protein
gi|18414376|ref|NP_567456.1| catalytic LigB subunit of
aromatic ring-opening dioxygenase family [Arabidopsis
thaliana]
Length = 269
Score = 376 bits (965), Expect = e-103
Identities = 180/267 (67%), Positives = 221/267 (82%), Gaps = 4/267 (1%)
Query: 3 LKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDS 62
+ TF++SHGSPTLSID+SL AR+F +SW ++V P +P SILVIS HWDT P+VN V
Sbjct: 4 VNQTFFLSHGSPTLSIDDSLEARQFFKSWTQKVLPQKPKSILVISAHWDTKFPSVNTV-L 62
Query: 63 TNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELL-KEGGFSRVDEDKKRGLDHGAWVP 121
N+TI+DF GFP PMY+LKY APGA L KRVKELL KEGG RVDED KRGLDHGAWVP
Sbjct: 63 RNNTIHDFSGFPDPMYKLKYEAPGAIELGKRVKELLMKEGGMKRVDEDTKRGLDHGAWVP 122
Query: 122 LLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHA 181
L+LMYPEADIP+CQLSVQSN +G++HYN+GKALA LKDEGVLI+GSGSA HNLR L+ +
Sbjct: 123 LMLMYPEADIPICQLSVQSNQNGSYHYNMGKALASLKDEGVLIIGSGSATHNLRKLDFNI 182
Query: 182 TVAA--PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGE 239
T + PWA+EFD+WL+++LL+GRY DVN +E+KAP+AK AHPWP+H YPLHV +GAAG
Sbjct: 183 TDGSPVPWALEFDHWLRDSLLQGRYGDVNEWEEKAPNAKMAHPWPEHLYPLHVVMGAAGG 242
Query: 240 NSKAKLIHSSIDLGSLSYASYQFTSDV 266
++KA+ IH+S LG+LSY+SY FTS +
Sbjct: 243 DAKAEQIHTSWQLGTLSYSSYSFTSSL 269
>emb|CAE47100.1| 4,5-DOPA dioxygenase extradiol [Beta vulgaris]
Length = 268
Score = 367 bits (942), Expect = e-100
Identities = 171/262 (65%), Positives = 211/262 (80%), Gaps = 1/262 (0%)
Query: 3 LKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDS 62
+K+TF+ISHG+P ++ID+S ++KFL+SW++++F +P +ILVIS HW+T P+VNVVD
Sbjct: 7 IKETFFISHGTPMMAIDDSKPSKKFLESWREKIFSKKPKAILVISAHWETDQPSVNVVD- 65
Query: 63 TNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPL 122
NDTIYDF GFP +YQ KY APG+P LA R+++LL GF V+ DKKRGLDHGAWVPL
Sbjct: 66 INDTIYDFRGFPARLYQFKYSAPGSPELANRIQDLLAGSGFKSVNTDKKRGLDHGAWVPL 125
Query: 123 LLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHAT 182
+LMYPEADIPVCQLSVQS+LDGTHHY +G+ALAPLKDEGVLI+GSGSA H +
Sbjct: 126 MLMYPEADIPVCQLSVQSHLDGTHHYKLGQALAPLKDEGVLIIGSGSATHPSNGTPPCSD 185
Query: 183 VAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSK 242
APWA FD+WL+ AL G YE+VN YE KAP+ K AHPWP+HFYPLHVA+GAAGENSK
Sbjct: 186 GVAPWAAAFDSWLETALTNGSYEEVNKYETKAPNWKLAHPWPEHFYPLHVAMGAAGENSK 245
Query: 243 AKLIHSSIDLGSLSYASYQFTS 264
A+LIH+S D G +SY SY+FTS
Sbjct: 246 AELIHNSWDGGIMSYGSYKFTS 267
>emb|CAB10288.1| hypothetical protein [Arabidopsis thaliana]
gi|7268255|emb|CAB78551.1| hypothetical protein
[Arabidopsis thaliana] gi|7485146|pir||F71414
hypothetical protein - Arabidopsis thaliana
Length = 1705
Score = 357 bits (915), Expect = 2e-97
Identities = 171/253 (67%), Positives = 209/253 (82%), Gaps = 4/253 (1%)
Query: 3 LKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDS 62
+ TF++SHGSPTLSID+SL AR+F +SW ++V P +P SILVIS HWDT P+VN V
Sbjct: 782 VNQTFFLSHGSPTLSIDDSLEARQFFKSWTQKVLPQKPKSILVISAHWDTKFPSVNTV-L 840
Query: 63 TNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELL-KEGGFSRVDEDKKRGLDHGAWVP 121
N+TI+DF GFP PMY+LKY APGA L KRVKELL KEGG RVDED KRGLDHGAWVP
Sbjct: 841 RNNTIHDFSGFPDPMYKLKYEAPGAIELGKRVKELLMKEGGMKRVDEDTKRGLDHGAWVP 900
Query: 122 LLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHA 181
L+LMYPEADIP+CQLSVQSN +G++HYN+GKALA LKDEGVLI+GSGSA HNLR L+ +
Sbjct: 901 LMLMYPEADIPICQLSVQSNQNGSYHYNMGKALASLKDEGVLIIGSGSATHNLRKLDFNI 960
Query: 182 TVAA--PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGE 239
T + PWA+EFD+WL+++LL+GRY DVN +E+KAP+AK AHPWP+H YPLHV +GAAG
Sbjct: 961 TDGSPVPWALEFDHWLRDSLLQGRYGDVNEWEEKAPNAKMAHPWPEHLYPLHVVMGAAGG 1020
Query: 240 NSKAKLIHSSIDL 252
++KA+ IH+S L
Sbjct: 1021 DAKAEQIHTSWQL 1033
>ref|NP_915030.1| P0471B04.17 [Oryza sativa (japonica cultivar-group)]
gi|22202675|dbj|BAC07333.1| putative 4,5-DOPA
dioxygenase extradiol [Oryza sativa (japonica
cultivar-group)] gi|21952792|dbj|BAC06208.1| putative
4,5-DOPA dioxygenase extradiol [Oryza sativa (japonica
cultivar-group)]
Length = 266
Score = 353 bits (907), Expect = 2e-96
Identities = 163/262 (62%), Positives = 203/262 (77%), Gaps = 2/262 (0%)
Query: 5 DTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPR--PTSILVISGHWDTAVPTVNVVDS 62
DTF++SHG+PTLSID+++ A+ F +SW P +ILV+SGHW+ A PTVNV+
Sbjct: 2 DTFFLSHGAPTLSIDDTIAAQGFFKSWLPAAVAGAELPRAILVVSGHWEAAAPTVNVIRG 61
Query: 63 TNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPL 122
NDTI+DFYGFPK MY+LKYPAPGAP LA + KELL++ GF V E+ RGLDHGAWVPL
Sbjct: 62 NNDTIHDFYGFPKAMYKLKYPAPGAPDLAMKTKELLEQAGFGPVKENHSRGLDHGAWVPL 121
Query: 123 LLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHAT 182
+ MYPEA++PVCQLS+QS DG +HY +G+ALAPL+D+GVL++GSGSA HNLR + T
Sbjct: 122 MFMYPEANVPVCQLSLQSGRDGAYHYELGRALAPLRDDGVLVLGSGSATHNLRRMGPEGT 181
Query: 183 VAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSK 242
WA EFD WL+EALL GR++DV YE+KAPH + AHP PDHF PLHVA+GAAGE +K
Sbjct: 182 PVPQWAAEFDGWLQEALLGGRHDDVKRYEEKAPHGRVAHPSPDHFLPLHVALGAAGEGAK 241
Query: 243 AKLIHSSIDLGSLSYASYQFTS 264
A+LIH S SLSYASY+FT+
Sbjct: 242 AELIHRSWSNASLSYASYRFTT 263
>ref|NP_915031.1| P0471B04.18 [Oryza sativa (japonica cultivar-group)]
gi|22202676|dbj|BAC07334.1| putative 4,5-DOPA
dioxygenase extradiol [Oryza sativa (japonica
cultivar-group)] gi|21952793|dbj|BAC06209.1| putative
4,5-DOPA dioxygenase extradiol [Oryza sativa (japonica
cultivar-group)]
Length = 280
Score = 335 bits (858), Expect = 1e-90
Identities = 162/269 (60%), Positives = 208/269 (77%), Gaps = 11/269 (4%)
Query: 5 DTFYISHGSPTLSIDESLVARKFLQSWKKEVF----PPRPTSILVISGHWDTAVPTVNVV 60
DTF++SHG+PTLSID+++ A+ F +SW PPR +IL++SGHW+TA PTVNVV
Sbjct: 2 DTFFLSHGTPTLSIDDTMPAQHFFRSWLPAAVAGAQPPR--AILIVSGHWETATPTVNVV 59
Query: 61 DSTNDTIYDF--YGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGA 118
NDTI+DF YGFPK M+QL+YPAPGAP +AK+ KELL++ GF V ED RGLDHGA
Sbjct: 60 RGNNDTIHDFEGYGFPKSMFQLEYPAPGAPDVAKKAKELLEQAGFGPVKEDHGRGLDHGA 119
Query: 119 WVPLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALE 178
WVPL+ MYPEA++PVCQLS+Q+ DG +HY++G+ALAPL+D+GVLI+GSG+A HNL +
Sbjct: 120 WVPLMFMYPEANVPVCQLSLQTGRDGAYHYDLGRALAPLRDDGVLILGSGNATHNLSCMA 179
Query: 179 --RHATVAAPWAVEFDNWLKEALLE-GRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIG 235
T WA EFD WL+EALL GR++DV YE+KAP+ K AHP PDHF PLHVA+G
Sbjct: 180 PVAEGTPVPQWAAEFDGWLQEALLAGGRHDDVKQYEEKAPNGKMAHPSPDHFLPLHVALG 239
Query: 236 AAGENSKAKLIHSSIDLGSLSYASYQFTS 264
AAGE++KA+LIH S +LS+ASY+FT+
Sbjct: 240 AAGEDAKAELIHHSWYNATLSHASYRFTT 268
>emb|CAE45178.1| 4,5-DOPA dioxygenase extradiol [Portulaca grandiflora]
gi|42558922|sp|Q7XA48|DODA_PORGR 4,5-DOPA dioxygenase
extradiol
Length = 271
Score = 327 bits (837), Expect = 3e-88
Identities = 158/265 (59%), Positives = 193/265 (72%), Gaps = 2/265 (0%)
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
++ K++F++SHG+P + DES +AR FL WKK VFP +P SILV+S HW+T VP V+
Sbjct: 7 VSFKESFFLSHGNPAMLADESFIARNFLLGWKKNVFPVKPKSILVVSAHWETDVPCVSAG 66
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
N IYDF P M+Q+KYPAPG P LAKRV+ELL GGF D++RG DH +WV
Sbjct: 67 QYPN-VIYDFTEVPASMFQMKYPAPGCPKLAKRVQELLIAGGFKSAKLDEERGFDHSSWV 125
Query: 121 PLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERH 180
PL +M PEADIPVCQLSVQ LD THH+N+G+ALAPLK EGVL +GSG AVH
Sbjct: 126 PLSMMCPEADIPVCQLSVQPGLDATHHFNVGRALAPLKGEGVLFIGSGGAVHPSDDTPHW 185
Query: 181 ATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHA-KKAHPWPDHFYPLHVAIGAAGE 239
APWA EFD WL++ALLEGRYEDVN+Y+ KAP K AHP P+HF PLHVA+GA GE
Sbjct: 186 FDGVAPWAAEFDQWLEDALLEGRYEDVNNYQTKAPEGWKLAHPIPEHFLPLHVAMGAGGE 245
Query: 240 NSKAKLIHSSIDLGSLSYASYQFTS 264
SKA+LI+ + D G+L YASY+FTS
Sbjct: 246 KSKAELIYRTWDHGTLGYASYKFTS 270
>emb|CAE47099.1| 4,5 dioxygenase extradiol [Physcomitrella patens]
Length = 264
Score = 287 bits (734), Expect = 2e-76
Identities = 139/248 (56%), Positives = 176/248 (70%), Gaps = 4/248 (1%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
TFY+SHGSP + ++++ + R+F +W E +P RP +IL IS HWDT P VN V S N
Sbjct: 9 TFYVSHGSPMMPLEDTPI-REFFSTWT-ERYPTRPKAILAISAHWDTREPAVNAV-SQNS 65
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +YQL+Y PGAP +AKRV LLK+ GF V ED KRGLDHGAW PL+LM
Sbjct: 66 TIHDFYGFPRELYQLQYTPPGAPDVAKRVTSLLKDAGFKTVLEDNKRGLDHGAWTPLMLM 125
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVAA 185
YP ADIPV Q+S+QSN DG HHY +G+ALAPLKDEGVLI SG+ VHNLR ++ A
Sbjct: 126 YPNADIPVLQVSIQSNKDGLHHYQLGRALAPLKDEGVLIFASGTTVHNLREIDFSAKKPT 185
Query: 186 PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKAKL 245
WA FD WL + LL ++++ +E KAP+A KAHP PDHF P+ V +GAAGE +A+
Sbjct: 186 VWAKAFDGWLTDVLLNSKHKEAMEWE-KAPYASKAHPHPDHFLPVLVGLGAAGEQCQAEK 244
Query: 246 IHSSIDLG 253
I+ G
Sbjct: 245 IYEEFAYG 252
>ref|ZP_00423589.1| Catalytic LigB subunit of aromatic ring-opening dioxygenase
[Burkholderia vietnamiensis G4]
gi|67532998|gb|EAM29771.1| Catalytic LigB subunit of
aromatic ring-opening dioxygenase [Burkholderia
vietnamiensis G4]
Length = 263
Score = 218 bits (556), Expect = 1e-55
Identities = 109/261 (41%), Positives = 159/261 (60%), Gaps = 8/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ Y+SHG+PTL ID SL + F + PRP ++L++S HW T P +V + +
Sbjct: 6 SLYLSHGAPTLPIDPSLPSAAFTHLGAEL---PRPRAVLMLSAHWGTQQPVASVA-AQPE 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y+++YPAPGAP +A+R ELL G + + GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPRALYEIRYPAPGAPEVAERAAELLNAAGIATATTE--HGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+P+AD+PV QLS+Q D HH+ +G+AL PL+DEGV+++GSG HNLRA + A
Sbjct: 120 FPDADVPVAQLSIQPRADAAHHFALGRALRPLRDEGVMVIGSGQITHNLRAADFGAAPED 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
A P EF +W + L + + Y ++APHA HP +H P+ A+GAA ++ +
Sbjct: 180 ADPRVAEFTDWFEARLAARNVDALLDYRRQAPHAALMHPTDEHLLPVFAALGAADDDYRL 239
Query: 244 KLIHSSIDLGSLSYASYQFTS 264
+ L+ +Y F S
Sbjct: 240 GIQSLGTYQRVLAMTNYVFAS 260
>ref|ZP_00448339.1| COG3384: Uncharacterized conserved protein [Burkholderia mallei
SAVP1] gi|67647474|ref|ZP_00445716.1| COG3384:
Uncharacterized conserved protein [Burkholderia mallei
NCTC 10247] gi|67639913|ref|ZP_00438741.1| COG3384:
Uncharacterized conserved protein [Burkholderia mallei
GB8 horse 4] gi|67634624|ref|ZP_00433593.1| COG3384:
Uncharacterized conserved protein [Burkholderia mallei
10399] gi|67632240|ref|ZP_00432096.1| COG3384:
Uncharacterized conserved protein [Burkholderia mallei
10229] gi|53717074|ref|YP_105271.1| class III
extradiol-type catecholic dioxygenase family protein
[Burkholderia mallei ATCC 23344]
gi|52423044|gb|AAU46614.1| class III extradiol-type
catecholic dioxygenase family protein [Burkholderia
mallei ATCC 23344]
Length = 262
Score = 217 bits (553), Expect = 2e-55
Identities = 108/261 (41%), Positives = 156/261 (59%), Gaps = 8/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ Y+SHG+PTL ID +L + F + + PRP ++L++S HW T P V+ + D
Sbjct: 6 SLYLSHGAPTLPIDPTLPSGAFTELGAEL---PRPRAVLMLSAHWLTQQPVVSTAERP-D 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y+++YPAPGAP +A LL+ G GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPRALYEIRYPAPGAPDVAAHAAALLRSAGIETAATP--HGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+P AD+PV QLS+Q D HH+ +G+AL PL+DEGV+++GSG HNLRA + A
Sbjct: 120 FPHADVPVAQLSIQPRADAAHHFRVGRALRPLRDEGVMVIGSGQITHNLRAADFSAAPDD 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
P EF W +EAL + + Y ++APHA HP +H P+ A+GAA ++ +
Sbjct: 180 VDPRVAEFTGWFEEALAARDVDALVDYRRRAPHAALMHPTDEHLLPVFAALGAANDDYRL 239
Query: 244 KLIHSSIDLGSLSYASYQFTS 264
++ L+ +Y F S
Sbjct: 240 RIQSLGTYQRMLAMTNYVFDS 260
>ref|ZP_00497919.1| COG3384: Uncharacterized conserved protein [Burkholderia
pseudomallei S13] gi|67754087|ref|ZP_00493004.1|
COG3384: Uncharacterized conserved protein [Burkholderia
pseudomallei Pasteur] gi|67715997|ref|ZP_00485358.1|
COG3384: Uncharacterized conserved protein [Burkholderia
pseudomallei 1710b] gi|67681691|ref|ZP_00476016.1|
COG3384: Uncharacterized conserved protein [Burkholderia
pseudomallei 1710a] gi|67669820|ref|ZP_00466641.1|
COG3384: Uncharacterized conserved protein [Burkholderia
pseudomallei 1655] gi|52211100|emb|CAH37088.1| conserved
hypothetical protein [Burkholderia pseudomallei K96243]
gi|53720686|ref|YP_109672.1| hypothetical protein
BPSL3077 [Burkholderia pseudomallei K96243]
Length = 262
Score = 217 bits (552), Expect = 3e-55
Identities = 108/261 (41%), Positives = 156/261 (59%), Gaps = 8/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ Y+SHG+PTL ID +L + F + + PRP ++L++S HW T P V+ + D
Sbjct: 6 SLYLSHGAPTLPIDPTLPSGAFTELGAEL---PRPRAVLMLSAHWLTQQPVVSTAERP-D 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y+++YPAPGAP +A LL+ G GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPRALYEIRYPAPGAPDVAAHAAALLRSAGIETAATP--HGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+P AD+PV QLS+Q D HH+ +G+AL PL+DEGV+++GSG HNLRA + A
Sbjct: 120 FPHADVPVAQLSIQPRADAAHHFRVGRALRPLRDEGVMVIGSGQITHNLRAADFSAAPDD 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
P EF W +EAL + + Y ++APHA HP +H P+ A+GAA ++ +
Sbjct: 180 VDPRVAEFTGWFEEALAARDVDALVDYRRRAPHAALMHPTDEHLLPVFAALGAADDDYRL 239
Query: 244 KLIHSSIDLGSLSYASYQFTS 264
++ L+ +Y F S
Sbjct: 240 RIQSLGTYQRMLAMTNYVFDS 260
>ref|ZP_00488716.1| COG3384: Uncharacterized conserved protein [Burkholderia
pseudomallei 668]
Length = 262
Score = 216 bits (551), Expect = 4e-55
Identities = 108/261 (41%), Positives = 156/261 (59%), Gaps = 8/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ Y+SHG+PTL ID +L + F + + PRP ++L++S HW T P V+ + D
Sbjct: 6 SLYLSHGAPTLPIDPTLPSGAFTELGAEL---PRPRAVLMLSAHWLTQQPVVSTAERP-D 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y+++YPAPGAP +A LL+ G GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPRALYEIRYPAPGAPDVAAHAAALLRSAGIETAATP--HGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+P AD+PV QLS+Q D HH+ +G+AL PL+DEGV+++GSG HNLRA + A
Sbjct: 120 FPHADVPVAQLSIQPRADAAHHFRVGRALRPLRDEGVMVIGSGQITHNLRAADFSAPPDD 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
P EF W +EAL + + Y ++APHA HP +H P+ A+GAA ++ +
Sbjct: 180 VDPRVAEFTGWFEEALAARDVDALVDYRRRAPHAALMHPTDEHLLPVFAALGAADDDYRL 239
Query: 244 KLIHSSIDLGSLSYASYQFTS 264
++ L+ +Y F S
Sbjct: 240 RIQSLGTYQRMLAMTNYVFDS 260
>ref|ZP_00464133.1| Catalytic LigB subunit of aromatic ring-opening dioxygenase
[Burkholderia cenocepacia HI2424]
gi|67655132|ref|ZP_00452519.1| Catalytic LigB subunit of
aromatic ring-opening dioxygenase [Burkholderia
cenocepacia AU 1054] gi|67097270|gb|EAM14792.1|
Catalytic LigB subunit of aromatic ring-opening
dioxygenase [Burkholderia cenocepacia AU 1054]
gi|67099578|gb|EAM16750.1| Catalytic LigB subunit of
aromatic ring-opening dioxygenase [Burkholderia
cenocepacia HI2424]
Length = 263
Score = 214 bits (544), Expect = 3e-54
Identities = 109/261 (41%), Positives = 157/261 (59%), Gaps = 8/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ Y+SHG+PTL ID +L + F + PRP ++L++S HW T P +V + D
Sbjct: 6 SLYLSHGAPTLPIDPALPSGAFTHLGAEL---PRPRAVLMLSAHWGTQRPVASVA-AHPD 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y ++YPAPGAP +A+R LL G + + GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPQALYDIRYPAPGAPDVAERAAALLNAAGIATATTE--HGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+PEAD+PV QLS+Q D HH+ +G+AL PL+DEGV+++GSG HNLRA + A
Sbjct: 120 FPEADVPVAQLSIQPRADAAHHFALGRALRPLRDEGVMVIGSGQITHNLRAADFGAAPED 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
A P EF +W + L + + Y ++APHA HP +H P+ A+GAA ++ +
Sbjct: 180 ADPRVAEFTDWFEAKLAARDVDALLDYRRQAPHAALMHPTDEHLLPVFTALGAADDDYQL 239
Query: 244 KLIHSSIDLGSLSYASYQFTS 264
+ L+ +Y F S
Sbjct: 240 GIQSLGTYQRVLAMTNYVFAS 260
>gb|AAQ61212.1| conserved hypothetical protein [Chromobacterium violaceum ATCC
12472] gi|34499005|ref|NP_903220.1| hypothetical protein
CV3550 [Chromobacterium violaceum ATCC 12472]
Length = 259
Score = 212 bits (540), Expect = 8e-54
Identities = 107/257 (41%), Positives = 164/257 (63%), Gaps = 5/257 (1%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ +ISHG+PTL I++S R FL+ ++ PRP +I+++S HW+ TVN+ +
Sbjct: 7 SLFISHGAPTLPIEDS-PTRHFLEQLGRDW--PRPRAIVIVSAHWEARALTVNL-QPRPE 62
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
T+YDF GFP + L+YPA LA+RV LL G V ++R LDHG W PL+LM
Sbjct: 63 TMYDFGGFPPVLRTLRYPAATDAALAERVIGLLDAAGLP-VAATEERPLDHGMWTPLMLM 121
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVAA 185
+P+ADIP+ +S++ HY +G+AL PL+D+ VL++GSGS HNL L + A
Sbjct: 122 FPQADIPLVMISLRHQAGVAEHYALGQALKPLRDDNVLVIGSGSYTHNLWELSPEGSEPA 181
Query: 186 PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKAKL 245
PWA +F W+ +ALL+ R +D+ H++Q+AP+ ++ HP +HF PL VA+GA+ +++
Sbjct: 182 PWAADFAGWMDQALLDNRRDDIVHWQQRAPYPRENHPSDEHFQPLLVALGASDDDAVPVK 241
Query: 246 IHSSIDLGSLSYASYQF 262
+H +GSLS A ++F
Sbjct: 242 LHDDWRMGSLSMACWRF 258
>ref|ZP_00278252.1| COG3384: Uncharacterized conserved protein [Burkholderia fungorum
LB400]
Length = 261
Score = 211 bits (538), Expect = 1e-53
Identities = 106/259 (40%), Positives = 159/259 (60%), Gaps = 8/259 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ ++SHG+PTL ID SL + +F + PRP S+L++S HW TA P + + +
Sbjct: 6 SLFLSHGAPTLPIDPSLPSAEFGVLGAQL---PRPESVLMLSAHWGTAQPVAST-STAPE 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y+++YPAPGAP +A+R LL + G + + + GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPRRLYEIQYPAPGAPDVARRAAVLLNDNGI--LTDVQPHGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+P AD+PV QLS+Q ++D HH+ +G+AL +D+GV+++GSG HNLRA + A
Sbjct: 120 FPHADVPVAQLSIQPHMDAAHHFRVGRALRSQRDDGVMVIGSGQITHNLRAADFSARPED 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
A P EF W +E L + + Y ++APHA HP +H P+ A+GAA ++
Sbjct: 180 ADPRVAEFTGWFEEKLAVHDIDALLDYRRQAPHAALMHPTDEHLLPVFAALGAADDDYSL 239
Query: 244 KLIHSSIDLGSLSYASYQF 262
+ SL+ +Y F
Sbjct: 240 AIQSLGTYQRSLAMTNYVF 258
>ref|ZP_00212419.1| COG3384: Uncharacterized conserved protein [Burkholderia cepacia
R18194]
Length = 263
Score = 211 bits (536), Expect = 2e-53
Identities = 107/261 (40%), Positives = 156/261 (58%), Gaps = 8/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ Y+SHG+PTL ID +L + F + PRP ++L++S HW T P ++ + +
Sbjct: 6 SLYLSHGAPTLPIDPTLPSGAFTHLAAEL---PRPRAVLMLSAHWGTQRPVASIA-AHPE 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DFYGFP+ +Y ++YPAPGAP +A+R LL G + GLDHGAWVP+LLM
Sbjct: 62 TIHDFYGFPQALYDIRYPAPGAPDVAERAAGLLNAAGIETATTE--HGLDHGAWVPMLLM 119
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATV-- 183
+PEAD+PV QLS+Q D HH+ +G+AL PL+DEGV+++GSG HNLRA + A
Sbjct: 120 FPEADVPVAQLSIQPRADAAHHFALGRALRPLRDEGVMVIGSGQITHNLRAADFGAAPED 179
Query: 184 AAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKA 243
A P EF +W + L + + Y ++APHA HP +H P+ A+GAA ++ +
Sbjct: 180 ADPRVTEFTDWFEAKLAARDVDALLDYRRQAPHAALMHPTDEHLLPVFTALGAADDDYQL 239
Query: 244 KLIHSSIDLGSLSYASYQFTS 264
+ L+ +Y F S
Sbjct: 240 GIQSLGTYQRVLAMTNYVFAS 260
>ref|NP_639012.1| hypothetical protein XCC3666 [Xanthomonas campestris pv. campestris
str. ATCC 33913] gi|21114949|gb|AAM42936.1| conserved
hypothetical protein [Xanthomonas campestris pv.
campestris str. ATCC 33913] gi|66770035|ref|YP_244797.1|
hypothetical protein XC_3737 [Xanthomonas campestris pv.
campestris str. 8004] gi|66575367|gb|AAY50777.1|
conserved hypothetical protein [Xanthomonas campestris
pv. campestris str. 8004]
Length = 272
Score = 203 bits (516), Expect = 5e-51
Identities = 113/265 (42%), Positives = 162/265 (60%), Gaps = 8/265 (3%)
Query: 1 MALKDTFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVV 60
M++ + YISHGSP ++ + + L + + + PRP +I+V S HW P V
Sbjct: 1 MSVLPSLYISHGSPMTALRPGELGTQ-LAALAQGL--PRPRAIVVASAHWLARRPLVGAA 57
Query: 61 DSTNDTIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWV 120
S TI+DF GFP P+Y+++YPAPGAP LA++V LL+ G +D +RGLDHG WV
Sbjct: 58 -SKPVTIHDFGGFPPPLYEIEYPAPGAPALAQQVTRLLELAGL-HPHQDPQRGLDHGIWV 115
Query: 121 PLLLMYPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALER- 179
PL ++YP+ADIPV LS+Q L H + +G+ALAPL+ EGVL++GSGS HNL L +
Sbjct: 116 PLRMLYPQADIPVVPLSIQPELGPAHQFAVGRALAPLRAEGVLVIGSGSITHNLHDLHQG 175
Query: 180 -HATVAAPWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAA- 237
A AP+ F +W + L + + Y ++APHA +AHP +H PL+VA+GAA
Sbjct: 176 YSAEREAPYVRPFIDWTERRLADDDVPALLDYRRQAPHAARAHPTDEHLLPLYVAMGAAG 235
Query: 238 GENSKAKLIHSSIDLGSLSYASYQF 262
G+ A+ I + ID G + Y+F
Sbjct: 236 GDRLGAQRIDAGIDQGLIGMDIYRF 260
>gb|AAM38549.1| conserved hypothetical protein [Xanthomonas axonopodis pv. citri
str. 306] gi|21244431|ref|NP_644013.1| hypothetical
protein XAC3706 [Xanthomonas axonopodis pv. citri str.
306]
Length = 272
Score = 202 bits (513), Expect = 1e-50
Identities = 113/261 (43%), Positives = 159/261 (60%), Gaps = 10/261 (3%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ YISHGSP ++ V + L + + PRP +I+V S HW P V+
Sbjct: 6 SLYISHGSPMTALQPGQVGER-LADLRHAL--PRPRAIVVASAHWLGRRPLVSGA-LRPP 61
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TI+DF GFP P+Y L YPAPG P LA+++ L++ G D +RG DHG WVPL L+
Sbjct: 62 TIHDFGGFPAPLYALDYPAPGEPALAQQIVRRLEQAGL-HPQLDPQRGFDHGVWVPLRLL 120
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVA- 184
YP+ADIPV LS+Q L H + +G+ALAPL+D+GVL++GSGS HNL + RH A
Sbjct: 121 YPDADIPVVSLSIQPELGPAHQFAVGRALAPLRDDGVLVIGSGSITHNLHDV-RHGYSAE 179
Query: 185 --APWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENS- 241
AP+ F W + LL+ + Y ++APHA++AHP +H+ PL+VA+GAAG++
Sbjct: 180 RQAPYVRPFIEWTERKLLDDDIPALLDYRRQAPHAERAHPSDEHWLPLYVAMGAAGDDRL 239
Query: 242 KAKLIHSSIDLGSLSYASYQF 262
A+ I + ID G L+ Y+F
Sbjct: 240 GAQRIDAGIDAGFLAMDLYRF 260
>gb|AAP08761.1| hypothetical protein [Bacillus cereus ATCC 14579]
gi|30019929|ref|NP_831560.1| hypothetical protein BC1787
[Bacillus cereus ATCC 14579]
Length = 253
Score = 200 bits (508), Expect = 4e-50
Identities = 105/257 (40%), Positives = 164/257 (62%), Gaps = 7/257 (2%)
Query: 6 TFYISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTND 65
+ +++HGSP L+I ++ R FL++ + +P +I++ + HW++ V T++ D+ +
Sbjct: 4 SLFLAHGSPMLAIQDTDYTR-FLKTLGETY---KPKAIVIFTAHWESEVLTISSSDNEYE 59
Query: 66 TIYDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLM 125
TIYDF GFP +Y++KY A G+ H+A ++ K G S V + RGLDHG+W L M
Sbjct: 60 TIYDFGGFPPELYEIKYRAKGSSHIASILETKFKNKGIS-VHHNMTRGLDHGSWTLLHRM 118
Query: 126 YPEADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRALERHATVAA 185
YPEA+IPV Q+SV L + IG+AL L E +L++GSG VHNLRAL+ + T
Sbjct: 119 YPEANIPVVQISVNPFLSAKEQFEIGEALKGLGQEDILVIGSGVTVHNLRALKWNQTTPE 178
Query: 186 PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKAKL 245
WA+EFD+W+ + + + + ++E+ APHA+ A P +HF PL +A+G +GENS ++
Sbjct: 179 KWAIEFDDWIIKHMQNNDKDALFNWEKNAPHAQLAVPRAEHFVPLFIAMG-SGENS-GEV 236
Query: 246 IHSSIDLGSLSYASYQF 262
IH S +LG+LSY +F
Sbjct: 237 IHRSYELGTLSYLCLRF 253
>ref|YP_147229.1| hypothetical protein GK1376 [Geobacillus kaustophilus HTA426]
gi|56379753|dbj|BAD75661.1| hypothetical conserved
protein [Geobacillus kaustophilus HTA426]
Length = 255
Score = 200 bits (508), Expect = 4e-50
Identities = 108/257 (42%), Positives = 156/257 (60%), Gaps = 10/257 (3%)
Query: 8 YISHGSPTLSIDESLVARKFLQSWKKEVFPPRPTSILVISGHWDTAVPTVNVVDSTNDTI 67
+++HGSP L+I+++ FL+ + + RP ++++ + HW T PTV+ V+ T D I
Sbjct: 7 FLAHGSPMLAIEDNAYTA-FLKKLGEGI---RPQAVVIFTAHWMTRQPTVSAVEGTYDMI 62
Query: 68 YDFYGFPKPMYQLKYPAPGAPHLAKRVKELLKEGGFSRVDEDKKRGLDHGAWVPLLLMYP 127
YDF GFP+ +Y++ YPA G+ A+RV+E L G + V D RGLDHG+WV L ++P
Sbjct: 63 YDFSGFPRELYEVVYPARGSVEWAERVRERL--AGVAEVAVDSTRGLDHGSWVLLHRLFP 120
Query: 128 EADIPVCQLSVQSNLDGTHHYNIGKALAPLKDEGVLIVGSGSAVHNLRAL--ERHATVAA 185
ADIP+ Q SV + +G+AL PL+DEG++++GSG VHNL AL E HA A
Sbjct: 121 NADIPIVQASVVPWWTPKQLFQLGEALRPLRDEGLMVIGSGGTVHNLMALRWEGHAE-AE 179
Query: 186 PWAVEFDNWLKEALLEGRYEDVNHYEQKAPHAKKAHPWPDHFYPLHVAIGAAGENSKAKL 245
WA FD+WL R EDV HYE+KAP+AK+A P P+H P +A GA + ++
Sbjct: 180 SWAAAFDDWLLRE-SAARSEDVFHYEEKAPYAKQAVPTPEHLAPYWIAYGAGDRDQAPRM 238
Query: 246 IHSSIDLGSLSYASYQF 262
+ GSLS + F
Sbjct: 239 LFREYQYGSLSLMAVSF 255
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 505,736,078
Number of Sequences: 2540612
Number of extensions: 22654259
Number of successful extensions: 51414
Number of sequences better than 10.0: 146
Number of HSP's better than 10.0 without gapping: 129
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 50859
Number of HSP's gapped (non-prelim): 165
length of query: 267
length of database: 863,360,394
effective HSP length: 126
effective length of query: 141
effective length of database: 543,243,282
effective search space: 76597302762
effective search space used: 76597302762
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0074.3


