

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0045.6
(214 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
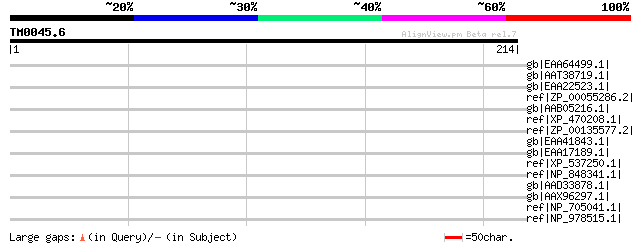
Score E
Sequences producing significant alignments: (bits) Value
gb|EAA64499.1| hypothetical protein AN2388.2 [Aspergillus nidula... 39 0.098
gb|AAT38719.1| putative F-Box protein [Solanum demissum] 38 0.17
gb|EAA22523.1| putative yir4 protein [Plasmodium yoelii yoelii] 37 0.37
ref|ZP_00055286.2| COG3263: NhaP-type Na+/H+ and K+/H+ antiporte... 37 0.49
gb|AAB05216.1| tail protein 35 1.1
ref|XP_470208.1| Putative retroelement [Oryza sativa (japonica c... 35 1.4
ref|ZP_00135577.2| COG1074: ATP-dependent exoDNAse (exonuclease ... 34 2.4
gb|EAA41843.1| GLP_158_8532_13532 [Giardia lamblia ATCC 50803] 34 3.2
gb|EAA17189.1| hypothetical protein [Plasmodium yoelii yoelii] 33 5.4
ref|XP_537250.1| PREDICTED: similar to death-associated protein ... 33 7.0
ref|NP_848341.1| enhancin-like [Choristoneura fumiferana MNPV] g... 33 7.0
gb|AAD33878.1| structural protein [Hepatitis E virus] 33 7.0
gb|AAX96297.1| retrotransposon protein, putative, unclassified [... 32 9.2
ref|NP_705041.1| methionyl-tRNA formyltransferase, putative [Pla... 32 9.2
ref|NP_978515.1| membrane-bound protease, putative [Bacillus cer... 32 9.2
>gb|EAA64499.1| hypothetical protein AN2388.2 [Aspergillus nidulans FGSC A4]
gi|67523865|ref|XP_659992.1| hypothetical protein
AN2388_2 [Aspergillus nidulans FGSC A4]
gi|49089872|ref|XP_406525.1| hypothetical protein
AN2388.2 [Aspergillus nidulans FGSC A4]
Length = 569
Score = 38.9 bits (89), Expect = 0.098
Identities = 22/79 (27%), Positives = 36/79 (44%), Gaps = 4/79 (5%)
Query: 27 HGYQNFQNMWRNIGAYEDG----SVRDMITLAKYNGGDLDTYFEHPDFEPLVEGPEALAQ 82
HG+ ++ +I Y+D ++R +I+ K N G D++ F P+ G L
Sbjct: 434 HGHGRGPEVYADITVYDDNDHLSNIRPVISCCKVNLGPNDSFLRAARFAPIFSGSHGLRI 493
Query: 83 VLGFSHVWHFAAPSNVKAF 101
+H W A P NV +F
Sbjct: 494 DYSDTHYWLIAWPVNVVSF 512
>gb|AAT38719.1| putative F-Box protein [Solanum demissum]
Length = 327
Score = 38.1 bits (87), Expect = 0.17
Identities = 34/112 (30%), Positives = 51/112 (45%), Gaps = 11/112 (9%)
Query: 99 KAFAWRFLLVK---IP-SRLNLWKLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNK 154
+A+ W + K +P SR NL G G FGY+ S+D++ VV I H +
Sbjct: 127 EAYIWNPTITKSKELPKSRSNLCSDGIKCG---FGYDESRDDYKVVFIDYPIHRHNHRTV 183
Query: 155 ITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQFLCDSLYWLKTKKATNVLV 206
+ I SLR + + + D F+ NL G+F+ LYW + N V
Sbjct: 184 VNIYSLR--TKSWTTLHDQLQGFFLLNLH--GRFVNGKLYWTSSSCINNYKV 231
>gb|EAA22523.1| putative yir4 protein [Plasmodium yoelii yoelii]
Length = 305
Score = 37.0 bits (84), Expect = 0.37
Identities = 32/121 (26%), Positives = 53/121 (43%), Gaps = 17/121 (14%)
Query: 69 DFEPLVEGPEAL-AQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLIS 127
DFE + G L Q+ G S ++ A SN+ ++L+ + LNL + G +
Sbjct: 45 DFERIDSGCLYLFKQIFGTSQLFKSVANSNINIVD--YILIWLSYMLNLKEDGKNVNNLQ 102
Query: 128 FGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQ 187
F YN + D K K +I + NY ++Y D+ D + YF+ + + I
Sbjct: 103 FFYNTTIDN--------------DKYKNSITGVENYYKNYKDLIDKKTYFFGMDRNIIFN 148
Query: 188 F 188
F
Sbjct: 149 F 149
>ref|ZP_00055286.2| COG3263: NhaP-type Na+/H+ and K+/H+ antiporters with a unique
C-terminal domain [Magnetospirillum magnetotacticum
MS-1]
Length = 597
Score = 36.6 bits (83), Expect = 0.49
Identities = 28/97 (28%), Positives = 47/97 (47%), Gaps = 1/97 (1%)
Query: 75 EGPEALAQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
+G LAQ++ F + A PS AFAW+ L+V L A L +G+NR++
Sbjct: 273 DGLTWLAQIVMFLTLGLLATPSEFPAFAWQSLMVAAVLIFVARPLAAWICLFPYGFNRNE 332
Query: 135 DEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIA 171
F V + L + + + + ++ + RDYF +A
Sbjct: 333 TAF-VAWVGLRGSVSVLLSIVPMVVGLPHGRDYFILA 368
>gb|AAB05216.1| tail protein
Length = 595
Score = 35.4 bits (80), Expect = 1.1
Identities = 42/163 (25%), Positives = 64/163 (38%), Gaps = 43/163 (26%)
Query: 34 NMWRNIGAYEDGSVRDMITLAKY---------NGGDLDTYFEHPDFE-PLVEGPEALAQV 83
N++R AY ++ TLA + N G++D F HP+++ P+ G
Sbjct: 167 NLYRPGRAYAAKAMGRAWTLAGFPAGASKVSINSGNVD--FWHPNWQAPIGAGKRG---- 220
Query: 84 LGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRS--KDEFLVVL 141
H W + SN++ +A K G T +S YN S D+ V
Sbjct: 221 ----HTWFYVCNSNIRGYA--------------GKKGTTPSNVSVLYNSSTYPDKVYVCR 262
Query: 142 IKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDY 184
NL A K YV+D ++I T+ +Y NL Y
Sbjct: 263 GSTSNLASQAAKK-------QYVQDSYNITPTKVIWYVLNLRY 298
>ref|XP_470208.1| Putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|27497213|gb|AAO17357.1| Putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|15451617|gb|AAK98741.1| Hypothetical protein similar
to putative retroelements [Oryza sativa]
Length = 387
Score = 35.0 bits (79), Expect = 1.4
Identities = 15/33 (45%), Positives = 21/33 (63%)
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWK 118
+S +W AAP VK FAW ++P+R+NL K
Sbjct: 226 WSAIWRSAAPPRVKFFAWLMSKNRLPTRVNLHK 258
>ref|ZP_00135577.2| COG1074: ATP-dependent exoDNAse (exonuclease V) beta subunit
(contains helicase and exonuclease domains)
[Actinobacillus pleuropneumoniae serovar 1 str. 4074]
Length = 1202
Score = 34.3 bits (77), Expect = 2.4
Identities = 42/149 (28%), Positives = 60/149 (40%), Gaps = 31/149 (20%)
Query: 58 GGDLDTYFEHPDFEPLVEGPEAL--AQVLGFSHVWH----------FAAPSNVKAFAWRF 105
G L ++FEH DF+ V+ + L + LG WH A P + FA +
Sbjct: 981 GNILHSFFEHSDFQQAVDFEQILTICEQLGLDENWHEPLQKWFEKVLATPFSETEFALKD 1040
Query: 106 LLVKIPSRLNLW----KLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLR 161
V I RLN W +L + GL N+ ++ V KL +L + L
Sbjct: 1041 --VPITKRLNEWQFYLRLKNSDGLRKL--NQLLKQYSAVSAKLPDL--------NLPQLE 1088
Query: 162 NYVRDYFD-IADTEYYFYKFNLDYIGQFL 189
Y+R + D I FY +DY FL
Sbjct: 1089 GYLRGFVDCIVQVNEKFYL--IDYKSNFL 1115
>gb|EAA41843.1| GLP_158_8532_13532 [Giardia lamblia ATCC 50803]
Length = 1666
Score = 33.9 bits (76), Expect = 3.2
Identities = 15/36 (41%), Positives = 25/36 (68%), Gaps = 1/36 (2%)
Query: 137 FLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIAD 172
F +V++KLCN L +N+++ S+ ++RDY IAD
Sbjct: 598 FFLVILKLCNF-RLVQNQVSYNSVLTFLRDYVKIAD 632
>gb|EAA17189.1| hypothetical protein [Plasmodium yoelii yoelii]
Length = 677
Score = 33.1 bits (74), Expect = 5.4
Identities = 18/55 (32%), Positives = 32/55 (57%), Gaps = 3/55 (5%)
Query: 160 LRNYVRDYFDIADTEYYFYKFNLDYIGQFLCD---SLYWLKTKKATNVLVVVRKL 211
+RN V +A+ YY+ KF +I + + + +L++ TK +NV++VVR L
Sbjct: 175 MRNVVDILISLANVNYYYDKFINFFIYEIIKNEKNNLFFFNTKNMSNVILVVRSL 229
>ref|XP_537250.1| PREDICTED: similar to death-associated protein 3 [Canis familiaris]
Length = 393
Score = 32.7 bits (73), Expect = 7.0
Identities = 21/73 (28%), Positives = 35/73 (47%), Gaps = 9/73 (12%)
Query: 62 DTYFEHPDFEPLVEGPEALAQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGA 121
+T F HP ++ G + + L HV HF A + +L++ IP +LW +
Sbjct: 109 NTNFAHPAVRYVLYGEKGTGKTLSLCHVVHFCAKQD-------WLILHIPD-AHLW-VKN 159
Query: 122 TRGLISFGYNRSK 134
RGL+ YN+ +
Sbjct: 160 CRGLLQSSYNKQR 172
>ref|NP_848341.1| enhancin-like [Choristoneura fumiferana MNPV]
gi|30270004|gb|AAP29820.1| enhancin-like [Choristoneura
fumiferana MNPV]
Length = 758
Score = 32.7 bits (73), Expect = 7.0
Identities = 21/66 (31%), Positives = 32/66 (47%), Gaps = 13/66 (19%)
Query: 25 IAHGYQNFQNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFEH-------PDFEPLVEGP 77
++H +F N+W +D +TL+ ++ D D YFEH P FE L+ G
Sbjct: 45 VSHNDVSF-NLWN-----DDSETEQNLTLSAHDMSDPDDYFEHVVEHDSVPFFECLLNGN 98
Query: 78 EALAQV 83
E AQ+
Sbjct: 99 EYTAQI 104
>gb|AAD33878.1| structural protein [Hepatitis E virus]
Length = 436
Score = 32.7 bits (73), Expect = 7.0
Identities = 16/66 (24%), Positives = 36/66 (54%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L ++ V P S ++ ++ F ++ + +L+ W+ G T+ + YN +
Sbjct: 282 GAQAVARSLDWTKVTLDGRPLSTIQQYSKTFFVLPLRGKLSFWEAGTTKAWYPYNYNTTA 341
Query: 135 DEFLVV 140
+ L+V
Sbjct: 342 SDQLLV 347
>gb|AAX96297.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)]
Length = 281
Score = 32.3 bits (72), Expect = 9.2
Identities = 13/36 (36%), Positives = 20/36 (55%)
Query: 89 VWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRG 124
+W F P +K F W F +I ++ NL+K G +G
Sbjct: 148 IWKFRTPLKIKVFLWLFTKNRILTKDNLYKRGWRKG 183
>ref|NP_705041.1| methionyl-tRNA formyltransferase, putative [Plasmodium falciparum
3D7] gi|23615286|emb|CAD52276.1| methionyl-tRNA
formyltransferase, putative [Plasmodium falciparum 3D7]
Length = 665
Score = 32.3 bits (72), Expect = 9.2
Identities = 21/73 (28%), Positives = 29/73 (38%), Gaps = 7/73 (9%)
Query: 130 YNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQFL 189
Y D+ ++ I L N L + K K NY+R Y YY YK NL +
Sbjct: 131 YKNQLDDINILYILLFNTLMIYKKKYDFFMNENYIRSY-------YYIYKNNLGRDKKIY 183
Query: 190 CDSLYWLKTKKAT 202
Y++ T T
Sbjct: 184 NTKNYFINTFSIT 196
>ref|NP_978515.1| membrane-bound protease, putative [Bacillus cereus ATCC 10987]
gi|42737190|gb|AAS41123.1| membrane-bound protease,
putative [Bacillus cereus ATCC 10987]
Length = 742
Score = 32.3 bits (72), Expect = 9.2
Identities = 21/69 (30%), Positives = 36/69 (51%), Gaps = 5/69 (7%)
Query: 150 LAKNKIT----IISLRNYVRDY-FDIADTEYYFYKFNLDYIGQFLCDSLYWLKTKKATNV 204
L K+K T ++++ NY D+ F T F N DY+ QFL D+ +T++
Sbjct: 428 LTKDKDTRYDKVLAIENYFTDHSFVYETTNVSFPTKNQDYVDQFLFDTKSGYCNNFSTSM 487
Query: 205 LVVVRKLGL 213
+V++R G+
Sbjct: 488 IVLLRSAGI 496
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.144 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 373,916,790
Number of Sequences: 2540612
Number of extensions: 15486053
Number of successful extensions: 39000
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 38993
Number of HSP's gapped (non-prelim): 18
length of query: 214
length of database: 863,360,394
effective HSP length: 123
effective length of query: 91
effective length of database: 550,865,118
effective search space: 50128725738
effective search space used: 50128725738
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0045.6


