

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0051b.7
(189 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
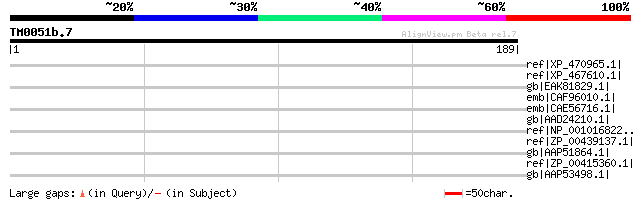
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_470965.1| OSJNBa0094O15.8 [Oryza sativa (japonica cultiva... 35 1.1
ref|XP_467610.1| hypothetical protein [Oryza sativa (japonica cu... 35 1.4
gb|EAK81829.1| hypothetical protein UM01222.1 [Ustilago maydis 5... 35 1.4
emb|CAF96010.1| unnamed protein product [Tetraodon nigroviridis] 33 3.1
emb|CAE56716.1| Hypothetical protein CBG24503 [Caenorhabditis br... 33 3.1
gb|AAD24210.1| CCCH zinc finger protein C3H-4 [Xenopus laevis] 33 5.3
ref|NP_001016822.1| hypothetical protein LOC549576 [Xenopus trop... 33 5.3
ref|ZP_00439137.1| COG3321: Polyketide synthase modules and rela... 32 6.9
gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sati... 32 6.9
ref|ZP_00415360.1| regulatory protein, DeoR [Azotobacter vinelan... 32 9.1
gb|AAP53498.1| putative polyprotein [Oryza sativa (japonica cult... 32 9.1
>ref|XP_470965.1| OSJNBa0094O15.8 [Oryza sativa (japonica cultivar-group)]
gi|38346201|emb|CAE04489.2| OSJNBa0094O15.8 [Oryza
sativa (japonica cultivar-group)]
Length = 716
Score = 35.0 bits (79), Expect = 1.1
Identities = 16/38 (42%), Positives = 23/38 (60%)
Query: 114 FGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWIVIT 151
F G VG+ Q MV+E++H F GA Y+W+V+T
Sbjct: 110 FMDGNVGYNQIFMVEEDIHNTAFRCPGAISLYEWVVMT 147
>ref|XP_467610.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|46390857|dbj|BAD16361.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|46390461|dbj|BAD15922.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 158
Score = 34.7 bits (78), Expect = 1.4
Identities = 31/98 (31%), Positives = 41/98 (41%), Gaps = 10/98 (10%)
Query: 12 GRVTGGSGRFHRNRLRQNEDEQAISD---QRSMEEEGKKLRRCSGGVARVVAAQRMAAQV 68
G G G R+R RQ E+ + D +R + G +GGVA+ V R+
Sbjct: 27 GGEAAGGGVVERDRHRQVGGERGVGDAVAERRGLDSGCGGGGVAGGVAKRVVQARLHVHG 86
Query: 69 MRRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPPSDP 106
+RR A VR A R P + P L PPS P
Sbjct: 87 CQRRARARVRGSAAMARRCP-------WHPPLPPPSPP 117
>gb|EAK81829.1| hypothetical protein UM01222.1 [Ustilago maydis 521]
gi|49069096|ref|XP_398837.1| hypothetical protein
UM01222.1 [Ustilago maydis 521]
Length = 967
Score = 34.7 bits (78), Expect = 1.4
Identities = 27/87 (31%), Positives = 39/87 (44%), Gaps = 6/87 (6%)
Query: 26 LRQNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVV---AVVRSDWA 82
L +DEQA++ R+ EE K RR GV +A +A +R V A S+W+
Sbjct: 127 LLDEDDEQALAGGRAREEWIKSARRARYGVGGSLARGSLAPSDIRNSTVGFEACTMSNWS 186
Query: 83 PNRNRPIQLIWSRFFPSLNPPSDPSEP 109
+ N S PS +P + S P
Sbjct: 187 SSSNNSPSTSSS---PSTSPSASTSSP 210
>emb|CAF96010.1| unnamed protein product [Tetraodon nigroviridis]
Length = 565
Score = 33.5 bits (75), Expect = 3.1
Identities = 27/90 (30%), Positives = 45/90 (50%), Gaps = 4/90 (4%)
Query: 24 NRLRQNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQR---MAAQVMRRRVVAVVRSD 80
NRL++ S++R ++EE K +R +AR+ AQ + RR+++A +
Sbjct: 414 NRLKEALKAARQSNERRLQEEQKAVRSAGVIMARLTRAQSHHCSDEERKRRKILARKVAP 473
Query: 81 WAPNRNRPIQLIWSRFFPSLNPPSDPSEPS 110
AP+R P S S +PP+ PS P+
Sbjct: 474 LAPSRKPPAPASGSGSGAS-SPPTCPSSPT 502
>emb|CAE56716.1| Hypothetical protein CBG24503 [Caenorhabditis briggsae]
Length = 545
Score = 33.5 bits (75), Expect = 3.1
Identities = 16/47 (34%), Positives = 29/47 (61%)
Query: 129 ENVHINIFDIQGAPLFYQWIVITSVVLGIIDYDISLEYISFYVVIML 175
++V IF + GAP FYQWI + + + +S+ Y+S+Y V+++
Sbjct: 97 QHVTAIIFCVCGAPEFYQWIRMDNEGTVSLGSALSVSYLSWYAVLVV 143
>gb|AAD24210.1| CCCH zinc finger protein C3H-4 [Xenopus laevis]
Length = 276
Score = 32.7 bits (73), Expect = 5.3
Identities = 13/27 (48%), Positives = 17/27 (62%)
Query: 92 IWSRFFPSLNPPSDPSEPSGPVFGRGP 118
++S FFP L+PP+DP P P F P
Sbjct: 10 LFSSFFPQLSPPADPETPLLPSFSAPP 36
>ref|NP_001016822.1| hypothetical protein LOC549576 [Xenopus tropicalis]
Length = 271
Score = 32.7 bits (73), Expect = 5.3
Identities = 14/27 (51%), Positives = 17/27 (62%)
Query: 92 IWSRFFPSLNPPSDPSEPSGPVFGRGP 118
++S FFP L+PPSDP P P F P
Sbjct: 10 LFSSFFPPLSPPSDPEIPLLPSFSAPP 36
>ref|ZP_00439137.1| COG3321: Polyketide synthase modules and related proteins
[Burkholderia mallei GB8 horse 4]
Length = 1516
Score = 32.3 bits (72), Expect = 6.9
Identities = 31/89 (34%), Positives = 40/89 (44%), Gaps = 13/89 (14%)
Query: 3 CVGLDRFRVGRVT-GGSGR---FHRNRL--------RQNEDEQAISDQRSMEEEGKKLRR 50
CV + F T GGSGR + R R+ R E+A +R E RR
Sbjct: 998 CVSMREFSARAETAGGSGRPNTYPRRRMARRGARRERAERAERAERGERGERGERAASRR 1057
Query: 51 CSGGVARVVAAQ-RMAAQVMRRRVVAVVR 78
+GG AR +AA AA+ RRR +A R
Sbjct: 1058 AAGGGARRIAASCARAARHRRRRALAAGR 1086
>gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37530550|ref|NP_919577.1| putative
retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|18581535|gb|AAK52540.2| Putative
retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 32.3 bits (72), Expect = 6.9
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Query: 114 FGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWIVITSVVLGI-IDYDISLEYI 167
F G G+ Q M +E++H F GA ++W+VIT V+ Y ++ YI
Sbjct: 673 FMDGNAGYNQIFMAEEDIHKTTFRCPGAIGLFEWVVITFVLKSAGATYQRAMNYI 727
>ref|ZP_00415360.1| regulatory protein, DeoR [Azotobacter vinelandii AvOP]
gi|67087748|gb|EAM07214.1| regulatory protein, DeoR
[Azotobacter vinelandii AvOP]
Length = 254
Score = 32.0 bits (71), Expect = 9.1
Identities = 28/108 (25%), Positives = 44/108 (39%), Gaps = 5/108 (4%)
Query: 14 VTGGSGRFHRNRLRQNEDEQAIS----DQRSMEEEGKKLRRCSGGVARVVAAQRMAAQVM 69
+TGG+ H + EQ + DQ + +G L R + ++ R+ A+V
Sbjct: 138 MTGGTWDPHSESFQGQVAEQVLRSYDFDQLFIGADGIDLARGTTTFNELLGLSRVMAEVA 197
Query: 70 RRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPPSDPSEPSGPVFGRG 117
R VV +V SD R ++L WS + P E G + G
Sbjct: 198 RE-VVVMVESDKVGRRIPNLELPWSSIHTLITDERLPPEARGQIAAHG 244
>gb|AAP53498.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37533818|ref|NP_921211.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|18652527|gb|AAL77160.1| Putative polyprotein [Oryza
sativa]
Length = 931
Score = 32.0 bits (71), Expect = 9.1
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 1/55 (1%)
Query: 114 FGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWIVITSVVLGI-IDYDISLEYI 167
F G VG+ Q M +E++H F GA ++W+V+T + + Y ++ YI
Sbjct: 437 FMDGNVGYNQIFMAEEDIHKTTFTCPGAIGLFEWVVMTFGLKSVGATYQRAMNYI 491
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.143 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 333,973,384
Number of Sequences: 2540612
Number of extensions: 13641265
Number of successful extensions: 51175
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 51169
Number of HSP's gapped (non-prelim): 12
length of query: 189
length of database: 863,360,394
effective HSP length: 121
effective length of query: 68
effective length of database: 555,946,342
effective search space: 37804351256
effective search space used: 37804351256
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0051b.7


