

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0050.15
(95 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
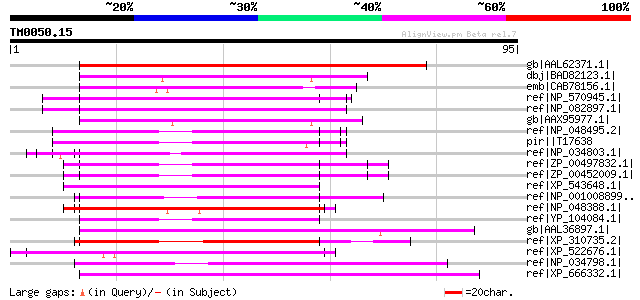
Score E
Sequences producing significant alignments: (bits) Value
gb|AAL62371.1| putative lipase [Arabidopsis thaliana] gi|2232982... 121 4e-27
dbj|BAD82123.1| hypothetical protein [Oryza sativa (japonica cul... 54 7e-07
emb|CAB78156.1| hypothetical protein [Arabidopsis thaliana] gi|4... 53 2e-06
ref|NP_570945.1| keratin associated protein 16-7 [Mus musculus] ... 52 3e-06
ref|NP_082897.1| keratin associated protein 21-1 [Mus musculus] ... 52 3e-06
gb|AAX95977.1| expressed protein [Oryza sativa (japonica cultiva... 52 3e-06
ref|NP_048495.2| Phe-, Gly-rich protein: RCGF 3X, GCGF 11X, RSGF... 52 4e-06
pir||T17638 glycine tyrosine-rich protein a147L - Chlorella viru... 52 4e-06
ref|NP_034803.1| keratin associated protein 6-2 [Mus musculus] g... 50 1e-05
ref|ZP_00497832.1| COG2378: Predicted transcriptional regulator ... 50 2e-05
ref|ZP_00452009.1| COG2378: Predicted transcriptional regulator ... 50 2e-05
ref|XP_543648.1| PREDICTED: similar to Keratin 5 [Canis familiaris] 50 2e-05
ref|NP_001008899.1| type II keratin Kb2 [Rattus norvegicus] gi|4... 49 3e-05
ref|NP_048388.1| contains Gly-rich motifs GAGLGTGF (5X); similar... 49 3e-05
ref|YP_104084.1| hypothetical protein BMA2541 [Burkholderia mall... 49 3e-05
gb|AAL36897.1| putative antimicrobial peptide [Litopenaeus setif... 49 3e-05
ref|XP_310735.2| ENSANGP00000007494 [Anopheles gambiae str. PEST... 49 4e-05
ref|XP_522676.1| PREDICTED: hypothetical protein XP_522676 [Pan ... 49 4e-05
ref|NP_034798.1| keratin complex 2, basic, gene 17 [Mus musculus... 48 5e-05
ref|XP_666332.1| hypothetical protein Chro.40165 [Cryptosporidiu... 48 5e-05
>gb|AAL62371.1| putative lipase [Arabidopsis thaliana]
gi|22329821|ref|NP_683330.1| glycine-rich protein
[Arabidopsis thaliana]
Length = 91
Score = 121 bits (304), Expect = 4e-27
Identities = 49/65 (75%), Positives = 60/65 (91%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
GFGVGCGFGVGWGFGGMP+N+LG+G GGGCGVG+GLGWGFG+AFGS YRSS++ FQG+E
Sbjct: 14 GFGVGCGFGVGWGFGGMPMNILGVGAGGGCGVGLGLGWGFGTAFGSHYRSSRLTFQGIEL 73
Query: 74 DSKEK 78
++ +K
Sbjct: 74 ETADK 78
>dbj|BAD82123.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|56784313|dbj|BAD82239.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 135
Score = 54.3 bits (129), Expect = 7e-07
Identities = 29/63 (46%), Positives = 34/63 (53%), Gaps = 9/63 (14%)
Query: 14 GFGVGCGFGVGWGF-------GGMPLNLLGLGVGGGCGVGVGLGWGFGS--AFGSKYRSS 64
G G GCG GV +G G P LGLG G GCG+G+G G+GFG A+ R S
Sbjct: 30 GIGAGCGVGVCFGLIGGAGIGAGFPGLQLGLGAGAGCGIGIGFGYGFGKGIAYDENGRYS 89
Query: 65 QIR 67
IR
Sbjct: 90 NIR 92
>emb|CAB78156.1| hypothetical protein [Arabidopsis thaliana]
gi|4538962|emb|CAB39786.1| hypothetical protein
[Arabidopsis thaliana] gi|22136796|gb|AAM91742.1|
unknown protein [Arabidopsis thaliana]
gi|19347875|gb|AAL85995.1| unknown protein [Arabidopsis
thaliana] gi|15235027|ref|NP_192771.1| glycine-rich
protein [Arabidopsis thaliana] gi|7486068|pir||T04048
hypothetical protein F24G24.130 - Arabidopsis thaliana
Length = 130
Score = 53.1 bits (126), Expect = 2e-06
Identities = 30/59 (50%), Positives = 35/59 (58%), Gaps = 9/59 (15%)
Query: 14 GFGVGCGFGVGWG------FG-GMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQ 65
G GVGCGFG G G FG G+P GLG G GCG+GVG G+G G G+ Y S+
Sbjct: 34 GLGVGCGFGFGAGLIGGVGFGPGVPGLQFGLGFGAGCGIGVGFGYGVGR--GAAYDHSR 90
>ref|NP_570945.1| keratin associated protein 16-7 [Mus musculus]
gi|14030413|gb|AAK52895.1| keratin-associated protein
16.7 [Mus musculus]
Length = 128
Score = 52.4 bits (124), Expect = 3e-06
Identities = 26/57 (45%), Positives = 32/57 (55%)
Query: 7 GFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G C + S +G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 11 GGCGYGSRYGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 67
Score = 50.8 bits (120), Expect = 8e-06
Identities = 25/51 (49%), Positives = 29/51 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G G GCG+G G+G G G G G GCG G G G G+GS +G Y SS
Sbjct: 26 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSS 76
Score = 49.7 bits (117), Expect = 2e-05
Identities = 24/50 (48%), Positives = 29/50 (58%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS++G Y S
Sbjct: 34 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSSYGCGYGS 83
Score = 48.9 bits (115), Expect = 3e-05
Identities = 23/50 (46%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+G+ +G Y S
Sbjct: 58 GSGYGCGYGSGYGCGYGSSYGCGYGSGYGCGYGSGYGCGYGTGYGCGYGS 107
Score = 45.1 bits (105), Expect = 4e-04
Identities = 21/45 (46%), Positives = 25/45 (54%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G GCG+G +G G G G G GCG G G G G+GS +G
Sbjct: 66 GSGYGCGYGSSYGCGYGSGYGCGYGSGYGCGYGTGYGCGYGSRYG 110
Score = 42.0 bits (97), Expect = 0.004
Identities = 21/48 (43%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G+G G G G G G+G G G GCG G G G G+GS +G Y
Sbjct: 56 GYGSGYGCGYGSGYG------CGYGSSYGCGYGSGYGCGYGSGYGCGY 97
Score = 36.2 bits (82), Expect = 0.21
Identities = 22/55 (40%), Positives = 27/55 (49%), Gaps = 7/55 (12%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG+ C + S +G G G G G G+G G G G G G G G G +G GS
Sbjct: 67 SGYGCGYGSSYGCGYGSGYGCGYGS------GYGCGYGTGYGCGYGSRYGCGCGS 115
>ref|NP_082897.1| keratin associated protein 21-1 [Mus musculus]
gi|26324323|dbj|BAB22970.2| unnamed protein product
[Mus musculus]
Length = 128
Score = 52.4 bits (124), Expect = 3e-06
Identities = 26/57 (45%), Positives = 32/57 (55%)
Query: 7 GFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G C + S +G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 11 GGCGYGSRYGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 67
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 50 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 99
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 58 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 107
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 26 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 75
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 66 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSRYGCGYGS 115
Score = 37.7 bits (86), Expect = 0.072
Identities = 23/53 (43%), Positives = 26/53 (48%), Gaps = 2/53 (3%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVG--VGLGWGFGSAFGSKYRSS 64
G G GCG+G G+G G G G G GCG G G G+G G K SS
Sbjct: 74 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSRYGCGYGSGCCSYRKCYSS 126
Score = 36.6 bits (83), Expect = 0.16
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 7/53 (13%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGG--GCGVGVGLGWGFGS 55
SG+ C + SG+G G G G G G+G G G G GCG G G G+GS
Sbjct: 67 SGYGCGYGSGYGCGYGSGYGCGYGSG----YGCGYGSGYGCGYGSRYGCGYGS 115
>gb|AAX95977.1| expressed protein [Oryza sativa (japonica cultivar-group)]
Length = 140
Score = 52.4 bits (124), Expect = 3e-06
Identities = 29/62 (46%), Positives = 32/62 (50%), Gaps = 9/62 (14%)
Query: 14 GFGVGCGFGVGWGFGG-------MPLNLLGLGVGGGCGVGVGLGWGFGS--AFGSKYRSS 64
G GVGCG G G GF G P LG GVG GCGVG G G+G G A+ R S
Sbjct: 38 GLGVGCGVGAGVGFFGGAGLGYGFPGLTLGFGVGAGCGVGFGFGYGLGKGIAYDQNKRYS 97
Query: 65 QI 66
+
Sbjct: 98 NV 99
Score = 32.0 bits (71), Expect = 3.9
Identities = 18/34 (52%), Positives = 19/34 (54%), Gaps = 6/34 (17%)
Query: 7 GFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVG 40
GF GFGVG G GVG+GFG GLG G
Sbjct: 60 GFPGLTLGFGVGAGCGVGFGFG------YGLGKG 87
>ref|NP_048495.2| Phe-, Gly-rich protein: RCGF 3X, GCGF 11X, RSGF 5X, GSGF 2X
[Paramecium bursaria Chlorella virus 1]
gi|52221429|gb|AAC96515.2| Phe-, Gly-rich protein: RCGF
3X, GCGF 11X, RSGF 5X, GSGF 2X [Paramecium bursaria
Chlorella virus 1]
Length = 260
Score = 52.0 bits (123), Expect = 4e-06
Identities = 27/54 (50%), Positives = 28/54 (51%), Gaps = 6/54 (11%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYR 62
C F GFG G G G G GFG G G G GCG G G G GFG FG +R
Sbjct: 68 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFGCGFR 115
Score = 50.8 bits (120), Expect = 8e-06
Identities = 26/50 (52%), Positives = 26/50 (52%), Gaps = 6/50 (12%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
C F GFG G G G G GFG G G G GCG G G G GFG FG
Sbjct: 64 CGFRCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFG 107
Score = 48.9 bits (115), Expect = 3e-05
Identities = 27/55 (49%), Positives = 27/55 (49%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C F GFG G G G G GFG G G G GCG G G G GFG F RS
Sbjct: 72 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFRCGLRS 120
Score = 47.0 bits (110), Expect = 1e-04
Identities = 26/55 (47%), Positives = 27/55 (48%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C F GFG G G G G GFG G G G GCG G G G GF S +RS
Sbjct: 76 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFRCGLRSGFRS 124
Score = 45.8 bits (107), Expect = 3e-04
Identities = 26/55 (47%), Positives = 27/55 (48%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C F GFG G G G G GFG G G G GCG G G G S F S +RS
Sbjct: 80 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFRCGLRSGFRSGFRS 128
Score = 42.7 bits (99), Expect = 0.002
Identities = 31/71 (43%), Positives = 33/71 (45%), Gaps = 9/71 (12%)
Query: 9 CLFLSGFGVGCGFGVGWGFG-GMPLNLL-----GLGVGGGCGVGVGLGWGFGSAFGSKYR 62
C F GFG G G G G GFG G L G G GC G G GFG FGS +
Sbjct: 92 CGFGCGFGCGFGCGFGCGFGCGFRCGLRSGFRSGFRSGFGCRFGCGFRSGFGCRFGSGFG 151
Query: 63 SSQIRFQGVEF 73
S F+GV F
Sbjct: 152 SG---FRGVLF 159
>pir||T17638 glycine tyrosine-rich protein a147L - Chlorella virus PBCV-1
Length = 260
Score = 52.0 bits (123), Expect = 4e-06
Identities = 27/54 (50%), Positives = 28/54 (51%), Gaps = 6/54 (11%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYR 62
C F GFG G G G G GFG G G G GCG G G G GFG FG +R
Sbjct: 64 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFGCGFR 111
Score = 50.8 bits (120), Expect = 8e-06
Identities = 26/50 (52%), Positives = 26/50 (52%), Gaps = 6/50 (12%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
C F GFG G G G G GFG G G G GCG G G G GFG FG
Sbjct: 60 CGFRCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFG 103
Score = 49.7 bits (117), Expect = 2e-05
Identities = 30/63 (47%), Positives = 31/63 (48%), Gaps = 14/63 (22%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFG--------SAFGSK 60
C F GFG G G G G GFG G G G GCG G G G GFG S F SK
Sbjct: 68 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFRCGLRSGFRSK 121
Query: 61 YRS 63
+RS
Sbjct: 122 FRS 124
Score = 39.3 bits (90), Expect = 0.025
Identities = 28/67 (41%), Positives = 30/67 (43%), Gaps = 5/67 (7%)
Query: 9 CLFLSGFGVG--CGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQI 66
C F GFG G CGF G G G G GCG G GFG FGS + S
Sbjct: 96 CGFGCGFGCGFGCGFRCGLRSGFRSKFRSGFGCRFGCGFRSGFRSGFGCRFGSGFGSG-- 153
Query: 67 RFQGVEF 73
F+GV F
Sbjct: 154 -FRGVLF 159
>ref|NP_034803.1| keratin associated protein 6-2 [Mus musculus]
gi|2130977|dbj|BAA20281.1| high-glycine tyrosine
keratin type II.4 [Mus musculus]
Length = 159
Score = 50.4 bits (119), Expect = 1e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 37 GSGYGCGYGSGYGCGSGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 86
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 69 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 118
Score = 49.3 bits (116), Expect = 2e-05
Identities = 28/59 (47%), Positives = 34/59 (57%), Gaps = 3/59 (5%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG+ C + SG+G G G G G G+G G G G GCG G G G G+GS +GS Y S
Sbjct: 70 SGYGCGYGSGYGCGYGSGYGCGYGSG--YGCGYGSGYGCGYGSGYGCGYGSGYGSGYGS 126
Score = 48.5 bits (114), Expect = 4e-05
Identities = 24/51 (47%), Positives = 28/51 (54%), Gaps = 8/51 (15%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG G GCG+G G+G G G G GCG G G G G+GS +G Y S
Sbjct: 52 SGSGYGCGYGSGYG--------CGYGSGYGCGYGSGYGCGYGSGYGCGYGS 94
Score = 48.1 bits (113), Expect = 5e-05
Identities = 28/61 (45%), Positives = 34/61 (54%), Gaps = 3/61 (4%)
Query: 4 SHSGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYR 62
S SG+ C + SG+G G G G G G+G G G G GCG G G G G+GS +G Y
Sbjct: 52 SGSGYGCGYGSGYGCGYGSGYGCGYGSG--YGCGYGSGYGCGYGSGYGCGYGSGYGCGYG 109
Query: 63 S 63
S
Sbjct: 110 S 110
Score = 45.8 bits (107), Expect = 3e-04
Identities = 25/55 (45%), Positives = 29/55 (52%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C + SG+G G G G G G+G G G G GCG G G G G GS +G Y S
Sbjct: 14 CGYGSGYGSGYGCGSGSGYG------CGYGSGYGCGYGSGYGCGSGSGYGCGYGS 62
Score = 44.7 bits (104), Expect = 6e-04
Identities = 23/50 (46%), Positives = 27/50 (54%), Gaps = 6/50 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G+G G G G G G+G G G G GCG G G G G+GS +G Y S
Sbjct: 35 GYGSGYGCGYGSGYG------CGSGSGYGCGYGSGYGCGYGSGYGCGYGS 78
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/44 (45%), Positives = 25/44 (56%)
Query: 15 FGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G CG+G G+G G G G G GCG G G G G+GS +G
Sbjct: 6 YGNSCGYGCGYGSGYGSGYGCGSGSGYGCGYGSGYGCGYGSGYG 49
Score = 38.5 bits (88), Expect = 0.042
Identities = 24/59 (40%), Positives = 30/59 (50%), Gaps = 11/59 (18%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG+ C + SG+G G G G G G+G + G G G GCG G +GS YRS
Sbjct: 94 SGYGCGYGSGYGCGYGSGYGCGYGSGYGSGYGSGYGSGCGCG----------YGSYYRS 142
Score = 31.2 bits (69), Expect = 6.7
Identities = 15/32 (46%), Positives = 16/32 (49%)
Query: 30 MPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
M N G G GCG G G G G+G GS Y
Sbjct: 1 MCCNYYGNSCGYGCGYGSGYGSGYGCGSGSGY 32
Score = 30.8 bits (68), Expect = 8.8
Identities = 15/31 (48%), Positives = 16/31 (51%)
Query: 33 NLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
N G G G G G G G G G GS +G Y S
Sbjct: 8 NSCGYGCGYGSGYGSGYGCGSGSGYGCGYGS 38
>ref|ZP_00497832.1| COG2378: Predicted transcriptional regulator [Burkholderia
pseudomallei S13]
Length = 337
Score = 49.7 bits (117), Expect = 2e-05
Identities = 25/54 (46%), Positives = 30/54 (55%), Gaps = 6/54 (11%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIR 67
GFG G GFG G+GFG G G G G G G G G+GFG FG + + + R
Sbjct: 21 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFGFGFGNIEAR 68
Score = 49.3 bits (116), Expect = 2e-05
Identities = 25/58 (43%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGV 71
GFG G GFG G+GFG G G G G G G G G+GFG FG + I + +
Sbjct: 19 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFGFGFGFGNIEARAI 70
Score = 48.5 bits (114), Expect = 4e-05
Identities = 24/48 (50%), Positives = 27/48 (56%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G GVG G G+G+GFG G G G G G G G G+GFG FG
Sbjct: 6 FAVGVGVGVGVGIGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 47
Score = 37.7 bits (86), Expect = 0.072
Identities = 19/42 (45%), Positives = 24/42 (56%), Gaps = 6/42 (14%)
Query: 17 VGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G F VG G G +G+G+G G G G G G+GFG FG
Sbjct: 2 LGIEFAVGVGVG------VGVGIGFGFGFGFGFGFGFGFGFG 37
Score = 32.0 bits (71), Expect = 3.9
Identities = 14/25 (56%), Positives = 17/25 (68%)
Query: 34 LLGLGVGGGCGVGVGLGWGFGSAFG 58
+LG+ G GVGVG+G GFG FG
Sbjct: 1 MLGIEFAVGVGVGVGVGIGFGFGFG 25
>ref|ZP_00452009.1| COG2378: Predicted transcriptional regulator [Burkholderia mallei
SAVP1]
Length = 335
Score = 49.7 bits (117), Expect = 2e-05
Identities = 25/54 (46%), Positives = 30/54 (55%), Gaps = 6/54 (11%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIR 67
GFG G GFG G+GFG G G G G G G G G+GFG FG + + + R
Sbjct: 19 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFGFGFGNIEAR 66
Score = 49.3 bits (116), Expect = 2e-05
Identities = 25/58 (43%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGV 71
GFG G GFG G+GFG G G G G G G G G+GFG FG + I + +
Sbjct: 17 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFGFGFGFGNIEARAI 68
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/45 (53%), Positives = 26/45 (57%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 11 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 49
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/45 (53%), Positives = 26/45 (57%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 13 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 51
Score = 47.4 bits (111), Expect = 9e-05
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G G GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 6 FAVGVGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 47
Score = 38.1 bits (87), Expect = 0.055
Identities = 20/42 (47%), Positives = 22/42 (51%), Gaps = 6/42 (14%)
Query: 17 VGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G F VG GFG G G G G G G G G+GFG FG
Sbjct: 2 LGIEFAVGVGFG------FGFGFGFGFGFGFGFGFGFGFGFG 37
>ref|XP_543648.1| PREDICTED: similar to Keratin 5 [Canis familiaris]
Length = 1609
Score = 49.7 bits (117), Expect = 2e-05
Identities = 24/48 (50%), Positives = 28/48 (58%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F + FG G G G G+GFGG + G G G G G G+G G GFG FG
Sbjct: 1120 FRNRFGAGAGAGGGYGFGGGAGSGFGFGGGAGAGFGLGGGAGFGGGFG 1167
Score = 39.3 bits (90), Expect = 0.025
Identities = 30/84 (35%), Positives = 38/84 (44%), Gaps = 22/84 (26%)
Query: 13 SGFGVGCGFGVGWGFGG----------MPLNLLGLGVGGG---------CGVGVGLGWGF 53
SGFG G G G+G G GG + G+G+GGG G G LG GF
Sbjct: 1525 SGFGGGLGGGLGGGLGGGLGGGGSGSYYSSSSGGVGLGGGLSVGGSGFSAGSGRSLGMGF 1584
Query: 54 GSAFGSKYRSSQIRFQGVEFDSKE 77
GS GS SS ++F S++
Sbjct: 1585 GSGGGS---SSSVKFVSTTSSSRK 1605
Score = 36.6 bits (83), Expect = 0.16
Identities = 23/45 (51%), Positives = 25/45 (55%), Gaps = 9/45 (20%)
Query: 20 GFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
GFG G GFGG G+GGG G G+G G G G GS Y SS
Sbjct: 1520 GFGGGSGFGG--------GLGGGLGGGLGGGLG-GGGSGSYYSSS 1555
Score = 35.4 bits (80), Expect = 0.36
Identities = 30/71 (42%), Positives = 34/71 (47%), Gaps = 11/71 (15%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGG--------CGVGVGLGWGFGSAFGSKYRSS 64
SGFG G GF G G GG LG G+GGG GVGLG G S GS + +
Sbjct: 1519 SGFGGGSGF--GGGLGGGLGGGLGGGLGGGGSGSYYSSSSGGVGLGGGL-SVGGSGFSAG 1575
Query: 65 QIRFQGVEFDS 75
R G+ F S
Sbjct: 1576 SGRSLGMGFGS 1586
>ref|NP_001008899.1| type II keratin Kb2 [Rattus norvegicus] gi|46485118|tpg|DAA02228.1|
TPA: type II keratin Kb2 [Rattus norvegicus]
Length = 685
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/45 (57%), Positives = 27/45 (59%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 104 GFGGGSGFGGGRGFGG------GSGFGGGSGFGGGSGFGGGSGFG 142
Score = 47.4 bits (111), Expect = 9e-05
Identities = 26/46 (56%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 97 SGFGGSGGFGGGSGFGG------GRGFGGGSGFGGGSGFGGGSGFG 136
Score = 46.2 bits (108), Expect = 2e-04
Identities = 27/58 (46%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQG 70
SGFG G GFG G GFGG G G GGG G G G G+G G G ++ F G
Sbjct: 109 SGFGGGRGFGGGSGFGG------GSGFGGGSGFGGGSGFGGGGFGGGRFGGGPGGFGG 160
Score = 37.0 bits (84), Expect = 0.12
Identities = 23/45 (51%), Positives = 23/45 (51%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG GFGG G G GGG G G G GFG G
Sbjct: 92 GFGGGSGFGGSGGFGG------GSGFGGGRGFGG--GSGFGGGSG 128
Score = 33.5 bits (75), Expect = 1.4
Identities = 20/45 (44%), Positives = 21/45 (46%), Gaps = 8/45 (17%)
Query: 22 GVGWGFGGMPLNLLG--------LGVGGGCGVGVGLGWGFGSAFG 58
G G GFGG G G GGG G G G G+G GS FG
Sbjct: 80 GYGGGFGGRSYGGFGGGSGFGGSGGFGGGSGFGGGRGFGGGSGFG 124
Score = 31.6 bits (70), Expect = 5.1
Identities = 24/61 (39%), Positives = 29/61 (47%), Gaps = 9/61 (14%)
Query: 1 KSLSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSK 60
KS+S S ++G+G G G GFGG G G GG G G G G G G GS
Sbjct: 72 KSISSS-----VAGYGGGFGGRSYGGFGGGS----GFGGSGGFGGGSGFGGGRGFGGGSG 122
Query: 61 Y 61
+
Sbjct: 123 F 123
>ref|NP_048388.1| contains Gly-rich motifs GAGLGTGF (5X); similar to Arabidopsis
Gly-rich protein, corresponds to Swiss-Prot Accession
Number P27843 [Paramecium bursaria Chlorella virus 1]
gi|624113|gb|AAC96408.1| contains Gly-rich motifs
GAGLGTGF (5X); similar to Arabidopsis Gly-rich protein,
corresponds to Swiss-Prot Accession Number P27843
[Paramecium bursaria Chlorella virus 1]
gi|7461240|pir||T17530 glycine-rich protein a40L -
Chlorella virus PBCV-1
Length = 129
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/51 (50%), Positives = 32/51 (61%), Gaps = 2/51 (3%)
Query: 11 FLSGFGVGCGFGVGWGFG-GMPLNL-LGLGVGGGCGVGVGLGWGFGSAFGS 59
F +G G G G G+G GFG G+ L G GVG G G+G G G GFG+ FG+
Sbjct: 20 FGAGLGTGFGAGLGTGFGTGLGAGLGAGFGVGFGAGLGTGFGAGFGTGFGA 70
Score = 48.5 bits (114), Expect = 4e-05
Identities = 24/49 (48%), Positives = 28/49 (56%), Gaps = 6/49 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
+GFG G G G G G G GLG G G G G GLG GFG+ FG+ +
Sbjct: 26 TGFGAGLGTGFGTGLGA------GLGAGFGVGFGAGLGTGFGAGFGTGF 68
Score = 47.4 bits (111), Expect = 9e-05
Identities = 24/46 (52%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G+G GFG GLG G G G+G GLG GFG FG+
Sbjct: 15 GLGTGFGAGLGTGFGA------GLGTGFGTGLGAGLGAGFGVGFGA 54
Score = 45.8 bits (107), Expect = 3e-04
Identities = 22/49 (44%), Positives = 28/49 (56%), Gaps = 6/49 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
+GFG G G G+G GFG +G G G G G G G G GFG+ G+ +
Sbjct: 34 TGFGTGLGAGLGAGFG------VGFGAGLGTGFGAGFGTGFGAGLGAGF 76
Score = 44.3 bits (103), Expect = 8e-04
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F +G G G G G G GFG GLG G G G G G G G G+ FG
Sbjct: 36 FGTGLGAGLGAGFGVGFGA------GLGTGFGAGFGTGFGAGLGAGFG 77
Score = 40.8 bits (94), Expect = 0.008
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G GFGVG+G G LG G G G G G G G G G G
Sbjct: 41 GAGLGAGFGVGFGAG------LGTGFGAGFGTGFGAGLGAGFGDG 79
Score = 39.7 bits (91), Expect = 0.019
Identities = 20/40 (50%), Positives = 23/40 (57%), Gaps = 6/40 (15%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
+GFGVG G G+G GFG G G G G G+G G G G
Sbjct: 46 AGFGVGFGAGLGTGFGA------GFGTGFGAGLGAGFGDG 79
Score = 37.7 bits (86), Expect = 0.072
Identities = 18/42 (42%), Positives = 23/42 (53%), Gaps = 6/42 (14%)
Query: 20 GFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G+G GFG GLG G G G+G G G G G+ G+ +
Sbjct: 13 GVGLGTGFGA------GLGTGFGAGLGTGFGTGLGAGLGAGF 48
>ref|YP_104084.1| hypothetical protein BMA2541 [Burkholderia mallei ATCC 23344]
gi|52429475|gb|AAU50068.1| hypothetical protein BMA2541
[Burkholderia mallei ATCC 23344]
Length = 188
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/45 (53%), Positives = 26/45 (57%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 77 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 115
Score = 41.6 bits (96), Expect = 0.005
Identities = 22/47 (46%), Positives = 25/47 (52%), Gaps = 6/47 (12%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+ G V GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 67 IRGRKVRFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 107
>gb|AAL36897.1| putative antimicrobial peptide [Litopenaeus setiferus]
Length = 188
Score = 48.9 bits (115), Expect = 3e-05
Identities = 30/81 (37%), Positives = 41/81 (50%), Gaps = 7/81 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRF----- 68
G G G G G+G G GG+ L GLG G G G+G GLG G G G + +S R+
Sbjct: 53 GLGGGLGGGLGGGLGGLGGGLGGLGGGLGGGLGGGLGGGLGGGLGGSHGTSDCRYWCKTP 112
Query: 69 --QGVEFDSKEKGDSPPLSKP 87
Q +S + ++P +KP
Sbjct: 113 EGQAYCCESAHEPETPVGTKP 133
Score = 40.8 bits (94), Expect = 0.008
Identities = 28/58 (48%), Positives = 31/58 (53%), Gaps = 4/58 (6%)
Query: 5 HSGFCLFLSGFGV---GCGFGVGWGFGGMPLNLLGLGVGGGCGVGV-GLGWGFGSAFG 58
+ GF L G GV G G GVG G GG LG G+GGG G G+ GLG G G G
Sbjct: 21 YRGFGQPLGGLGVPGGGVGVGVGGGLGGGLGGGLGGGLGGGLGGGLGGLGGGLGGLGG 78
Score = 39.7 bits (91), Expect = 0.019
Identities = 24/53 (45%), Positives = 27/53 (50%), Gaps = 8/53 (15%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCG--------VGVGLGWGFGSAFG 58
G GVG G G+G G GG LG G+GGG G +G GLG G G G
Sbjct: 37 GVGVGVGGGLGGGLGGGLGGGLGGGLGGGLGGLGGGLGGLGGGLGGGLGGGLG 89
Score = 39.3 bits (90), Expect = 0.025
Identities = 24/46 (52%), Positives = 27/46 (58%), Gaps = 2/46 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGG-GCGVGVGLGWGFGSAFG 58
G G G G G+G G GG L LG G+GG G G+G GLG G G G
Sbjct: 49 GLGGGLGGGLGGGLGG-GLGGLGGGLGGLGGGLGGGLGGGLGGGLG 93
>ref|XP_310735.2| ENSANGP00000007494 [Anopheles gambiae str. PEST]
gi|55243434|gb|EAA06609.2| ENSANGP00000007494
[Anopheles gambiae str. PEST]
Length = 171
Score = 48.5 bits (114), Expect = 4e-05
Identities = 21/46 (45%), Positives = 30/46 (64%), Gaps = 8/46 (17%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG+G GF +G GFG +G+G G G+G G+ +GFG ++G
Sbjct: 1 SGFGIGSGFEIGSGFG--------IGIGPGIGIGFGISFGFGVSYG 38
Score = 44.7 bits (104), Expect = 6e-04
Identities = 25/62 (40%), Positives = 30/62 (48%), Gaps = 4/62 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
GFG+ GFGV +G G G+G G G G G G+G GF GS S G+E
Sbjct: 26 GFGISFGFGVSYGIGSGIRIGSGIGTGSGIGTGSGIGTGFRIEIGSGIEIS----SGIEI 81
Query: 74 DS 75
S
Sbjct: 82 SS 83
Score = 40.8 bits (94), Expect = 0.008
Identities = 24/63 (38%), Positives = 31/63 (49%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVE 72
SG G+G G G+G GFG G V G G G+ +G G G+ G + S I G+E
Sbjct: 83 SGIGIGSGIGIGSGFGIGCGIRFGFRVSYGIGSGIIIGSGIGTGSGIENGSGIIVSSGIE 142
Query: 73 FDS 75
S
Sbjct: 143 IGS 145
Score = 39.3 bits (90), Expect = 0.025
Identities = 21/57 (36%), Positives = 30/57 (51%), Gaps = 12/57 (21%)
Query: 13 SGFGVGCGFGVGWGF-----------GGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G+G G G+G GF G+ ++ G+G+G G G+G G G G G FG
Sbjct: 51 TGSGIGTGSGIGTGFRIEIGSGIEISSGIEISS-GIGIGSGIGIGSGFGIGCGIRFG 106
>ref|XP_522676.1| PREDICTED: hypothetical protein XP_522676 [Pan troglodytes]
Length = 325
Score = 48.5 bits (114), Expect = 4e-05
Identities = 25/63 (39%), Positives = 33/63 (51%), Gaps = 2/63 (3%)
Query: 1 KSLSHSGFCLFLSGFG--VGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
K+ S G L+ G+G GCG+G +G G G G G GCG G G G G+G +G
Sbjct: 238 KAFSACGSHLYGCGYGSGYGCGYGSSYGCGYGTGYSCGYGSGSGCGYGTGYGCGYGCGYG 297
Query: 59 SKY 61
+ Y
Sbjct: 298 TGY 300
Score = 45.4 bits (106), Expect = 3e-04
Identities = 25/62 (40%), Positives = 31/62 (49%), Gaps = 6/62 (9%)
Query: 4 SHSGFCLFLSGFGVG------CGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAF 57
SH C + SG+G G CG+G G+ G + G G G GCG G G G G+G
Sbjct: 245 SHLYGCGYGSGYGCGYGSSYGCGYGTGYSCGYGSGSGCGYGTGYGCGYGCGYGTGYGCGC 304
Query: 58 GS 59
GS
Sbjct: 305 GS 306
Score = 40.0 bits (92), Expect = 0.014
Identities = 21/50 (42%), Positives = 25/50 (50%), Gaps = 6/50 (12%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
C + +G+ G G G G G+G G G G GCG G G G G GS G
Sbjct: 266 CGYGTGYSCGYGSGSGCGYG------TGYGCGYGCGYGTGYGCGCGSGSG 309
>ref|NP_034798.1| keratin complex 2, basic, gene 17 [Mus musculus]
gi|398168|emb|CAA52788.1| keratin 2 epidermis [Mus
musculus]
Length = 707
Score = 48.1 bits (113), Expect = 5e-05
Identities = 31/70 (44%), Positives = 32/70 (45%), Gaps = 6/70 (8%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVE 72
SGFG G GFG G GFGG G GGG G G G G+G GS FG F G
Sbjct: 94 SGFGGGRGFGGGQGFGGSG------GFGGGSGFGGGQGFGGGSRFGGGSGFGGGGFGGGS 147
Query: 73 FDSKEKGDSP 82
F G P
Sbjct: 148 FGGGRFGGGP 157
Score = 36.2 bits (82), Expect = 0.21
Identities = 26/57 (45%), Positives = 27/57 (46%), Gaps = 8/57 (14%)
Query: 4 SHSGFCLFLSGFGVGC--GFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
S G F G G G GFG GFGG G G GGG G G G +G GS FG
Sbjct: 89 SFGGSSGFGGGRGFGGGQGFGGSGGFGG------GSGFGGGQGFGGGSRFGGGSGFG 139
Score = 32.7 bits (73), Expect = 2.3
Identities = 22/47 (46%), Positives = 22/47 (46%), Gaps = 6/47 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G G G GG G G GGG G G G G GS GS
Sbjct: 596 SGGGYGSGGGSRGGSGG------GYGSGGGSGSGGGYSSGGGSRGGS 636
Score = 32.7 bits (73), Expect = 2.3
Identities = 24/57 (42%), Positives = 28/57 (49%), Gaps = 2/57 (3%)
Query: 4 SHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSK 60
S SG + SG G G G G+G GG G G G G G G G G G+ S GS+
Sbjct: 579 SGSGGGGYSSGGGSRGGSGGGYGSGGGSRG--GSGGGYGSGGGSGSGGGYSSGGGSR 633
>ref|XP_666332.1| hypothetical protein Chro.40165 [Cryptosporidium hominis]
gi|54657307|gb|EAL36102.1| hypothetical protein
Chro.40165 [Cryptosporidium hominis]
Length = 920
Score = 48.1 bits (113), Expect = 5e-05
Identities = 29/75 (38%), Positives = 36/75 (47%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
G G+G G G+G G G P +G+G+G G G+GVGLG G GS GS
Sbjct: 698 GPGIGLGIGMGPGLGMGPGLGIGMGMGIGMGMGVGLGLGLGSGSGSDSGLEMKMGSETSL 757
Query: 74 DSKEKGDSPPLSKPT 88
DS S S P+
Sbjct: 758 DSNSNPISKSSSSPS 772
Score = 43.9 bits (102), Expect = 0.001
Identities = 22/45 (48%), Positives = 26/45 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G G P LG+G+G G G+G GLG G G G
Sbjct: 682 GIGMGAGIGTGIGAGIGPGIGLGIGMGPGLGMGPGLGIGMGMGIG 726
Score = 42.4 bits (98), Expect = 0.003
Identities = 20/46 (43%), Positives = 27/46 (58%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G+G G G P +G G+G G G+G+G+G G G G
Sbjct: 691 TGIGAGIGPGIGLGIGMGPGLGMGPGLGIGMGMGIGMGMGVGLGLG 736
Score = 42.0 bits (97), Expect = 0.004
Identities = 24/60 (40%), Positives = 31/60 (51%), Gaps = 9/60 (15%)
Query: 2 SLSHSGFCL---FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
S+S SG F G G+G G G G G G +G G+G G G+G+G+G G G G
Sbjct: 663 SISDSGLATNPGFGPGLGIGIGMGAGIGTG------IGAGIGPGIGLGIGMGPGLGMGPG 716
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.147 0.481
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 196,936,766
Number of Sequences: 2540612
Number of extensions: 9323112
Number of successful extensions: 86720
Number of sequences better than 10.0: 1386
Number of HSP's better than 10.0 without gapping: 809
Number of HSP's successfully gapped in prelim test: 620
Number of HSP's that attempted gapping in prelim test: 52337
Number of HSP's gapped (non-prelim): 14587
length of query: 95
length of database: 863,360,394
effective HSP length: 71
effective length of query: 24
effective length of database: 682,976,942
effective search space: 16391446608
effective search space used: 16391446608
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0050.15


