

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0067a.13
(212 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
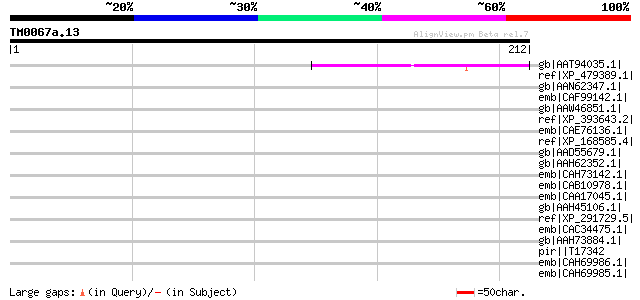
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT94035.1| hypothetical protein [Oryza sativa (japonica cult... 46 6e-04
ref|XP_479389.1| hypothetical protein [Oryza sativa (japonica cu... 45 0.002
gb|AAN62347.1| CTV.20 [Poncirus trifoliata] 45 0.002
emb|CAF99142.1| unnamed protein product [Tetraodon nigroviridis] 41 0.026
gb|AAW46851.1| hypothetical protein CNM00300 [Cryptococcus neofo... 40 0.057
ref|XP_393643.2| PREDICTED: similar to ENSANGP00000004283 [Apis ... 40 0.057
emb|CAE76136.1| related to sec7-domain protein [Neurospora crass... 39 0.075
ref|XP_168585.4| PREDICTED: similar to mucin 11 [Homo sapiens] 39 0.075
gb|AAD55679.1| mucin 11 [Homo sapiens] 39 0.075
gb|AAH62352.1| TAF3 protein [Homo sapiens] 39 0.098
emb|CAH73142.1| TAF3 RNA polymerase II, TATA box binding protein... 39 0.098
emb|CAB10978.1| scr1 [Schizosaccharomyces pombe] gi|19112509|ref... 39 0.098
emb|CAA17045.1| SPBC6B1.02 [Schizosaccharomyces pombe] gi|191128... 39 0.098
gb|AAH45106.1| TAF3 protein [Homo sapiens] 39 0.098
ref|XP_291729.5| PREDICTED: RNA polymerase II transcription fact... 39 0.098
emb|CAC34475.1| TAFII140 protein [Homo sapiens] 39 0.098
gb|AAH73884.1| TAF3 protein [Homo sapiens] 39 0.098
pir||T17342 hypothetical protein DKFZp586K1924.1 - human (fragme... 39 0.098
emb|CAH69986.1| SAMD6 [Homo sapiens] gi|55663435|emb|CAH71296.1|... 39 0.13
emb|CAH69985.1| SAMD6 [Homo sapiens] gi|55663434|emb|CAH71295.1|... 39 0.13
>gb|AAT94035.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 950
Score = 46.2 bits (108), Expect = 6e-04
Identities = 25/92 (27%), Positives = 49/92 (53%), Gaps = 4/92 (4%)
Query: 124 IAKQVFPPNFHYMSGHPSKGRLFYEFILVDTDSIEVTHNKDKNGEIICSKVKILKVLNMQ 183
I + FP + +++ +P K + +YE IL +T S ++ H ++ E+ +K+ I K++++
Sbjct: 675 IMGKYFPQHQYFIPEYPGKDQNYYETILCETRSAQIFHTRN-GDELGFTKLLIQKIISID 733
Query: 184 DW---KHPLFQSKQFSREFTPTGHNYFDYMDA 212
DW +P +S +NY+DY A
Sbjct: 734 DWDKSSNPYVARTIYSTSCANKRYNYWDYQKA 765
>ref|XP_479389.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|23617112|dbj|BAC20794.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 376
Score = 44.7 bits (104), Expect = 0.002
Identities = 25/92 (27%), Positives = 49/92 (53%), Gaps = 4/92 (4%)
Query: 124 IAKQVFPPNFHYMSGHPSKGRLFYEFILVDTDSIEVTHNKDKNGEIICSKVKILKVLNMQ 183
I + FP + +++ +P K + +YE IL +T S ++ H ++ ++ +K+ I K +++
Sbjct: 69 IMGKYFPQHQYFIPEYPGKDQNYYETILCETRSAQIFHTRN-GDDLGFTKLLIQKNISID 127
Query: 184 DW---KHPLFQSKQFSREFTPTGHNYFDYMDA 212
DW +P +S T +NY+DY A
Sbjct: 128 DWDKSSNPYVARTIYSTSCTNKRYNYWDYQKA 159
>gb|AAN62347.1| CTV.20 [Poncirus trifoliata]
Length = 3148
Score = 44.7 bits (104), Expect = 0.002
Identities = 40/192 (20%), Positives = 83/192 (42%), Gaps = 12/192 (6%)
Query: 28 SSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSPLVPLSNKYSILDYQNTI 87
+ +P + L+ I + ++P +I + ++S++ ++ + S Q +
Sbjct: 1483 TKSPELMFALQGISKSKALSQIPEEEKPISKEITKLSSQSQNVVISGESSSS----QIVL 1538
Query: 88 SSKTPGKTEQYEYQEKPECLPSVMIEKAWLNCNPRDIAKQVFPPNFHYMSGHPSKGRLFY 147
S TP K ++ +K + +E + + +P + FP ++ + +K + +Y
Sbjct: 1539 SQPTPSKKTS-DWFDKSHFQNVLTMEHGFYHSDPFQAISKFFPQSWFFNPWDLTKPQSYY 1597
Query: 148 EFILVDTDSIEVTH--NKDKNGEIICSKVKILKVLNMQDWKHPLFQSKQFSREFTP---- 201
+ IL T+S++ H + + E S ILKVL+ W L + K F F
Sbjct: 1598 QSILEATESVKFKHFFLSETHSEPAYSTATILKVLSPNQWGDQLHKYKSFPPNFQMRLPH 1657
Query: 202 -TGHNYFDYMDA 212
++Y+DY A
Sbjct: 1658 CLVYSYWDYQQA 1669
>emb|CAF99142.1| unnamed protein product [Tetraodon nigroviridis]
Length = 2314
Score = 40.8 bits (94), Expect = 0.026
Identities = 38/139 (27%), Positives = 52/139 (37%), Gaps = 17/139 (12%)
Query: 13 PPLGSSRKLPSSITGSSAPVAVTPLKTI----KPPGPSTPTKSSQRPTVSQIVQSPAKSK 68
P +S S T SSA VA+ P + + +PP + T+SS PT P K K
Sbjct: 1760 PARPNSAHADHSSTSSSAHVAIAPQQPVGTISQPPIQPSKTESSAAPT-------PGKDK 1812
Query: 69 SPLVPLSNKYSILDYQNTISSKTPGKTEQ------YEYQEKPECLPSVMIEKAWLNCNPR 122
L + S+ D N++ P T + P LPS A LN
Sbjct: 1813 PALAAENQPASVTDGMNSVGFSAPAMTMSAKAEPLQQLPPPPASLPSSEAPPALLNPQIS 1872
Query: 123 DIAKQVFPPNFHYMSGHPS 141
V PP + HP+
Sbjct: 1873 SHLPTVPPPVLSHSVAHPN 1891
>gb|AAW46851.1| hypothetical protein CNM00300 [Cryptococcus neoformans var.
neoformans JEC21] gi|58261916|ref|XP_568368.1|
hypothetical protein CNM00300 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 1701
Score = 39.7 bits (91), Expect = 0.057
Identities = 28/93 (30%), Positives = 37/93 (39%), Gaps = 11/93 (11%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PT PP S RK P G + P + L T P G +TP +
Sbjct: 466 STTPTESQSQPPPRKSPRKPPPPNLGITPPNPASLLSTPNPSGSATPAAT---------- 515
Query: 62 QSPAKSKSPLVPLSNKYSILDYQNTISSKTPGK 94
P KSPL +S S+ Y + S + PG+
Sbjct: 516 -PPVPLKSPLRSISGGSSVTSYNSYSSVEIPGE 547
>ref|XP_393643.2| PREDICTED: similar to ENSANGP00000004283 [Apis mellifera]
Length = 2004
Score = 39.7 bits (91), Expect = 0.057
Identities = 39/122 (31%), Positives = 47/122 (37%), Gaps = 16/122 (13%)
Query: 12 GPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVS-QIVQSPAKSKSP 70
GPP G LP GS A +PL T S P PT+S Q S A + P
Sbjct: 483 GPPTGGDC-LPVPSVGSPGSPAPSPLPTPHSEPASVPPAEPTMPTLSPQPPPSHANTAPP 541
Query: 71 LVPLSNKYSILDY-QNTISSKTPGKT------------EQY-EYQEKPECLPSVMIEKAW 116
L P S N + S PG T E Y E ++ CL I++AW
Sbjct: 542 LTPSQGPKSTSSACNNQVHSPVPGPTLKRPVLVSRECEEIYLENEQSLRCLYDYSIQEAW 601
Query: 117 LN 118
LN
Sbjct: 602 LN 603
>emb|CAE76136.1| related to sec7-domain protein [Neurospora crassa]
gi|32414849|ref|XP_327904.1| hypothetical protein
[Neurospora crassa] gi|28917818|gb|EAA27506.1|
hypothetical protein [Neurospora crassa]
Length = 1448
Score = 39.3 bits (90), Expect = 0.075
Identities = 27/91 (29%), Positives = 40/91 (43%), Gaps = 2/91 (2%)
Query: 20 KLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSS--QRPTVSQIVQSPAKSKSPLVPLSNK 77
++P S S VTP + P PT SS +PT + S + S P S +
Sbjct: 54 EVPISSVSPSPVTPVTPATPVMPTVDLEPTMSSTAPQPTGDPVDHSLSPSVPAETPKSKR 113
Query: 78 YSILDYQNTISSKTPGKTEQYEYQEKPECLP 108
+S+L ++N S+ K + EKP LP
Sbjct: 114 FSMLRFRNASDSQLAAKAKLQAASEKPPPLP 144
>ref|XP_168585.4| PREDICTED: similar to mucin 11 [Homo sapiens]
Length = 3927
Score = 39.3 bits (90), Expect = 0.075
Identities = 24/91 (26%), Positives = 37/91 (40%)
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSP 64
P+S G L +R S + G S P ++P T P +PT S + SP
Sbjct: 2108 PSSSGSTGTTLSPARSTTSGLVGESTPSRLSPSSTETTTLPGSPTTPSLSEKSTTFYTSP 2167
Query: 65 AKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
+ L P + S + +++ S PG T
Sbjct: 2168 RSPDATLSPATTTSSGVSEESSTSHSQPGST 2198
Score = 39.3 bits (90), Expect = 0.075
Identities = 24/91 (26%), Positives = 37/91 (40%)
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSP 64
P+S G L +R S + G S P ++P T P +PT S + SP
Sbjct: 1025 PSSSGSTGTKLSPARSTTSGLVGESTPSRLSPSSTETTTLPGSPTTPSLSEKSTTFYTSP 1084
Query: 65 AKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
+ L P + S + +++ S PG T
Sbjct: 1085 RSPDATLSPATTTSSGVSEESSTSHSQPGST 1115
Score = 39.3 bits (90), Expect = 0.075
Identities = 24/91 (26%), Positives = 37/91 (40%)
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSP 64
P+S G L +R S + G S P ++P T P +PT S + SP
Sbjct: 3665 PSSSGSTGTKLSPARSTTSGLVGESTPSRLSPSSTETTTLPGSPTTPSLSEKSTTFYTSP 3724
Query: 65 AKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
+ L P + S + +++ S PG T
Sbjct: 3725 RSPDATLSPATTTSSGVSEESSTSHSQPGST 3755
Score = 39.3 bits (90), Expect = 0.075
Identities = 24/91 (26%), Positives = 37/91 (40%)
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSP 64
P+S G L +R S + G S P ++P T P +PT S + SP
Sbjct: 2582 PSSSGSTGTTLSPARSTTSGLVGESTPSRLSPSSTETTTLPGSPTTPSLSEKSTTFYTSP 2641
Query: 65 AKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
+ L P + S + +++ S PG T
Sbjct: 2642 RSPDATLSPATTTSSGVSEESSTSHSQPGST 2672
Score = 32.7 bits (73), Expect = 7.0
Identities = 25/92 (27%), Positives = 42/92 (45%), Gaps = 4/92 (4%)
Query: 6 TSRDQYGPPLGSSRKLPSSITGSS--APVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQS 63
+S+D G + +R S + G S +P++ ++T PG +T S++ T S
Sbjct: 3192 SSQDATGTIVLPARSTTSVLLGESTTSPISSGSMETTALPGSTTTPGLSEKSTTFH--SS 3249
Query: 64 PAKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
P + L P S S + ++T S PG T
Sbjct: 3250 PRSPATTLSPASTTSSGVSEESTTSHSRPGST 3281
>gb|AAD55679.1| mucin 11 [Homo sapiens]
Length = 957
Score = 39.3 bits (90), Expect = 0.075
Identities = 24/91 (26%), Positives = 37/91 (40%)
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSP 64
P+S G L +R S + G S P ++P T P +PT S + SP
Sbjct: 746 PSSSGSTGTTLSPARSTTSGLVGESTPSRLSPSSTETTTLPGSPTTPSLSEKSTTFYTSP 805
Query: 65 AKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
+ L P + S + +++ S PG T
Sbjct: 806 RSPDATLSPATTTSSGVSEESSTSHSQPGST 836
>gb|AAH62352.1| TAF3 protein [Homo sapiens]
Length = 771
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 312 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 364
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 365 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 408
>emb|CAH73142.1| TAF3 RNA polymerase II, TATA box binding protein (TBP)-associated
factor, 140kDa (TAF3)(TAF140,TAFII140) [Homo sapiens]
gi|55662526|emb|CAH73451.1| TAF3 RNA polymerase II, TATA
box binding protein (TBP)-associated factor, 140kDa
(TAF3)(TAF140,TAFII140) [Homo sapiens]
gi|55958032|emb|CAI15661.1| TAF3 RNA polymerase II, TATA
box binding protein (TBP)-associated factor, 140kDa
(TAF3)(TAF140,TAFII140) [Homo sapiens]
Length = 929
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 255 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 307
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 308 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 351
>emb|CAB10978.1| scr1 [Schizosaccharomyces pombe] gi|19112509|ref|NP_595717.1|
hypothetical protein SPBC1D7.02c [Schizosaccharomyces
pombe 972h-] gi|7493740|pir||T39863 zinc finger protein
- fission yeast (Schizosaccharomyces pombe)
gi|12230559|sp|O14335|SCR1_SCHPO DNA-binding protein
scr1
Length = 565
Score = 38.9 bits (89), Expect = 0.098
Identities = 27/75 (36%), Positives = 38/75 (50%), Gaps = 4/75 (5%)
Query: 1 MSEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQI 60
M KP S+ GP SS + + SS V+V+ L PP PS+ TKS+ + S
Sbjct: 493 MPHKPASQSNVGPVRISSNRRSRKFSSSSR-VSVSNLLAGSPPSPSSSTKSA---SSSYS 548
Query: 61 VQSPAKSKSPLVPLS 75
+PA S PL P++
Sbjct: 549 TTTPAFSIGPLTPMT 563
>emb|CAA17045.1| SPBC6B1.02 [Schizosaccharomyces pombe] gi|19112873|ref|NP_596081.1|
hypothetical protein SPBC6B1.02 [Schizosaccharomyces
pombe 972h-] gi|7492952|pir||T40643 probable serine
threonine-protein kinase - fission yeast
(Schizosaccharomyces pombe)
Length = 953
Score = 38.9 bits (89), Expect = 0.098
Identities = 29/77 (37%), Positives = 38/77 (48%), Gaps = 8/77 (10%)
Query: 11 YGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSP 70
Y PP ++ PS GS P++ P T PP P+ T SS P V+ + KSKSP
Sbjct: 346 YNPPRAPLQRTPS---GSLTPLSSRPAHTSLPPIPTVQTTSSNVPPVN---RPSLKSKSP 399
Query: 71 LVP--LSNKYSILDYQN 85
V LSN+ S + N
Sbjct: 400 SVSNILSNQLSPISSAN 416
>gb|AAH45106.1| TAF3 protein [Homo sapiens]
Length = 539
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 255 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 307
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 308 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 351
>ref|XP_291729.5| PREDICTED: RNA polymerase II transcription factor TAFII140 [Homo
sapiens]
Length = 1248
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 522 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 574
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 575 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 618
>emb|CAC34475.1| TAFII140 protein [Homo sapiens]
Length = 727
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 255 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 307
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 308 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 351
>gb|AAH73884.1| TAF3 protein [Homo sapiens]
Length = 601
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 320 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 372
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 373 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 416
>pir||T17342 hypothetical protein DKFZp586K1924.1 - human (fragment)
gi|5912256|emb|CAB56032.1| hypothetical protein [Homo
sapiens]
Length = 681
Score = 38.9 bits (89), Expect = 0.098
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 233 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 285
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 286 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 329
>emb|CAH69986.1| SAMD6 [Homo sapiens] gi|55663435|emb|CAH71296.1| SAMD6 [Homo
sapiens]
Length = 585
Score = 38.5 bits (88), Expect = 0.13
Identities = 25/61 (40%), Positives = 35/61 (56%), Gaps = 3/61 (4%)
Query: 1 MSEKPTSRDQ-YGPPLGSS-RKLPSSITGSSAPVAVTPLKTIK-PPGPSTPTKSSQRPTV 57
+ +KP+ R GP GSS +LP+S G SAPV L+T K PP ++ T S PT+
Sbjct: 378 LEQKPSHRSSPVGPAPGSSPSELPASPAGGSAPVGKQKLETSKRPPSGTSTTSKSTSPTL 437
Query: 58 S 58
+
Sbjct: 438 T 438
>emb|CAH69985.1| SAMD6 [Homo sapiens] gi|55663434|emb|CAH71295.1| SAMD6 [Homo
sapiens]
Length = 334
Score = 38.5 bits (88), Expect = 0.13
Identities = 25/61 (40%), Positives = 35/61 (56%), Gaps = 3/61 (4%)
Query: 1 MSEKPTSRDQ-YGPPLGSS-RKLPSSITGSSAPVAVTPLKTIK-PPGPSTPTKSSQRPTV 57
+ +KP+ R GP GSS +LP+S G SAPV L+T K PP ++ T S PT+
Sbjct: 141 LEQKPSHRSSPVGPAPGSSPSELPASPAGGSAPVGKQKLETSKRPPSGTSTTSKSTSPTL 200
Query: 58 S 58
+
Sbjct: 201 T 201
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.130 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 416,140,824
Number of Sequences: 2540612
Number of extensions: 18574406
Number of successful extensions: 66165
Number of sequences better than 10.0: 496
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 445
Number of HSP's that attempted gapping in prelim test: 65108
Number of HSP's gapped (non-prelim): 1344
length of query: 212
length of database: 863,360,394
effective HSP length: 122
effective length of query: 90
effective length of database: 553,405,730
effective search space: 49806515700
effective search space used: 49806515700
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0067a.13


