

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0034b.2
(494 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
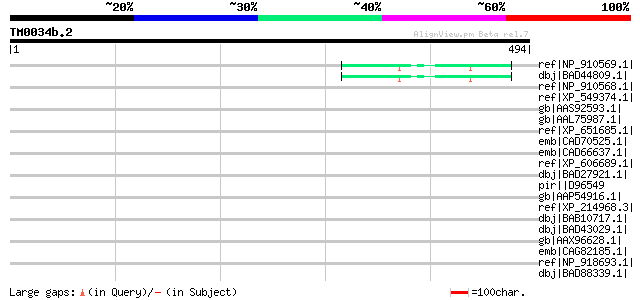
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_910569.1| Similar to Zea mays putative AC9 transposase. (... 52 6e-05
dbj|BAD44809.1| hypothetical protein [Oryza sativa (japonica cul... 52 6e-05
ref|NP_910568.1| Similar to Arabidopsis thaliana chromosome II B... 45 0.005
ref|XP_549374.1| PREDICTED: similar to putative protein (5I806) ... 40 0.13
gb|AAS92593.1| excretory/secretory protein Juv-p120 precursor [L... 39 0.29
gb|AAL75987.1| putative gypsy-type retrotransposon RIRE2 protein... 39 0.37
ref|XP_651685.1| hypothetical protein 152.t00013 [Entamoeba hist... 37 1.1
emb|CAD70525.1| hypothetical protein [Neurospora crassa] 36 2.4
emb|CAD66637.1| phytocyanin protein, PUP2 [Arabidopsis thaliana] 36 2.4
ref|XP_606689.1| PREDICTED: similar to glucocorticoid receptor D... 36 2.4
dbj|BAD27921.1| translational activator protein-like [Oryza sati... 36 2.4
pir||D96549 protein hypothetical protein F11M15.4 [imported] - A... 36 2.4
gb|AAP54916.1| putative gypsy-type retrotransposon protein [Oryz... 36 3.2
ref|XP_214968.3| PREDICTED: similar to microtubule-associated pr... 36 3.2
dbj|BAB10717.1| unnamed protein product [Arabidopsis thaliana] g... 36 3.2
dbj|BAD43029.1| predicted GPI-anchored protein [Arabidopsis thal... 36 3.2
gb|AAX96628.1| retrotransposon protein, putative, Ty3-gypsy sub-... 36 3.2
emb|CAG82185.1| unnamed protein product [Yarrowia lipolytica CLI... 35 4.1
ref|NP_918693.1| P0560B06.7 [Oryza sativa (japonica cultivar-gro... 35 4.1
dbj|BAD88339.1| membrane-associated protein-like [Oryza sativa (... 35 4.1
>ref|NP_910569.1| Similar to Zea mays putative AC9 transposase. (P03010) [Oryza
sativa (japonica cultivar-group)]
Length = 947
Score = 51.6 bits (122), Expect = 6e-05
Identities = 49/171 (28%), Positives = 66/171 (37%), Gaps = 24/171 (14%)
Query: 317 MFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVGIAPSP------- 369
M E +RLP PF VL AP QL PNAW + F L G+AP P
Sbjct: 92 MEEERLMRLPLQPFFAAVLAHFGLAPSQLTPNAWRLLAGFAALCRRAGVAPPPPPPLTVF 151
Query: 370 TSFFYLYDVDPKSIKNKGWISLKARAGRKCLHPHKSNAKSSFARKYFRVAVHPAYPEAFT 429
FF P + W S +AR S + S + V ++P + E F
Sbjct: 152 RHFFTAIGCPPGA-----WYSFRARG---------SASTGSLFTRLSNVTMNPWWKEQFI 197
Query: 430 LRDGTALF--PLYWTEKPNGITDPSEESLSANDKAFLNLLA-PLPILDCTT 477
L A + P+ W + P + + D A LLA ++D TT
Sbjct: 198 LVSSPAPWPCPVRWGKPRKHANFPPVLTTAEKDMATKLLLARGSSLIDLTT 248
>dbj|BAD44809.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 569
Score = 51.6 bits (122), Expect = 6e-05
Identities = 49/171 (28%), Positives = 66/171 (37%), Gaps = 24/171 (14%)
Query: 317 MFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVGIAPSP------- 369
M E +RLP PF VL AP QL PNAW + F L G+AP P
Sbjct: 92 MEEERLMRLPLQPFFAAVLAHFGLAPSQLTPNAWRLLAGFAALCRRAGVAPPPPPPLTVF 151
Query: 370 TSFFYLYDVDPKSIKNKGWISLKARAGRKCLHPHKSNAKSSFARKYFRVAVHPAYPEAFT 429
FF P + W S +AR S + S + V ++P + E F
Sbjct: 152 RHFFTAIGCPPGA-----WYSFRARG---------SASTGSLFTRLSNVTMNPWWKEQFI 197
Query: 430 LRDGTALF--PLYWTEKPNGITDPSEESLSANDKAFLNLLA-PLPILDCTT 477
L A + P+ W + P + + D A LLA ++D TT
Sbjct: 198 LVSSPAPWPCPVRWGKPRKHANFPPVLTTAEKDMATKLLLARGSSLIDLTT 248
>ref|NP_910568.1| Similar to Arabidopsis thaliana chromosome II BAC F13B15 genomic
sequence, putative non-LTR retroelement reverse
transcriptase. (AC006300) [Oryza sativa (japonica
cultivar-group)]
Length = 1285
Score = 45.1 bits (105), Expect = 0.005
Identities = 27/80 (33%), Positives = 36/80 (44%), Gaps = 8/80 (10%)
Query: 320 ELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVGIAPSPT---SFFYLY 376
E G+RLP PF +LR + AP Q+ PN W + F +LS PS FF L+
Sbjct: 306 EAGMRLPLHPFACELLRHLGVAPSQITPNGWRVVAGFLLLSHHAAAPPSLAVFRRFFRLF 365
Query: 377 DVDPKSIKNKGWISLKARAG 396
K GW + + G
Sbjct: 366 -----LSKLNGWYHFRGKRG 380
>ref|XP_549374.1| PREDICTED: similar to putative protein (5I806) [Canis familiaris]
Length = 1745
Score = 40.4 bits (93), Expect = 0.13
Identities = 29/111 (26%), Positives = 48/111 (43%), Gaps = 8/111 (7%)
Query: 55 PHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHAS----N 110
PH A + S + H P V NSP H H+P ++ ++PH +
Sbjct: 1097 PHQTAHPPSDSASPRIPCHTAHAPLERSSPVSQNSP-HRQHIP---VSQHIPHQTAYPRQ 1152
Query: 111 TRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFR 161
T HP S SPSP + ++P RP + + ++ P + + + PF+
Sbjct: 1153 TSHPLSDSPSPDREVIPRQRGHSRPDRSFHVRHRIPCQTADTPSDSTSPFQ 1203
>gb|AAS92593.1| excretory/secretory protein Juv-p120 precursor [Litomosoides
sigmodontis]
Length = 719
Score = 39.3 bits (90), Expect = 0.29
Identities = 55/217 (25%), Positives = 82/217 (37%), Gaps = 17/217 (7%)
Query: 54 PPHVAAPRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHA----S 109
P H PR +S ++ + F HFP+ + +P TH P I +S A +
Sbjct: 64 PTHFPYPRFSSTSTAAPFPPT-HFPYPRFSSTSTAAPFSPTHFPYPIFSSTSTAAPFPPT 122
Query: 110 NTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHF-------RSGAVAYHPFRY 162
+ +PS S S P P+ R TS A +P+ HF S A + P +
Sbjct: 123 HFPYPSFSSTSTAAPFPPTHFPYPRFSSTSTAAPFSPT-HFPYPIFSSTSTAAPFPPTHF 181
Query: 163 PLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPTSPNHSSSSSGSDPDATAAGGL 222
P + S+S S TH SF S + S T P + + + +
Sbjct: 182 P-YPSFSSTSTAAPFPPTHFPYPSFSSTS--TAVTSPRTFPYPRTFGTTPVFNHSVTFPT 238
Query: 223 RLEAPEIPPYRLKTYATLPLPVQIAPTHEAIDQPSIF 259
P IP + L T+ T P PT + P+ F
Sbjct: 239 AFPYPGIPTFPLLTFPT-AFPYPGIPTFPPVTFPTAF 274
>gb|AAL75987.1| putative gypsy-type retrotransposon RIRE2 protein [Sorghum bicolor]
Length = 447
Score = 38.9 bits (89), Expect = 0.37
Identities = 37/119 (31%), Positives = 52/119 (43%), Gaps = 7/119 (5%)
Query: 318 FSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAA-VGIAPSPTSFFYLY 376
F + G +P PF+Q +L C LH N+ I F LS A VGI P F YL+
Sbjct: 66 FLKRGFGVPVHPFLQGLLLYYEIGICNLHRNSILLISTFIHLSEAYVGIEPHFDLFRYLF 125
Query: 377 DVDPKSI----KNKGWISLKARAGRK--CLHPHKSNAKSSFARKYFRVAVHPAYPEAFT 429
+ K K G + L R G K LH + + + + RK+ + A P + T
Sbjct: 126 CLRKKGAVGGSKVAGGVYLNLRDGMKNRYLHCPWNTSLTEWYRKWSTSGKNRAAPRSAT 184
>ref|XP_651685.1| hypothetical protein 152.t00013 [Entamoeba histolytica HM-1:IMSS]
gi|56468454|gb|EAL46298.1| hypothetical protein
152.t00013 [Entamoeba histolytica HM-1:IMSS]
Length = 492
Score = 37.4 bits (85), Expect = 1.1
Identities = 35/149 (23%), Positives = 66/149 (43%), Gaps = 7/149 (4%)
Query: 65 VTSISLFLRNRHFPFHNR--PIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPI 122
V S S+ + + P H+ P+ S+ P+H++ +P + +S+ P S++ HP S P+
Sbjct: 235 VHSSSVPVHSSSHPVHSSSTPVQSSSHPVHSSSVP--VHSSSTPVQSSS-HPVHSSSHPV 291
Query: 123 KTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYS--GESHCPYLCIT 180
++ H P+ +S + + S S + H P+ +S S P +
Sbjct: 292 QSSSHPVHSSSVPVHSSSVPVHSSSVPVHSSSHPVHSSSVPVHSSSHPVHSSSTPVQSSS 351
Query: 181 HILIFSFLSLHCLLKMASNPTSPNHSSSS 209
H + S + + + +SP H SSS
Sbjct: 352 HPVHSSSVPVQSSSTPVHSSSSPKHGSSS 380
Score = 37.0 bits (84), Expect = 1.4
Identities = 35/154 (22%), Positives = 64/154 (40%), Gaps = 20/154 (12%)
Query: 70 LFLRNRHFPFHNR----PIVRSNSPLHATHLP-----KTIMTSNLPHASNTR------HP 114
+ +R RH P S+SP+H++ +P + +S++P S++ HP
Sbjct: 98 VIIRYRHIVIKEESSYIPTTSSSSPVHSSSVPVHSSSHPVRSSSIPIRSSSTPVKSSSHP 157
Query: 115 SSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGESHC 174
S +P+K+ H P+ +S Q+ S S + H +P+ + S
Sbjct: 158 VHSSSTPVKSSSVPIHSSSHPVHSSSTPVQSSSHPVHSSSHPVHSSSHPVHS-----SSV 212
Query: 175 PYLCITHILIFSFLSLHCLLKMASNPTSPNHSSS 208
P +H + S + +H + + P HSSS
Sbjct: 213 PVHSSSHPVHSSSVPVHSSSHPVHSSSVPVHSSS 246
>emb|CAD70525.1| hypothetical protein [Neurospora crassa]
Length = 550
Score = 36.2 bits (82), Expect = 2.4
Identities = 32/118 (27%), Positives = 45/118 (38%), Gaps = 12/118 (10%)
Query: 97 PKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVA 156
P T +S+ P +T P S S + T P +P + + S H S +
Sbjct: 89 PSTSGSSSHPRPPSTSKPPSISSTTTTTSTSDLKPNPKPKAFLRLLHIPHSPHRSSSLLI 148
Query: 157 YHPFRYPLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPTSPNHSSSSSGSDP 214
P LF + + H YL TH F L+ H NHSS+S+ S P
Sbjct: 149 PPPLLLRLFCLLNLDPHALYLHHTHTPGFHVLNCH------------NHSSTSTSSSP 194
>emb|CAD66637.1| phytocyanin protein, PUP2 [Arabidopsis thaliana]
Length = 370
Score = 36.2 bits (82), Expect = 2.4
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 3/92 (3%)
Query: 59 APRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHA-THLPKTIMTSNLPHASN--TRHPS 115
+P+ S S S H P H+ S+SP HA +H P + + HA + H
Sbjct: 210 SPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSX 269
Query: 116 SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPS 147
S SP+ K+ P S P P S + Q+PS
Sbjct: 270 SHSPATPKSPSPSSSPAQSPATPSPMTPQSPS 301
>ref|XP_606689.1| PREDICTED: similar to glucocorticoid receptor DNA binding factor 1,
partial [Bos taurus]
Length = 153
Score = 36.2 bits (82), Expect = 2.4
Identities = 25/74 (33%), Positives = 38/74 (50%), Gaps = 5/74 (6%)
Query: 66 TSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTL 125
T I LF++ F FHNRPI + P AT + + S +P ++T P + PSP ++
Sbjct: 77 TIIELFIQQCPFFFHNRPI---SEPPGATPSSPSALASTVPFLTST--PITSQPSPPQSP 131
Query: 126 LPHSHPFLRPLFTS 139
P ++PL S
Sbjct: 132 PPTPQSPMQPLLPS 145
>dbj|BAD27921.1| translational activator protein-like [Oryza sativa (japonica
cultivar-group)] gi|50252663|dbj|BAD28832.1|
translational activator protein-like [Oryza sativa
(japonica cultivar-group)]
Length = 920
Score = 36.2 bits (82), Expect = 2.4
Identities = 34/133 (25%), Positives = 53/133 (39%), Gaps = 11/133 (8%)
Query: 64 SVTSISLFLRNRHFPFHNRPIVRSNSPLHATHLPKTIMTSNLPHASNTRHPSS---FSPS 120
S+ +++ L N P RS P A + P T+ ++P SN HP + P+
Sbjct: 508 SLRTVAAILPNNGLPEFE---ARSAEPSEAANAPPTV---SVPAGSNATHPVTPVPQPPA 561
Query: 121 PIKTLLPHSHPFLRPLFTSLIAYQTPSAHFRSGAVAYHPFRYPLFTSYSGESHCPYLCIT 180
P L P + F SL+ + SA + +A YP+ + + Y I
Sbjct: 562 PAADQLQDMIPMQKEAFISLLKAISTSAASATATIAIFVVSYPI--KWPDKVSVNYQMIL 619
Query: 181 HILIFSFLSLHCL 193
L+F F L L
Sbjct: 620 EALLFVFFPLALL 632
>pir||D96549 protein hypothetical protein F11M15.4 [imported] - Arabidopsis
thaliana gi|4836929|gb|AAD30631.1| Hypothetical protein
[Arabidopsis thaliana]
Length = 663
Score = 36.2 bits (82), Expect = 2.4
Identities = 35/141 (24%), Positives = 54/141 (37%), Gaps = 12/141 (8%)
Query: 313 AYEYMFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEILSAAVGIAPSPTSF 372
A+E FSE G+ P + ++ + A QL PN + C L+ G + + F
Sbjct: 44 AFEIFFSECGLFFPLPDILIDLMHEFGIALPQLCPNVIRTVLCLLTLAEEDGFRLALSDF 103
Query: 373 FYLYDVDPKSIKNKGWISLKARAGRKCLH--PHKSNAKSSFARKYFRVAVHPAYPEAFTL 430
LY V K + L R G + P K + + YF V+ T
Sbjct: 104 LQLYAV--KKSRTNNTFFLSPRKGFRVFEDFPEKD---EHWRKSYFFFPVN-----ELTY 153
Query: 431 RDGTALFPLYWTEKPNGITDP 451
+ + LF WT + +T P
Sbjct: 154 EEKSDLFVSDWTTRAVKVTIP 174
>gb|AAP54916.1| putative gypsy-type retrotransposon protein [Oryza sativa (japonica
cultivar-group)] gi|37536654|ref|NP_922629.1| putative
gypsy-type retrotransposon protein [Oryza sativa
(japonica cultivar-group)] gi|13876534|gb|AAK43510.1|
putative gypsy-type retrotransposon protein [Oryza
sativa (japonica cultivar-group)]
Length = 742
Score = 35.8 bits (81), Expect = 3.2
Identities = 32/128 (25%), Positives = 58/128 (45%), Gaps = 5/128 (3%)
Query: 322 GIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCF-EILSAAVGIAPSPTSFFYLYDVD- 379
G P S F+ VL + L+PN+ AF+ F + A +G+ P F + Y++
Sbjct: 27 GFLPPPSEFLLLVLNFYGLSLLHLNPNSIAFLSIFVHLCEAYIGVEPFLDLFRFYYELRW 86
Query: 380 PKSIKNKGWISLKARAGRKCLH-PHK-SNAKSSFARKYFRVAVHPAYPEAFTLRDGTALF 437
+S + G + + R G K + P + +++S + ++F + + + P D
Sbjct: 87 MESNRVSGCVGFRLRDGMKSRYIPFQCPSSRSKWRNRWFYLEIKDSNPVFVVPEDQPDKI 146
Query: 438 PLYWTEKP 445
P WT KP
Sbjct: 147 PA-WTAKP 153
>ref|XP_214968.3| PREDICTED: similar to microtubule-associated protein 7 [Rattus
norvegicus]
Length = 804
Score = 35.8 bits (81), Expect = 3.2
Identities = 31/99 (31%), Positives = 44/99 (44%), Gaps = 15/99 (15%)
Query: 44 PYFKTLLNLYPPHVAAPRDASVTSISLFLRNR-HFPFHNRPIVR---------SNSPLHA 93
P + L + PP +A R ++ IS + R R H PFH P VR + SP A
Sbjct: 343 PVDRPKLFITPPEGSARR-RTIHGISSYKREREHIPFHVSPGVRRTLSPSNLKARSPAPA 401
Query: 94 THLPKTIMTSNLPHASNTRHP-SSFSPSPIKTLLPHSHP 131
P + ++PH T P SS +P P++ S P
Sbjct: 402 RLWPP---SKSMPHLPGTPRPASSLTPGPVRAAASSSSP 437
>dbj|BAB10717.1| unnamed protein product [Arabidopsis thaliana]
gi|15238868|ref|NP_200198.1| plastocyanin-like
domain-containing protein [Arabidopsis thaliana]
Length = 370
Score = 35.8 bits (81), Expect = 3.2
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 3/92 (3%)
Query: 59 APRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHA-THLPKTIMTSNLPHA--SNTRHPS 115
+P+ S S S H P H+ S+SP HA +H P + + HA + H
Sbjct: 210 SPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSP 269
Query: 116 SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPS 147
S SP+ K+ P S P P S + Q+PS
Sbjct: 270 SHSPATPKSPSPSSSPAQSPATPSPMTPQSPS 301
>dbj|BAD43029.1| predicted GPI-anchored protein [Arabidopsis thaliana]
Length = 370
Score = 35.8 bits (81), Expect = 3.2
Identities = 29/92 (31%), Positives = 41/92 (44%), Gaps = 3/92 (3%)
Query: 59 APRDASVTSISLFLRNRHFPFHNRPIVRSNSPLHA-THLPKTIMTSNLPHA--SNTRHPS 115
+P+ S S S H P H+ S+SP HA +H P + + HA + H
Sbjct: 210 SPKSPSPVSHSPSHSPAHTPSHSPAHTPSHSPAHAPSHSPAHAPSHSPAHAPSHSPAHSP 269
Query: 116 SFSPSPIKTLLPHSHPFLRPLFTSLIAYQTPS 147
S SP+ K+ P S P P S + Q+PS
Sbjct: 270 SHSPATPKSPSPSSSPAQSPATPSPMTPQSPS 301
>gb|AAX96628.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 546
Score = 35.8 bits (81), Expect = 3.2
Identities = 33/130 (25%), Positives = 59/130 (45%), Gaps = 9/130 (6%)
Query: 322 GIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIRCFEIL-SAAVGIAPSPTSFFYLYD--- 377
G P S F+ VL + L+PN+ AF+ F L A +G+ P F + Y+
Sbjct: 165 GFLPPPSEFLLLVLNFYGLSLLHLNPNSIAFLSIFSYLCEAYIGVEPFLDLFRFYYELRW 224
Query: 378 VDPKSIKNKGWISLKARAGRKCLH-PHK-SNAKSSFARKYFRVAVHPAYPEAFTLRDGTA 435
++PK + G + + R G K + P + +++S + ++F + + + P +
Sbjct: 225 MEPKKV--SGCVGFRLRDGMKSRYIPFQCPSSRSKWRGRWFYLQIENSDPVFIVPEEQPD 282
Query: 436 LFPLYWTEKP 445
P WT KP
Sbjct: 283 KIP-EWTTKP 291
>emb|CAG82185.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50548805|ref|XP_501872.1| hypothetical protein
[Yarrowia lipolytica]
Length = 812
Score = 35.4 bits (80), Expect = 4.1
Identities = 55/213 (25%), Positives = 91/213 (41%), Gaps = 22/213 (10%)
Query: 85 VRSNSPLHATHLPKTIMTSNLPHASNTRHPSSFSPSPIKTLLPHSHPFLRPLFTSLIAYQ 144
V S+SP+ ++ +S+ P +S++ PSS SPS + S P +S +
Sbjct: 388 VSSSSPVSSSS-----PSSSNPGSSSSSSPSSSSPSSSSSSPSSSSSSSSPSSSSSSSSS 442
Query: 145 TPSAHFRSGAVAYHPFRYPLFTSYSGESHCPYLCITHILIFSFLSLHCLLKMASNPTSPN 204
+PS+ S + + +S S S P + S S +S+ TS +
Sbjct: 443 SPSSSSSSSSPS---------SSSSFSSSSPSSSSSSSSSSSSSSSPSASSSSSSSTSLS 493
Query: 205 HSSSSSGSDPDATAAGGLRLEAPEIPPY---RLKTYATLPLPVQIAPTHEAIDQPSIFQN 261
SSSSS S ++++ L L + P+ R+ + P Q E I Q + N
Sbjct: 494 SSSSSSSS---SSSSAPLPLVTQQGMPWGLSRISHQSPEMAPYQDPMVGEYIHQQFVDPN 550
Query: 262 SDLIVQFVNSLGGLSNDDEFSAKLRIWICSPGD 294
++V V+S G N D F+ K IW+ + D
Sbjct: 551 PKVVVYVVDS-GVNINHDNFATK-PIWLANYAD 581
>ref|NP_918693.1| P0560B06.7 [Oryza sativa (japonica cultivar-group)]
gi|15528678|dbj|BAB64744.1| P0560B06.7 [Oryza sativa
(japonica cultivar-group)]
Length = 917
Score = 35.4 bits (80), Expect = 4.1
Identities = 29/89 (32%), Positives = 38/89 (42%), Gaps = 9/89 (10%)
Query: 295 RPWLHKPDGDVKRSHFFFAYEYMFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIR 354
RP H P RS FF F+ G+ LPFS F VL + L PNA +
Sbjct: 47 RPAPHYPG----RSVFFLP----FAMAGLVLPFSSFFMDVLEFYDLQMAHLTPNAVMTLA 98
Query: 355 CF-EILSAAVGIAPSPTSFFYLYDVDPKS 382
F + +G+ PS F + + V P S
Sbjct: 99 IFAHLCEMFIGVRPSLRLFRWFFTVQPVS 127
>dbj|BAD88339.1| membrane-associated protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 853
Score = 35.4 bits (80), Expect = 4.1
Identities = 29/89 (32%), Positives = 38/89 (42%), Gaps = 9/89 (10%)
Query: 295 RPWLHKPDGDVKRSHFFFAYEYMFSELGIRLPFSPFVQTVLRDINAAPCQLHPNAWAFIR 354
RP H P RS FF F+ G+ LPFS F VL + L PNA +
Sbjct: 47 RPAPHYPG----RSVFFLP----FAMAGLVLPFSSFFMDVLEFYDLQMAHLTPNAVMTLA 98
Query: 355 CF-EILSAAVGIAPSPTSFFYLYDVDPKS 382
F + +G+ PS F + + V P S
Sbjct: 99 IFAHLCEMFIGVRPSLRLFRWFFTVQPVS 127
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 928,643,539
Number of Sequences: 2540612
Number of extensions: 41857605
Number of successful extensions: 101361
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 31
Number of HSP's that attempted gapping in prelim test: 101300
Number of HSP's gapped (non-prelim): 76
length of query: 494
length of database: 863,360,394
effective HSP length: 132
effective length of query: 362
effective length of database: 527,999,610
effective search space: 191135858820
effective search space used: 191135858820
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0034b.2


