

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0066.12
(163 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
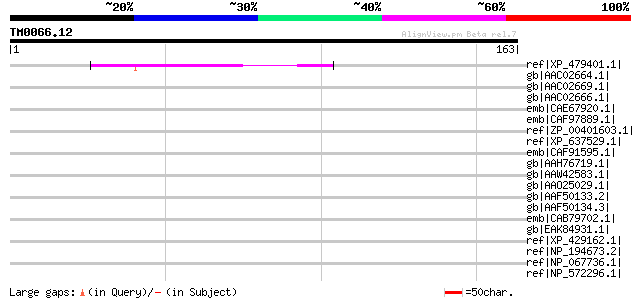
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_479401.1| hypothetical protein [Oryza sativa (japonica cu... 62 4e-09
gb|AAC02664.1| polyprotein [Arabidopsis thaliana] 42 0.006
gb|AAC02669.1| polyprotein [Arabidopsis thaliana] 42 0.006
gb|AAC02666.1| polyprotein [Arabidopsis thaliana] 42 0.006
emb|CAE67920.1| Hypothetical protein CBG13518 [Caenorhabditis br... 36 0.32
emb|CAF97889.1| unnamed protein product [Tetraodon nigroviridis] 36 0.42
ref|ZP_00401603.1| HhH-GPD [Anaeromyxobacter dehalogenans 2CP-C]... 34 1.6
ref|XP_637529.1| hypothetical protein DDB0191406 [Dictyostelium ... 34 1.6
emb|CAF91595.1| unnamed protein product [Tetraodon nigroviridis] 34 1.6
gb|AAH76719.1| LOC398539 protein [Xenopus laevis] 33 2.1
gb|AAW42583.1| hypothetical protein CNC06330 [Cryptococcus neofo... 33 2.7
gb|AAO25029.1| LD16921p [Drosophila melanogaster] 33 2.7
gb|AAF50133.2| CG7958-PB, isoform B [Drosophila melanogaster] gi... 33 2.7
gb|AAF50134.3| CG7958-PA, isoform A [Drosophila melanogaster] gi... 33 2.7
emb|CAB79702.1| putative protein [Arabidopsis thaliana] gi|25407... 33 3.6
gb|EAK84931.1| hypothetical protein UM03901.1 [Ustilago maydis 5... 33 3.6
ref|XP_429162.1| PREDICTED: similar to RIKEN cDNA A230083H22 gen... 33 3.6
ref|NP_194673.2| expressed protein [Arabidopsis thaliana] 33 3.6
ref|NP_067736.1| overlapping protein/movement protein [Chayote m... 32 4.7
ref|NP_572296.1| CG15899-PB [Drosophila melanogaster] gi|2283179... 32 4.7
>ref|XP_479401.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50510134|dbj|BAD31099.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|22775617|dbj|BAC15471.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 501
Score = 62.4 bits (150), Expect = 4e-09
Identities = 37/79 (46%), Positives = 42/79 (52%), Gaps = 18/79 (22%)
Query: 27 DMELPLPTTTSSE-TEPFRLDQAVCSHGLFMMAPNSWDPLSNTLTRPLRLHDQDTDPSSS 85
++ELPL PF L+ AVCSHGLFMMAPN WDP S L RPLRL
Sbjct: 20 ELELPLGGAPPYPGAAPFDLEAAVCSHGLFMMAPNRWDPASRALVRPLRL---------- 69
Query: 86 SPSFIVTVSQRSESIAVRV 104
S R+ S+AVRV
Sbjct: 70 -------ASDRAASVAVRV 81
>gb|AAC02664.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 8/91 (8%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLP-------TTTSSETEPFRLDQAVCSHGLFM 56
L PP +P+CS P SPS + ++ P P + TS T P ++++ H
Sbjct: 784 LTTAPPSSPSCSAPHRSPSQSE-NLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSET 842
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSP 87
S PLS RP Q T P SSSP
Sbjct: 843 GPTGSSPPLSPQPQRPQPQSPQSTSPHSSSP 873
>gb|AAC02669.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 8/91 (8%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLP-------TTTSSETEPFRLDQAVCSHGLFM 56
L PP +P+CS P SPS + ++ P P + TS T P ++++ H
Sbjct: 784 LTTAPPSSPSCSAPHRSPSQSE-NLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSET 842
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSP 87
S PLS RP Q T P SSSP
Sbjct: 843 GPTGSSPPLSPQPQRPQPQSPQSTSPHSSSP 873
>gb|AAC02666.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 8/91 (8%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLP-------TTTSSETEPFRLDQAVCSHGLFM 56
L PP +P+CS P SPS + ++ P P + TS T P ++++ H
Sbjct: 784 LTTAPPSSPSCSAPHRSPSQSE-NLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSET 842
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSP 87
S PLS RP Q T P SSSP
Sbjct: 843 GPTGSSPPLSPQPQRPQPQSPQSTSPHSSSP 873
>emb|CAE67920.1| Hypothetical protein CBG13518 [Caenorhabditis briggsae]
Length = 1032
Score = 36.2 bits (82), Expect = 0.32
Identities = 40/141 (28%), Positives = 58/141 (40%), Gaps = 13/141 (9%)
Query: 3 ELEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSET-EPFRLDQAVCSHGLFMMAPNS 61
+L+K C P S P +S ST P P+ TSS T +L + V HG+ + S
Sbjct: 506 QLDKKAMC-PTSSSPSTSRGSTTTSQASPAPSHTSSSTSSSVQLRENVARHGVSLKETMS 564
Query: 62 WDPLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQ-----RSESIAVRVHH----GTHLLS 112
+ + L + + P + +PS + SQ RS S HH G S
Sbjct: 565 LRDVGPLSRDAISLRETVSPPITRAPSQPPSYSQPRPPPRSVSSVNNHHHSLPNGDQFSS 624
Query: 113 PHEVRALMVPTS--LTASLNS 131
R + PT+ LT+S S
Sbjct: 625 LDRTRGSVTPTAPPLTSSAAS 645
>emb|CAF97889.1| unnamed protein product [Tetraodon nigroviridis]
Length = 828
Score = 35.8 bits (81), Expect = 0.42
Identities = 36/133 (27%), Positives = 51/133 (38%), Gaps = 16/133 (12%)
Query: 15 SKPESSPSSTWFDMELPLPTTTSSETEPFRLDQA---------VCSHGLFMMAPNSWDPL 65
S P+S+ S DM+L + + EP++LD A H F P S D
Sbjct: 544 SSPDSNASYAVEDMDL---NGSYYQKEPYQLDVARERALTISSPFRHSPFSYKPKSLDQN 600
Query: 66 SNTLTRPLRL----HDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALMV 121
NT RPL L H P SS S ++ + S H ++LL
Sbjct: 601 QNTRPRPLSLLQHSHSNSLPPKLSSLSLSLSPTPPSSPSCASPHSSSYLLPRPNTPGSTS 660
Query: 122 PTSLTASLNSWVF 134
P+ S +S+ F
Sbjct: 661 PSPSVRSSSSFNF 673
>ref|ZP_00401603.1| HhH-GPD [Anaeromyxobacter dehalogenans 2CP-C]
gi|66776651|gb|EAL77761.1| HhH-GPD [Anaeromyxobacter
dehalogenans 2CP-C]
Length = 309
Score = 33.9 bits (76), Expect = 1.6
Identities = 17/35 (48%), Positives = 19/35 (53%)
Query: 41 EPFRLDQAVCSHGLFMMAPNSWDPLSNTLTRPLRL 75
EPF L V SHG + + P WD L RPLRL
Sbjct: 5 EPFDLVLTVRSHGWYDLPPWRWDEARRVLGRPLRL 39
>ref|XP_637529.1| hypothetical protein DDB0191406 [Dictyostelium discoideum]
gi|60465942|gb|EAL64010.1| hypothetical protein
DDB0191406 [Dictyostelium discoideum]
gi|5834504|dbj|BAA84094.1| lagC2 [Dictyostelium
discoideum]
Length = 895
Score = 33.9 bits (76), Expect = 1.6
Identities = 26/99 (26%), Positives = 37/99 (37%), Gaps = 6/99 (6%)
Query: 10 CNPNCSKPESSPSSTWF-DMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSNT 68
C S P STW D P P P V GLF+ + L+N+
Sbjct: 97 CTRTVSDPTEDCQSTWITDNAFPEPLNVKFSGYPSTSGGDVVFTGLFLRLAGGPNTLANS 156
Query: 69 LTRP-----LRLHDQDTDPSSSSPSFIVTVSQRSESIAV 102
T P +++H +DP+ + +F T S I V
Sbjct: 157 FTNPSKTFLIKVHGNFSDPNFNCNNFTATFPPSSGYIKV 195
>emb|CAF91595.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1146
Score = 33.9 bits (76), Expect = 1.6
Identities = 27/108 (25%), Positives = 49/108 (45%), Gaps = 12/108 (11%)
Query: 10 CNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSNTL 69
C+ CS+P P S P P+++S+ E S G + P+S +S+ +
Sbjct: 813 CSCCCSEPPQPPPSPGLG---PYPSSSSAIKER-------PSDGSLPVPPSSCSEISSEI 862
Query: 70 TRPLRLHDQDTDPSSSSP--SFIVTVSQRSESIAVRVHHGTHLLSPHE 115
+ + + SS P S +V + +++ + +HH TH LS H+
Sbjct: 863 STSEMSSEVGSTASSDEPPQSSLVLLHEQAHHPHLHLHHQTHPLSTHQ 910
>gb|AAH76719.1| LOC398539 protein [Xenopus laevis]
Length = 2414
Score = 33.5 bits (75), Expect = 2.1
Identities = 31/115 (26%), Positives = 54/115 (46%), Gaps = 19/115 (16%)
Query: 3 ELEKPPPCNPNCSK----PESSPSSTWFDMELPLPTTTSSETEPFR-LDQAVCS-HGLFM 56
E+E+P P +P C + P+++PS+T +P TT E + + VC + +
Sbjct: 2135 EIEEPNPFDPCCPRYRCEPKTTPSTT-------MPITTKQECDNVKCYKDVVCGPNDNEV 2187
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQ--RSESIAVRVHHGTH 109
PN +DP R + T PS+ + S I+T ++ S+ I V++ H
Sbjct: 2188 EEPNPFDP----CCPKYRCEPKSTTPSTITTSTIITTTKIDCSKRICVKISCRLH 2238
>gb|AAW42583.1| hypothetical protein CNC06330 [Cryptococcus neoformans var.
neoformans JEC21] gi|50259275|gb|EAL21948.1|
hypothetical protein CNBC0880 [Cryptococcus neoformans
var. neoformans B-3501A] gi|58265468|ref|XP_569890.1|
hypothetical protein CNC06330 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 438
Score = 33.1 bits (74), Expect = 2.7
Identities = 28/111 (25%), Positives = 44/111 (39%), Gaps = 8/111 (7%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWD 63
L +PP P + P SSP + P +T+S+ T Q+V + M P S
Sbjct: 52 LSRPPNSGPASTAPSSSPRFQPYSRPQPQRSTSSNST------QSVQQNNNMGMPPPSLP 105
Query: 64 PLSNTLTRPLRLH-DQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSP 113
P SN++ P ++ P S + + S + H HL +P
Sbjct: 106 P-SNSIRPPTQISPSHQPSPLHQSALSFLPPQELSRGYSESTHSPNHLSNP 155
>gb|AAO25029.1| LD16921p [Drosophila melanogaster]
Length = 1109
Score = 33.1 bits (74), Expect = 2.7
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 13/136 (9%)
Query: 8 PPCNPNCSKPESSPS-------STWFDMELPLPTTTSSETEPFRL--DQAVCSHGLFMMA 58
PP P CS P SP+ +M L P PFRL + +V +H +F +
Sbjct: 507 PPLTPACSVPYVSPNPDIKPPMDNSEEMRLTFPVRDGIILAPFRLLHNLSVSNH-VFHLK 565
Query: 59 PNSWDPL--SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAV-RVHHGTHLLSPHE 115
N ++ L N L L+ QD +++ VTVS + + + R + L P
Sbjct: 566 QNVYNTLMCRNDLELQLKCFHQDDRQMNTNWPHTVTVSANATPLNIERSEKNSTALRPLY 625
Query: 116 VRALMVPTSLTASLNS 131
++A+ P T L +
Sbjct: 626 LKAVCQPGRNTLQLTA 641
>gb|AAF50133.2| CG7958-PB, isoform B [Drosophila melanogaster]
gi|45551544|ref|NP_729629.2| CG7958-PB, isoform B
[Drosophila melanogaster]
Length = 1149
Score = 33.1 bits (74), Expect = 2.7
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 13/136 (9%)
Query: 8 PPCNPNCSKPESSPS-------STWFDMELPLPTTTSSETEPFRL--DQAVCSHGLFMMA 58
PP P CS P SP+ +M L P PFRL + +V +H +F +
Sbjct: 547 PPLTPACSVPYVSPNPDIKPPMDNSEEMRLTFPVRDGIILAPFRLLHNLSVSNH-VFHLK 605
Query: 59 PNSWDPL--SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAV-RVHHGTHLLSPHE 115
N ++ L N L L+ QD +++ VTVS + + + R + L P
Sbjct: 606 QNVYNTLMCRNDLELQLKCFHQDDRQMNTNWPHTVTVSANATPLNIERSEKNSTALRPLY 665
Query: 116 VRALMVPTSLTASLNS 131
++A+ P T L +
Sbjct: 666 LKAVCQPGRNTLQLTA 681
>gb|AAF50134.3| CG7958-PA, isoform A [Drosophila melanogaster]
gi|45550589|ref|NP_648412.2| CG7958-PA, isoform A
[Drosophila melanogaster]
Length = 1135
Score = 33.1 bits (74), Expect = 2.7
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 13/136 (9%)
Query: 8 PPCNPNCSKPESSPS-------STWFDMELPLPTTTSSETEPFRL--DQAVCSHGLFMMA 58
PP P CS P SP+ +M L P PFRL + +V +H +F +
Sbjct: 533 PPLTPACSVPYVSPNPDIKPPMDNSEEMRLTFPVRDGIILAPFRLLHNLSVSNH-VFHLK 591
Query: 59 PNSWDPL--SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAV-RVHHGTHLLSPHE 115
N ++ L N L L+ QD +++ VTVS + + + R + L P
Sbjct: 592 QNVYNTLMCRNDLELQLKCFHQDDRQMNTNWPHTVTVSANATPLNIERSEKNSTALRPLY 651
Query: 116 VRALMVPTSLTASLNS 131
++A+ P T L +
Sbjct: 652 LKAVCQPGRNTLQLTA 667
>emb|CAB79702.1| putative protein [Arabidopsis thaliana] gi|25407614|pir||E85343
hypothetical protein AT4g29440 [imported] - Arabidopsis
thaliana
Length = 1071
Score = 32.7 bits (73), Expect = 3.6
Identities = 32/104 (30%), Positives = 43/104 (40%), Gaps = 14/104 (13%)
Query: 12 PNCSKPESSPSSTWFDMELPLPTT-------TSSETEPFRLDQAVCSHGLFMMAPNSWDP 64
P+ P+SS S DMELP + T S T P L V L P
Sbjct: 730 PSSIPPDSSSSDDESDMELPKRVSFRYQEKRTESRTRPTHLHSGVSHKDLEEEIPTR--- 786
Query: 65 LSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGT 108
++T ++ R H T P+S+S S+ T+S E VH T
Sbjct: 787 -ASTRSQDRRTH--KTTPASASASYFHTMSSDDED-EKEVHRDT 826
>gb|EAK84931.1| hypothetical protein UM03901.1 [Ustilago maydis 521]
gi|49074818|ref|XP_401516.1| hypothetical protein
UM03901.1 [Ustilago maydis 521]
Length = 1444
Score = 32.7 bits (73), Expect = 3.6
Identities = 16/49 (32%), Positives = 24/49 (48%)
Query: 64 PLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLS 112
PL+ + TRP LH TDP S+ I T R + + ++ H L+
Sbjct: 889 PLATSSTRPAGLHRSSTDPQLSACKPIATQPPRQQQLHAQIQASEHQLA 937
>ref|XP_429162.1| PREDICTED: similar to RIKEN cDNA A230083H22 gene [Gallus gallus]
Length = 3148
Score = 32.7 bits (73), Expect = 3.6
Identities = 21/72 (29%), Positives = 33/72 (45%)
Query: 64 PLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALMVPT 123
PLS + + L+ D +P SS +FI + SI V L+SP A ++ +
Sbjct: 390 PLSQGSSGIMELYGSDVEPQPSSVNFIENPQDLNGSIQAHVDLNVDLVSPDSGLATIISS 449
Query: 124 SLTASLNSWVFI 135
+S S VF+
Sbjct: 450 RSRSSKESSVFL 461
>ref|NP_194673.2| expressed protein [Arabidopsis thaliana]
Length = 1090
Score = 32.7 bits (73), Expect = 3.6
Identities = 32/104 (30%), Positives = 43/104 (40%), Gaps = 14/104 (13%)
Query: 12 PNCSKPESSPSSTWFDMELPLPTT-------TSSETEPFRLDQAVCSHGLFMMAPNSWDP 64
P+ P+SS S DMELP + T S T P L V L P
Sbjct: 749 PSSIPPDSSSSDDESDMELPKRVSFRYQEKRTESRTRPTHLHSGVSHKDLEEEIPTR--- 805
Query: 65 LSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGT 108
++T ++ R H T P+S+S S+ T+S E VH T
Sbjct: 806 -ASTRSQDRRTH--KTTPASASASYFHTMSSDDED-EKEVHRDT 845
>ref|NP_067736.1| overlapping protein/movement protein [Chayote mosaic virus]
gi|6456717|gb|AAF09239.1| overlapping protein [chayote
mosaic tymovirus]
Length = 680
Score = 32.3 bits (72), Expect = 4.7
Identities = 30/101 (29%), Positives = 39/101 (37%), Gaps = 21/101 (20%)
Query: 7 PPPCNPNCSKPESSPSSTWFD-MELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPL 65
P P P S+P S PSS+ LPL + + + P Q C PL
Sbjct: 520 PHPSIPTMSQPPSLPSSSSGPPRHLPLRSPSPPVSAPMEAPQLPCVES---------PPL 570
Query: 66 SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHH 106
S L+ P SSP IV +RS S++ HH
Sbjct: 571 SPPLSHP-----------PSSPELIVYRPRRSASLSPHSHH 600
>ref|NP_572296.1| CG15899-PB [Drosophila melanogaster] gi|22831794|gb|AAF46127.2|
CG15899-PB [Drosophila melanogaster]
Length = 2893
Score = 32.3 bits (72), Expect = 4.7
Identities = 24/88 (27%), Positives = 36/88 (40%), Gaps = 8/88 (9%)
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPF----RLDQAVCSHGLFMMAPNSW 62
PPP + P S+P+ M +P+P PF +DQ L + P S
Sbjct: 2707 PPPLSLPIVTPTSTPTPLQLPMPMPMPMAHPPPRRPFGWSQSVDQGCLRSNLLLSVPRSM 2766
Query: 63 DPLSNT-LTRPL---RLHDQDTDPSSSS 86
P S + T+ L + D+D D +S
Sbjct: 2767 PPRSRSGSTKQLFKQQALDEDADMDENS 2794
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.132 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 299,119,378
Number of Sequences: 2540612
Number of extensions: 11529942
Number of successful extensions: 38551
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 38522
Number of HSP's gapped (non-prelim): 47
length of query: 163
length of database: 863,360,394
effective HSP length: 118
effective length of query: 45
effective length of database: 563,568,178
effective search space: 25360568010
effective search space used: 25360568010
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0066.12


