

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.14
(267 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
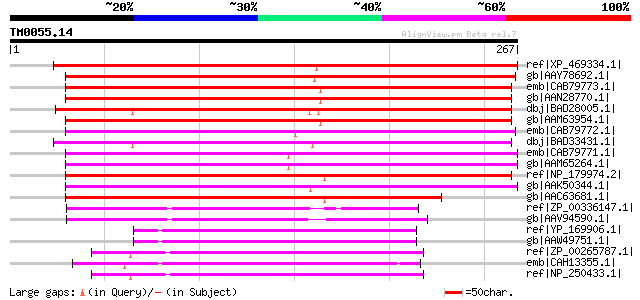
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_469334.1| putative glutamine synthetase [Oryza sativa] gi... 246 6e-64
gb|AAY78692.1| putative defense-related protein [Arabidopsis tha... 232 9e-60
emb|CAB79773.1| putative protein [Arabidopsis thaliana] gi|15234... 218 1e-55
gb|AAN28770.1| At4g30550/F17I23_110 [Arabidopsis thaliana] gi|17... 218 2e-55
dbj|BAD28005.1| putative glutamine amidotransferase class-I doma... 215 1e-54
gb|AAM63954.1| unknown [Arabidopsis thaliana] 213 4e-54
emb|CAB79772.1| putative protein [Arabidopsis thaliana] gi|23296... 208 1e-52
dbj|BAD33431.1| glutamine synthetase-like protein [Oryza sativa ... 208 1e-52
emb|CAB79771.1| putative protein [Arabidopsis thaliana] gi|20147... 204 2e-51
gb|AAM65264.1| defense-related protein [Arabidopsis thaliana] 204 3e-51
ref|NP_179974.2| defense-related protein, putative [Arabidopsis ... 201 2e-50
gb|AAK50344.1| defense-related protein [Brassica carinata] 199 9e-50
gb|AAC63681.1| unknown protein [Arabidopsis thaliana] gi|2541217... 182 8e-45
ref|ZP_00336147.1| COG0518: GMP synthase - Glutamine amidotransf... 117 4e-25
gb|AAV94590.1| glutamine amidotransferase, class I [Silicibacter... 116 6e-25
ref|YP_169906.1| glutamine amidotransferase, class I [Francisell... 110 4e-23
gb|AAW49751.1| hypothetical protein FTT0909 [synthetic construct] 110 4e-23
ref|ZP_00265787.1| COG0518: GMP synthase - Glutamine amidotransf... 110 5e-23
emb|CAH13355.1| hypothetical protein [Legionella pneumophila str... 109 9e-23
ref|NP_250433.1| probable amidotransferase [Pseudomonas aerugino... 108 2e-22
>ref|XP_469334.1| putative glutamine synthetase [Oryza sativa]
gi|13174241|gb|AAK14415.1| putative glutamine synthetase
[Oryza sativa]
Length = 293
Score = 246 bits (627), Expect = 6e-64
Identities = 118/247 (47%), Positives = 161/247 (64%), Gaps = 3/247 (1%)
Query: 24 GGGGKGKRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELP 83
GG G G YA+L CGEDSEY+ K +GG F F +LAE+GERW +Y+ V GE P +E
Sbjct: 25 GGVGVGGSYAVLQCGEDSEYVRKAYGGYFEVFRALLAEDGERWRVYRAVRGELPGEEEAA 84
Query: 84 LYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSS 143
DGFVI+GSC DAHA+ PWI L+ L+ ++ K+ILG+CFGHQ+L R LGGK GRS
Sbjct: 85 GIDGFVISGSCSDAHADDPWIVALVDLIRRQNAAGKRILGVCFGHQVLCRALGGKTGRSK 144
Query: 144 TGWDIGVTTMKLASSSS---IALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMF 200
GWDIGV + ++ + + LP + I + H+DEV LPP+AE++ S+ TG+EMF
Sbjct: 145 KGWDIGVNCIHPTAAMARLFSPIKLPVHMPIIEFHQDEVWELPPQAEVLARSDMTGVEMF 204
Query: 201 RYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCV 260
R GD +G+QGHPE++ DIL DRL +L+ + K L++P+ + WK++C
Sbjct: 205 RLGDRAMGVQGHPEYSKDILMSIADRLLRNDLILDHQVDKAKASFDLRQPDKDLWKKVCR 264
Query: 261 NFLKDRL 267
FLK RL
Sbjct: 265 GFLKGRL 271
>gb|AAY78692.1| putative defense-related protein [Arabidopsis thaliana]
gi|3738324|gb|AAC63665.1| unknown protein [Arabidopsis
thaliana] gi|25412179|pir||A84631 hypothetical protein
At2g23970 [imported] - Arabidopsis thaliana
gi|15224079|ref|NP_179975.1| defense-related protein,
putative [Arabidopsis thaliana]
Length = 251
Score = 232 bits (591), Expect = 9e-60
Identities = 110/242 (45%), Positives = 156/242 (64%), Gaps = 5/242 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+AL + DS ++ K +GG F F E+GE+WDL++V++GEFP +L YDGFV
Sbjct: 6 KRFALFLATSDSTFVKKAYGGYFNVFVSTFGEDGEQWDLFRVIDGEFPDDKDLDKYDGFV 65
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS +DA + WI L SL LD + KK+LGICFGHQIL R+ GGKVGR+S G D+G
Sbjct: 66 ISGSLNDAFGDDDWIVKLCSLCQKLDDMKKKVLGICFGHQILSRIKGGKVGRASRGLDMG 125
Query: 150 VTTMKLASSS-----SIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
+ ++ + + + +P L+I KCH+DEV LP A ++ +S+K +EM YG+
Sbjct: 126 LRSITMVTDAVKPGGYFGSQIPKSLAIIKCHQDEVLELPESATLLAYSDKYNVEMCSYGN 185
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
H+LGIQGHPE+ +ILF IDR+ + L+++ FA K EP+ + W+ LC NFLK
Sbjct: 186 HLLGIQGHPEYNKEILFEIIDRVVNLKLMEQDFADKAKATMENAEPDRKQWQTLCKNFLK 245
Query: 265 DR 266
R
Sbjct: 246 GR 247
>emb|CAB79773.1| putative protein [Arabidopsis thaliana]
gi|15234767|ref|NP_194784.1| glutamine amidotransferase
class-I domain-containing protein [Arabidopsis thaliana]
gi|25407659|pir||D85357 hypothetical protein AT4g30550
[imported] - Arabidopsis thaliana
Length = 249
Score = 218 bits (556), Expect = 1e-55
Identities = 105/241 (43%), Positives = 152/241 (62%), Gaps = 6/241 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+AL + DSE++ K +GG F F EEGE+WDL++V++G+FP ++L YDGFV
Sbjct: 8 KRFALFLATCDSEFVKKTYGGYFNVFVSTFGEEGEQWDLFRVIDGQFPDENDLDKYDGFV 67
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L + LD + KK+LGICFGHQI+ RV GGK+GR+ G D+G
Sbjct: 68 ISGSPHDAFGDADWIVKLCEVCQKLDHMKKKVLGICFGHQIITRVKGGKIGRALKGADMG 127
Query: 150 VTTMKLASSSSIA------LNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYG 203
+ ++ +A + + +P+ L+I KCH+DEV LP A ++ SE +EMF G
Sbjct: 128 LRSITIAKDNEKLRGYFGDVEVPASLAIIKCHQDEVLELPESATLLASSEVCNVEMFSIG 187
Query: 204 DHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFL 263
DH IQGHPE+ +ILF +DR+ + L+++ FA K +P+ W++LC NFL
Sbjct: 188 DHFFCIQGHPEYNKEILFEIVDRVLNMKLMEQEFADKAKSTMETAQPDRILWQKLCKNFL 247
Query: 264 K 264
K
Sbjct: 248 K 248
>gb|AAN28770.1| At4g30550/F17I23_110 [Arabidopsis thaliana]
gi|17381294|gb|AAL36065.1| AT4g30550/F17I23_110
[Arabidopsis thaliana]
Length = 249
Score = 218 bits (554), Expect = 2e-55
Identities = 105/241 (43%), Positives = 152/241 (62%), Gaps = 6/241 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+AL + DSE++ K +GG F F EEGE+WDL++V++G+FP ++L YDGFV
Sbjct: 8 KRFALFLATCDSEFVKKTYGGYFNVFVSTFGEEGEQWDLFRVIDGQFPDENDLDKYDGFV 67
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L + LD + KK+LGICFGHQI+ RV GGK+GR+ G D+G
Sbjct: 68 ISGSPHDAFGDADWIVKLCEVCQKLDHMKKKVLGICFGHQIITRVKGGKIGRALKGADMG 127
Query: 150 VTTMKLASSSSIA------LNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYG 203
+ ++ +A + + +P+ L+I KCH+DEV LP A ++ SE +EMF G
Sbjct: 128 LRSITIAKDNEKLRGYFGDVEVPAYLAIIKCHQDEVLELPESATLLASSEVCNVEMFSIG 187
Query: 204 DHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFL 263
DH IQGHPE+ +ILF +DR+ + L+++ FA K +P+ W++LC NFL
Sbjct: 188 DHFFCIQGHPEYNKEILFEIVDRVLNMKLMEQEFADKAKSTMETAQPDRILWQKLCKNFL 247
Query: 264 K 264
K
Sbjct: 248 K 248
>dbj|BAD28005.1| putative glutamine amidotransferase class-I domain-containing
protein [Oryza sativa (japonica cultivar-group)]
Length = 299
Score = 215 bits (547), Expect = 1e-54
Identities = 108/251 (43%), Positives = 152/251 (60%), Gaps = 11/251 (4%)
Query: 25 GGGKGKRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEG-----ERWDLYKVVNGEFPRH 79
GG G+RYALL+ DSEY K +GG F L G ERWD ++V++GEFP
Sbjct: 10 GGVGGRRYALLLALNDSEYARKVYGGYGNVFVSALGGGGGGGEEERWDCFRVIDGEFPAA 69
Query: 80 DELPLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKV 139
+E+ Y+GFV++GS HDA+ + WI L SL+ L ++ K+ILGICFGHQ+L R LGG++
Sbjct: 70 EEVGRYEGFVVSGSPHDAYGDERWILRLCSLLRALHAMGKRILGICFGHQVLCRALGGRI 129
Query: 140 GRSSTGWDIGVTTMKLA---SSSSI---ALNLPSELSIFKCHRDEVRNLPPKAEIIGWSE 193
G++ +GW+IGV M S + +P SI + H+DEV +PP ++ +S+
Sbjct: 130 GKARSGWNIGVKKMTFVRDFEGSKLFGDLKEIPQSASIIEVHQDEVLEVPPMGRVLAYSD 189
Query: 194 KTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTE 253
KT +EMF GD++LGIQGHPE+T DIL + IDRL + N + + + EP+
Sbjct: 190 KTPVEMFAVGDNVLGIQGHPEYTSDILLNLIDRLVNNNTITSGIGEEARRTVEASEPDRR 249
Query: 254 AWKRLCVNFLK 264
W LC FLK
Sbjct: 250 FWTGLCKGFLK 260
>gb|AAM63954.1| unknown [Arabidopsis thaliana]
Length = 249
Score = 213 bits (542), Expect = 4e-54
Identities = 104/241 (43%), Positives = 151/241 (62%), Gaps = 6/241 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+AL + DSE++ K +GG F F EEGE+ DL++V++G+FP ++L YDGFV
Sbjct: 8 KRFALFLATCDSEFVKKTYGGYFNVFVSTFGEEGEQSDLFRVIDGQFPDENDLDKYDGFV 67
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L + LD + KK+LGICFGHQI+ RV GGK+GR+ G D+G
Sbjct: 68 ISGSPHDAFGDADWIVKLCEVCQKLDHMKKKVLGICFGHQIITRVKGGKIGRALKGADMG 127
Query: 150 VTTMKLASSSSIA------LNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYG 203
+ ++ +A + + +P+ L+I KCH+DEV LP A ++ SE +EMF G
Sbjct: 128 LRSITIAKDNEKLRGYFGDVEVPASLAIIKCHQDEVLELPESATLLASSEVCNVEMFSIG 187
Query: 204 DHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFL 263
DH IQGHPE+ +ILF +DR+ + L+++ FA K +P+ W++LC NFL
Sbjct: 188 DHFFCIQGHPEYNKEILFEIVDRVLNMKLMEQEFADKAKSTMETAQPDRILWQKLCKNFL 247
Query: 264 K 264
K
Sbjct: 248 K 248
>emb|CAB79772.1| putative protein [Arabidopsis thaliana] gi|23296437|gb|AAN13059.1|
unknown protein [Arabidopsis thaliana]
gi|15234765|ref|NP_194783.1| glutamine amidotransferase
class-I domain-containing protein [Arabidopsis thaliana]
gi|25407657|pir||C85357 hypothetical protein AT4g30540
[imported] - Arabidopsis thaliana
Length = 248
Score = 208 bits (530), Expect = 1e-52
Identities = 107/240 (44%), Positives = 145/240 (59%), Gaps = 3/240 (1%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
+RYAL DSE++ + +GG F F +EGE+WDL++V++GEFPR ++L Y+GFV
Sbjct: 6 RRYALFQATPDSEFVKEMYGGYFNVFVSAFGDEGEQWDLFRVIDGEFPRDEDLEKYEGFV 65
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA WI +L S+ LD + KKILGICFGHQI+ RV GGKVGR+ G DIG
Sbjct: 66 ISGSLHDAFTEEDWIIELCSVCKKLDVMKKKILGICFGHQIICRVRGGKVGRARKGPDIG 125
Query: 150 ---VTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHM 206
+T ++ + LSI +CHRDEV P A +IG+S+K +E+F DH+
Sbjct: 126 LGNITIVQDVIKPGDYFDQIESLSIIQCHRDEVLEPPESARVIGFSDKCDVEIFSVEDHL 185
Query: 207 LGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLKDR 266
L QGHPE+ +IL IDR+ ++E K EP+T+ LC NFLK R
Sbjct: 186 LCFQGHPEYNKEILLEIIDRVHKIKFVEEEILEKAKDSIKKFEPDTQRLHMLCKNFLKGR 245
>dbj|BAD33431.1| glutamine synthetase-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 272
Score = 208 bits (530), Expect = 1e-52
Identities = 110/259 (42%), Positives = 154/259 (58%), Gaps = 18/259 (6%)
Query: 24 GGGGKGKRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEG-------ERWDLYKVVNGEF 76
GG + +RYALL+ DS+Y+ K +GG F R ++G E WD+++ V+GE
Sbjct: 10 GGVRRRRRYALLLAARDSDYVRKVYGGYLEVFIRAFGDDGDVGDGGGEEWDMFRAVDGEL 69
Query: 77 PRHDELPLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLG 136
P DE+ YDGFVI+GS HDA+A+ WI L LV L ++ K++LGICFGHQ++ R LG
Sbjct: 70 PGADEVDGYDGFVISGSPHDAYADDLWILRLCLLVRDLVAMRKRLLGICFGHQVICRALG 129
Query: 137 GKVGRSSTGWDIGVTTMKLASS-----------SSIALNLPSELSIFKCHRDEVRNLPPK 185
G+VG++ GWDIG+ + +A S I I + H+DEV LP
Sbjct: 130 GRVGKARGGWDIGIREVAMAESLPPYRFLDDALQGITAAAAPYAKITEVHQDEVWELPAG 189
Query: 186 AEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKA 245
AE++ S KTG+EMF GD +LGIQGHPE+T DIL + +DRL+S + A V+ +
Sbjct: 190 AEVLASSSKTGVEMFCAGDRVLGIQGHPEYTADILLNLVDRLSSAGSITMAVAEGVRRQL 249
Query: 246 ALQEPNTEAWKRLCVNFLK 264
P+ E W +LC +FLK
Sbjct: 250 EDTGPDREFWIKLCKSFLK 268
>emb|CAB79771.1| putative protein [Arabidopsis thaliana] gi|20147269|gb|AAM10348.1|
AT4g30530/F17I23_130 [Arabidopsis thaliana]
gi|15234763|ref|NP_194782.1| defense-related protein,
putative [Arabidopsis thaliana]
gi|15294152|gb|AAK95253.1| AT4g30530/F17I23_130
[Arabidopsis thaliana] gi|25407655|pir||B85357
hypothetical protein AT4g30530 [imported] - Arabidopsis
thaliana
Length = 250
Score = 204 bits (520), Expect = 2e-51
Identities = 101/243 (41%), Positives = 144/243 (58%), Gaps = 5/243 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KRYAL + DSE++ K +GG F +EGE WD ++VV+GEFP +L YDGFV
Sbjct: 5 KRYALFLATLDSEFVKKTYGGYHNVFVTTFGDEGEHWDSFRVVSGEFPDEKDLEKYDGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTG---- 145
I+GS HDA N WI L +V +D + KKILGICFGHQI+ RV GG VGR+ G
Sbjct: 65 ISGSSHDAFENDDWILKLCDIVKKIDEMKKKILGICFGHQIIARVRGGTVGRAKKGPELK 124
Query: 146 -WDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
DI + + S +P ++I KCH+DEV LP A+++ +S+ +EM+ D
Sbjct: 125 LGDITIVKDAITPGSYFGNEIPDSIAIIKCHQDEVLVLPETAKVLAYSKNYEVEMYSIED 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
H+ IQGHPE+ +ILF +DR+ + +++ FA K + + + W+ +C NFLK
Sbjct: 185 HLFCIQGHPEYNKEILFEIVDRVLALGYVKQEFADAAKATMENRGADRKLWETICKNFLK 244
Query: 265 DRL 267
R+
Sbjct: 245 GRV 247
>gb|AAM65264.1| defense-related protein [Arabidopsis thaliana]
Length = 250
Score = 204 bits (518), Expect = 3e-51
Identities = 101/243 (41%), Positives = 144/243 (58%), Gaps = 5/243 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KRYAL + DSE++ K +GG F +EGE WD ++VV+GEFP +L YDGFV
Sbjct: 5 KRYALFLATLDSEFVKKTYGGYHNVFVTTFGDEGEHWDSFRVVSGEFPDEKDLEKYDGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTG---- 145
I+GS HDA N WI L +V +D + KKILGICFGHQI+ RV GG VGR+ G
Sbjct: 65 ISGSSHDAFENDDWILKLCDIVKKIDEMKKKILGICFGHQIIARVRGGTVGRAKKGPELK 124
Query: 146 -WDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
DI + + S +P ++I KCH+DEV LP A+++ +S+ +EM+ D
Sbjct: 125 LGDITIVKDAITPGSYFGNEIPDSIAIIKCHQDEVLVLPETAKVLAYSKNYEVEMYSIED 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
H+ IQGHPE+ +ILF +DR+ + +++ FA K + + + W+ +C NFLK
Sbjct: 185 HLFCIQGHPEYNKEILFEIVDRVLALGYVKQEFADAAKATMENRGADRKLWETICKNFLK 244
Query: 265 DRL 267
R+
Sbjct: 245 VRV 247
>ref|NP_179974.2| defense-related protein, putative [Arabidopsis thaliana]
Length = 251
Score = 201 bits (511), Expect = 2e-50
Identities = 97/240 (40%), Positives = 146/240 (60%), Gaps = 5/240 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
K+Y L + DSE+ K +GG F +L +EGE+WD ++VV+GEFP +L Y+GFV
Sbjct: 5 KKYLLFLATPDSEFAKKTYGGYHNVFVSLLGDEGEQWDSFRVVDGEFPEEKDLEKYEGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L ++ LD +NKK+LGICFGHQ++ R GGKV R+ G ++
Sbjct: 65 ISGSSHDAFQDTDWILKLCDIIKKLDDMNKKVLGICFGHQLIARAKGGKVARARKGPELC 124
Query: 150 VTTMKLASSSSIALN-----LPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
+ + + + + N +P+ L I KCH+DEV LP A+++ +S +EM+ D
Sbjct: 125 LGNITIVKEAVMPENYFGEEVPANLRIIKCHQDEVLELPENAKLLAYSSMYEVEMYSIKD 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
+ L IQGHPE+ DILF IDR+ + +++ FA K E + + W+++C NFLK
Sbjct: 185 NFLCIQGHPEYNRDILFDIIDRVLAGGHIKQNFAETSKATMEKNEADRKFWQKICKNFLK 244
>gb|AAK50344.1| defense-related protein [Brassica carinata]
Length = 250
Score = 199 bits (505), Expect = 9e-50
Identities = 100/243 (41%), Positives = 143/243 (58%), Gaps = 5/243 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
K++AL + DSE++ K++GG F +EGE WD ++VV GEFP +L YDGFV
Sbjct: 5 KKFALFLATPDSEFVKKEYGGYHNVFVSTFGDEGEHWDSFRVVEGEFPDEKDLDKYDGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HD+ N PWI L +V LD KKILGICFGHQI+ RV GG VGR+ G ++
Sbjct: 65 ISGSSHDSFENDPWILRLCEIVKILDEKKKKILGICFGHQIIARVRGGTVGRARKGPELK 124
Query: 150 VTTMKLAS-----SSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
+T + + S +P ++I K H+DEV LP A+++ +SEK +EMF D
Sbjct: 125 LTDITIVKDAIKPGSFFGNEIPDSIAILKLHQDEVLVLPESAKVLAYSEKYEVEMFSIED 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
H+ IQGHPE+ +IL +DR+ ++E FA K + + + + +C NFLK
Sbjct: 185 HLFCIQGHPEYNREILHEIVDRVLRLGFIKEDFADAAKASMENRGADRKLLETICKNFLK 244
Query: 265 DRL 267
R+
Sbjct: 245 GRV 247
>gb|AAC63681.1| unknown protein [Arabidopsis thaliana] gi|25412177|pir||H84630
hypothetical protein At2g23960 [imported] - Arabidopsis
thaliana
Length = 217
Score = 182 bits (462), Expect = 8e-45
Identities = 87/203 (42%), Positives = 127/203 (61%), Gaps = 5/203 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
K+Y L + DSE+ K +GG F +L +EGE+WD ++VV+GEFP +L Y+GFV
Sbjct: 5 KKYLLFLATPDSEFAKKTYGGYHNVFVSLLGDEGEQWDSFRVVDGEFPEEKDLEKYEGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L ++ LD +NKK+LGICFGHQ++ R GGKV R+ G ++
Sbjct: 65 ISGSSHDAFQDTDWILKLCDIIKKLDDMNKKVLGICFGHQLIARAKGGKVARARKGPELC 124
Query: 150 VTTMKLASSSSIALN-----LPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
+ + + + + N +P+ L I KCH+DEV LP A+++ +S +EM+ D
Sbjct: 125 LGNITIVKEAVMPENYFGEEVPANLRIIKCHQDEVLELPENAKLLAYSSMYEVEMYSIKD 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRL 227
+ L IQGHPE+ DILF IDR+
Sbjct: 185 NFLCIQGHPEYNRDILFDIIDRV 207
>ref|ZP_00336147.1| COG0518: GMP synthase - Glutamine amidotransferase domain
[Silicibacter sp. TM1040]
Length = 226
Score = 117 bits (292), Expect = 4e-25
Identities = 67/185 (36%), Positives = 98/185 (52%), Gaps = 11/185 (5%)
Query: 31 RYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFVI 90
+ +L G + L G F ++LA G +D Y V++G FP E DG++I
Sbjct: 2 KIGILQTGHAPDELKPTSGNYDAMFRKLLAGHGFSFDTYPVLDGVFPEGAEAA--DGWLI 59
Query: 91 TGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGV 150
TGS H A+ +H WI L L+ + + ++GICFGHQI+ + LGGKV + S GW +G
Sbjct: 60 TGSKHGAYEDHDWIPPLEDLIRQIHARKMPLVGICFGHQIIAQALGGKVEKFSGGWSVGH 119
Query: 151 TTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQ 210
T +L + P EL+ + H+D+V P +A +IG S+ YGDH+ Q
Sbjct: 120 TQYRLHGA-------PVELNAW--HQDQVVTRPAEARVIGESDFCANAFLAYGDHIWTSQ 170
Query: 211 GHPEF 215
HPEF
Sbjct: 171 PHPEF 175
>gb|AAV94590.1| glutamine amidotransferase, class I [Silicibacter pomeroyi DSS-3]
gi|56696187|ref|YP_166544.1| glutamine amidotransferase,
class I [Silicibacter pomeroyi DSS-3]
Length = 226
Score = 116 bits (291), Expect = 6e-25
Identities = 62/190 (32%), Positives = 101/190 (52%), Gaps = 11/190 (5%)
Query: 31 RYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFVI 90
+ +L+ G + L+ G FF R+L + ++ Y VV+G+FP + DG++I
Sbjct: 2 KIGILLTGHAPDTLVDATGDYDAFFVRLLGPQNFEFETYSVVDGQFPSGADAA--DGWLI 59
Query: 91 TGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGV 150
TGS H + +HPWI L +L+ + + ++G+CFGHQI+ + LGGKV + GW IG
Sbjct: 60 TGSRHGVYEDHPWIPPLEALIRQIRDQGQPLIGVCFGHQIIAQALGGKVEKFQGGWAIGP 119
Query: 151 TTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQ 210
T + S +++ H+D+V LP AE++ ++ M YGD + IQ
Sbjct: 120 TEYDMGS---------ERVTVNAWHQDQVVALPEGAEVLASNDFCRNAMVAYGDTIWTIQ 170
Query: 211 GHPEFTIDIL 220
HPE+ D +
Sbjct: 171 AHPEYDNDFI 180
>ref|YP_169906.1| glutamine amidotransferase, class I [Francisella tularensis subsp.
tularensis Schu 4] gi|56604502|emb|CAG45542.1| glutamine
amidotransferase, class I [Francisella tularensis subsp.
tularensis SCHU S4]
Length = 235
Score = 110 bits (275), Expect = 4e-23
Identities = 60/149 (40%), Positives = 88/149 (58%), Gaps = 2/149 (1%)
Query: 66 WDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGIC 125
+D++ V E+P+ + +YDGF+ITGS A N WI L + + L +KKI+GIC
Sbjct: 40 FDIFDVTMQEYPQ--DYDIYDGFIITGSKATAFDNLAWIIKLKTEIVKLHDNHKKIIGIC 97
Query: 126 FGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPK 185
FGHQIL + LGG+V R G+ +GV +++ + S LS+ H+D V LP
Sbjct: 98 FGHQILAQALGGRVERGPKGFAVGVRNVEILTRKPWMEPFHSYLSLLFYHQDMVVKLPQG 157
Query: 186 AEIIGWSEKTGIEMFRYGDHMLGIQGHPE 214
AE+I S+ ++MF +H+LGIQ HPE
Sbjct: 158 AELISTSDYCKVQMFCINNHILGIQAHPE 186
>gb|AAW49751.1| hypothetical protein FTT0909 [synthetic construct]
Length = 270
Score = 110 bits (275), Expect = 4e-23
Identities = 60/149 (40%), Positives = 88/149 (58%), Gaps = 2/149 (1%)
Query: 66 WDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGIC 125
+D++ V E+P+ + +YDGF+ITGS A N WI L + + L +KKI+GIC
Sbjct: 66 FDIFDVTMQEYPQ--DYDIYDGFIITGSKATAFDNLAWIIKLKTEIVKLHDNHKKIIGIC 123
Query: 126 FGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPK 185
FGHQIL + LGG+V R G+ +GV +++ + S LS+ H+D V LP
Sbjct: 124 FGHQILAQALGGRVERGPKGFAVGVRNVEILTRKPWMEPFHSYLSLLFYHQDMVVKLPQG 183
Query: 186 AEIIGWSEKTGIEMFRYGDHMLGIQGHPE 214
AE+I S+ ++MF +H+LGIQ HPE
Sbjct: 184 AELISTSDYCKVQMFCINNHILGIQAHPE 212
>ref|ZP_00265787.1| COG0518: GMP synthase - Glutamine amidotransferase domain
[Pseudomonas fluorescens PfO-1]
Length = 240
Score = 110 bits (274), Expect = 5e-23
Identities = 58/177 (32%), Positives = 95/177 (52%), Gaps = 4/177 (2%)
Query: 44 LLKKHGGCFGFFTRMLAEE--GERWDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHANH 101
L+ ++ G F R+ +++ + +Y V+ G++P D +D +++TGS D+
Sbjct: 17 LVDQYQGYGQMFQRLFSQQPIAAEFTVYNVMQGDYPSDDLT--FDAYLVTGSKADSFGTD 74
Query: 102 PWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSI 161
PWI L + T K+LG+CFGHQ+L +LGGK R++ GW +G+ KLA+ +
Sbjct: 75 PWIQTLKQYLLTRYERGDKLLGVCFGHQLLALLLGGKSERATQGWGVGIHNYKLAAKAPW 134
Query: 162 ALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTID 218
+ EL++ H+D+V LP A +I S+ + D +L QGHPEF D
Sbjct: 135 MSPVREELTLLISHQDQVTALPENATVIASSDFCPFAAYHINDQVLCFQGHPEFIHD 191
>emb|CAH13355.1| hypothetical protein [Legionella pneumophila str. Paris]
gi|54298146|ref|YP_124515.1| hypothetical protein
lpp2203 [Legionella pneumophila str. Paris]
Length = 232
Score = 109 bits (272), Expect = 9e-23
Identities = 61/185 (32%), Positives = 100/185 (53%), Gaps = 5/185 (2%)
Query: 34 LLMCGEDSEYLLKKHGGCFGFFTRML--AEEGERWDLYKVVNGEFPRHDELPLYDGFVIT 91
+L+C + SE + HG F +L A+ + ++ +GE P ++ + D ++I+
Sbjct: 5 ILLCDKVSEVFVADHGQYPEMFANLLRPADSTLEFTVFDAEHGELPT--DVHVVDAYLIS 62
Query: 92 GSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVT 151
GS H + N+PWI L V TL + KK++GICFGHQ++ + LGGKV +S GW +G++
Sbjct: 63 GSRHGVNDNYPWIRKLEEFVCTLHASQKKLIGICFGHQLIAKALGGKVIKSPKGWGVGMS 122
Query: 152 TMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQG 211
++ + ++ H+D+V LP AEI+ S+ M + G +QG
Sbjct: 123 QNQIYQFKEWMRPSLNCFNLLVSHQDQVIELPTGAEILAGSDFCPNYMMQIGS-FFSVQG 181
Query: 212 HPEFT 216
HPEFT
Sbjct: 182 HPEFT 186
>ref|NP_250433.1| probable amidotransferase [Pseudomonas aeruginosa PAO1]
gi|9947719|gb|AAG05131.1| probable amidotransferase
[Pseudomonas aeruginosa PAO1] gi|11350950|pir||C83428
probable amidotransferase PA1742 [imported] -
Pseudomonas aeruginosa (strain PAO1)
Length = 240
Score = 108 bits (270), Expect = 2e-22
Identities = 58/177 (32%), Positives = 94/177 (52%), Gaps = 4/177 (2%)
Query: 44 LLKKHGGCFGFFTRMLAEE--GERWDLYKVVNGEFPRHDELPLYDGFVITGSCHDAHANH 101
L++++ G F ++ A++ + +Y VV G +P DE +D +++TGS D+
Sbjct: 17 LIERYEGYGRMFQQLFAKQPIAAEFVIYNVVEGRYPADDEC--FDAYLVTGSKADSFGPD 74
Query: 102 PWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIGVTTMKLASSSSI 161
PWI L + + K+LG+CFGHQ+L +LGGK R+S GW +G+ +L +
Sbjct: 75 PWIQTLKTFLLDRYERGDKLLGVCFGHQLLALLLGGKAERASQGWGMGIHQYQLNERADW 134
Query: 162 ALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHMLGIQGHPEFTID 218
+L++ H+D+V LP A +I S+ + GD +L QGHPEF D
Sbjct: 135 MSPALPDLTLLISHQDQVTRLPENARVIASSDFCPYAAYAIGDQVLCFQGHPEFVHD 191
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.143 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 501,678,766
Number of Sequences: 2540612
Number of extensions: 22822921
Number of successful extensions: 194925
Number of sequences better than 10.0: 981
Number of HSP's better than 10.0 without gapping: 346
Number of HSP's successfully gapped in prelim test: 637
Number of HSP's that attempted gapping in prelim test: 193251
Number of HSP's gapped (non-prelim): 1343
length of query: 267
length of database: 863,360,394
effective HSP length: 126
effective length of query: 141
effective length of database: 543,243,282
effective search space: 76597302762
effective search space used: 76597302762
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0055.14


