

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
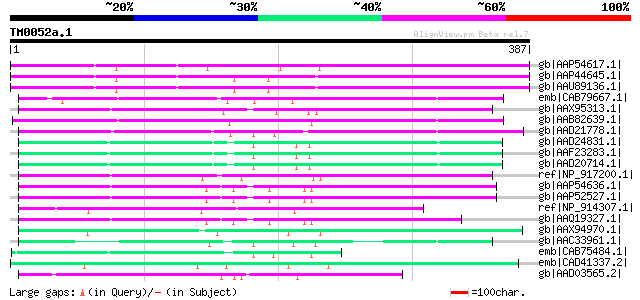
Query= TM0052a.1
(387 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP54617.1| putative non-LTR retroelement reverse transcripta... 165 2e-39
gb|AAP44645.1| putative reverse transcriptase [Oryza sativa (jap... 160 5e-38
gb|AAU89136.1| F-box domain containing protein [Oryza sativa (ja... 160 5e-38
emb|CAB79667.1| putative protein [Arabidopsis thaliana] gi|49720... 137 4e-31
gb|AAX95313.1| hypothetical protein [Oryza sativa (japonica cult... 136 1e-30
gb|AAB82639.1| putative non-LTR retroelement reverse transcripta... 127 8e-28
gb|AAD21778.1| putative non-LTR retroelement reverse transcripta... 118 3e-25
gb|AAD24831.1| putative non-LTR retroelement reverse transcripta... 114 4e-24
gb|AAF23283.1| putative non-LTR reverse transcriptase [Arabidops... 114 4e-24
gb|AAD20714.1| putative non-LTR retroelement reverse transcripta... 113 9e-24
ref|NP_917200.1| P0707D10.17 [Oryza sativa (japonica cultivar-gr... 112 1e-23
gb|AAP54636.1| putative reverse transcriptase [Oryza sativa (jap... 108 3e-22
gb|AAP52527.1| putative reverse transcriptase. [Oryza sativa (ja... 108 3e-22
ref|NP_914307.1| reverse transcriptase -like protein [Oryza sati... 104 4e-21
gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultiva... 102 2e-20
gb|AAX94970.1| retrotransposon protein, putative, unclassified [... 102 2e-20
gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam... 102 3e-20
emb|CAB75484.1| putative protein [Arabidopsis thaliana] gi|11357... 98 4e-19
emb|CAD41337.2| OJ991113_30.21 [Oryza sativa (japonica cultivar-... 98 5e-19
gb|AAD03565.2| putative non-LTR retroelement reverse transcripta... 94 5e-18
>gb|AAP54617.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|37536056|ref|NP_922330.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
gi|10140689|gb|AAG13524.1| putative non-LTR retroelement
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1382
Score = 165 bits (417), Expect = 2e-39
Identities = 121/404 (29%), Positives = 183/404 (44%), Gaps = 19/404 (4%)
Query: 1 DHDRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKL 60
+ D CW DLI +F A+ IL IP+S +G D WP G Y+ ++ Y R
Sbjct: 980 NEDARCWDGDLIRSLFPVDIAKEILQIPISRHGDADFASWPHDKLGLYSVRSAYNLARSE 1039
Query: 61 ASIEAPSSSTAVGFSPRF------WKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHV 114
A A S++ G + R WKG W +A + K T WRA L +L RRH+
Sbjct: 1040 AFF-ADQSNSGRGMASRLLESQKDWKGLWKINAPGKMKITLWRAAHECLATGFQLRRRHI 1098
Query: 115 DVDPGCCLCSGENETAEHLFMLCPAAKQV*YA----SSLSLRVDSFIAFADFWAWVVEQG 170
GC C+ ++T EH+F+ CP A Q+ ++ L + F + +++G
Sbjct: 1099 PSTDGCVFCN-RDDTVEHVFLFCPFAAQIWEEIKGKCAVKLGRNGFSTMRQWIFDFLKRG 1157
Query: 171 DSEFISTVQTIIYAIWEARNQVQFQQRTFS----VTGVLRRVAAMNSASQKSDGGPQTGS 226
S + + + IWEARN + T V +L V + + K+ G + G+
Sbjct: 1158 SSHANTLLAVTFWHIWEARNNTKNNNGTVHPQRVVIKILSYVDMILKHNTKTVDGQRGGN 1217
Query: 227 RPA--RWRRPQPGTIKCNFDAS-FRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLA 283
A RW+ P N DA+ F T G+G + R++ G+ + A + P LA
Sbjct: 1218 TQAIPRWQPPPASVWMINSDAAIFSSSRTMGVGALIRDNTGKCLVACSEMISDVVLPELA 1277
Query: 284 EAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFV 343
EA+A+R + LA + G IV+ +DCL + +T DRS + +I D + L S F
Sbjct: 1278 EALAIRRALGLAKEEGLEHIVMASDCLTVIRRIQTSGRDRSGVGCVIEDIKKLASTFVLC 1337
Query: 344 SLCFVRRTGNAVADFLARNANTYANMVWVEEVPLAVLPLFDADV 387
S V R N A LARNA V+ +P + + DV
Sbjct: 1338 SFMHVNRLSNLAAHSLARNAELSTCTVYRSVIPDYIRDILCDDV 1381
>gb|AAP44645.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|50917607|ref|XP_469200.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 404
Score = 160 bits (406), Expect = 5e-38
Identities = 119/405 (29%), Positives = 180/405 (44%), Gaps = 21/405 (5%)
Query: 1 DHDRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKL 60
+ D CW DLI +F A+ IL IP+S +G D WP G Y+ ++ Y +R
Sbjct: 2 NEDARCWDGDLIRSLFPVDIAKEILQIPISRHGDADFASWPHEKLGLYSVRSAYNMVRSE 61
Query: 61 ASIEAPSSSTAVGFSPRF------WKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHV 114
A A S++ G + R WKG W +A + K WRA L +L RRH+
Sbjct: 62 AFF-ADQSNSGRGMASRLLESQKDWKGLWKINAPGKMKIILWRAAHECLATGFQLPRRHI 120
Query: 115 DVDPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVD-SFIAFADFWAWV---VEQG 170
GC C+ ++T EH+F+ CP A Q+ V F+ W+ +++G
Sbjct: 121 PSTDGCVFCN-RDDTVEHVFLFCPFAAQIWEEIKRKCAVKLGRNGFSTMRHWIFDFLKRG 179
Query: 171 DSEFISTVQTIIYAIWEARNQ-------VQFQQRTFSVTGVLRRVAAMNSASQKSDGGPQ 223
S + + + IWEARN V Q+ + + + N+ + G
Sbjct: 180 SSHTNTLLAVTFWHIWEARNNTKNNNGTVHPQRVVIKILSYVDMILKHNTKTVDGQRGEN 239
Query: 224 TGSRPARWRRPQPGTIKCNFDAS-FRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLL 282
T + P RW+ G N DA+ F T G+G + R++ G+ + A + P L
Sbjct: 240 TQAIP-RWQPLPAGVWMINSDAAIFSSSRTMGVGALIRDNTGKCLVACSEMISDVVLPEL 298
Query: 283 AEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEF 342
AEA+A+R + LA + G IV+ +DCL + +T DRS + +I D + L S F
Sbjct: 299 AEALAIRRALGLAKEEGLQHIVMASDCLTVIRRIQTSGRDRSGVGCVIEDIKKLASTFVL 358
Query: 343 VSLCFVRRTGNAVADFLARNANTYANMVWVEEVPLAVLPLFDADV 387
S V R N A LARNA V+ +P + + DV
Sbjct: 359 CSFMHVNRLSNLAAHSLARNAELSTCTVYRSVIPDYIRDILCDDV 403
>gb|AAU89136.1| F-box domain containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 855
Score = 160 bits (406), Expect = 5e-38
Identities = 119/405 (29%), Positives = 180/405 (44%), Gaps = 21/405 (5%)
Query: 1 DHDRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKL 60
+ D CW DLI +F A+ IL IP+S +G D WP G Y+ ++ Y +R
Sbjct: 453 NEDARCWDGDLIRSLFPVDIAKEILQIPISRHGDADFASWPHEKLGLYSVRSAYNMVRSE 512
Query: 61 ASIEAPSSSTAVGFSPRF------WKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHV 114
A A S++ G + R WKG W +A + K WRA L +L RRH+
Sbjct: 513 AFF-ADQSNSGRGMASRLLESQKDWKGLWKINAPGKMKIILWRAAHECLATGFQLPRRHI 571
Query: 115 DVDPGCCLCSGENETAEHLFMLCPAAKQV*YASSLSLRVD-SFIAFADFWAWV---VEQG 170
GC C+ ++T EH+F+ CP A Q+ V F+ W+ +++G
Sbjct: 572 PSTDGCVFCN-RDDTVEHVFLFCPFAAQIWEEIKRKCAVKLGRNGFSTMRHWIFDFLKRG 630
Query: 171 DSEFISTVQTIIYAIWEARNQ-------VQFQQRTFSVTGVLRRVAAMNSASQKSDGGPQ 223
S + + + IWEARN V Q+ + + + N+ + G
Sbjct: 631 SSHTNTLLAVTFWHIWEARNNTKNNNGTVHPQRVVIKILSYVDMILKHNTKTVDGQRGEN 690
Query: 224 TGSRPARWRRPQPGTIKCNFDAS-FRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLL 282
T + P RW+ G N DA+ F T G+G + R++ G+ + A + P L
Sbjct: 691 TQAIP-RWQPLPAGVWMINSDAAIFSSSRTMGVGALIRDNTGKCLVACSEMISDVVLPEL 749
Query: 283 AEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEF 342
AEA+A+R + LA + G IV+ +DCL + +T DRS + +I D + L S F
Sbjct: 750 AEALAIRRALGLAKEEGLQHIVMASDCLTVIRRIQTSGRDRSGVGCVIEDIKKLASTFVL 809
Query: 343 VSLCFVRRTGNAVADFLARNANTYANMVWVEEVPLAVLPLFDADV 387
S V R N A LARNA V+ +P + + DV
Sbjct: 810 CSFMHVNRLSNLAAHSLARNAELSTCTVYRSVIPDYIRDILCDDV 854
>emb|CAB79667.1| putative protein [Arabidopsis thaliana] gi|4972055|emb|CAB43923.1|
putative protein [Arabidopsis thaliana]
gi|67633766|gb|AAY78807.1| putative reverse
transcriptase/RNA-dependent DNA polymerase [Arabidopsis
thaliana] gi|15233451|ref|NP_194638.1| reverse
transcriptase, putative / RNA-dependent DNA polymerase,
putative [Arabidopsis thaliana] gi|7485741|pir||T08964
hypothetical protein F19B15.120 - Arabidopsis thaliana
Length = 575
Score = 137 bits (346), Expect = 4e-31
Identities = 105/383 (27%), Positives = 176/383 (45%), Gaps = 28/383 (7%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDIL---FWPESSNGHYTSKAGYVFLRKLASI 63
WR+D+IE +F ++I + GGR IL W +S+G YT K+GY L ++ +
Sbjct: 182 WRKDVIEMLFPEVERKLIGELR---PGGRRILDSYTWDYTSSGDYTVKSGYWVLTQIINK 238
Query: 64 EA-PSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCL 122
+ P + +P + K W + P+ + W+ +S LP+ L RH+ + C
Sbjct: 239 RSSPQEVSEPSLNPIYQK-IWKSQTSPKIQHFLWKCLSNSLPVAGALAYRHLSKESACIR 297
Query: 123 CSGENETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFAD------FWAWVVEQGDSEFIS 176
C ET HL C A+ SS+ + + +AD +W + + G+ ++
Sbjct: 298 CPSCKETVNHLLFKCTFARLTWAISSIPIPLGG--EWADSIYVNLYWVFNLGNGNPQWEK 355
Query: 177 TVQTI---IYAIWEARNQVQFQQRTFSVTGVLRRVA------AMNSASQKSDGGPQTG-S 226
Q + ++ +W+ RN++ F+ R F+ VLRR + + ++ PQ S
Sbjct: 356 ASQLVPWLLWRLWKNRNELVFRGREFNAQEVLRRAEDDLEEWRIRTEAESCGTKPQVNRS 415
Query: 227 RPARWRRPQPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEA 285
RWR P +KCN DA++ R G+G V RN +GE + S L AE
Sbjct: 416 SCGRWRPPPHQWVKCNTDATWNRDNERCGIGWVLRNEKGEVKWMGARALPKLKSVLEAEL 475
Query: 286 MAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSL 345
AMRW + ++ ++ E+D L +E L I+D + L+S F V
Sbjct: 476 EAMRWAVLSLSRFQYNYVIFESDSQVLIEILNN-DEIWPSLKPTIQDLQRLLSQFTEVKF 534
Query: 346 CFVRRTGNAVADFLARNANTYAN 368
F+ R GN +A+ +AR + ++ N
Sbjct: 535 VFIPREGNTLAERVARESLSFLN 557
>gb|AAX95313.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 1063
Score = 136 bits (342), Expect = 1e-30
Identities = 110/386 (28%), Positives = 165/386 (42%), Gaps = 37/386 (9%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W D I F P ++IL I LS D L W G ++ ++ Y LA ++
Sbjct: 652 WDSDRISQAFLPIDVDLILGIRLSARQEEDFLVWHPDRLGRFSIRSAYHLATSLAESDSS 711
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
SSS+ V S + W L + K W+A S YLP + +R+++ C +C E
Sbjct: 712 SSSSGVILS-KTWDCLRKCKVLQKVKIHAWKAASNYLPTMENKKKRNLEASEVCKICGIE 770
Query: 127 NETAEHLFMLCPAAKQV*YASSLSLRVDSFIA--------FADFWAWVVEQGDSEFISTV 178
E H CP A+Q+ + +D + D + E F+ T
Sbjct: 771 KEDTIHALYRCPQARQLWILMRDVIYLDPAVVGRCTGPNRVLDLLMLLPEDNRMFFLMT- 829
Query: 179 QTIIYAIWEARNQVQFQQR----TFSVTGVLRRVAAMNSASQKSDGG-----------PQ 223
++ +W RN++ + T S +L V ++ SQ DG PQ
Sbjct: 830 ---LWRVWFIRNEIIHGKEPPPATVSKRFLLSYVRSLLDLSQHPDGSLLKGKHVASSSPQ 886
Query: 224 TGSR--------PARWRRPQPGTIKCNFDASFRGG-GTAGLGMVARNSEGEAMAAACSFP 274
T A W +PQPG +K N D SF G+G++ RNS G + +AC F
Sbjct: 887 TNGMVKPVNVQASAPWIKPQPGWMKLNIDGSFDASLNQGGIGVIPRNSTGSVIFSACGFI 946
Query: 275 VPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCR 334
+ L AE +A++ ++ LA+ I +ETDCL N ++ RS L+ LIRD
Sbjct: 947 DRCNCALEAELLALKESIILALHWTLLPICIETDCLEAVNLIQSARSVRSELAHLIRDIG 1006
Query: 335 LLISAFEFVSLCFVRRTGNAVADFLA 360
LIS + + V R N V+ FLA
Sbjct: 1007 ALISGNREMQIKKVLRCQNTVSHFLA 1032
>gb|AAB82639.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408936|pir||A84888 hypothetical protein
At2g45230 [imported] - Arabidopsis thaliana
Length = 1374
Score = 127 bits (318), Expect = 8e-28
Identities = 101/380 (26%), Positives = 162/380 (42%), Gaps = 16/380 (4%)
Query: 3 DRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLAS 62
D W +L+ +F +T E IL++ RD W S +GHY+ K+GY + ++ +
Sbjct: 975 DGRDWNWNLVSLLFPDNTQENILALRPGGKETRDRFTWEYSRSGHYSVKSGYWVMTEIIN 1034
Query: 63 IEA-PSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCC 121
P P F + W P+ WR ++ L + L RH+ + C
Sbjct: 1035 QRNNPQEVLQPSLDPIFQQ-IWKLDVPPKIHHFLWRCVNNCLSVASNLAYRHLAREKSCV 1093
Query: 122 LCSGENETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFADF-------WAWVVEQGDSEF 174
C ET HL CP A+ S L A + F + +S+
Sbjct: 1094 RCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAESLFRNMHHVLSVHKSQPEESDH 1153
Query: 175 ISTVQTIIYAIWEARNQVQFQQRTFSVTGV-LRRVAAMNSASQKSDGGPQ----TGSRPA 229
+ + I++ +W+ RN + F+ R F+ V L+ M++ + + + PQ T R
Sbjct: 1154 HALIPWILWRLWKNRNDLVFKGREFTAPQVILKATEDMDAWNNRKEPQPQVTSSTRDRCV 1213
Query: 230 RWRRPQPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAM 288
+W+ P G +KCN D ++ + G G+G V RN G + S L E A+
Sbjct: 1214 KWQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALPSQQSVLETEVEAL 1273
Query: 289 RWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSLCFV 348
RW + + ++ E+D L + + E D L+ I+D R L+ FE V F
Sbjct: 1274 RWAVLSLSRFNYRRVIFESDSQYLVSLIQN-EMDIPSLAPRIQDIRNLLRHFEEVKFQFT 1332
Query: 349 RRTGNAVADFLARNANTYAN 368
RR GN VAD AR + + N
Sbjct: 1333 RREGNNVADRTARESLSLMN 1352
>gb|AAD21778.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25410938|pir||G84429 hypothetical protein
At2g01840 [imported] - Arabidopsis thaliana
Length = 1715
Score = 118 bits (296), Expect = 3e-25
Identities = 102/402 (25%), Positives = 168/402 (41%), Gaps = 32/402 (7%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W + E V +P ++ S+ LS RD W + N YT ++GY + E
Sbjct: 1314 WDPVIFEGVLNPEDQQLAKSLYLSNYAARDSYKWAYTRNTQYTVRSGYWVATHVNLTEEE 1373
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
+ G P + W P+ K WR +SG L +L R++ DP C C
Sbjct: 1374 IINPLEGDVP-LKQEIWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNA 1432
Query: 127 NETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFADFW---AWVVEQGDSEFISTVQT--- 180
+ET H+ C A+ V +++ S + + F D ++ QG +
Sbjct: 1433 DETINHIIFTCSYAQVVWRSANFS--GSNRLCFTDNLEENIRLILQGKKNQNLPILNGLM 1490
Query: 181 ---IIYAIWEARNQVQFQQ---------------RTFSVTGVLRRVAAMNSASQKSDGGP 222
I++ +W++RN+ FQQ T V ++ A ++ +Q +D
Sbjct: 1491 PFWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVETMVNDTAISHNTAQSND--- 1547
Query: 223 QTGSRPARWRRPQPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSFPVPISSPL 281
+ SR +W P G +KCNFD+ + +G G + R+ G + + C+ S L
Sbjct: 1548 RPLSRSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSAL 1607
Query: 282 LAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFE 341
AEA+ +Q+ G+ + E D L L N ED L L+ D R ++
Sbjct: 1608 QAEALGFLHALQMVWIRGYCYVWFEGDNLELTNLINK-TEDHHLLETLLYDIRFWMTKLP 1666
Query: 342 FVSLCFVRRTGNAVADFLARNANTYANMVWVEEVPLAVLPLF 383
F S+ +V R N AD L + AN+ +++ VP L L+
Sbjct: 1667 FSSIGYVNRERNLAADKLTKYANSMSSLYETFHVPPRWLQLY 1708
>gb|AAD24831.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408166|pir||G84721 hypothetical protein
At2g31520 [imported] - Arabidopsis thaliana
Length = 1524
Score = 114 bits (286), Expect = 4e-24
Identities = 106/383 (27%), Positives = 154/383 (39%), Gaps = 30/383 (7%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W I S I I L+ + D + W ++ G YT ++GY L S P
Sbjct: 1128 WDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIP 1187
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
+ + G S W +P+ K WRA+S L +RL R + +DP C C E
Sbjct: 1188 AINPPHG-SIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPICPRCHRE 1246
Query: 127 NETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFADFWAWVVEQGDSEFISTVQT------ 180
NE+ H CP A + S SL + + + DF E+ S ++ VQ
Sbjct: 1247 NESINHALFTCPFATMAWWLSDSSL-IRNQLMSNDF-----EENISNILNFVQDTTMSDF 1300
Query: 181 -------IIYAIWEARNQVQFQQRTFSVTGVLRRVAAMN------SASQKSDGGP--QTG 225
+I+ IW+ARN V F + S + + A + S K P Q
Sbjct: 1301 HKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIA 1360
Query: 226 SRPARWRRPQPGTIKCNFDASFRGGG-TAGLGMVARNSEGEAMAAACSFPVPISSPLLAE 284
WR P +KCNFDA F A G + RN G ++ S+PL AE
Sbjct: 1361 ENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEAE 1420
Query: 285 AMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVS 344
A+ +Q G+ + +E DC L N I S L+ + D + F +
Sbjct: 1421 TKALLAALQQTWIRGYTQVFMEGDCQTLINLINGI-SFHSSLANHLEDISFWANKFASIQ 1479
Query: 345 LCFVRRTGNAVADFLARNANTYA 367
F+RR GN +A LA+ TY+
Sbjct: 1480 FGFIRRKGNKLAHVLAKYGCTYS 1502
>gb|AAF23283.1| putative non-LTR reverse transcriptase [Arabidopsis thaliana]
gi|15232695|ref|NP_187562.1| hypothetical protein
[Arabidopsis thaliana]
Length = 484
Score = 114 bits (286), Expect = 4e-24
Identities = 106/383 (27%), Positives = 153/383 (39%), Gaps = 30/383 (7%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W I S I I L+ + D + W ++ G YT ++GY L S P
Sbjct: 88 WDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIP 147
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
+ + G S W +P+ K WRA+S L +RL R + +DP C C E
Sbjct: 148 AINPPHG-SIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRE 206
Query: 127 NETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFADFWAWVVEQGDSEFISTVQT------ 180
NE+ H CP A S SL + + + DF E+ S ++ VQ
Sbjct: 207 NESINHALFTCPFATMAWRLSDSSL-IRNQLMSNDF-----EENISNILNFVQDTTMSDF 260
Query: 181 -------IIYAIWEARNQVQFQQRTFSVTGVLRRVAAMN------SASQKSDGGP--QTG 225
+I+ IW+ARN V F + S + + A + S K P Q
Sbjct: 261 HKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIA 320
Query: 226 SRPARWRRPQPGTIKCNFDASFRGGG-TAGLGMVARNSEGEAMAAACSFPVPISSPLLAE 284
WR P +KCNFDA F A G + RN G ++ S+PL AE
Sbjct: 321 ENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEAE 380
Query: 285 AMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVS 344
A+ +Q G+ + +E DC L N I S L+ + D + F +
Sbjct: 381 TKALLAALQQTWIRGYTQVFMEGDCQTLINLINGI-SFHSSLANHLEDISFWANKFASIQ 439
Query: 345 LCFVRRTGNAVADFLARNANTYA 367
F+RR GN +A LA+ TY+
Sbjct: 440 FGFIRRKGNKLAHVLAKYGCTYS 462
>gb|AAD20714.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25412331|pir||G84649 hypothetical protein
At2g25550 [imported] - Arabidopsis thaliana
Length = 1750
Score = 113 bits (283), Expect = 9e-24
Identities = 105/383 (27%), Positives = 153/383 (39%), Gaps = 30/383 (7%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W I S I I L+ + D + W ++ G YT ++GY L S P
Sbjct: 1354 WDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIP 1413
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
+ + G S W +P+ K WRA+S L +RL R + +DP C C E
Sbjct: 1414 AINPPHG-SIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPRCHRE 1472
Query: 127 NETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFADFWAWVVEQGDSEFISTVQT------ 180
NE+ H CP A S SL + + + DF E+ S ++ VQ
Sbjct: 1473 NESINHALFTCPFATMAWRLSDSSL-IRNQLMSNDF-----EENISNILNFVQDTTMSDF 1526
Query: 181 -------IIYAIWEARNQVQFQQRTFSVTGVLRRVAAMN------SASQKSDGGP--QTG 225
+I+ IW+ARN V F + S + + A + S K P Q
Sbjct: 1527 HKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIA 1586
Query: 226 SRPARWRRPQPGTIKCNFDASFRGGG-TAGLGMVARNSEGEAMAAACSFPVPISSPLLAE 284
WR P +KCNFDA F A G + RN G ++ S+PL AE
Sbjct: 1587 ENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEAE 1646
Query: 285 AMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVS 344
A+ +Q G+ + +E DC L N I S L+ + D + F +
Sbjct: 1647 TKALLAALQQTWIRGYTQVFMEGDCQTLINLINGI-SFHSSLANHLEDISFWANKFASIQ 1705
Query: 345 LCFVRRTGNAVADFLARNANTYA 367
F+R+ GN +A LA+ TY+
Sbjct: 1706 FGFIRKKGNKLAHVLAKYGCTYS 1728
>ref|NP_917200.1| P0707D10.17 [Oryza sativa (japonica cultivar-group)]
Length = 892
Score = 112 bits (281), Expect = 1e-23
Identities = 93/385 (24%), Positives = 168/385 (43%), Gaps = 35/385 (9%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W + VF P AE I SI +S D + W +G ++ ++ Y L ++E
Sbjct: 481 WDETRVNQVFLPVDAEAICSIRVSSRQEDDFVAWHPDKHGKFSVRSAYGLACNLVNMETS 540
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
SSS++ R W W T+A + + WRA + L + +R ++ C +C E
Sbjct: 541 SSSSSARCK-RMWNLIWQTNAPQKVRIFAWRAATNCLATMENKAKRKLEQSDICLVCGLE 599
Query: 127 NETAEHLFMLCPAAKQ----V*YASSLSLRVDSFIAFADFWAWVVEQGD--SEFISTV-Q 179
E +H+ CP A+ + A ++SL + + W+ +Q + ++ TV
Sbjct: 600 KEDTKHVLCRCPDARNLWGAMCEAGNISLNGGN---LCNDPGWIFDQLEILLDYERTVFL 656
Query: 180 TIIYAIWEARNQVQFQQRTFSVTGVLRRVAAMNSASQKSDGGPQTG-------------- 225
I++ IW RN++ ++ + R + + + + PQ
Sbjct: 657 LILWRIWFVRNEIVHGKQPPPIEASKRFLESYVKSLLEIRQHPQANVEKGKHVISFVNGK 716
Query: 226 SRPAR---------WRRPQPGTIKCNFDASFRG-GGTAGLGMVARNSEGEAMAAACSFPV 275
+ PAR WR+P G +K N D SF G+ G+G+V R+ G+ + A C
Sbjct: 717 TNPARRPPEDKNQKWRKPCEGWMKVNVDGSFDAQTGSGGIGVVMRDWSGKVIFACCKHLP 776
Query: 276 PISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRL 335
S L +E +A + + + + V+ETDCL + + E + S ++ L+++
Sbjct: 777 RCGSALESEILACKEGILVGLQWSLRQFVVETDCLMVVQMLESKEREMSEVAFLVKEATE 836
Query: 336 LISAFEFVSLCFVRRTGNAVADFLA 360
L+S + L + R+ N+V+ LA
Sbjct: 837 LLSGDREIQLRKIHRSQNSVSHLLA 861
>gb|AAP54636.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37536094|ref|NP_922349.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|13786450|gb|AAK39575.1| putative
reverse transcriptase [Oryza sativa]
Length = 791
Score = 108 bits (270), Expect = 3e-22
Identities = 97/390 (24%), Positives = 162/390 (40%), Gaps = 39/390 (10%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W I F AE+IL+I +S D + W NG ++ ++ Y +L +IE
Sbjct: 380 WDVPKIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEES 439
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
SSS + + W+ W + K WR S L +R ++ C +C E
Sbjct: 440 SSSGTNNIN-KAWEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRE 498
Query: 127 NETAEHLFMLCPAAKQV*Y----ASSLSLRVDSFIAFADFWAW-----VVEQGDSEFIST 177
NE H C A Q+ + S+S+ + + + FW + + E + F+ T
Sbjct: 499 NEDDAHALCRCIQASQLWSCMHKSGSVSVDIKASV-LGRFWLFDCLEKIPEYEQAMFLMT 557
Query: 178 VQTIIYAIWEARNQV-------------QFQQRTFSVTGVLRRVAAMNSASQKS------ 218
++ W RN++ +F Q + +R+ + K
Sbjct: 558 ----LWRNWYVRNELIHGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVP 613
Query: 219 -DGGPQ---TGSRPARWRRPQPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSF 273
GGP+ + W RP+ G +K N D SF G GLGM+ RNS G+ + +C
Sbjct: 614 LKGGPKYRVLNNHQPCWERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKP 673
Query: 274 PVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDC 333
++PL +E A ++LAI I +ETDC ++ + D S L+ +I +
Sbjct: 674 LERCNNPLESELRACVEGLKLAIHWTLLPIQVETDCASVVQLLQGTGRDFSVLANIIHEA 733
Query: 334 RLLISAFEFVSLCFVRRTGNAVADFLARNA 363
R L+ +S+ + R+ N ++ LA A
Sbjct: 734 RHLLVGERVISIKKICRSQNCISHHLANTA 763
>gb|AAP52527.1| putative reverse transcriptase. [Oryza sativa (japonica
cultivar-group)] gi|37531876|ref|NP_920240.1| putative
reverse transcriptase. [Oryza sativa (japonica
cultivar-group)]
Length = 493
Score = 108 bits (270), Expect = 3e-22
Identities = 97/390 (24%), Positives = 162/390 (40%), Gaps = 39/390 (10%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W I F AE+IL+I +S D + W NG ++ ++ Y +L +IE
Sbjct: 82 WDVPKIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEES 141
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
SSS + + W+ W + K WR S L +R ++ C +C E
Sbjct: 142 SSSGTNNIN-KAWEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRE 200
Query: 127 NETAEHLFMLCPAAKQV*Y----ASSLSLRVDSFIAFADFWAW-----VVEQGDSEFIST 177
NE H C A Q+ + S+S+ + + + FW + + E + F+ T
Sbjct: 201 NEDDAHALCRCIQASQLWSCMHKSGSVSVDIKASV-LGRFWLFDCLEKIPEYEQAMFLMT 259
Query: 178 VQTIIYAIWEARNQV-------------QFQQRTFSVTGVLRRVAAMNSASQKS------ 218
++ W RN++ +F Q + +R+ + K
Sbjct: 260 ----LWRNWYVRNELIHGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVP 315
Query: 219 -DGGPQ---TGSRPARWRRPQPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSF 273
GGP+ + W RP+ G +K N D SF G GLGM+ RNS G+ + +C
Sbjct: 316 LKGGPKYRVLNNHQPCWERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKP 375
Query: 274 PVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDC 333
++PL +E A ++LAI I +ETDC ++ + D S L+ +I +
Sbjct: 376 LERCNNPLESELRACVEGLKLAIHWTLLPIQVETDCASVVQLLQGTGRDFSVLANIIHEA 435
Query: 334 RLLISAFEFVSLCFVRRTGNAVADFLARNA 363
R L+ +S+ + R+ N ++ LA A
Sbjct: 436 RHLLVGERVISIKKICRSQNCISHHLANTA 465
>ref|NP_914307.1| reverse transcriptase -like protein [Oryza sativa (japonica
cultivar-group)]
Length = 727
Score = 104 bits (260), Expect = 4e-21
Identities = 82/313 (26%), Positives = 139/313 (44%), Gaps = 13/313 (4%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFL-----RKLA 61
W + L+E +F S A IL+IP+ L+ D + W + G Y+ K+ Y L R
Sbjct: 414 WDKPLVEEIFWESDARNILAIPVRLDTN-DFVAWHFDTRGMYSVKSAYHVLEDQRERSAK 472
Query: 62 SIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCC 121
E SSS AV W W+ +P+ K+ WR LPL+ + RR D + C
Sbjct: 473 KQEGGSSSGAVNQRNFEWDKIWSLQCIPKVKQFIWRLAHNSLPLKLNIKRRVPDAETLCP 532
Query: 122 LCSGENETAEHLFMLCPAAK---QV*YASSLSLRVDSFIAFADFWAWVVEQGDSEFISTV 178
+C +E H F+ C K ++ + L + ++ D +++ +++ V
Sbjct: 533 VCKRFDENGGHCFLQCKPMKLCWRILCLEDIRLSLTQLVSARDVVQTILKL-ENDRRMEV 591
Query: 179 QTIIYAIWEARNQVQFQQRTFSVTGVLRRVAAM--NSASQKSDGGPQTGSRPARWRRPQP 236
+++ W ARN+V + V V+ +V + + AS + P+ + +W P
Sbjct: 592 FFLLWVWWYARNKVNSGEDVIRVEEVVHKVKQLICDHASLRKGKHPKVNVQRNKWVPPVD 651
Query: 237 GTIKCNFDASFRG-GGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQLA 295
G +K NFD +FR + G + R+ G A+ A + AEA A ++ A
Sbjct: 652 GNLKLNFDGAFRAVNKSGGYDFLVRDHRGCAVLAGAGCLEHVHDAFAAEAEAGLAGLKSA 711
Query: 296 IDLGFH*IVLETD 308
I G + + +ETD
Sbjct: 712 ISHGINSVQVETD 724
>gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultivar-group)]
Length = 2367
Score = 102 bits (254), Expect = 2e-20
Identities = 92/364 (25%), Positives = 151/364 (41%), Gaps = 39/364 (10%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W I F AE+IL+I +S D + W NG ++ ++ Y +L +IE
Sbjct: 1704 WDVPKIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEES 1763
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
SSS + + W+ W + K WR S L +R ++ C +C E
Sbjct: 1764 SSSGTNNIN-KAWEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRE 1822
Query: 127 NETAEHLFMLCPAAKQV*Y----ASSLSLRVDSFIAFADFWAW-----VVEQGDSEFIST 177
NE H C A Q+ + S+S+ + + + FW + + E + F+ T
Sbjct: 1823 NEDDAHALCRCIQASQLWSCMHKSGSVSVDIKASV-LGRFWLFDCLEKIPEYEQAMFLMT 1881
Query: 178 VQTIIYAIWEARNQV-------------QFQQRTFSVTGVLRRVAAMNSASQKS------ 218
++ W RN++ +F Q + +R+ + K
Sbjct: 1882 ----LWRNWYVRNELIHGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVP 1937
Query: 219 -DGGPQ---TGSRPARWRRPQPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSF 273
GGP+ + W RP+ G +K N D SF G GLGM+ RNS G+ + +C
Sbjct: 1938 LKGGPKYRVLNNHQPCWERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKP 1997
Query: 274 PVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDC 333
++PL +E A ++LAI I +ETDC ++ + I D S L+ +I +
Sbjct: 1998 LERCNNPLESELRACVEGLKLAIHWTLLPIQVETDCASVVQLLQGIGRDFSVLANIIHEA 2057
Query: 334 RLLI 337
R L+
Sbjct: 2058 RHLL 2061
>gb|AAX94970.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)]
Length = 816
Score = 102 bits (254), Expect = 2e-20
Identities = 100/408 (24%), Positives = 165/408 (39%), Gaps = 36/408 (8%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVF--LRKLASIE 64
W LI +F P + IL I L + D L W G +T ++ Y + + E
Sbjct: 241 WDEGLIRQIFYPPYVDEILKIKLPESPTDDFLAWHPERLGVFTVRSAYKLGLAEEHRANE 300
Query: 65 APSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCS 124
A + S+ WK W+ + WR LP + +R++ C +C
Sbjct: 301 AGACSSNGNGDRVLWKNIWSAQVPQKVCIFAWRLAVERLPTQHNKQKRNIVPSARCEVCG 360
Query: 125 GENETAEHLFMLCPAAKQV*YASSLSLRV-------DSFIAFADFWAWVV-EQGDSEFIS 176
E H ++C AK A LR D F+ W ++ + D +
Sbjct: 361 APREDGFHATVVCTKAK----ALRDELRAFWPIPPEDKFVRQGPDWLLLLLDTLDMRNKA 416
Query: 177 TVQTIIYAIWEARNQVQFQQRTFSVTGVLR----RVAAMNSASQKSDG-GPQTGSRPAR- 230
+ +++ W RN T +++G +R V + QK D G P R
Sbjct: 417 QMLLMLWRAWHLRNNSVHDTGTLTISGSIRFLQSYVQLLFPIRQKEDPKGKCAAQVPGRL 476
Query: 231 --------------WRRPQPGTIKCNFDASFR-GGGTAGLGMVARNSEGEAMAAACSFPV 275
W P G K N D +F T G+G++AR+SEG+A+ ++ +
Sbjct: 477 NQNPGVTKADCLQIWEPPPEGWAKINVDGAFSMTDNTGGIGVIARDSEGKALLSSWKYLH 536
Query: 276 PISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRL 335
+ E +A M+LA + I+LE+DC+ + +E+RS + LIR +
Sbjct: 537 RCADAEQVEILACYEGMKLAAEWIRKPIILESDCVTVIGRMTAEDEERSRWTFLIRSAKA 596
Query: 336 LISAFEFVSLCFVRRTGNAVADFLARNANTYAN-MVWVEEVPLAVLPL 382
++ + + V + +R N VA LA+ A A+ VW + VP V+ L
Sbjct: 597 VMRSLQEVRIQHRKRECNRVAHELAQLAKRTAHCAVWRDHVPSCVMQL 644
>gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam: rvt.hmm, score:
42.57) [Arabidopsis thaliana] gi|7486711|pir||T01893
hypothetical protein F8M12.22 - Arabidopsis thaliana
Length = 1662
Score = 102 bits (253), Expect = 3e-20
Identities = 96/375 (25%), Positives = 141/375 (37%), Gaps = 82/375 (21%)
Query: 7 WRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEAP 66
W LI + SP +++ LS +D L W SS+G YT
Sbjct: 1319 WNAHLISQLVSPDDHRFVMNHHLSRIVHQDKLVWNYSSSGDYT----------------- 1361
Query: 67 SSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSGE 126
W +P+ K WR IS LP R RL R +D+DP C C E
Sbjct: 1362 ---------------LWKLPIIPKIKYMLWRTISKALPTRSRLLTRGMDIDPHCPRCPTE 1406
Query: 127 NETAEHLFMLCPAAKQV*YAS--------SLSLRVDSFIAFADFWAWVVEQGDSEFISTV 178
ET H+ CP A + S + S + I+F ++ + ++T
Sbjct: 1407 EETINHVLFTCPYAASIWGLSNFPWLPGHTFSQDTEENISF------LINSFSNNTLNTE 1460
Query: 179 QT-----IIYAIWEARNQVQFQQRTFSVTGVLRRVAA-----MNSASQKSDGGPQTGS-- 226
Q +I+ +W+ARN + F + + S + V+ + A + S +++ D T S
Sbjct: 1461 QRLAPFWLIWRLWKARNNLVFNKFSESCSRVVTQTEAEVNEWLQSVNRREDASVLTRSSH 1520
Query: 227 -RPARWRRPQPGTIKCNFDASFRGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEA 285
RW++P +KCNFDA F G T G ++ E
Sbjct: 1521 RANVRWKKPVFPLVKCNFDAGFTGNNTQSTG----------------------GWIIPET 1558
Query: 286 MAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLSILIRDCRLLISAFEFVSL 345
A+ MQ A G+ + E DC L A R ++ L+RD S F V
Sbjct: 1559 KALLIAMQQAWVRGYKCVQFEGDCEILIKAING-AISRCEITSLLRDVDFWASRFSTVVF 1617
Query: 346 CFVRRTGNAVADFLA 360
F R N A LA
Sbjct: 1618 TFTNRLCNNTAHLLA 1632
>emb|CAB75484.1| putative protein [Arabidopsis thaliana] gi|11357921|pir||T47495
hypothetical protein F9K21.130 - Arabidopsis thaliana
Length = 851
Score = 98.2 bits (243), Expect = 4e-19
Identities = 77/267 (28%), Positives = 108/267 (39%), Gaps = 29/267 (10%)
Query: 2 HDRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLA 61
H R CW ++ S + I I LS D L W +S G YT ++GY
Sbjct: 589 HSR-CWDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYNSTGDYTVRSGYWLSTHDP 647
Query: 62 SIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCC 121
S P+ + G S W +P+ K WR +S LP DRL R + +DPGC
Sbjct: 648 SNTIPTMAKPHG-SVDLKTKIWNLPIMPKLKHFLWRILSKALPTTDRLTTRGMRIDPGCP 706
Query: 122 LCSGENETAEHLFMLCPAAKQV*YASSLSLRVDSFIAFADFWAWVVEQGDSEFISTVQT- 180
C ENE+ H CP A S L S ++ +E S + +Q
Sbjct: 707 RCRRENESINHALFTCPFATMAWRLSDTPLYRSSILSNN------IEDNISNILLLLQNT 760
Query: 181 ------------IIYAIWEARNQVQFQ--QRTFSVTGVLRRVAAMNSASQKSDGGPQ--- 223
+++ IW+ARN V F + + S+T V + + GP+
Sbjct: 761 TITDSQKLIPFWLLWRIWKARNNVVFNNLRESPSITVVRAKAETNEWLNATQTQGPRRLP 820
Query: 224 ---TGSRPARWRRPQPGTIKCNFDASF 247
T + W +PQ IKCNFDASF
Sbjct: 821 KRTTAAGNTTWVKPQMPYIKCNFDASF 847
>emb|CAD41337.2| OJ991113_30.21 [Oryza sativa (japonica cultivar-group)]
gi|50925420|ref|XP_472970.1| OJ991113_30.21 [Oryza
sativa (japonica cultivar-group)]
Length = 541
Score = 97.8 bits (242), Expect = 5e-19
Identities = 101/413 (24%), Positives = 158/413 (37%), Gaps = 34/413 (8%)
Query: 1 DHDRHCWRRDLIEFVFSPSTAEIILSIPLSLNGGRDILFWPESSNGHYTSKAGY--VFLR 58
+ D + W DLI+ V SP IL I L G D L W G +T ++ Y
Sbjct: 116 NQDTNSWDADLIKNVCSPLDVNEILKIKLPHRGMEDFLAWHFEKTGVFTVRSAYHLALHN 175
Query: 59 KLASIEAPSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDP 118
+L + E SSS++ + W W + K WR L + +R +++D
Sbjct: 176 QLKANELGSSSSSTSGERKIWASLWTAPVPQKVKIFAWRLARECLATMENRKKRKLEIDS 235
Query: 119 GCCLCSGENETAEHLFMLCP--AAKQV*YASSLSLRVDSFIAFA--DFWAWVVEQGDSEF 174
C +C E H + C AA + L + F+ + D+ +++ ++E
Sbjct: 236 TCRICGLGGEDGFHAVISCTKAAALRSLIREVWELPDERFLIRSGPDWLLIILDSINAES 295
Query: 175 ISTVQTIIYAIWEARNQVQFQQRTFSVTGVLR----------------------RVAAMN 212
+ +++ W RN SV G L + M
Sbjct: 296 RAKFLLLLWRAWFLRNDSVHCSGKASVLGSLLFLQSLWESLFMSTSQGKVDDKGKKPVMQ 355
Query: 213 SASQKSDGGPQTGSRPARWRRPQP----GTIKCNFDASF-RGGGTAGLGMVARNSEGEAM 267
S D + + W P P G +K N D SF + A G V R+ G +
Sbjct: 356 SRLSSPDSHERNHTTTMGWTPPPPPRPMGWVKVNVDGSFIQSSEPASAGFVIRDHTGSVL 415
Query: 268 AAACSFPVPISSPLLAEAMAMRWTMQLAIDLGFH*IVLETDCLNLFNAWRTIEEDRSYLS 327
+ S AEA A + LA + ++LETDC NL + + DR+ L
Sbjct: 416 LTSWRIISHCGSAEEAEATACWEGVNLAAEWVKKPLILETDCANLVSMLTSSGFDRAQLC 475
Query: 328 ILIRDCRLLISAFEFVSLCFVRRTGNAVADFLARNA-NTYANMVWVEEVPLAV 379
++R +LL+ A + +RR N VA LA+ A T + VW + P V
Sbjct: 476 HVLRSIKLLLQALPDFRVQRIRRECNRVAHDLAKFAMRTNHSAVWRMQAPSCV 528
>gb|AAD03565.2| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411819|pir||H84557 hypothetical protein
At2g17910 [imported] - Arabidopsis thaliana
Length = 1344
Score = 94.4 bits (233), Expect = 5e-18
Identities = 76/300 (25%), Positives = 130/300 (43%), Gaps = 17/300 (5%)
Query: 7 WRRDLIEFVFSPSTAEIILSI-PLSLNGGRDILFWPESSNGHYTSKAGYVFLRKLASIEA 65
W +++ +F EIIL PL D W S NG Y+ K GY FL K
Sbjct: 950 WNLNMLRDLFPWKDVEIILKQRPLFFK--EDSFCWLHSHNGLYSVKTGYEFLSKQVHHRL 1007
Query: 66 PSSSTAVGFSPRFWKGFWATSALPRCKETCWRAISGYLPLRDRLFRRHVDVDPGCCLCSG 125
+ + W P+ + W+A+ G +P+ DRL R + D GC +C
Sbjct: 1008 YQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCDT 1067
Query: 126 ENETAEHLFMLCPAAKQV*YASSLSLRVDSF--IAFADFWAWV--VEQGD----SEFIST 177
ENET H+ CP A+QV + LS F + + + +Q D F+S
Sbjct: 1068 ENETINHILFECPLARQVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVS- 1126
Query: 178 VQTIIYAIWEARNQVQFQQR-TFSVTGVLRRVAAMNS--ASQKSDGGPQTGSRPARWRRP 234
I++ +W+ RN + F+ + + + T V + A + ++Q + + +W P
Sbjct: 1127 -PWILWFLWKNRNALLFEGKGSITTTLVDKAYEAYHEWFSAQTHMQNDEKHLKITKWCPP 1185
Query: 235 QPGTIKCNFDASF-RGGGTAGLGMVARNSEGEAMAAACSFPVPISSPLLAEAMAMRWTMQ 293
PG +KCN ++ + +G V R+S+G+ + + + SP A+ + W ++
Sbjct: 1186 LPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSAKIRSWEWALE 1245
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.138 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 655,206,354
Number of Sequences: 2540612
Number of extensions: 26528790
Number of successful extensions: 67170
Number of sequences better than 10.0: 305
Number of HSP's better than 10.0 without gapping: 169
Number of HSP's successfully gapped in prelim test: 136
Number of HSP's that attempted gapping in prelim test: 66661
Number of HSP's gapped (non-prelim): 374
length of query: 387
length of database: 863,360,394
effective HSP length: 130
effective length of query: 257
effective length of database: 533,080,834
effective search space: 137001774338
effective search space used: 137001774338
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0052a.1


