

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.12
(125 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
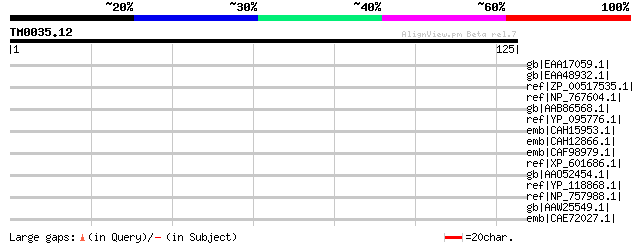
Score E
Sequences producing significant alignments: (bits) Value
gb|EAA17059.1| putative copper efflux ATPase [Plasmodium yoelii ... 36 0.24
gb|EAA48932.1| hypothetical protein MG00590.4 [Magnaporthe grise... 34 0.70
ref|ZP_00517535.1| 1-(5-Phosphoribosyl)-5-amino-4-imidazole-carb... 33 2.0
ref|NP_767604.1| probable flavin-binding family monooxygenase [B... 32 2.7
gb|AAB86568.1| unknown [Schistosoma mansoni] 32 2.7
ref|YP_095776.1| ClpB protein [Legionella pneumophila subsp. pne... 32 3.5
emb|CAH15953.1| endopeptidase Clp ATP-binding chain B (ClpB) [Le... 32 3.5
emb|CAH12866.1| endopeptidase Clp ATP-binding chain B (ClpB) [Le... 32 3.5
emb|CAF98979.1| unnamed protein product [Tetraodon nigroviridis] 32 3.5
ref|XP_601686.1| PREDICTED: similar to Olfactory receptor 11A1 (... 32 4.6
gb|AAO52454.1| hypothetical protein [Dictyostelium discoideum] g... 31 6.0
ref|YP_118868.1| putative NADH dehydrogenase I chain L [Nocardia... 31 7.8
ref|NP_757988.1| hypothetical protein MYPE6020 [Mycoplasma penet... 31 7.8
gb|AAW25549.1| unknown [Schistosoma japonicum] 31 7.8
emb|CAE72027.1| Hypothetical protein CBG19109 [Caenorhabditis br... 31 7.8
>gb|EAA17059.1| putative copper efflux ATPase [Plasmodium yoelii yoelii]
Length = 1976
Score = 35.8 bits (81), Expect = 0.24
Identities = 21/69 (30%), Positives = 35/69 (50%), Gaps = 4/69 (5%)
Query: 2 IHNPGQHHHSHRRSFFTHINWAPDQH-PHEVLAFMPSLFKSN---DLARRKRASDGDKSL 57
+HN H +FF H N + Q+ P V M + + SN D ++K+ +D ++ +
Sbjct: 541 VHNLNNEHKVFNNNFFEHDNISLSQNIPFSVSHKMDNNYNSNENLDFRKKKKKNDDEEKI 600
Query: 58 NKTQEEKAS 66
K Q+E AS
Sbjct: 601 KKKQQEIAS 609
>gb|EAA48932.1| hypothetical protein MG00590.4 [Magnaporthe grisea 70-15]
gi|39974527|ref|XP_368654.1| hypothetical protein
MG00590.4 [Magnaporthe grisea 70-15]
Length = 1073
Score = 34.3 bits (77), Expect = 0.70
Identities = 24/70 (34%), Positives = 33/70 (46%), Gaps = 6/70 (8%)
Query: 13 RRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSL-----NKTQEEKASI 67
R S+ + IN + P EV F P LF + D A S+G +L + Q++ SI
Sbjct: 267 RGSWLSEINLIGHEGPTEVTQFSPRLFHTTDPALANGNSNGSSNLVTVIASAGQDKTLSI 326
Query: 68 WNKTQIKTPL 77
WN T PL
Sbjct: 327 WN-TNTSRPL 335
>ref|ZP_00517535.1| 1-(5-Phosphoribosyl)-5-amino-4-imidazole-carboxylate (AIR)
carboxylase [Crocosphaera watsonii WH 8501]
gi|67854049|gb|EAM49361.1|
1-(5-Phosphoribosyl)-5-amino-4-imidazole-carboxylate
(AIR) carboxylase [Crocosphaera watsonii WH 8501]
Length = 264
Score = 32.7 bits (73), Expect = 2.0
Identities = 23/109 (21%), Positives = 45/109 (41%), Gaps = 14/109 (12%)
Query: 9 HHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIW 68
HH H R+ F + W P + P +++ M ++ + + + R +A+I
Sbjct: 43 HHRHLRTGFPEVIWGPGKTPEQIIQIMNTMARHSPVVMATRI-------------EAAIA 89
Query: 69 NKTQIKTP-LNKASTFEATKLRSGTCIFRSGAAVRFVILGFAERREMEE 116
+ + + P L T L+SG F++ + + G A+ EE
Sbjct: 90 EQLEAEIPSLIYYPTARICALQSGQTSFQASGVISILTAGTADLPVAEE 138
>ref|NP_767604.1| probable flavin-binding family monooxygenase [Bradyrhizobium
japonicum USDA 110] gi|27349214|dbj|BAC46229.1| blr0964
[Bradyrhizobium japonicum USDA 110]
Length = 524
Score = 32.3 bits (72), Expect = 2.7
Identities = 18/59 (30%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Query: 30 EVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKL 88
E+LA+M + + ND+ARR R S + + ++ ++W + T +A TF A L
Sbjct: 116 EILAYMNEVIEDNDIARRIRYKHKINSASWSSDQ--NLWTIEAVTTDTGEARTFTANFL 172
>gb|AAB86568.1| unknown [Schistosoma mansoni]
Length = 406
Score = 32.3 bits (72), Expect = 2.7
Identities = 26/106 (24%), Positives = 55/106 (51%), Gaps = 7/106 (6%)
Query: 20 INWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNK 79
IN +QH +E+L + DL R R S+ + +LN ++E+A+ +N ++ LN+
Sbjct: 243 INKQDEQHKNELLHNNSNDNIYEDL--RNRLSEVENNLNAAEKERATFYN--LYRSALNE 298
Query: 80 ASTFEATKLRSGTCIFRSGAAVRFVILGFAE---RREMEEEEKRKQ 122
+++T +++ + + G +V+ + + E E+ + RKQ
Sbjct: 299 VDHYKSTLIQTEEVLAKLGXSVKQTESKWRQLLSESEFEQIQLRKQ 344
>ref|YP_095776.1| ClpB protein [Legionella pneumophila subsp. pneumophila str.
Philadelphia 1] gi|52629088|gb|AAU27829.1| ClpB protein
[Legionella pneumophila subsp. pneumophila str.
Philadelphia 1]
Length = 858
Score = 32.0 bits (71), Expect = 3.5
Identities = 22/66 (33%), Positives = 33/66 (49%), Gaps = 11/66 (16%)
Query: 37 SLFKSNDLARRKRASDGDKSLNKTQE-----------EKASIWNKTQIKTPLNKASTFEA 85
+L K ND A +KR D KS+++ ++ EKA++ TQIK L +A
Sbjct: 428 ALKKENDEASKKRLVDLQKSIDELEQNYSDLEEIWKAEKATMQGSTQIKEALEQAKLEME 487
Query: 86 TKLRSG 91
T R+G
Sbjct: 488 TARRAG 493
>emb|CAH15953.1| endopeptidase Clp ATP-binding chain B (ClpB) [Legionella
pneumophila str. Lens] gi|54294637|ref|YP_127052.1|
endopeptidase Clp ATP-binding chain B (ClpB) [Legionella
pneumophila str. Lens]
Length = 858
Score = 32.0 bits (71), Expect = 3.5
Identities = 22/66 (33%), Positives = 33/66 (49%), Gaps = 11/66 (16%)
Query: 37 SLFKSNDLARRKRASDGDKSLNKTQE-----------EKASIWNKTQIKTPLNKASTFEA 85
+L K ND A +KR D KS+++ ++ EKA++ TQIK L +A
Sbjct: 428 ALKKENDEASKKRLVDLQKSIDELEQNYSDLEEIWKAEKATMQGSTQIKEALEQAKLEME 487
Query: 86 TKLRSG 91
T R+G
Sbjct: 488 TARRAG 493
>emb|CAH12866.1| endopeptidase Clp ATP-binding chain B (ClpB) [Legionella
pneumophila str. Paris] gi|54297663|ref|YP_124032.1|
endopeptidase Clp ATP-binding chain B (ClpB) [Legionella
pneumophila str. Paris]
Length = 858
Score = 32.0 bits (71), Expect = 3.5
Identities = 22/66 (33%), Positives = 33/66 (49%), Gaps = 11/66 (16%)
Query: 37 SLFKSNDLARRKRASDGDKSLNKTQE-----------EKASIWNKTQIKTPLNKASTFEA 85
+L K ND A +KR D KS+++ ++ EKA++ TQIK L +A
Sbjct: 428 ALKKENDEASKKRLVDLQKSIDELEQNYSDLEEIWKAEKATMQGSTQIKEALEQAKLEME 487
Query: 86 TKLRSG 91
T R+G
Sbjct: 488 TARRAG 493
>emb|CAF98979.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1792
Score = 32.0 bits (71), Expect = 3.5
Identities = 16/45 (35%), Positives = 25/45 (55%)
Query: 47 RKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSG 91
RK+AS D + +E A + KT+ + PL + S+ E+ K SG
Sbjct: 1018 RKKASTEDDRGSSDEETPAELVKKTESRKPLRRGSSMESDKPESG 1062
>ref|XP_601686.1| PREDICTED: similar to Olfactory receptor 11A1 (Hs6M1-18), partial
[Bos taurus]
Length = 378
Score = 31.6 bits (70), Expect = 4.6
Identities = 22/77 (28%), Positives = 31/77 (39%), Gaps = 2/77 (2%)
Query: 40 KSNDLARRKRASDGD--KSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRS 97
K DL +RK GD K + +E + W K Q T + T E T +
Sbjct: 9 KGKDLQQRKAKGIGDLEKEMESGRENDSGKWKKMQRSTRICHLITAETWMTFMETVSIGN 68
Query: 98 GAAVRFVILGFAERREM 114
FV+LGF + E+
Sbjct: 69 QTITEFVLLGFYVKAEL 85
>gb|AAO52454.1| hypothetical protein [Dictyostelium discoideum]
gi|66821455|ref|XP_644203.1| hypothetical protein
DDB0167628 [Dictyostelium discoideum]
gi|60472151|gb|EAL70104.1| hypothetical protein
DDB0167628 [Dictyostelium discoideum]
Length = 696
Score = 31.2 bits (69), Expect = 6.0
Identities = 21/86 (24%), Positives = 44/86 (50%), Gaps = 13/86 (15%)
Query: 49 RASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFE----ATKLRSGTCIFRS------- 97
R + + ++N Q++++ N +QIK NK+S + A K+R F++
Sbjct: 594 RNENENTNINHEQQQQSKTVNISQIKNDANKSSEIDDDGIARKIRDSFTHFQTREELLDA 653
Query: 98 --GAAVRFVILGFAERREMEEEEKRK 121
A++FV A ++E+++E++ K
Sbjct: 654 IQNLAIKFVKQTSALKKELQQEQQEK 679
>ref|YP_118868.1| putative NADH dehydrogenase I chain L [Nocardia farcinica IFM
10152] gi|54016134|dbj|BAD57504.1| putative NADH
dehydrogenase I chain L [Nocardia farcinica IFM 10152]
Length = 628
Score = 30.8 bits (68), Expect = 7.8
Identities = 12/21 (57%), Positives = 13/21 (61%)
Query: 15 SFFTHINWAPDQHPHEVLAFM 35
+FF WAPD HPHE A M
Sbjct: 440 TFFGERRWAPDTHPHEAPAVM 460
>ref|NP_757988.1| hypothetical protein MYPE6020 [Mycoplasma penetrans HF-2]
gi|26454062|dbj|BAC44392.1| hypothetical protein
[Mycoplasma penetrans HF-2]
Length = 630
Score = 30.8 bits (68), Expect = 7.8
Identities = 22/57 (38%), Positives = 30/57 (52%), Gaps = 7/57 (12%)
Query: 36 PSLFKSNDLARRKRASDGDKSLN--KTQEEKASIWNKTQIKTPLNKASTFEATKLRS 90
PSL K + L K+ D+SLN + EK S+ +KT LNK S+ E LR+
Sbjct: 50 PSLIKPDSLDVSKQDKKSDESLNLENKENEKPSL-----LKTKLNKESSSENEDLRT 101
>gb|AAW25549.1| unknown [Schistosoma japonicum]
Length = 175
Score = 30.8 bits (68), Expect = 7.8
Identities = 21/84 (25%), Positives = 41/84 (48%), Gaps = 7/84 (8%)
Query: 36 PSLFKSNDLARRKRASDGDKSLNKTQEE-----KASIWNKTQIKTPLNKASTFEATKLRS 90
PSL+ N + S+G + L K + KA + ++++ +K + E T L
Sbjct: 5 PSLYSINSILILD--SEGKRVLTKYYDSSLPTVKAQLEFESKLFKKTSKTNGAEITLLDG 62
Query: 91 GTCIFRSGAAVRFVILGFAERREM 114
TC++R+ + F ++G A+ E+
Sbjct: 63 ATCVYRNVGDLYFYVIGDAKENEL 86
>emb|CAE72027.1| Hypothetical protein CBG19109 [Caenorhabditis briggsae]
Length = 1139
Score = 30.8 bits (68), Expect = 7.8
Identities = 15/46 (32%), Positives = 23/46 (49%), Gaps = 7/46 (15%)
Query: 1 MIHNPGQH-------HHSHRRSFFTHINWAPDQHPHEVLAFMPSLF 39
M+H P QH ++ H +FF +++P Q PH+ F P F
Sbjct: 1078 MLHRPPQHPSQSAFYNNPHDNNFFRPRDFSPPQPPHQSRMFSPPSF 1123
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.129 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 210,963,789
Number of Sequences: 2540612
Number of extensions: 7371705
Number of successful extensions: 26780
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 26775
Number of HSP's gapped (non-prelim): 17
length of query: 125
length of database: 863,360,394
effective HSP length: 101
effective length of query: 24
effective length of database: 606,758,582
effective search space: 14562205968
effective search space used: 14562205968
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0035.12


