

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.10
(150 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
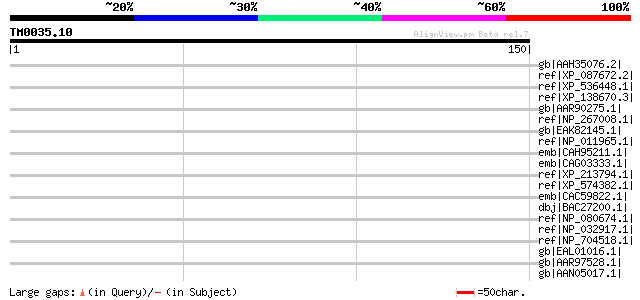
Sequences producing significant alignments: (bits) Value
gb|AAH35076.2| KIAA1935 protein [Homo sapiens] 33 1.1
ref|XP_087672.2| PREDICTED: KIAA1935 protein [Homo sapiens] 33 1.1
ref|XP_536448.1| PREDICTED: similar to KIAA1935 protein [Canis f... 33 1.8
ref|XP_138670.3| PREDICTED: similar to hypothetical protein 4932... 33 1.8
gb|AAR90275.1| polyketide synthase [Cochliobolus heterostrophus] 33 1.8
ref|NP_267008.1| hypothetical protein L73264 [Lactococcus lactis... 32 2.4
gb|EAK82145.1| hypothetical protein UM01282.1 [Ustilago maydis 5... 32 3.1
ref|NP_011965.1| Hypothetical ORF; Yhr097cp [Saccharomyces cerev... 32 4.1
emb|CAH95211.1| ubiquitin carboxyl-terminal hydrolase, putative ... 32 4.1
emb|CAG03333.1| unnamed protein product [Tetraodon nigroviridis] 31 5.3
ref|XP_213794.1| PREDICTED: similar to processing of precursor 5... 31 7.0
ref|XP_574382.1| PREDICTED: similar to zinc finger protein 108 [... 31 7.0
emb|CAC59822.1| Pop5 protein [Homo sapiens] gi|15214743|gb|AAH12... 30 9.1
dbj|BAC27200.1| unnamed protein product [Mus musculus] 30 9.1
ref|NP_080674.1| processing of precursor 5, ribonuclease P/MRP f... 30 9.1
ref|NP_032917.1| pinin [Mus musculus] gi|2570049|emb|CAA69960.1|... 30 9.1
ref|NP_704518.1| hypothetical protein [Plasmodium falciparum 3D7... 30 9.1
gb|EAL01016.1| possible secreted protein [Candida albicans SC531... 30 9.1
gb|AAR97528.1| zinc finger protein 183 [Caenorhabditis briggsae]... 30 9.1
gb|AAN05017.1| pinin [Mus musculus] 30 9.1
>gb|AAH35076.2| KIAA1935 protein [Homo sapiens]
Length = 571
Score = 33.5 bits (75), Expect = 1.1
Identities = 27/118 (22%), Positives = 50/118 (41%), Gaps = 2/118 (1%)
Query: 12 KSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPS 71
K+ +H + + ++ KK+ + +T N + +PSS S + L P
Sbjct: 432 KNMDHAKAREVLKIAKEKAQKKQNETSTSKNAHSKVHFTRRYQ-NPSSGSLPPRVRLKPQ 490
Query: 72 LHQCGWMDGCMPTRNSYDTSGSLS-HADTRATSAKATCLLSWFLLHQSLSRSLTYPQC 128
++ D + T SG ++ +R TS ++ L W LLHQ+++ QC
Sbjct: 491 RYRNEENDSSLKTGLEKMRSGKMAPKPQSRCTSTRSAGLNKWQLLHQTVTSPAAPLQC 548
>ref|XP_087672.2| PREDICTED: KIAA1935 protein [Homo sapiens]
Length = 560
Score = 33.5 bits (75), Expect = 1.1
Identities = 27/118 (22%), Positives = 50/118 (41%), Gaps = 2/118 (1%)
Query: 12 KSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPS 71
K+ +H + + ++ KK+ + +T N + +PSS S + L P
Sbjct: 421 KNMDHAKAREVLKIAKEKAQKKQNETSTSKNAHSKVHFTRRYQ-NPSSGSLPPRVRLKPQ 479
Query: 72 LHQCGWMDGCMPTRNSYDTSGSLS-HADTRATSAKATCLLSWFLLHQSLSRSLTYPQC 128
++ D + T SG ++ +R TS ++ L W LLHQ+++ QC
Sbjct: 480 RYRNEENDSSLKTGLEKMRSGKMAPKPQSRCTSTRSAGLNKWQLLHQTVTSPAAPLQC 537
>ref|XP_536448.1| PREDICTED: similar to KIAA1935 protein [Canis familiaris]
Length = 530
Score = 32.7 bits (73), Expect = 1.8
Identities = 27/118 (22%), Positives = 49/118 (40%), Gaps = 2/118 (1%)
Query: 12 KSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPS 71
K+ +H + + ++ KK+ + +T N + PSS S + L P
Sbjct: 391 KNMDHAKAREVLKIAKEKAQKKQSETSTSKNAHSKVHFTRRYQ-SPSSGSLPPRVRLKPQ 449
Query: 72 LHQCGWMDGCMPTRNSYDTSGSLS-HADTRATSAKATCLLSWFLLHQSLSRSLTYPQC 128
++ D + T SG ++ +R TS ++ L W LLHQ+++ QC
Sbjct: 450 RYRNEENDSSLKTGLEKMRSGKMAPKPQSRCTSTRSAGLNKWQLLHQTVTSPAAPLQC 507
>ref|XP_138670.3| PREDICTED: similar to hypothetical protein 4932411G14 [Mus
musculus]
Length = 1006
Score = 32.7 bits (73), Expect = 1.8
Identities = 33/115 (28%), Positives = 48/115 (41%), Gaps = 19/115 (16%)
Query: 25 LHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCMPT 84
LH + L K + Q D+++ S+H S + I T+L SL + GC
Sbjct: 529 LHTPQHLSYSKTSEDQLEDKNTQSFWGLPSLHSESMNSIATVLTDSSL-----ISGCF-- 581
Query: 85 RNSYDTSGSLSHADTRATSAKATCLLSWFLLHQSLSRSLTYP------QCKVQTP 133
N + S L+H T S LS QSLS+S + P Q ++Q P
Sbjct: 582 -NRFSDSPMLTHRTTLPLSESQLHNLS-----QSLSQSQSQPVSQANLQAQLQLP 630
>gb|AAR90275.1| polyketide synthase [Cochliobolus heterostrophus]
Length = 2275
Score = 32.7 bits (73), Expect = 1.8
Identities = 31/109 (28%), Positives = 46/109 (41%), Gaps = 12/109 (11%)
Query: 43 DRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCMPTRNSY-DTSGSLSHADTRA 101
D + + L+DQ SIH ++ I+ LH+C W + ++SY TS S R
Sbjct: 398 DSNDATLKDQTSIHKNALQSIL-------LHRCEWYNIVSELKSSYFSTSTSFVAFGERC 450
Query: 102 TSAKATCLLSWFLLHQSLSRSLTYPQCKVQTPKKYQVVAVGLEMGSAPS 150
A LLS F + T P +V P V+ V + + A S
Sbjct: 451 MPAS---LLSGFSGRVAYFDDPTTPALRV-NPNDVAVIGVAINVAGASS 495
>ref|NP_267008.1| hypothetical protein L73264 [Lactococcus lactis subsp. lactis Il1403]
gi|12723779|gb|AAK04950.1| UNKNOWN PROTEIN [Lactococcus
lactis subsp. lactis Il1403] gi|25401280|pir||D86731
hypothetical protein yihD [imported] - Lactococcus lactis
subsp. lactis (strain IL1403)
Length = 1063
Score = 32.3 bits (72), Expect = 2.4
Identities = 30/97 (30%), Positives = 46/97 (46%), Gaps = 8/97 (8%)
Query: 32 KKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCMPTRNSYDTS 91
K ++ N+ S+ L S ++SS +ML P+L Q +G T NS TS
Sbjct: 921 KNSSHSSLSSNESSSASLTSSESSGSTTSS---SMLTNPNL-QSNDKNGSSLTSNSSYTS 976
Query: 92 ----GSLSHADTRATSAKATCLLSWFLLHQSLSRSLT 124
+LSH++ + AK + L+S SLS SL+
Sbjct: 977 EVKLNNLSHSNLSSNEAKVSTLISKVSSSPSLSSSLS 1013
>gb|EAK82145.1| hypothetical protein UM01282.1 [Ustilago maydis 521]
gi|49069216|ref|XP_398897.1| hypothetical protein
UM01282.1 [Ustilago maydis 521]
Length = 3023
Score = 32.0 bits (71), Expect = 3.1
Identities = 25/86 (29%), Positives = 41/86 (47%), Gaps = 10/86 (11%)
Query: 24 TLHMQRKLKKKKKNTTQHNDRDSSELRDQV--SIHPSSSSFIITMLLGPSLHQCGWMDGC 81
T H +L +++ + D S+ L D+ + PSS + + + SLH + C
Sbjct: 406 TAHTSSRLTRERDDPDAREDASSTGLGDKFPEAFMPSS----LCLAISTSLHI---LYPC 458
Query: 82 MP-TRNSYDTSGSLSHADTRATSAKA 106
P TR+ +D S+ HAD + SA A
Sbjct: 459 FPPTRSPFDMPPSIDHADLASASAAA 484
>ref|NP_011965.1| Hypothetical ORF; Yhr097cp [Saccharomyces cerevisiae]
gi|487941|gb|AAB68935.1| Yhr097cp [Saccharomyces
cerevisiae] gi|731687|sp|P38809|YHP7_YEAST Hypothetical
40.7 kDa protein in HXT5-NRK1 intergenic region
gi|626644|pir||S46727 hypothetical protein YHR097c -
yeast (Saccharomyces cerevisiae)
Length = 366
Score = 31.6 bits (70), Expect = 4.1
Identities = 15/57 (26%), Positives = 25/57 (43%), Gaps = 5/57 (8%)
Query: 35 KKNTTQHNDRDSSELRDQVSIHPSSS-----SFIITMLLGPSLHQCGWMDGCMPTRN 86
++ + H D+D + + + + P + +T L G S H G D C P RN
Sbjct: 104 RRTSHGHRDKDKQKSKSRTKVKPPKNVDTIDKMDVTGLFGGSFHHDGPFDACTPQRN 160
>emb|CAH95211.1| ubiquitin carboxyl-terminal hydrolase, putative [Plasmodium berghei]
Length = 1838
Score = 31.6 bits (70), Expect = 4.1
Identities = 13/47 (27%), Positives = 30/47 (63%)
Query: 28 QRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQ 74
++K KKKKKN Q+ ++++++ D+ S P++ ++ + L P + +
Sbjct: 1036 KKKKKKKKKNYAQNTKQNNADISDETSQLPNNINYFDSNLCFPKIEK 1082
>emb|CAG03333.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1417
Score = 31.2 bits (69), Expect = 5.3
Identities = 17/50 (34%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Query: 45 DSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCM-PTRNSYDTSGS 93
+ S R + S +P + S ++ GP+LHQ DGC P R+ Y G+
Sbjct: 436 EESPARRRKSPNPKAPSGLVERRHGPALHQQTGRDGCYHPRRDDYSDRGT 485
>ref|XP_213794.1| PREDICTED: similar to processing of precursor 5, ribonuclease P/MRP
family [Rattus norvegicus]
Length = 169
Score = 30.8 bits (68), Expect = 7.0
Identities = 13/44 (29%), Positives = 25/44 (56%)
Query: 1 MWRLVPSLTRMKSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDR 44
+W +P +T +++ HR C + TLH+ ++ +K Q+N R
Sbjct: 77 VWSALPFITCLENKGHRYSCFFNTLHVGGTIRTCQKFLIQYNRR 120
>ref|XP_574382.1| PREDICTED: similar to zinc finger protein 108 [Rattus norvegicus]
Length = 627
Score = 30.8 bits (68), Expect = 7.0
Identities = 17/43 (39%), Positives = 22/43 (50%), Gaps = 1/43 (2%)
Query: 21 SYITLHMQRKLKKKKKNTTQHND-RDSSELRDQVSIHPSSSSF 62
S++ LH Q L+ K T H D R SS + Q SIHP +
Sbjct: 231 SFLELHQQVLLEMKSSGHTAHEDTRRSSSVPIQQSIHPGEKRY 273
>emb|CAC59822.1| Pop5 protein [Homo sapiens] gi|15214743|gb|AAH12505.1| Processing
of precursor 5, ribonuclease P/MRP subunit, isoform a
[Homo sapiens] gi|20270249|ref|NP_057002.2| processing
of precursor 5, ribonuclease P/MRP subunit isoform a
[Homo sapiens]
Length = 163
Score = 30.4 bits (67), Expect = 9.1
Identities = 13/44 (29%), Positives = 25/44 (56%)
Query: 1 MWRLVPSLTRMKSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDR 44
+W +P +T +++ HR C + TLH+ ++ +K Q+N R
Sbjct: 77 VWSALPFITYLENKGHRYPCFFNTLHVGGTIRTCQKFLIQYNRR 120
>dbj|BAC27200.1| unnamed protein product [Mus musculus]
Length = 726
Score = 30.4 bits (67), Expect = 9.1
Identities = 22/84 (26%), Positives = 37/84 (43%)
Query: 23 ITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCM 82
+T+H + K K K ++ ++ R+ + S SSSS T S G
Sbjct: 555 LTVHSENKSKNKTRSRSRGRARNKTSKSRSRSSSSSSSSSSSTSSSSGSSSSSGSSSSRS 614
Query: 83 PTRNSYDTSGSLSHADTRATSAKA 106
+ +S TSGS S + +TS+ +
Sbjct: 615 SSSSSSSTSGSSSRDSSSSTSSSS 638
>ref|NP_080674.1| processing of precursor 5, ribonuclease P/MRP family [Mus musculus]
gi|18490752|gb|AAH22670.1| Processing of precursor 5,
ribonuclease P/MRP family [Mus musculus]
gi|12837743|dbj|BAB23935.1| unnamed protein product [Mus
musculus]
Length = 169
Score = 30.4 bits (67), Expect = 9.1
Identities = 13/44 (29%), Positives = 25/44 (56%)
Query: 1 MWRLVPSLTRMKSGNHRPCCSYITLHMQRKLKKKKKNTTQHNDR 44
+W +P +T +++ HR C + TLH+ ++ +K Q+N R
Sbjct: 77 VWSALPFITYLENKGHRYPCFFNTLHVGGTIRTCQKFLIQYNRR 120
>ref|NP_032917.1| pinin [Mus musculus] gi|2570049|emb|CAA69960.1| Pinin [Mus
musculus]
Length = 725
Score = 30.4 bits (67), Expect = 9.1
Identities = 22/84 (26%), Positives = 37/84 (43%)
Query: 23 ITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCM 82
+T+H + K K K ++ ++ R+ + S SSSS T S G
Sbjct: 554 LTVHSENKSKNKTRSRSRGRARNKTSKSRSRSSSSSSSSSSSTSSSSGSSSSSGSSSSRS 613
Query: 83 PTRNSYDTSGSLSHADTRATSAKA 106
+ +S TSGS S + +TS+ +
Sbjct: 614 SSSSSSSTSGSSSRDSSSSTSSSS 637
>ref|NP_704518.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23499257|emb|CAD51337.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 2245
Score = 30.4 bits (67), Expect = 9.1
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 2/32 (6%)
Query: 27 MQRKLKKKKKNTTQHNDRDSS--ELRDQVSIH 56
++ K KKKKKN TQ+ND D L+D + H
Sbjct: 180 LKNKKKKKKKNGTQNNDSDDEHVNLKDNKTSH 211
>gb|EAL01016.1| possible secreted protein [Candida albicans SC5314]
gi|46441595|gb|EAL00891.1| possible secreted protein
[Candida albicans SC5314]
Length = 430
Score = 30.4 bits (67), Expect = 9.1
Identities = 24/67 (35%), Positives = 35/67 (51%), Gaps = 8/67 (11%)
Query: 37 NTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCMPTRNSYDTSGSLSH 96
++T+ ND DSS R SI P+SS F+ +++ G L G +G NSY +H
Sbjct: 241 SSTKDNDSDSSSARGIKSI-PNSSEFLASLING--LGSGGGGNGSGSNTNSYK-----NH 292
Query: 97 ADTRATS 103
+ T TS
Sbjct: 293 STTSTTS 299
>gb|AAR97528.1| zinc finger protein 183 [Caenorhabditis briggsae]
gi|39591826|emb|CAE71404.1| Hypothetical protein
CBG18314 [Caenorhabditis briggsae]
Length = 385
Score = 30.4 bits (67), Expect = 9.1
Identities = 17/45 (37%), Positives = 25/45 (54%), Gaps = 1/45 (2%)
Query: 27 MQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPS 71
M + KKK+ N QH++ DS + D +I ++ SF T GPS
Sbjct: 45 MIQSTKKKEMNNAQHSEDDSDD-SDDANIAVATHSFAATGEAGPS 88
>gb|AAN05017.1| pinin [Mus musculus]
Length = 725
Score = 30.4 bits (67), Expect = 9.1
Identities = 22/84 (26%), Positives = 37/84 (43%)
Query: 23 ITLHMQRKLKKKKKNTTQHNDRDSSELRDQVSIHPSSSSFIITMLLGPSLHQCGWMDGCM 82
+T+H + K K K ++ ++ R+ + S SSSS T S G
Sbjct: 554 LTVHSENKSKNKTRSRSRGRARNKTSKSRSRSSSSSSSSSSSTSSSSGSSSSSGSSSSRS 613
Query: 83 PTRNSYDTSGSLSHADTRATSAKA 106
+ +S TSGS S + +TS+ +
Sbjct: 614 SSSSSSSTSGSSSRDSSSSTSSSS 637
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.128 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 252,287,645
Number of Sequences: 2540612
Number of extensions: 8899481
Number of successful extensions: 22733
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 22721
Number of HSP's gapped (non-prelim): 26
length of query: 150
length of database: 863,360,394
effective HSP length: 126
effective length of query: 24
effective length of database: 543,243,282
effective search space: 13037838768
effective search space used: 13037838768
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0035.10


