

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047b.8
(171 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
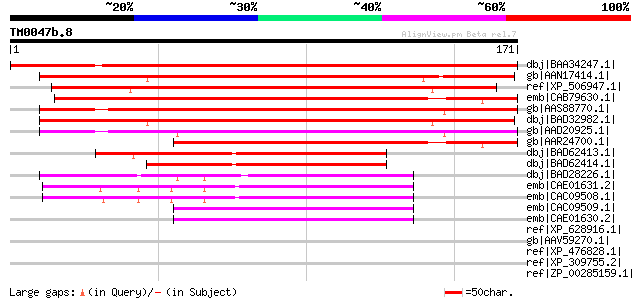
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAA34247.1| GPI-anchored protein [Vigna radiata] 278 3e-74
gb|AAN17414.1| putative protein [Arabidopsis thaliana] gi|975863... 207 1e-52
ref|XP_506947.1| PREDICTED P0654B04.18 gene product [Oryza sativ... 182 4e-45
emb|CAB79630.1| putative GPI-anchored protein [Arabidopsis thali... 157 1e-37
gb|AAS88770.1| At2g20700 [Arabidopsis thaliana] gi|45752642|gb|A... 153 2e-36
dbj|BAD32982.1| putative GPI-anchored protein [Oryza sativa (jap... 150 1e-35
gb|AAD20925.1| hypothetical protein [Arabidopsis thaliana] gi|25... 142 3e-33
gb|AAR24700.1| At4g28280 [Arabidopsis thaliana] gi|44681402|gb|A... 140 1e-32
dbj|BAD62413.1| putative GPI-anchored protein [Oryza sativa (jap... 97 2e-19
dbj|BAD62414.1| putative GPI-anchored protein [Oryza sativa (jap... 96 3e-19
dbj|BAD28226.1| unknown protein [Oryza sativa (japonica cultivar... 77 2e-13
emb|CAE01631.2| OSJNBa0029H02.12 [Oryza sativa (japonica cultiva... 72 6e-12
emb|CAC09508.1| hypothetical protein [Oryza sativa (indica culti... 72 8e-12
emb|CAC09509.1| hypothetical protein [Oryza sativa (indica culti... 70 3e-11
emb|CAE01630.2| OSJNBa0029H02.11 [Oryza sativa (japonica cultiva... 66 4e-10
ref|XP_628916.1| hypothetical protein DDB0192167 [Dictyostelium ... 35 1.1
gb|AAV59270.1| At3g19300 [Arabidopsis thaliana] gi|53828541|gb|A... 34 1.8
ref|XP_476828.1| hypothetical protein [Oryza sativa (japonica cu... 33 3.1
ref|XP_309755.2| ENSANGP00000012454 [Anopheles gambiae str. PEST... 33 4.1
ref|ZP_00285159.1| COG3464: Transposase and inactivated derivati... 33 4.1
>dbj|BAA34247.1| GPI-anchored protein [Vigna radiata]
Length = 169
Score = 278 bits (712), Expect = 3e-74
Identities = 136/171 (79%), Positives = 147/171 (85%), Gaps = 2/171 (1%)
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNF 60
M FSL++RF +SSS LLL ALSVSS + TFLSD VF SQAH GRNLLQAKKGCSVNF
Sbjct: 1 MVFSLHKRFWVSSSFLLLFWALSVSSLTP--TFLSDHVFDSQAHAGRNLLQAKKGCSVNF 58
Query: 61 EFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPP 120
EFLNYTIITSKCKGP YPPK+CCGAFKEFACPY DVLNDLTN+CASTMFSYINLYGKYPP
Sbjct: 59 EFLNYTIITSKCKGPQYPPKECCGAFKEFACPYADVLNDLTNECASTMFSYINLYGKYPP 118
Query: 121 GLFAHECREGKEGLACPALPPSALADDTSSQVVHFPSLVLVLTACFLILLF 171
GLFA EC + K GL CPALPPS ADDTS+Q +H PSL+L+LT CFLILLF
Sbjct: 119 GLFASECHDTKRGLECPALPPSVSADDTSNQFLHCPSLLLLLTTCFLILLF 169
>gb|AAN17414.1| putative protein [Arabidopsis thaliana] gi|9758637|dbj|BAB09299.1|
unnamed protein product [Arabidopsis thaliana]
gi|15241141|ref|NP_200428.1| expressed protein
[Arabidopsis thaliana] gi|24899659|gb|AAN65044.1|
putative protein [Arabidopsis thaliana]
Length = 168
Score = 207 bits (526), Expect = 1e-52
Identities = 104/164 (63%), Positives = 123/164 (74%), Gaps = 5/164 (3%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHT-GRNLLQAKKGCSVNFEFLNYTIIT 69
L S L L LSV SS SSS+F+SD VF SQ+ GRNLLQ KK C VNFEF+NYTIIT
Sbjct: 3 LLSRALFFFLLLSVLSSFSSSSFISDGVFESQSLVLGRNLLQTKKTCPVNFEFMNYTIIT 62
Query: 70 SKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECRE 129
SKCKGP YPPK+CCGAFK+FACPY D LNDL++DCA+TMFSYINLYGKYPPGLFA++C+E
Sbjct: 63 SKCKGPKYPPKECCGAFKDFACPYTDQLNDLSSDCATTMFSYINLYGKYPPGLFANQCKE 122
Query: 130 GKEGLACPA---LPPSALADDTSSQVVHFPSLVLVLTACFLILL 170
GKEGL CPA LPP A + ++ L L ++A L+ +
Sbjct: 123 GKEGLECPAGSQLPPETSA-EVNAATTSSSRLWLTVSAALLVFV 165
>ref|XP_506947.1| PREDICTED P0654B04.18 gene product [Oryza sativa (japonica
cultivar-group)] gi|50912235|ref|XP_467525.1| unknown
protein [Oryza sativa (japonica cultivar-group)]
gi|45735979|dbj|BAD13008.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 167
Score = 182 bits (461), Expect = 4e-45
Identities = 92/156 (58%), Positives = 111/156 (70%), Gaps = 6/156 (3%)
Query: 15 LLLLILALSVSSSSSSSTFLSDAVF-GSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCK 73
LLL++ A + +S+S F+SD+VF GS TGR+LLQAKK C VNFEF NYTIITSKCK
Sbjct: 8 LLLVVSAAVLVGLASASPFISDSVFLGSVGSTGRSLLQAKKNCPVNFEFQNYTIITSKCK 67
Query: 74 GPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECREGKEG 133
GP +P K CC AFKEFACP+ + +ND +NDCASTMFSYINLYGKYPPGLFA+ECREGK G
Sbjct: 68 GPRFPAKQCCDAFKEFACPFNEYINDESNDCASTMFSYINLYGKYPPGLFANECREGKLG 127
Query: 134 LACPALPP-----SALADDTSSQVVHFPSLVLVLTA 164
L+C + S+ S ++ F L L A
Sbjct: 128 LSCEGVSQKDSVVSSAGQQAQSSLLAFIMLTFGLAA 163
>emb|CAB79630.1| putative GPI-anchored protein [Arabidopsis thaliana]
gi|15235244|ref|NP_194557.1| expressed protein
[Arabidopsis thaliana] gi|7460059|pir||T09044
hypothetical protein F26K10.160 - Arabidopsis thaliana
Length = 160
Score = 157 bits (396), Expect = 1e-37
Identities = 83/159 (52%), Positives = 103/159 (64%), Gaps = 9/159 (5%)
Query: 16 LLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCKGP 75
L+ +L++ + S + S +S F S T R LLQAK C +F NYTIITSKCKGP
Sbjct: 8 LVSLLSILLLSGFAFSHHISLDEFESHPSTSRALLQAKATCKEDFAAKNYTIITSKCKGP 67
Query: 76 SYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECREGKEGLA 135
+YP K CC AFK+FACP+ +VLND DCASTMFSYINLYG+YPPG+FA+ C+EGKEGL
Sbjct: 68 NYPAKVCCSAFKDFACPFAEVLNDEKTDCASTMFSYINLYGRYPPGIFANMCKEGKEGLD 127
Query: 136 CPALPPSALADDTSSQVVHFPSL---VLVLTACFLILLF 171
C + P TSS P + VL++T L LF
Sbjct: 128 CTDVTP------TSSSHASIPLVSTHVLLITVSILFHLF 160
>gb|AAS88770.1| At2g20700 [Arabidopsis thaliana] gi|45752642|gb|AAS76219.1|
At2g20700 [Arabidopsis thaliana]
gi|30681101|ref|NP_179662.2| expressed protein
[Arabidopsis thaliana]
Length = 163
Score = 153 bits (386), Expect = 2e-36
Identities = 83/162 (51%), Positives = 103/162 (63%), Gaps = 5/162 (3%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITS 70
+S LL +L + + S S LS F A T R LLQ + C +F NYTIITS
Sbjct: 3 ISPYCLLSLLPIFLLSGFS----LSYDEFDGHAATSRALLQTRTTCKEDFANKNYTIITS 58
Query: 71 KCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECREG 130
+CKGP+YP CC AFK+FACP+ +VLND NDCASTMFSYINLYG+YPPG+FA+ C+EG
Sbjct: 59 RCKGPNYPANVCCSAFKDFACPFAEVLNDEKNDCASTMFSYINLYGRYPPGIFANMCKEG 118
Query: 131 KEGLACPALPPSALA-DDTSSQVVHFPSLVLVLTACFLILLF 171
KEGL C + SA A D+ + SL ++ T L LLF
Sbjct: 119 KEGLDCTDVTQSASATSDSIPRASTTASLAVLSTFLVLCLLF 160
>dbj|BAD32982.1| putative GPI-anchored protein [Oryza sativa (japonica
cultivar-group)] gi|50725088|dbj|BAD33221.1| putative
GPI-anchored protein [Oryza sativa (japonica
cultivar-group)]
Length = 174
Score = 150 bits (379), Expect = 1e-35
Identities = 77/170 (45%), Positives = 103/170 (60%), Gaps = 10/170 (5%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHT----GRNLLQAKKGCSVNFEFLNYT 66
L + + L AL+ ++S++ F+S+ G+ + GR+LLQAKK C VNFE NYT
Sbjct: 3 LHARVALFAAALAAVLAASTAGFISNEAVGASSAASGGAGRSLLQAKKDCPVNFEEANYT 62
Query: 67 IITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHE 126
+ITS+CKGP YPP CC A K+ ACP+ +ND CA++MFSYINLYGKYPPGLFA+
Sbjct: 63 VITSRCKGPMYPPALCCQALKDLACPFTAYINDAQTTCAASMFSYINLYGKYPPGLFANT 122
Query: 127 CREGKEGLACPALPP------SALADDTSSQVVHFPSLVLVLTACFLILL 170
C+EG GL CP P A ++ V VL + FL+L+
Sbjct: 123 CKEGANGLECPEDTPQMKPGEDKAASSAAAIVAAVARPVLAAVSAFLMLI 172
>gb|AAD20925.1| hypothetical protein [Arabidopsis thaliana] gi|25337807|pir||C84592
hypothetical protein At2g20700 [imported] - Arabidopsis
thaliana
Length = 181
Score = 142 bits (359), Expect = 3e-33
Identities = 83/180 (46%), Positives = 103/180 (57%), Gaps = 23/180 (12%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKG--------------- 55
+S LL +L + + S S LS F A T R LLQ +
Sbjct: 3 ISPYCLLSLLPIFLLSGFS----LSYDEFDGHAATSRALLQTRTNTLIFKYGSSVFFVGI 58
Query: 56 ---CSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYI 112
C +F NYTIITS+CKGP+YP CC AFK+FACP+ +VLND NDCASTMFSYI
Sbjct: 59 ETACKEDFANKNYTIITSRCKGPNYPANVCCSAFKDFACPFAEVLNDEKNDCASTMFSYI 118
Query: 113 NLYGKYPPGLFAHECREGKEGLACPALPPSALA-DDTSSQVVHFPSLVLVLTACFLILLF 171
NLYG+YPPG+FA+ C+EGKEGL C + SA A D+ + SL ++ T L LLF
Sbjct: 119 NLYGRYPPGIFANMCKEGKEGLDCTDVTQSASATSDSIPRASTTASLAVLSTFLVLCLLF 178
>gb|AAR24700.1| At4g28280 [Arabidopsis thaliana] gi|44681402|gb|AAS47641.1|
At4g28280 [Arabidopsis thaliana]
Length = 134
Score = 140 bits (354), Expect = 1e-32
Identities = 69/119 (57%), Positives = 82/119 (67%), Gaps = 9/119 (7%)
Query: 56 CSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLY 115
C +F NYTIITSKCKGP+YP K CC AFK+FACP+ +VLND DCASTMFSYINLY
Sbjct: 22 CKEDFAAKNYTIITSKCKGPNYPAKVCCSAFKDFACPFAEVLNDEKTDCASTMFSYINLY 81
Query: 116 GKYPPGLFAHECREGKEGLACPALPPSALADDTSSQVVHFPSL---VLVLTACFLILLF 171
G+YPPG+FA+ C+EGKEGL C + P TSS P + VL++T L LF
Sbjct: 82 GRYPPGIFANMCKEGKEGLDCTDVTP------TSSSHASIPLVSTHVLLITVSILFHLF 134
>dbj|BAD62413.1| putative GPI-anchored protein [Oryza sativa (japonica
cultivar-group)]
Length = 163
Score = 96.7 bits (239), Expect = 2e-19
Identities = 49/100 (49%), Positives = 63/100 (63%), Gaps = 3/100 (3%)
Query: 30 SSTFLSDAVFG--SQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFK 87
S LSD V + R+LLQAK GC V+FE NYT ITSKCK P +P CC A
Sbjct: 15 SPPILSDGVLQVMGTSMNRRSLLQAKGGCPVSFENQNYTTITSKCKSP-WPADLCCPALN 73
Query: 88 EFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHEC 127
EFAC + +ND + +CA +M+ Y+N +G YP GLF++EC
Sbjct: 74 EFACNFSQYINDESTNCAESMWVYLNAHGSYPAGLFSNEC 113
>dbj|BAD62414.1| putative GPI-anchored protein [Oryza sativa (japonica
cultivar-group)]
Length = 137
Score = 96.3 bits (238), Expect = 3e-19
Identities = 44/81 (54%), Positives = 57/81 (70%), Gaps = 1/81 (1%)
Query: 47 RNLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCAS 106
R+LLQAK GC V+FE NYT ITSKCK P +P CC A EFAC + +ND + +CA
Sbjct: 8 RSLLQAKGGCPVSFENQNYTTITSKCKSP-WPADLCCPALNEFACNFSQYINDESTNCAE 66
Query: 107 TMFSYINLYGKYPPGLFAHEC 127
+M+ Y+N +G YP GLF++EC
Sbjct: 67 SMWVYLNAHGSYPAGLFSNEC 87
>dbj|BAD28226.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|50251295|dbj|BAD28075.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 147
Score = 76.6 bits (187), Expect = 2e-13
Identities = 46/136 (33%), Positives = 69/136 (49%), Gaps = 13/136 (9%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKG--------CSVNFEF 62
LS +++ L + + S +++ F S + + T R L+ A C V F+
Sbjct: 15 LSMVIVMTQLPPTEADSVAAAEFASSDLKADKL-TSRKLMGAANAPAPAPLGMCPVRFDE 73
Query: 63 LN--YTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPP 120
+ +T + KCK S +CC AFKE ACP+ +LNDL N C MF +I+ YG+ PP
Sbjct: 74 MKGPFTELGKKCKAASVT--ECCDAFKEIACPHNTLLNDLNNGCGDDMFYFIHTYGRLPP 131
Query: 121 GLFAHECREGKEGLAC 136
G +C EG G+ C
Sbjct: 132 GTIFKKCVEGPYGMKC 147
>emb|CAE01631.2| OSJNBa0029H02.12 [Oryza sativa (japonica cultivar-group)]
gi|50926125|ref|XP_473056.1| OSJNBa0029H02.12 [Oryza
sativa (japonica cultivar-group)]
Length = 150
Score = 72.0 bits (175), Expect = 6e-12
Identities = 50/149 (33%), Positives = 71/149 (47%), Gaps = 25/149 (16%)
Query: 12 SSSLLLLILALSVSSSSS---------------SSTFLSDAVFGSQ---AHTGRNLLQAK 53
SS++L +ALSV ++++ + FLS A S A T R L ++
Sbjct: 3 SSTILFYCVALSVVAAAAVVSSAAEEAEGPQDEAGRFLSAATLASSDSDAKTSRRALTSQ 62
Query: 54 -----KGCSVNFEFLN-YTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCAST 107
K C V FE + + + +KC K+CC FK+ ACPY +LND+TN CA+
Sbjct: 63 EEIIAKPCPVEFEQVKGFGELGAKCNDKQ-TMKECCELFKKIACPYNHLLNDITNVCANE 121
Query: 108 MFSYINLYGKYPPGLFAHECREGKEGLAC 136
F I+ GK PG C EG G+ C
Sbjct: 122 FFYLIHTKGKLQPGTILENCNEGPMGINC 150
>emb|CAC09508.1| hypothetical protein [Oryza sativa (indica cultivar-group)]
Length = 151
Score = 71.6 bits (174), Expect = 8e-12
Identities = 50/150 (33%), Positives = 71/150 (47%), Gaps = 26/150 (17%)
Query: 12 SSSLLLLILALSVSSSSSS----------------STFLSDAVFGSQ---AHTGRNLLQA 52
SS++L +ALSV +++++ FLS A S A T R L +
Sbjct: 3 SSTILFYCVALSVVAAAAAVVSSAAEEAEGPQDEAGRFLSAATLASSDSDAKTSRRALTS 62
Query: 53 K-----KGCSVNFEFLN-YTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCAS 106
+ K C V FE + + + +KC K+CC FK+ ACPY +LND+TN CA+
Sbjct: 63 QEEIIAKPCPVEFEQVKGFGELGAKCNDKQ-TMKECCELFKKIACPYNHLLNDITNVCAN 121
Query: 107 TMFSYINLYGKYPPGLFAHECREGKEGLAC 136
F I+ GK PG C EG G+ C
Sbjct: 122 EFFYLIHTKGKLQPGTILENCNEGPMGINC 151
>emb|CAC09509.1| hypothetical protein [Oryza sativa (indica cultivar-group)]
Length = 149
Score = 69.7 bits (169), Expect = 3e-11
Identities = 33/81 (40%), Positives = 44/81 (53%)
Query: 56 CSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLY 115
C V F+ + I K + K CC AFK FACP+ ++ND+ N CA MF I+ Y
Sbjct: 69 CPVRFDKMKGPAIELGKKCKTTGVKVCCEAFKTFACPHNKLINDVNNGCADEMFYTIHTY 128
Query: 116 GKYPPGLFAHECREGKEGLAC 136
G+ PPG +C EG G+ C
Sbjct: 129 GQLPPGTIFKKCLEGPHGMKC 149
>emb|CAE01630.2| OSJNBa0029H02.11 [Oryza sativa (japonica cultivar-group)]
gi|50926113|ref|XP_473055.1| OSJNBa0029H02.11 [Oryza
sativa (japonica cultivar-group)]
Length = 149
Score = 65.9 bits (159), Expect = 4e-10
Identities = 32/81 (39%), Positives = 43/81 (52%)
Query: 56 CSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLY 115
C V F+ + I K + K CC AFK FACP+ ++ND+ N CA MF I+ Y
Sbjct: 69 CPVRFDKMKGPAIELGKKCKTTGVKVCCEAFKTFACPHNKLINDVNNGCADEMFYTIHTY 128
Query: 116 GKYPPGLFAHECREGKEGLAC 136
G+ PG +C EG G+ C
Sbjct: 129 GQLLPGTIFKKCLEGPHGMKC 149
>ref|XP_628916.1| hypothetical protein DDB0192167 [Dictyostelium discoideum]
gi|60462278|gb|EAL60504.1| hypothetical protein
DDB0192167 [Dictyostelium discoideum]
Length = 563
Score = 34.7 bits (78), Expect = 1.1
Identities = 29/101 (28%), Positives = 46/101 (44%), Gaps = 10/101 (9%)
Query: 25 SSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCG 84
SSSSSS+T + + + T +Q + + NF+ N I SK GP Y K
Sbjct: 300 SSSSSSTTTTTTPATKTSSSTSSPSIQPQTKINENFDESNSNI--SKIIGPKYRKKRLTD 357
Query: 85 AFKE--FACPYVDV------LNDLTNDCASTMFSYINLYGK 117
A KE A P++ ++D T A+ + Y++ + K
Sbjct: 358 ALKERSLAFPFISTSTFGFNIDDATEISANAISEYLHFHEK 398
>gb|AAV59270.1| At3g19300 [Arabidopsis thaliana] gi|53828541|gb|AAU94380.1|
At3g19300 [Arabidopsis thaliana]
gi|11994452|dbj|BAB02454.1| unnamed protein product
[Arabidopsis thaliana] gi|18402188|ref|NP_566630.1|
protein kinase family protein [Arabidopsis thaliana]
Length = 663
Score = 33.9 bits (76), Expect = 1.8
Identities = 22/77 (28%), Positives = 33/77 (42%), Gaps = 11/77 (14%)
Query: 50 LQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYV----------DVLND 99
L + GC ++F N+T++ S C + K CC F V V +D
Sbjct: 24 LLTEAGCPLDFTSSNFTLVASVCSNNTERAK-CCRYMNAFVAISVARYANYTADLGVTSD 82
Query: 100 LTNDCASTMFSYINLYG 116
LT C +T+ + LYG
Sbjct: 83 LTEICITTISRTMELYG 99
>ref|XP_476828.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|34393837|dbj|BAC83441.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 619
Score = 33.1 bits (74), Expect = 3.1
Identities = 15/32 (46%), Positives = 20/32 (61%)
Query: 128 REGKEGLACPALPPSALADDTSSQVVHFPSLV 159
REG+E +A P +PP SS+ +H PSLV
Sbjct: 40 REGEESVAQPPVPPPRRIATCSSEEIHLPSLV 71
>ref|XP_309755.2| ENSANGP00000012454 [Anopheles gambiae str. PEST]
gi|55244367|gb|EAA05540.2| ENSANGP00000012454 [Anopheles
gambiae str. PEST]
Length = 234
Score = 32.7 bits (73), Expect = 4.1
Identities = 15/41 (36%), Positives = 20/41 (48%)
Query: 121 GLFAHECREGKEGLACPALPPSALADDTSSQVVHFPSLVLV 161
G + EC G +G C LPPS + TSS PS ++
Sbjct: 152 GHYRCECPSGYKGRNCEILPPSMMTSTTSSSTTEQPSTTMM 192
>ref|ZP_00285159.1| COG3464: Transposase and inactivated derivatives [Enterococcus
faecium]
Length = 145
Score = 32.7 bits (73), Expect = 4.1
Identities = 29/115 (25%), Positives = 44/115 (38%), Gaps = 17/115 (14%)
Query: 17 LLILALSVSSSSSSSTFLSDAVFGSQAHTGRN--------------LLQAKKGCSVNFEF 62
L+I +S + T + DAVF HT RN + KK V FE
Sbjct: 16 LMITEVSYETFQKKKTLIVDAVFSPAPHTCRNCGSTVVDGNGKVIVVKNGKKETIVRFEQ 75
Query: 63 LNYTIITSKCKGPSYPPKDC---CGAFKEFACPYVDVLNDLTNDCASTMFSYINL 114
N+ + ++ K Y K+C A F P + N + AS + ++L
Sbjct: 76 YNHMPLVTRLKKQRYTCKNCRTHWTAQSYFVQPRHSIANHVRYKIASLLTEKVSL 130
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.137 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 273,189,100
Number of Sequences: 2540612
Number of extensions: 10534096
Number of successful extensions: 29396
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 29365
Number of HSP's gapped (non-prelim): 29
length of query: 171
length of database: 863,360,394
effective HSP length: 119
effective length of query: 52
effective length of database: 561,027,566
effective search space: 29173433432
effective search space used: 29173433432
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0047b.8


