

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
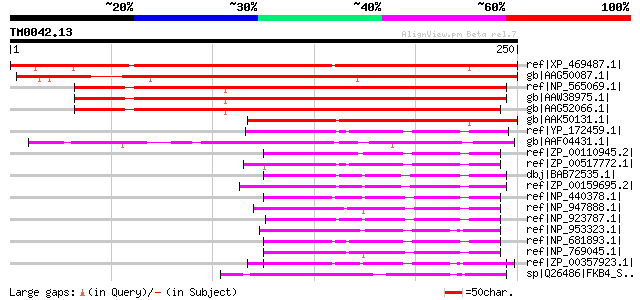
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_469487.1| putative FK506-binding protein [Oryza sativa] g... 315 6e-85
gb|AAG50087.1| unknown protein [Arabidopsis thaliana] gi|2159373... 287 2e-76
ref|NP_565069.1| immunophilin / FKBP-type peptidyl-prolyl cis-tr... 238 9e-62
gb|AAW38975.1| At1g73655 [Arabidopsis thaliana] gi|21592999|gb|A... 237 2e-61
gb|AAG52066.1| unknown protein; 19725-16797 [Arabidopsis thalian... 235 1e-60
gb|AAK50131.1| putative FK506-binding protein, 5'-partial [Oryza... 192 9e-48
ref|YP_172459.1| FKBP-type peptidyl-prolyl cis-trans isomerase [... 75 2e-12
gb|AAF04431.1| unknown protein [Arabidopsis thaliana] gi|2168961... 73 6e-12
ref|ZP_00110945.2| COG0545: FKBP-type peptidyl-prolyl cis-trans ... 73 8e-12
ref|ZP_00517772.1| Peptidylprolyl isomerase [Crocosphaera watson... 72 2e-11
dbj|BAB72535.1| FKBP-type peptidyl-prolyl cis-trans isomerase [N... 69 9e-11
ref|ZP_00159695.2| COG0545: FKBP-type peptidyl-prolyl cis-trans ... 69 2e-10
ref|NP_440378.1| FKBP-type peptidyl-prolyl cis-trans isomerase [... 66 1e-09
ref|NP_947888.1| FKBP-type peptidyl-prolyl cis-trans isomerase [... 64 4e-09
ref|NP_923787.1| FKBP-type peptidyl-prolyl cis-trans isomerase [... 64 4e-09
ref|NP_953323.1| FKBP-type peptidyl-prolyl cis-trans isomerase [... 64 5e-09
ref|NP_681893.1| FKBP-type peptidyl-prolyl cis-trans isomerase [... 63 7e-09
ref|NP_769045.1| Peptidylprolyl isomerase [Bradyrhizobium japoni... 63 7e-09
ref|ZP_00357923.1| COG0545: FKBP-type peptidyl-prolyl cis-trans ... 63 7e-09
sp|Q26486|FKB4_SPOFR 46 kDa FK506-binding nuclear protein (Pepti... 63 9e-09
>ref|XP_469487.1| putative FK506-binding protein [Oryza sativa]
gi|62733558|gb|AAX95675.1| peptidyl-prolyl cis-trans
isomerase, FKBP-type, putative [Oryza sativa (japonica
cultivar-group)]
Length = 252
Score = 315 bits (808), Expect = 6e-85
Identities = 160/255 (62%), Positives = 200/255 (77%), Gaps = 8/255 (3%)
Query: 1 MATFFGSPPVF-SLPLTRNSHHIASSSPTPP---PIPPQQSQSPTASSSQSVQGTAKAQP 56
MATF GS P F + P+ H + SP PP P P QQ +PT + QS+Q +A+P
Sbjct: 1 MATFLGSSPAFLARPVAAKPHLSCAQSPRPPSAQPPPEQQPPTPTPTQQQSMQAQPQARP 60
Query: 57 QNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEAN 116
+ P +++ DSTDW+ATSLTRRFG+GAGLAWVGFLAFGVVSEQ+KTR EV+QQ AN
Sbjct: 61 RRA--PAAAASGADSTDWVATSLTRRFGIGAGLAWVGFLAFGVVSEQLKTRFEVAQQLAN 118
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
T+DVE+E+EV LPNGIRY E++VGGG PRPGDLVVID+ G+V + GE FV+TF K+P
Sbjct: 119 TKDVEQEQEVVLPNGIRYYEMRVGGGDVPRPGDLVVIDLKGRV-TGGEAFVDTFGDGKRP 177
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLG-SGVQIPPLAT 235
LALVMGSRPY++G+CEG+EYVL++M+AGGKR+V+VPP LGF ++GAD G + Q+PP AT
Sbjct: 178 LALVMGSRPYTRGMCEGVEYVLRSMRAGGKRRVVVPPALGFGDDGADFGDAAAQVPPGAT 237
Query: 236 LEYIVEVDKVSIAPA 250
LEY+VEVDKVSIAPA
Sbjct: 238 LEYVVEVDKVSIAPA 252
>gb|AAG50087.1| unknown protein [Arabidopsis thaliana] gi|21593730|gb|AAM65697.1|
putative FK506-binding protein [Arabidopsis thaliana]
gi|8671768|gb|AAF78374.1| T10O22.14 [Arabidopsis
thaliana] gi|18394580|ref|NP_564048.1| immunophilin /
FKBP-type peptidyl-prolyl cis-trans isomerase family
protein [Arabidopsis thaliana] gi|9719731|gb|AAF97833.1|
Contains weak similarity to immunophilin FKBP46 from
Spodoptera frugiperda gb|U15038 and contains a FKBP-type
peptidyl-prolyl cis-trans isomerase PF|00254 domain.
ESTs gb|T42383, gb|N38271, gb|R90399, gb|AA605386,
gb|AA394960 come from this gene. [Arabidopsis thaliana]
Length = 247
Score = 287 bits (735), Expect = 2e-76
Identities = 152/258 (58%), Positives = 192/258 (73%), Gaps = 26/258 (10%)
Query: 4 FFGSPPVFSL--PLTRN-SHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI 60
F + P SL P T+ S+H +S + PP P S P P P
Sbjct: 5 FTATAPFLSLSKPFTKTASYHQCYASSSNPPEPESSSPPP---------------PPPPP 49
Query: 61 KPVTSSTK----VDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEAN 116
+P+ S K V++TDW+A+SLTRRFG+GAGLAW GFLAFGV+SEQIKTR+EVSQ+ AN
Sbjct: 50 QPLASQQKRKKNVETTDWVASSLTRRFGIGAGLAWAGFLAFGVISEQIKTRIEVSQEVAN 109
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTF----EG 172
TRDVE+E+E+ LPNGIRY + +VGGGA+PR GDLVVID+ G+V+ TG+VFV+TF +
Sbjct: 110 TRDVEEEKEIVLPNGIRYYDQRVGGGATPRAGDLVVIDLKGQVQGTGQVFVDTFGTKDKK 169
Query: 173 DKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPP 232
KPLALV+GS+PYSKG+CEGI+YVL++MKAGGKR+VIVPP LGF +GA+L SG+QIPP
Sbjct: 170 KMKPLALVVGSKPYSKGLCEGIDYVLRSMKAGGKRRVIVPPSLGFGVDGAELESGLQIPP 229
Query: 233 LATLEYIVEVDKVSIAPA 250
A+LEYIVE+D+VSIAPA
Sbjct: 230 NASLEYIVEIDRVSIAPA 247
>ref|NP_565069.1| immunophilin / FKBP-type peptidyl-prolyl cis-trans isomerase family
protein [Arabidopsis thaliana]
Length = 234
Score = 238 bits (608), Expect = 9e-62
Identities = 121/214 (56%), Positives = 154/214 (71%), Gaps = 5/214 (2%)
Query: 33 PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWV 92
P Q + S S SV T QP V + + TDWI + +TRRFG+GAG W
Sbjct: 18 PSQHPKQSLLSQSLSVTFTENPQPT----AVVTLQEQQLTDWITSPVTRRFGIGAGFTWA 73
Query: 93 GFLAFGVVSEQIK-TRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
GFLAFGVVSEQ+K +RL+V Q+E NTR +EK+EE+ LPNGIRY +L+VG GA+P G LV
Sbjct: 74 GFLAFGVVSEQMKKSRLDVFQEEDNTRGLEKQEEIILPNGIRYYDLQVGSGATPSSGYLV 133
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIV 211
V D+ G+V T +VFV+TF G K LA+VM SRPYSKG+C+GIE+VL++MKAGGKRKVI+
Sbjct: 134 VFDVKGQVHGTEQVFVDTFGGKGKSLAMVMDSRPYSKGLCQGIEHVLRSMKAGGKRKVII 193
Query: 212 PPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
PP LGF + + G G++IPP ATL+YI+EVD V
Sbjct: 194 PPSLGFGDRNVEFGQGLEIPPSATLDYIIEVDTV 227
>gb|AAW38975.1| At1g73655 [Arabidopsis thaliana] gi|21592999|gb|AAM64948.1|
putative FK506-binding protein [Arabidopsis thaliana]
gi|28393660|gb|AAO42248.1| putative peptidylprolyl
isomerase [Arabidopsis thaliana]
gi|60543363|gb|AAX22279.1| At1g73655 [Arabidopsis
thaliana]
Length = 234
Score = 237 bits (605), Expect = 2e-61
Identities = 120/214 (56%), Positives = 154/214 (71%), Gaps = 5/214 (2%)
Query: 33 PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWV 92
P Q + S S SV T QP V + + TDWI + +TRRFG+GAG W
Sbjct: 18 PSQHPKQSLLSQSLSVTFTENPQPT----AVVTLQEQQLTDWITSPVTRRFGIGAGFTWA 73
Query: 93 GFLAFGVVSEQIK-TRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
GFLAFGVVSEQ+K +RL+V Q+E NTR +EK+EE+ LPNGIRY +L+VG GA+P G LV
Sbjct: 74 GFLAFGVVSEQMKKSRLDVFQEEDNTRGLEKQEEIILPNGIRYYDLQVGSGATPSSGYLV 133
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIV 211
V D+ G+V T +VFV+TF G K LA+VM SRPYSKG+C+GIE+VL++MKAGGKR+VI+
Sbjct: 134 VFDVKGQVHGTEQVFVDTFGGKGKSLAMVMDSRPYSKGLCQGIEHVLRSMKAGGKRRVII 193
Query: 212 PPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
PP LGF + + G G++IPP ATL+YI+EVD V
Sbjct: 194 PPSLGFGDRNVEFGQGLEIPPSATLDYIIEVDTV 227
>gb|AAG52066.1| unknown protein; 19725-16797 [Arabidopsis thaliana]
gi|25406281|pir||E96763 unknown protein F25P22.7
[imported] - Arabidopsis thaliana
Length = 451
Score = 235 bits (599), Expect = 1e-60
Identities = 119/211 (56%), Positives = 152/211 (71%), Gaps = 5/211 (2%)
Query: 33 PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWV 92
P Q + S S SV T QP V + + TDWI + +TRRFG+GAG W
Sbjct: 18 PSQHPKQSLLSQSLSVTFTENPQPT----AVVTLQEQQLTDWITSPVTRRFGIGAGFTWA 73
Query: 93 GFLAFGVVSEQIK-TRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
GFLAFGVVSEQ+K +RL+V Q+E NTR +EK+EE+ LPNGIRY +L+VG GA+P G LV
Sbjct: 74 GFLAFGVVSEQMKKSRLDVFQEEDNTRGLEKQEEIILPNGIRYYDLQVGSGATPSSGYLV 133
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIV 211
V D+ G+V T +VFV+TF G K LA+VM SRPYSKG+C+GIE+VL++MKAGGKRKVI+
Sbjct: 134 VFDVKGQVHGTEQVFVDTFGGKGKSLAMVMDSRPYSKGLCQGIEHVLRSMKAGGKRKVII 193
Query: 212 PPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
PP LGF + + G G++IPP ATL+YI+EV
Sbjct: 194 PPSLGFGDRNVEFGQGLEIPPSATLDYIIEV 224
>gb|AAK50131.1| putative FK506-binding protein, 5'-partial [Oryza sativa]
Length = 149
Score = 192 bits (487), Expect = 9e-48
Identities = 93/134 (69%), Positives = 115/134 (85%), Gaps = 2/134 (1%)
Query: 118 RDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPL 177
RDVE+E+EV LPNGIRY E++VGGG PRPGDLVVID+ G+V GE FV+TF K+PL
Sbjct: 17 RDVEQEQEVVLPNGIRYYEMRVGGGDVPRPGDLVVIDLKGRVTG-GEAFVDTFGDGKRPL 75
Query: 178 ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLG-SGVQIPPLATL 236
ALVMGSRPY++G+CEG+EYVL++M+AGGKR+V+VPP LGF ++GAD G + Q+PP ATL
Sbjct: 76 ALVMGSRPYTRGMCEGVEYVLRSMRAGGKRRVVVPPALGFGDDGADFGDAAAQVPPGATL 135
Query: 237 EYIVEVDKVSIAPA 250
EY+VEVDKVSIAPA
Sbjct: 136 EYVVEVDKVSIAPA 149
>ref|YP_172459.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Synechococcus
elongatus PCC 6301] gi|56686717|dbj|BAD79939.1|
FKBP-type peptidyl-prolyl cis-trans isomerase
[Synechococcus elongatus PCC 6301]
gi|46130465|ref|ZP_00165335.2| COG0545: FKBP-type
peptidyl-prolyl cis-trans isomerases 1 [Synechococcus
elongatus PCC 7942]
Length = 174
Score = 75.1 bits (183), Expect = 2e-12
Identities = 48/130 (36%), Positives = 73/130 (55%), Gaps = 10/130 (7%)
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
++ + E VT +G++Y +L+VG GA+P+PG VV+ G +E G F ++ +P
Sbjct: 54 SQTTQSETIVTTASGLQYVDLEVGTGATPQPGQTVVVHYTGTLED-GTQF-DSSRDRNRP 111
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATL 236
+G KG EGI TMK GG+RK+ +PP L + E GA G IPP ATL
Sbjct: 112 FQFKLGVGQVIKGWDEGI----ATMKVGGRRKLTIPPTLAYGERGA----GGVIPPNATL 163
Query: 237 EYIVEVDKVS 246
+ VE+ +++
Sbjct: 164 IFDVELIRIA 173
>gb|AAF04431.1| unknown protein [Arabidopsis thaliana] gi|21689613|gb|AAM67428.1|
AT3g10060/T22K18_11 [Arabidopsis thaliana]
gi|20453145|gb|AAM19814.1| AT3g10060/T22K18_11
[Arabidopsis thaliana] gi|15232828|ref|NP_187617.1|
immunophilin, putative / FKBP-type peptidyl-prolyl
cis-trans isomerase, putative [Arabidopsis thaliana]
Length = 230
Score = 73.2 bits (178), Expect = 6e-12
Identities = 67/249 (26%), Positives = 110/249 (43%), Gaps = 39/249 (15%)
Query: 10 VFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKA-QPQNPIKPVTSSTK 68
+ ++ L HH SSS P P ++ QS S G K + + + P S +
Sbjct: 2 ILTMKLVHPLHHSLSSSI---PFPSRKRQSKPYRCSLPSPGCEKVIRTETVLPPAPVSCE 58
Query: 69 VDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTL 128
RR LG LA A G++S + S++ + + + TL
Sbjct: 59 -----------GRRVLLGCLLA----TASGILSTGSAEAVSTSRRALRASKLPESDFTTL 103
Query: 129 PNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYS- 187
PNG++Y ++KVG GA G V + + K + G F+ + +G V G PY
Sbjct: 104 PNGLKYYDIKVGNGAEAVKGSRVAVHYVAKWK--GITFMTSRQG-----LGVGGGTPYGF 156
Query: 188 -------KGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV 240
V +G++ ++ M+ GG+R VIVPP+L + + G +IPP AT+E +
Sbjct: 157 DVGQSERGNVLKGLDLGVEGMRVGGQRLVIVPPELAYGKKGVQ-----EIPPNATIELDI 211
Query: 241 EVDKVSIAP 249
E+ + +P
Sbjct: 212 ELLSIKQSP 220
>ref|ZP_00110945.2| COG0545: FKBP-type peptidyl-prolyl cis-trans isomerases 1 [Nostoc
punctiforme PCC 73102]
Length = 163
Score = 72.8 bits (177), Expect = 8e-12
Identities = 45/117 (38%), Positives = 67/117 (56%), Gaps = 10/117 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT P+G++Y ELK G GA+P+PG V + +G +E + + G +P + +G
Sbjct: 53 VTTPSGLKYVELKEGTGATPQPGQTVEVHYVGTLEDGTKFDSSRDRG--QPFSFKIGVGQ 110
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG+ T+K GG+RK+I+P +LG+ GA G IPP ATL + VE+
Sbjct: 111 VIKGWDEGV----STIKVGGRRKLIIPSELGYGARGA----GGVIPPNATLIFDVEL 159
>ref|ZP_00517772.1| Peptidylprolyl isomerase [Crocosphaera watsonii WH 8501]
gi|67853825|gb|EAM49151.1| Peptidylprolyl isomerase
[Crocosphaera watsonii WH 8501]
Length = 175
Score = 71.6 bits (174), Expect = 2e-11
Identities = 48/130 (36%), Positives = 74/130 (56%), Gaps = 13/130 (10%)
Query: 116 NTRDVEKEE---EVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEG 172
N R++ +EE VT +G++Y +LK G G SP+ G V +D G +E+ G+ F ++
Sbjct: 52 NNREITQEEIDNAVTTESGLKYIDLKEGDGESPQKGQTVTVDYTGTLEN-GKKF-DSSRD 109
Query: 173 DKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPP 232
K+P + +G KG EG+ +MK GG+R +I+PP+LG+ GA G IP
Sbjct: 110 RKQPFSFKIGVGQVIKGWDEGV----ASMKVGGQRILIIPPELGYGSRGA----GGVIPG 161
Query: 233 LATLEYIVEV 242
ATL + VE+
Sbjct: 162 NATLIFDVEL 171
>dbj|BAB72535.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Nostoc sp. PCC 7120]
gi|17228073|ref|NP_484621.1| FKBP-type peptidyl-prolyl
cis-trans isomerase [Nostoc sp. PCC 7120]
gi|25292526|pir||AH1878 FKBP-type peptidyl-prolyl
cis-trans isomerase [imported] - Nostoc sp. (strain PCC
7120)
Length = 165
Score = 69.3 bits (168), Expect = 9e-11
Identities = 45/120 (37%), Positives = 70/120 (57%), Gaps = 10/120 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT +G++Y E++ G GA+P+ G VV+ G +E G F ++ + ++ P + +G
Sbjct: 55 VTTDSGLKYVEIEEGTGATPKSGQTVVVHYTGTLED-GTKFDSSRDRNR-PFSFTIGVGQ 112
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
KG EG L TMK GG+R++I+P +LG+ GA G IPP ATL + VE+ +V
Sbjct: 113 VIKGWDEG----LSTMKVGGRRQLIIPSELGYGARGA----GGVIPPYATLLFDVELLEV 164
>ref|ZP_00159695.2| COG0545: FKBP-type peptidyl-prolyl cis-trans isomerases 1 [Anabaena
variabilis ATCC 29413]
Length = 165
Score = 68.6 bits (166), Expect = 2e-10
Identities = 45/132 (34%), Positives = 74/132 (55%), Gaps = 10/132 (7%)
Query: 114 EANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGD 173
++N + VT +G++Y E++ G GA+P+ G VV+ G +E G F ++ + +
Sbjct: 43 KSNIMSDASKPTVTTDSGLKYVEIEEGTGATPKSGQTVVVHYTGTLED-GTKFDSSRDRN 101
Query: 174 KKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPL 233
+ P + +G KG EG L TMK GG+R++I+P +LG+ GA G IPP
Sbjct: 102 R-PFSFTIGVGQVIKGWDEG----LSTMKVGGRRQLIIPSELGYGARGA----GGVIPPY 152
Query: 234 ATLEYIVEVDKV 245
+TL + VE+ +V
Sbjct: 153 STLLFDVELLEV 164
>ref|NP_440378.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Synechocystis sp.
PCC 6803] gi|1652134|dbj|BAA17058.1| FKBP-type
peptidyl-prolyl cis-trans isomerase [Synechocystis sp.
PCC 6803] gi|7428434|pir||S75144 FKBP-type
peptidyl-prolyl cis-trans isomerase - Synechocystis sp.
(strain PCC 6803)
Length = 201
Score = 65.9 bits (159), Expect = 1e-09
Identities = 44/117 (37%), Positives = 64/117 (54%), Gaps = 10/117 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT +G++Y + VG G SP G V + G++ + G F ++ + +K P +G
Sbjct: 91 VTTESGLQYIDEVVGEGPSPTKGQKVEVHYTGRL-TDGTKFDSSVDRNK-PFTFTIGVGQ 148
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG+ TM+ GGKRK+I+PP L + GA G IPP ATLE+ VE+
Sbjct: 149 VIKGWDEGVA----TMQVGGKRKLIIPPDLAYGSRGA----GGVIPPNATLEFEVEL 197
>ref|NP_947888.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Rhodopseudomonas
palustris CGA009] gi|39649465|emb|CAE27987.1| FKBP-type
peptidyl-prolyl cis-trans isomerase [Rhodopseudomonas
palustris CGA009]
Length = 152
Score = 63.9 bits (154), Expect = 4e-09
Identities = 44/125 (35%), Positives = 63/125 (50%), Gaps = 12/125 (9%)
Query: 121 EKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGD---KKPL 177
E + VT P+G++ + +VG GASP G + V+ G + G F+ +P
Sbjct: 33 ESAKSVTTPSGLQIVDTQVGTGASPARGQICVMHYTGWLYENGAK-TRKFDSSVDRNEPF 91
Query: 178 ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLE 237
+G KG EG+ +MK GGKR +I+PP LG+ GA G IPP ATL
Sbjct: 92 EFPIGMGRVIKGWDEGVA----SMKVGGKRTLIIPPDLGYGARGA----GGVIPPNATLV 143
Query: 238 YIVEV 242
+ VE+
Sbjct: 144 FDVEL 148
>ref|NP_923787.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Gloeobacter
violaceus PCC 7421] gi|35211403|dbj|BAC88782.1|
FKBP-type peptidyl-prolyl cis-trans isomerase
[Gloeobacter violaceus PCC 7421]
Length = 161
Score = 63.9 bits (154), Expect = 4e-09
Identities = 43/116 (37%), Positives = 64/116 (55%), Gaps = 10/116 (8%)
Query: 127 TLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPY 186
T +G++Y + VG GASP+ G V + G +E G+ F ++ + + P + +G
Sbjct: 52 TTTSGLKYLDETVGNGASPQKGQRVTVHYTGTLED-GKKFDSSRDRGQ-PFSFTIGVGQV 109
Query: 187 SKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
+G EG+ TMK GGKRK++VP LG+ GA G IPP ATL + VE+
Sbjct: 110 IQGWDEGVA----TMKVGGKRKLVVPANLGYGARGA----GGVIPPNATLLFDVEL 157
>ref|NP_953323.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Geobacter
sulfurreducens PCA] gi|39984263|gb|AAR35650.1| FKBP-type
peptidyl-prolyl cis-trans isomerase [Geobacter
sulfurreducens PCA]
Length = 138
Score = 63.5 bits (153), Expect = 5e-09
Identities = 44/119 (36%), Positives = 65/119 (53%), Gaps = 10/119 (8%)
Query: 124 EEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGS 183
+ VT +G+ Y +L G GA+P G V + G +E+ + + G+ P +G+
Sbjct: 25 DAVTTASGLSYVDLAAGSGAAPVAGKPVKVHYTGWLENGTKFDSSVDRGE--PFVFTIGA 82
Query: 184 RPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
G EG+ +MK GGKR++IVPPQLG+ GA G+G IPP ATL + VE+
Sbjct: 83 GEVIPGWDEGV----MSMKVGGKRRLIVPPQLGY---GA-AGAGGVIPPNATLIFEVEL 133
>ref|NP_681893.1| FKBP-type peptidyl-prolyl cis-trans isomerase [Thermosynechococcus
elongatus BP-1] gi|22294826|dbj|BAC08655.1| FKBP-type
peptidyl-prolyl cis-trans isomerase [Thermosynechococcus
elongatus BP-1]
Length = 162
Score = 63.2 bits (152), Expect = 7e-09
Identities = 41/117 (35%), Positives = 64/117 (54%), Gaps = 10/117 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
VT P+G++Y ++ VG GA P+ GD V + G + + G +F ++ +P +G
Sbjct: 52 VTTPSGLQYEDIVVGSGAQPQVGDRVTVHYTGML-TDGRIF-DSSRDRGQPFQFQIGVGQ 109
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
KG EG+ +M GG+R++I+PP LG+ G G IPP ATL + VE+
Sbjct: 110 VIKGWDEGV----GSMHVGGQRRLIIPPNLGYGARGV----GGVIPPNATLIFDVEL 158
>ref|NP_769045.1| Peptidylprolyl isomerase [Bradyrhizobium japonicum USDA 110]
gi|27350660|dbj|BAC47670.1| Peptidylprolyl isomerase
[Bradyrhizobium japonicum USDA 110]
Length = 154
Score = 63.2 bits (152), Expect = 7e-09
Identities = 43/120 (35%), Positives = 63/120 (51%), Gaps = 12/120 (10%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGD---KKPLALVMG 182
+T +G++ + VG GASP+PG + V+ G + G+ F+ K+P +G
Sbjct: 40 MTTASGLQIIDTAVGTGASPQPGQICVMHYTGWLYENGQKG-KKFDSSVDRKEPFEFPIG 98
Query: 183 SRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
G EG+ +MK GGKR +I+PPQLG+ GA G IPP ATL + VE+
Sbjct: 99 KGRVIAGWDEGVA----SMKVGGKRTLIIPPQLGYGARGA----GGVIPPNATLMFDVEL 150
>ref|ZP_00357923.1| COG0545: FKBP-type peptidyl-prolyl cis-trans isomerases 1
[Chloroflexus aurantiacus]
Length = 237
Score = 63.2 bits (152), Expect = 7e-09
Identities = 43/124 (34%), Positives = 69/124 (54%), Gaps = 10/124 (8%)
Query: 126 VTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRP 185
+T +G++Y E++ G G P+PG +V + G + + G VF +++E +P+ +G
Sbjct: 1 MTTASGLQYVEIQAGDGEQPQPGAIVAVHYRGML-ADGSVFDSSYERG-EPIRFPLGVGM 58
Query: 186 YSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
G EGI M+ GGK ++I+PP LG+ GA +G IPP ATL + VE+ +V
Sbjct: 59 VIPGWDEGI----GLMRVGGKARLIIPPHLGY---GA-MGYPPVIPPNATLTFDVELVEV 110
Query: 246 SIAP 249
P
Sbjct: 111 LPGP 114
Score = 52.8 bits (125), Expect = 9e-06
Identities = 38/128 (29%), Positives = 66/128 (50%), Gaps = 10/128 (7%)
Query: 118 RDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPL 177
+D+ + T +G++Y +L VG GA+ G V + G + + G +F ++ + P
Sbjct: 119 QDLPADRYTTSASGLQYADLTVGDGATAMAGRTVTVHYTGWL-TDGSMFDSSLSRGE-PF 176
Query: 178 ALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLE 237
+G+ +G EG+ M+ GG+R++I+P L + GA G IPP ATL
Sbjct: 177 VFPLGAGRVIRGWDEGVA----GMRVGGRRQLIIPAALAYGNRGA----GGVIPPGATLI 228
Query: 238 YIVEVDKV 245
+ VE+ +V
Sbjct: 229 FEVELLEV 236
>sp|Q26486|FKB4_SPOFR 46 kDa FK506-binding nuclear protein (Peptidyl-prolyl cis-trans
isomerase) (PPIase) (Rotamase) gi|1079010|pir||A55320
immunophilin FKBP46 - fall armyworm
gi|595845|gb|AAA58962.1| immunophilin FKBP46
Length = 412
Score = 62.8 bits (151), Expect = 9e-09
Identities = 45/141 (31%), Positives = 73/141 (50%), Gaps = 13/141 (9%)
Query: 105 KTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGE 164
K + E ++EA VEK+E+ + G+ +LKVG G + G +V++ G+++ +
Sbjct: 284 KKKPEAKKEEA---PVEKKEKKQIAGGVSIEDLKVGSGPVAKAGKVVMVYYEGRLKQNNK 340
Query: 165 VFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADL 224
+F N +G L SK V G + + MK GGKRK++ PP + + GA
Sbjct: 341 MFDNCVKGPGFKFRL------GSKEVISGWDVGIAGMKVGGKRKIVCPPAMAY---GAK- 390
Query: 225 GSGVQIPPLATLEYIVEVDKV 245
GS IPP +TL + V++ V
Sbjct: 391 GSPPVIPPNSTLVFEVDLKNV 411
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 451,552,145
Number of Sequences: 2540612
Number of extensions: 19822838
Number of successful extensions: 138156
Number of sequences better than 10.0: 1063
Number of HSP's better than 10.0 without gapping: 278
Number of HSP's successfully gapped in prelim test: 811
Number of HSP's that attempted gapping in prelim test: 135400
Number of HSP's gapped (non-prelim): 2581
length of query: 250
length of database: 863,360,394
effective HSP length: 125
effective length of query: 125
effective length of database: 545,783,894
effective search space: 68222986750
effective search space used: 68222986750
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0042.13


