

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036b.1
(122 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
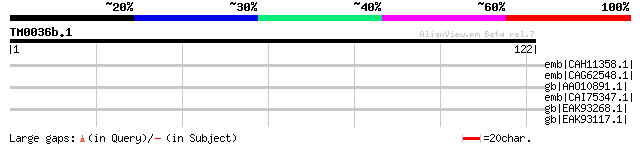
emb|CAH11358.1| hypothetical protein [Legionella pneumophila str... 31 6.0
emb|CAG62548.1| unnamed protein product [Candida glabrata CBS138... 31 6.0
gb|AAO10891.1| Unknown [Vibrio vulnificus CMCP6] gi|27365836|ref... 31 7.9
emb|CAI75347.1| hypothetical protein [Theileria annulata] 31 7.9
gb|EAK93268.1| hypothetical protein CaO19.8781 [Candida albicans... 31 7.9
gb|EAK93117.1| hypothetical protein CaO19.1190 [Candida albicans... 31 7.9
>emb|CAH11358.1| hypothetical protein [Legionella pneumophila str. Paris]
gi|54296185|ref|YP_122554.1| hypothetical protein
lpp0211 [Legionella pneumophila str. Paris]
Length = 269
Score = 31.2 bits (69), Expect = 6.0
Identities = 13/39 (33%), Positives = 23/39 (58%)
Query: 7 WYEPNDSLRSDRFIVSLKDHSSTCNFSELLEILSRHVVA 45
W PN+ + D ++S+ D TC+ + LE LS+ ++A
Sbjct: 223 WTTPNEVVEQDLEMISIIDEVRTCDLKQSLENLSQPLIA 261
>emb|CAG62548.1| unnamed protein product [Candida glabrata CBS138]
gi|50294321|ref|XP_449572.1| unnamed protein product
[Candida glabrata]
Length = 547
Score = 31.2 bits (69), Expect = 6.0
Identities = 16/39 (41%), Positives = 24/39 (61%)
Query: 25 DHSSTCNFSELLEILSRHVVAGIFYKGLNIVLCILILQK 63
D ST NF+ LLE+LS ++ KGL ++ CI+ +K
Sbjct: 18 DRISTKNFTILLEVLSNNINKVDSSKGLPLIQCIINFEK 56
>gb|AAO10891.1| Unknown [Vibrio vulnificus CMCP6] gi|27365836|ref|NP_761364.1|
hypothetical protein VV12535 [Vibrio vulnificus CMCP6]
Length = 207
Score = 30.8 bits (68), Expect = 7.9
Identities = 23/88 (26%), Positives = 40/88 (45%), Gaps = 20/88 (22%)
Query: 1 MSDELDWYEPNDSLRSDRFIVSLKDH----------------SSTCNFSELLEILSRHVV 44
M D++E S+ +R I+S++DH SS N S + E+LSRH+
Sbjct: 108 MGKAQDYFEGVKSIAVERGILSIRDHIEKVANIRNTILYASDSSLPNVSNVEEMLSRHIG 167
Query: 45 AGIFYKGLNIVLCILILQKDKLIRTHTC 72
A + + ++ +L++Q K C
Sbjct: 168 AALIIQ----MIYLLVVQNKKQNLVEQC 191
>emb|CAI75347.1| hypothetical protein [Theileria annulata]
Length = 248
Score = 30.8 bits (68), Expect = 7.9
Identities = 12/25 (48%), Positives = 17/25 (68%)
Query: 55 VLCILILQKDKLIRTHTCTLLHQLM 79
+L +LI KDK +RT+TC LH +
Sbjct: 158 ILLLLIYSKDKRLRTNTCQFLHHFV 182
>gb|EAK93268.1| hypothetical protein CaO19.8781 [Candida albicans SC5314]
Length = 943
Score = 30.8 bits (68), Expect = 7.9
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Query: 47 IFYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSLKI 102
+F ++ +CIL+ + H+ L H + SK F Y+Y P F+S+ I
Sbjct: 889 LFAMWFSLTVCILVFMEGTSAMLHSLRL-HWVEAMSKFFQGEGYAYEPFTFKSIDI 943
>gb|EAK93117.1| hypothetical protein CaO19.1190 [Candida albicans SC5314]
Length = 943
Score = 30.8 bits (68), Expect = 7.9
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Query: 47 IFYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKSKVFDANCYSYPPPLFRSLKI 102
+F ++ +CIL+ + H+ L H + SK F Y+Y P F+S+ I
Sbjct: 889 LFAMWFSLTVCILVFMEGTSAMLHSLRL-HWVEAMSKFFQGEGYAYEPFTFKSIDI 943
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.324 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 200,329,182
Number of Sequences: 2540612
Number of extensions: 6733848
Number of successful extensions: 17383
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 17380
Number of HSP's gapped (non-prelim): 6
length of query: 122
length of database: 863,360,394
effective HSP length: 98
effective length of query: 24
effective length of database: 614,380,418
effective search space: 14745130032
effective search space used: 14745130032
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0036b.1


