

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0029b.1
(1377 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
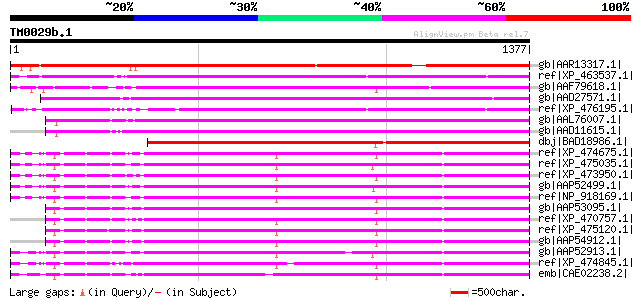
Sequences producing significant alignments: (bits) Value
gb|AAR13317.1| gag-pol polyprotein [Phaseolus vulgaris] 1264 0.0
ref|XP_463537.1| putative polyprotein [Oryza sativa (japonica cu... 1033 0.0
gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||... 1011 0.0
gb|AAD27571.1| polyprotein [Sorghum bicolor] gi|4378066|gb|AAD19... 957 0.0
ref|XP_476195.1| putative polyprotein [Oryza sativa (japonica cu... 947 0.0
gb|AAL76007.1| prpol [Zea mays] 929 0.0
gb|AAD11615.1| prpol [Zea mays] gi|7489805|pir||T14595 polyprote... 917 0.0
dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera] 912 0.0
ref|XP_474675.1| OSJNBa0027O01.4 [Oryza sativa (japonica cultiva... 857 0.0
ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultiva... 856 0.0
ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultiv... 856 0.0
gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonic... 855 0.0
ref|NP_918169.1| putative gag-pol polyprotein [Oryza sativa (jap... 855 0.0
gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cul... 849 0.0
ref|XP_470757.1| putative gag-pol precursor [Oryza sativa] gi|18... 848 0.0
ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cu... 843 0.0
gb|AAP54912.1| gag-pol precursor [Oryza sativa (japonica cultiva... 841 0.0
gb|AAP52913.1| putative retroelement [Oryza sativa (japonica cul... 838 0.0
ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultiva... 836 0.0
emb|CAE02238.2| OSJNBb0054B09.2 [Oryza sativa (japonica cultivar... 832 0.0
>gb|AAR13317.1| gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 1264 bits (3271), Expect = 0.0
Identities = 649/1433 (45%), Positives = 919/1433 (63%), Gaps = 95/1433 (6%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGY------------------------QGNRQGQ 37
C++HR+ GH T+ C+ L+ +I++L++AG+ G R +
Sbjct: 464 CQYHRNYGHTTEGCQALKDKIEELVQAGHLRKFVKTTITAPRSPQRDHDPRERSGRRDDR 523
Query: 38 WRNGGDQNKAHKREE----------ERADTKGKKKQESAAIATKGADDTFAQHSGPPVGT 87
R+ ++ KR E E + + + KQ+ + A G P
Sbjct: 524 TRDNHYRSSRRKRSESPIRRTRPKSESPERRSRTKQKVRTVINTIAGPVSL---GQPPQE 580
Query: 88 INTIAGGFGGGGDTHAARKRHVRAVNSVHEVAFGFVH--PDITISMADFEGIKPHKDDPI 145
IN IAGGF GGG +++ARK+H+RA+ SVH P IT + DF I P +DDP+
Sbjct: 581 INYIAGGFAGGGCSNSARKKHLRAIQSVHSTPTQRRPHIPPITFTDEDFTAIDPSQDDPM 640
Query: 146 VVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMVRGY 205
V+ + ++ F I +VL+DQGSS DI+Y + F K+ + + ++ PY +VGF+ E+V +G+
Sbjct: 641 VITVEIDKFAIAKVLVDQGSSVDILYWETFKKMKIPEAEIQPYNEQIVGFSRERVDTKGF 700
Query: 206 IDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGK 265
IDL T FG+D ++ + +RYL++ SYN+++GR ++NRL A++ST HLA+K+P +G
Sbjct: 701 IDLYTTFGDDYLSKTINIRYLLVNANTSYNILLGRPSINRLKAIVSTPHLAMKFPSVNGD 760
Query: 266 VGRLKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRP------ 319
+ + +DQK ARECY L + + E + + R GR +R
Sbjct: 761 IATVHIDQKTARECYVASLKVEPTRRLYTTSAERTTERRGRSTERRSRGRESRRHLVALV 820
Query: 320 --TPIEETKALKFG-----------DRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKD 366
P + ++ G DR +GT L + + K L +N DLFAW+ D
Sbjct: 821 DLDPRLDDPRMEAGEDLQPIFLRDKDRKTYMGTSLKPDDRETIGKTLTKNADLFAWTAAD 880
Query: 367 MPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLA 426
MPG+ + I HRL++ +P++Q +R+LG ++ KA ++E DKL+ A FI++ Y TWLA
Sbjct: 881 MPGVKSDVITHRLSVYTEARPIAQKKRKLGEERRKAAREETDKLIQAGFIQKAHYTTWLA 940
Query: 427 NVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQI 486
NVVMVKK NGKWRMCVDYTDLNKACPKDSYPLP+ID LVDGA+G+++LS +DAYSGY+QI
Sbjct: 941 NVVMVKKTNGKWRMCVDYTDLNKACPKDSYPLPTIDRLVDGAAGHQILSFLDAYSGYNQI 1000
Query: 487 RMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIV 546
+M+ D +KTAF T N+ Y MPFGLKNAGATYQRLMD VF +GRN+EVYVDD++V
Sbjct: 1001 QMYHRDREKTAFRTDSDNFFYEVMPFGLKNAGATYQRLMDHVFHDMIGRNVEVYVDDIVV 1060
Query: 547 KSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAI 606
KS H DL+E F +R++ MRLNPEKC+FGV+GGKFLGFM+T RGIE NPEKCKAI
Sbjct: 1061 KSDSCEQHVSDLKEVFQALRQYRMRLNPEKCAFGVEGGKFLGFMLTHRGIEANPEKCKAI 1120
Query: 607 QQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLK 666
+M+SP +KE+QRL R+ +LSRF+PK +R+ P K L+K FEWT ECE+ F +LK
Sbjct: 1121 TEMRSPKGLKEIQRLVSRLTSLSRFVPKLAERTRPIIKLLKKTSKFEWTDECEQNFQQLK 1180
Query: 667 ELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQK 726
L+SPP++ KP P+ +Y AVS+ A+SS ++QEI E R VYFVS L AE RYQ
Sbjct: 1181 AFLASPPVIQKPNAREPIVVYLAVSNEAVSSALVQEIKAEERPVYFVSRVLHDAETRYQM 1240
Query: 727 IEKAALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYD 786
+EK A A+++TARR+R YFQ+ V VRT+ P+ ++L KPDL+GR++ W+VELSE+ ++Y
Sbjct: 1241 VEKVAFALVITARRMRMYFQNHKVIVRTNYPIMKILTKPDLAGRMIGWAVELSEFHIEYQ 1300
Query: 787 KRGTVGAQSLADFVVELTPDRFERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSL 846
RG + +Q+LADF ELTP ER +WTL+VDGSSNS SGAGV LEGPGE+V+EQ++
Sbjct: 1301 PRGAIKSQALADFTAELTPYLTERT-PRWTLYVDGSSNSRSSGAGVVLEGPGEIVVEQAM 1359
Query: 847 KFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLE 906
KFEFK +NNQAEYEA+IAGL LA E+++ ++ ++DS+LV Q+ G ++V++ L +Y
Sbjct: 1360 KFEFKTSNNQAEYEAIIAGLHLAIELEVTNITCKSDSRLVVGQLTGEYEVRETLLQQYFH 1419
Query: 907 RVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSC 966
V+ L+ F+E+ ++V R N RADAL++LA+ +K G ++S I TLA PS+ E
Sbjct: 1420 FVKNLLNRFKEISFQHVRRENNTRADALSRLATLKKKGAHRSAIHVTLAKPSVGTEECMA 1479
Query: 967 VNRGRTWMDPIISILAGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEK 1026
+ WM PI L + K + +A+ Y LI LYRRG+S PLLKC+ PE+
Sbjct: 1480 TDTQPNWMTPIKQYLTDGVCD-PHLEKTMKLQAARYILIGEDLYRRGYSRPLLKCLGPEQ 1538
Query: 1027 YEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAP 1086
+M+E+HEG+C +H G R+++ K+LRAG+YWPTL+ DC ++
Sbjct: 1539 VTYVMTELHEGICGTHSGARTMSAKILRAGYYWPTLQGDCTEY----------------- 1581
Query: 1087 PKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVN 1146
G+D++GPF + Q KF+LV +DYFTKWIEAEPL IT+ + +
Sbjct: 1582 ----------------GMDIIGPFTPGKGQCKFLLVGIDYFTKWIEAEPLTAITARNVQS 1625
Query: 1147 FYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRV 1206
F WK IVCRFG+P+ I++DNG QF+ EF +++ I+ +SVEHPQTNGQ E+AN+V
Sbjct: 1626 FVWKNIVCRFGLPQIIITDNGRQFTDRGLAEFYEKLHIKHITSSVEHPQTNGQAEAANKV 1685
Query: 1207 ILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWR 1266
IL L++RL +KG W +EL VLW+Y T QSTT+ETP+ +TYG +AM+PVEI + R
Sbjct: 1686 ILNELKKRLGPSKGNWTEELLEVLWAYRCTPQSTTQETPYSLTYGTEAMIPVEIGEPSLR 1745
Query: 1267 TRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLKW 1326
+ + N+ ++ V LDL++E RD+ IRE A K R A ++NS+V+ R Q GDLV +
Sbjct: 1746 RQTLDLDLNKESLLVGLDLINELRDKCKIREEACKIRAARRYNSKVKPRSYQKGDLVWRM 1805
Query: 1327 RSGA--PGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
RS A G K + NWEGP+RI GAY+LE L G+ PR++N L+++Y
Sbjct: 1806 RSDARKDGGKFSSNWEGPFRISNTATGGAYYLEYLSGKSAPRTWNATHLKFYY 1858
>ref|XP_463537.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2001
Score = 1033 bits (2671), Expect = 0.0
Identities = 554/1392 (39%), Positives = 834/1392 (59%), Gaps = 58/1392 (4%)
Query: 1 WCEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGK 60
+C H GH T++CR + ++K + N+ + E D +G+
Sbjct: 652 FCHIHGPGGHSTEECRQMTHLLEKHV------------------NRYENKYEGARDQRGQ 693
Query: 61 KKQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHVRAVNSVHEVAF 120
E+ + A + P IN I GG G ++ RK +VR VN V
Sbjct: 694 NAIEAPQVMKIEAIEEV------PKRVINAITGGSSLGVESKRQRKAYVRQVNHVGTSYQ 747
Query: 121 G----FVHPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFD 176
+ I+ D EGI DP+VV + + ++RVL+D GSSAD+++ DAF
Sbjct: 748 SNPPVYSKTVISFGPEDAEGILFPHQDPLVVSVEIAQCEVQRVLIDGGSSADVLFYDAFK 807
Query: 177 KLGLTDNDLTPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNV 236
K+ + ++ LT L GF G+QV G I L +FG+ R ++ + V+ + YN
Sbjct: 808 KMQIPEDRLTNAGVPLQGFGGQQVHAIGKISLQVVFGKGTNVRKEEIVFDVVDMPYQYNA 867
Query: 237 IIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNNCLNLYGKKSALVGH 296
I+GR+T+N A+I ++ +K P G + ++ +Q AR+ L G S H
Sbjct: 868 ILGRSTINIFEAIIHHNYICMKLPGLRGVI-TVRGEQLAARK-----YELQGTPSVKGVH 921
Query: 297 RCYEIEASDENLDPRGEG-RVNRPTPIEETKALKFGDRT----LKIGTRLTEEQETRLTK 351
D+ +GE ++ +P P +TK ++ + + IG L + E + K
Sbjct: 922 ------VVDQK---QGEYIKIQKPIPEGKTKKVQLDEHNPGKFILIGENLEKHIEEEILK 972
Query: 352 LLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLL 411
++ EN+ +FAWS ++ G+D + I H LA+ KP Q RR+ D+ +A + E++KLL
Sbjct: 973 VVKENMAVFAWSPDELQGVDRSLIEHNLAIKSGYKPKKQKLRRMSTDRQQAAKIELEKLL 1032
Query: 412 AAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGN 471
A+ IREV +P WLAN V+VKKANGKWRMC+D+TDLNKACPKD +PLP ID LVD +G
Sbjct: 1033 KAKVIREVMHPEWLANPVLVKKANGKWRMCIDFTDLNKACPKDDFPLPRIDQLVDATAGC 1092
Query: 472 ELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAR 531
EL+S +DAYSGYHQ+ M DE+KT+F+T YC+ MPFGLKNAGAT+ RL+ +V A+
Sbjct: 1093 ELMSFLDAYSGYHQVFMVKEDEEKTSFITPFGTYCFIRMPFGLKNAGATFARLIGKVLAK 1152
Query: 532 QVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMI 591
Q+GRN+E Y+DD++VKS + H +DL+E F +RK S++LNPEKC FGV+ GK LGF++
Sbjct: 1153 QLGRNVEAYIDDIVVKSKQAFTHGKDLQETFENLRKCSVKLNPEKCVFGVRAGKLLGFLV 1212
Query: 592 TSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVV 651
+ RGIE NP+K AI QM+ P N +EVQRLTGR+A+LSRFL KS ++ PFFK LR
Sbjct: 1213 SKRGIEANPDKIAAIHQMEPPKNTREVQRLTGRMASLSRFLSKSAEKGLPFFKTLRGANT 1272
Query: 652 FEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVY 711
FEWTAEC++AF LK+ L P L+ P +G PL +Y A + + +S+V++QE + VY
Sbjct: 1273 FEWTAECQQAFDDLKKYLHEMPTLASPPKGQPLLMYVAATPATVSAVLVQEEENRQVPVY 1332
Query: 712 FVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRL 771
FVS LQG + RY ++EK A+++ +R+LR YF S + + + P+ +VL +++GR+
Sbjct: 1333 FVSEALQGPKTRYSEVEKLIYAIVMASRKLRHYFLSHDITIPSAYPIGEVLTNKEVAGRI 1392
Query: 772 VAWSVELSEYGLQYDKRGTVGAQSLADFVVELTPDRFER---VDTQWTLFVDGSSNSSGS 828
W++EL + L+Y R + +Q LADFV E TP+ E+ V W +F DG+ N++G+
Sbjct: 1393 AKWAMELLPFDLKYISRTAIKSQVLADFVAEWTPNEVEQQEEVKKPWIVFSDGACNAAGA 1452
Query: 829 GAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVEN 888
GA ++ P + L+ S++ F +TNN AEYE ++ ++ AR + R L+++TDS+LV
Sbjct: 1453 GAAAVVKTPMKQTLKYSVQLVFPSTNNTAEYEGVLLAMRKARALGARRLIVKTDSKLVAG 1512
Query: 889 QVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKS 948
+F+ K+ + KYLE R F + V+ + R EN AD LAK A+T +P N
Sbjct: 1513 HFSKSFEAKEETMAKYLEEARLNEKHFLGITVKAITREENGEADELAKAAATGQPLENS- 1571
Query: 949 VIQETLAYPSIEGELMSCVNRGRTWMDPIIS-ILAGDPAEVEQCTKEQQREASHYTLIDG 1007
+ + PS E + ++C+ R W +PI+ +++ E E+ K Q + Y +++G
Sbjct: 1572 -FFDIITQPSYEKKEVACIQREGDWREPILKYLVSAQLPEKEEEAKRIQLMSKKYKVVEG 1630
Query: 1008 HLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCM 1067
LY+ G + PLLKCV+ E+ ++ E+HEG+C +H S+A KV+R G YWPT+ KD
Sbjct: 1631 QLYKSGVTAPLLKCVTREEGMKMVVEIHEGLCGAHQAPWSVASKVIRQGIYWPTIMKDTE 1690
Query: 1068 DFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYF 1127
++K CK CQ F ++KAPPKEL + WPF WG+D+VGP P A+ ++F++VA++YF
Sbjct: 1691 KYIKTCKACQKFGPMTKAPPKELQPIPPVWPFYRWGIDIVGPLPRAKGDLRFVIVAIEYF 1750
Query: 1128 TKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMR 1187
++WIEAE +A+ITSA + F WK I+CRFGIP+ IV DNG QF S + ++ CK + +Q+
Sbjct: 1751 SRWIEAEAVARITSAAVQKFVWKNIICRFGIPKEIVCDNGKQFESGKFQDMCKGLNLQIN 1810
Query: 1188 FASVEHPQTNGQVESANRVILRGLRRRL-AEAKGAWLDELPAVLWSYNTTEQSTTRETPF 1246
FASV HPQTNG VE AN I+ +++RL AKG W ++L +VLW+ TT +T TPF
Sbjct: 1811 FASVGHPQTNGVVERANGKIMEAIKKRLEGSAKGKWPEDLLSVLWALRTTVVRSTGMTPF 1870
Query: 1247 RMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAA 1306
R+ YG +AM P E+ + R F+++++ + L++L E R EA + + + +
Sbjct: 1871 RLVYGDEAMTPSEVGAHS--PRMIFDQKDEEGREITLEMLDEIRVEALEKMASYTEGTKS 1928
Query: 1307 KFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLP 1365
+N +V+ R ++ GDLVL K + KL WEGP+ + K GA+ L LDG L
Sbjct: 1929 YYNQKVKTRPIEEGDLVLKKVLNEVAVGKLESKWEGPFIVKKKTETGAFKLAYLDGEELK 1988
Query: 1366 RSFNGLSLRYFY 1377
++N +SL+ FY
Sbjct: 1989 HTWNAVSLKKFY 2000
>gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||H86337 protein
F5M15.26 [imported] - Arabidopsis thaliana
Length = 1838
Score = 1011 bits (2613), Expect = 0.0
Identities = 584/1426 (40%), Positives = 822/1426 (56%), Gaps = 94/1426 (6%)
Query: 1 WCEFHRSAGHDTDDCRTLQREIDKLIRAGYQ-GNRQGQWRNGGDQNKAHKREEERA---- 55
+CE+H+S H T++CR LQ L+ A Y+ G + NK +R E A
Sbjct: 457 YCEYHKSRAHSTENCRFLQG----LLMAKYKSGGITIECDRPPINNKNQRRNETTARQYL 512
Query: 56 --DTKGKKKQESAAIATKGADDTFA--QHSGPPVGT-------INTIAGGFGGGGDTHAA 104
TK E I + ADD A Q +G + ++ I GG D+ +
Sbjct: 513 NDQTKPPTPAEQGIITS--ADDPAAKRQRNGKAIAAEPVVVRQVHVIMGGLQNCSDSVRS 570
Query: 105 RKRHVRAVNSVHEVAFGFV-----HPD----ITISMADFEGIKPHKDDPIVVQLRMNSFN 155
K++ + V VA+ +P+ I+ + D EG+ +DP+VV+L ++
Sbjct: 571 IKQYRKKAEMV--VAWPSSTSTTRNPNQSAPISFTDVDLEGLDTPHNDPLVVELIISDSR 628
Query: 156 IRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMVRGYIDLDTIFGED 215
+ RVL+D GSS D+I+ D + +TD + P + L GF G+ VM G I L G
Sbjct: 629 VTRVLIDTGSSVDLIFKDVLTAMNITDRQIKPVSKPLAGFDGDFVMTIGTIKLPIFVG-- 686
Query: 216 ECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKM 275
+ V+++V+ A YNVI+G ++++ A+ ST H VK+P
Sbjct: 687 --GLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIPSTYHQCVKFPT-------------- 730
Query: 276 ARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGDRTL 335
+N L K A R YE + L ++ P R +
Sbjct: 731 ----HNGIFTLRAPKEAKTPSRSYE----ESELCRTEMVNIDESDPT----------RCV 772
Query: 336 KIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRL 395
+G ++ L LL N FAWS +DM GIDP H L ++P+ KPV Q RR+L
Sbjct: 773 GVGAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNVDPTFKPVKQKRRKL 832
Query: 396 GGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDS 455
G ++ +AV +EV+KLL A I EVKYP WLAN V+VKK NGKWR+CVDYTDLNKACPKDS
Sbjct: 833 GPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDS 892
Query: 456 YPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLK 515
YPLP ID LV+ SGN LLS MDA+SGY+QI MH D++KT+F+T R YCY+ M FGLK
Sbjct: 893 YPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTYCYKVMSFGLK 952
Query: 516 NAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPE 575
NAGATYQR ++++ A Q+GR +EVY+DDM+VKS++ DH + L + F + + M+LNP
Sbjct: 953 NAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPT 1012
Query: 576 KCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKS 635
KC+FGV G+FLG+++T RGIE NP++ +AI ++ SP N +EVQRLTGRIAAL+RF+ +S
Sbjct: 1013 KCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFISRS 1072
Query: 636 GDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSAL 695
D+ PF+ L++ F+W + EEAF +LK+ LS+PPIL KP G L+LY AVSD A+
Sbjct: 1073 TDKCLPFYNLLKRRAQFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAV 1132
Query: 696 SSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTD 755
SSV+++E GE R +++ S +L AE RY IEKAALAV+ +AR+LRPYFQS + V TD
Sbjct: 1133 SSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTD 1192
Query: 756 LPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELTPDRFERV---- 811
PLR L P SGR+ W+VELSEY + + R + +Q LADF++EL ER
Sbjct: 1193 QPLRVALHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQVLADFLIELPLQSAERAVSGN 1252
Query: 812 -DTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAR 870
+W+L+VDGSS++ GSG G+ L P VLEQS + F ATNN AEYE LIAGL+LA
Sbjct: 1253 RGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRFVATNNVAEYEVLIAGLRLAA 1312
Query: 871 EVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQR 930
++I ++ TDSQL+ Q+ G ++ K+ + YL+ V+ + F+ + +PR +N
Sbjct: 1313 GMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQLMTKDFENFKLSKIPRGDNAP 1372
Query: 931 ADALAKLASTRKPGNNKSVIQETLAYPSIEG-ELMSCVNRGRT-----------WMDPII 978
ADALA LA T + + E++ PSI+ + + VN R+ W I
Sbjct: 1373 ADALAALALTSDSDLRRIIPVESIDKPSIDSTDAVEIVNTIRSSNAPDPADPTDWRVEIR 1432
Query: 979 SILAGDPAEVEQCTKEQQR-EASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEG 1037
L+ ++ T + R +A+ YTL+ HL + +L C+ + IM E HEG
Sbjct: 1433 DYLSDGTLPSDKWTARRLRIKAAKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKETHEG 1492
Query: 1038 VCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPW 1097
+H GGR+LA K+ + GFYWPT+ DC F +C++CQ A P + L AP+
Sbjct: 1493 AAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQPTELLRAGVAPY 1552
Query: 1098 PFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFG 1157
PF W +D+VGP P +R Q +FILV DYFTKW+EAE A I + + NF WK I+CR G
Sbjct: 1553 PFMRWAMDIVGPMPASR-QKRFILVMTDYFTKWVEAESYATIRANDVQNFVWKFIICRHG 1611
Query: 1158 IPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAE 1217
+P I++DNG+QF S FC I++ ++ +PQ NGQ E+ N+ IL GL++RL E
Sbjct: 1612 LPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEATNKTILSGLKKRLDE 1671
Query: 1218 AKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFE--EEN 1275
KGAW DEL VLWSY TT +S T +TPF YG++AM P E+ + R + E N
Sbjct: 1672 KKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKNPELN 1731
Query: 1276 QANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLK--WRSGAPGN 1333
M LD L E R+ A R + A +N +V R VGDLVL+ + + A N
Sbjct: 1732 DRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGDLVLRKVFENTAEIN 1791
Query: 1334 --KLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
KL NWEG Y++ K++ G Y L + G +PR++N + L+ +Y
Sbjct: 1792 AGKLGANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSMHLKRYY 1837
>gb|AAD27571.1| polyprotein [Sorghum bicolor] gi|4378066|gb|AAD19359.1| polyprotein
[Sorghum bicolor]
Length = 1877
Score = 957 bits (2475), Expect = 0.0
Identities = 537/1319 (40%), Positives = 760/1319 (56%), Gaps = 38/1319 (2%)
Query: 83 PPVGTINTIAGGFGGGGDTHAARKRHVRAVNSVHEVA----FGFVHPDITISMADFEGIK 138
P +G I IAGG T +K H+R VN+V + IT + D
Sbjct: 572 PTIGMILPIAGGSSMEFQTKKQKKDHLRLVNNVAVQGPVRCTDWSRTPITFTEEDLRLES 631
Query: 139 PHKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGE 198
D +V+ + + + R+L+D GSSADII+ FD++ L+ + L P L+GF G+
Sbjct: 632 YPHTDALVITTNVAGWGVSRILVDSGSSADIIFAGTFDQMKLSRSQLQPSESPLIGFGGK 691
Query: 199 QVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVK 258
Q+ G + L FG E AR V + V+ + YN I GR LN+ A +L +K
Sbjct: 692 QIHALGKVALPVSFGTTENARTEYVTFDVVDLHYPYNAIFGRGFLNKFNAAAHMGYLCMK 751
Query: 259 YPLSSGKV---GRLKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGR 315
P G + G K + + + Y + N+ S + EAS PRG+
Sbjct: 752 IPALHGVITVHGSQKEARNIEKAIYKSFRNINSVDST-------QHEASQPPDMPRGKTN 804
Query: 316 VNRPTPIEETKALKFG----DRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGID 371
+ EETK + D+ + I L+ E+E L +L +N D+FAWS D+ G+
Sbjct: 805 L---ADQEETKCIPLQEAVPDKKVTISATLSREEELELLDVLQKNQDIFAWSAADLQGVS 861
Query: 372 PNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMV 431
+ I H L ++P ++P Q +R++ ++ A + EV +LL A+ IREV YP WLANVV+V
Sbjct: 862 RDIIEHSLDIDPRMRPKKQRQRKMSEERTLAAKAEVQRLLDAKVIREVIYPEWLANVVLV 921
Query: 432 KKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPA 491
K NGK RMC+D+TDLNKAC KDS+PLP ID+ VD A+G + SL+D +SGYHQI +
Sbjct: 922 PKKNGKMRMCIDFTDLNKACVKDSFPLPRIDTSVDKAAGCQRFSLLDCFSGYHQIWLKKE 981
Query: 492 DEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRG 551
DE K +F T YCY MP GLKNAGAT+ R+M +V Q+ RN+ YVDD++V S R
Sbjct: 982 DEGKASFTTPFGTYCYTRMPEGLKNAGATFSRMMGKVLGSQLQRNIIAYVDDVVVMSKRK 1041
Query: 552 LDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKS 611
DH +DL+E F +R ++LNPEKC FGV GK LG++I+S GI NP+K KAI M
Sbjct: 1042 EDHIKDLQETFVNLRSAGLKLNPEKCVFGVSKGKMLGYIISSEGIRANPDKTKAIMSMAE 1101
Query: 612 PSNVKEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSS 671
PSN KEVQRLTGRIAAL+RF+ +S +RS PFFK LR+ EW E EAF +LK +++
Sbjct: 1102 PSNKKEVQRLTGRIAALNRFISRSAERSLPFFKVLREGKT-EWGPEQSEAFRQLKNYIAT 1160
Query: 672 PPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGE----HRIVYFVSHTLQGAEVRYQKI 727
+++ P PL LY A S+ A+S V++ E R VY+VS L GA++ Y +I
Sbjct: 1161 NLLVTVPEPDTPLLLYVAASEHAVSGVLVHETSDTKGTVQRPVYYVSEALSGAKLNYTEI 1220
Query: 728 EKAALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDK 787
EK A AVL +R+L+ YFQS +KV T PL +L+ + SGR+ W+ ELS++ + Y
Sbjct: 1221 EKIAYAVLCASRKLKHYFQSHEIKVPTSQPLGDILRNKEASGRIGKWAAELSQFDITYVP 1280
Query: 788 RGTVGAQSLADFVVELTPDRFER---VDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQ 844
R ++ +Q+LADF+ + TP +D WTL+ DG+ +G+GA L P L L+
Sbjct: 1281 RTSIKSQALADFMADWTPSNKNEEKVIDQPWTLYTDGAWGQAGAGAAAVLIAPSGLKLKF 1340
Query: 845 SLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKY 904
+++ EFKATNN AEYE LI GL A+ ++L+I+TDSQ+V QV+ + +P L +Y
Sbjct: 1341 AIRLEFKATNNIAEYEGLILGLNKAKASGAKTLVIKTDSQVVAGQVEKEYLAHNPELARY 1400
Query: 905 LERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKP---GNNKSVIQETLAYPSIEG 961
L VR L F+ ++Y+PRAEN AD LAK A+ P G ++ +
Sbjct: 1401 LAVVRGLERRFKGFTLQYIPRAENYEADELAKAAANNTPLPEGTFHQIVTTPATETLPKA 1460
Query: 962 ELMSCVNRGRTWMDPIISILAGDPAEVEQCT-KEQQREASHYTLIDGHLYRRGFSTPLLK 1020
+ W I L G ++ T K + A +YT IDG LY++G PLLK
Sbjct: 1461 FRSVLLTESEDWRQAIADCLKGKTTVDDEATAKRMEARARNYTSIDGILYKKGVVQPLLK 1520
Query: 1021 CVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFA 1080
C+S + ++ E+H G+C SHIG R+L+ K LR GFYWPT +D + VK CK CQ F+
Sbjct: 1521 CISQSEGRELLREIHSGMCGSHIGPRALSAKALRQGFYWPTHIRDAEEIVKTCKACQTFS 1580
Query: 1081 DLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKIT 1140
+ P + A WP WG+DLVGP PTA+ KF +VA++YFT+WIEA+PL IT
Sbjct: 1581 PIQSGPSALTQLIPASWPLQRWGMDLVGPMPTAQGGNKFAVVAIEYFTRWIEAKPLTTIT 1640
Query: 1141 SAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQV 1200
S I F+W+ IVCRFG+PR + DNG QF S +EFC +G ++ FASV HP++NG V
Sbjct: 1641 SETIRKFFWQNIVCRFGVPRLLTVDNGKQFDSDNFKEFCHLIGTKIAFASVYHPESNGAV 1700
Query: 1201 ESANRVILRGLRRRLAE-AKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVE 1259
E ANR I + + L KG W++ELP V+WS+NTT T TPF++ YG +AMLP E
Sbjct: 1701 ERANRTIFSAISKTLLNLRKGKWVEELPRVVWSHNTTVSRATGFTPFKLLYGEEAMLPEE 1760
Query: 1260 IDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQV 1319
I + + R+ E++ + L R EA T +Q + +V +D+Q
Sbjct: 1761 IKHQSLRSMKQQLAEDEEYCK---ETLESIRLEAVENITRYQQETKNWRDRKVVRKDIQN 1817
Query: 1320 GDLVLKWRSGAP-GNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
GDLVL+ + P KL P WEGPY ++ +G+++L++L+GR ++N +LR FY
Sbjct: 1818 GDLVLRKKGDHPNAGKLQPKWEGPYTAIQAGRSGSFYLKDLEGRTSTHTWNVDNLRRFY 1876
>ref|XP_476195.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46981311|gb|AAT07629.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|46981243|gb|AAT07561.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1459
Score = 947 bits (2448), Expect = 0.0
Identities = 542/1384 (39%), Positives = 793/1384 (57%), Gaps = 90/1384 (6%)
Query: 5 HRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKKKQE 64
H H T++CR + ++K +R Y G +G G QN +G+K +
Sbjct: 154 HGPCNHATEECRQMASLVEKHVRQ-YDGKYEGVQ---GGQNML----------EGQKVMK 199
Query: 65 SAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHVRAVNSV----HEVAF 120
AI P IN I GG G ++ RK +VR ++ V V
Sbjct: 200 IEAIEEA------------PKRVINAITGGSSLGVESKRQRKAYVRQIHHVGTSYQSVPP 247
Query: 121 GFVHPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGL 180
+ + I+ D EGI DP+V+ + + ++RVL+D GSSAD+++ DAF K+ +
Sbjct: 248 AYSNTVISFGPEDAEGILFPHQDPLVISVEIAQCEVQRVLVDGGSSADVLFYDAFKKMQI 307
Query: 181 TDNDLTPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGR 240
++ LT L GF G QV G I L +FG E R +V + V+ + YN ++GR
Sbjct: 308 PEDRLTHAGIPLQGFGGHQVHTIGKISLQVVFGGGENKRREEVVFDVVDMPYQYNAVLGR 367
Query: 241 NTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNNCLNLYGKKSALVGHRCYE 300
+T+N A+I ++ +K P G + ++ Q AR+ L G + Y
Sbjct: 368 STINIFEAIIHHNYICMKLPGPKGVIS-VRGGQLAARK-----FELQGTPNM---KGVYI 418
Query: 301 IEASDENLDPRGEGRVNRPTPIEETKALKFGDRTLKIGTRLTEEQETRLTKLLGENLDLF 360
IE +GE N+ PI E K T K+ E +E
Sbjct: 419 IEQK------QGEYNKNQK-PIPEGK-------TKKVVLDKNEPEE-------------- 450
Query: 361 AWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVK 420
+ G++ + I H LA+ P KP Q RR+ D+ +A + E++KLL A+ IREV
Sbjct: 451 ------LEGVERSLIEHNLAIKPEHKPKKQKLRRMSIDRQQAAKAELEKLLKAKVIREVL 504
Query: 421 YPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAY 480
+P WLAN V+VK G LNKACPKD +PLP ID LVD +G EL+S +DAY
Sbjct: 505 HPEWLANPVLVKMQMGSGEY------LNKACPKDDFPLPRIDQLVDATAGCELMSFLDAY 558
Query: 481 SGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVY 540
SGYHQ+ M DE+K + +T YCY MPFGLKNAGAT+ RL+ +V A Q+GRN+E Y
Sbjct: 559 SGYHQVFMVKEDEEKPSLITPFGTYCYIRMPFGLKNAGATFARLICKVLANQLGRNVEAY 618
Query: 541 VDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINP 600
VDD++VKS + H +DL+E F +RK S++LNPEKC FGV+ GK LGF+++ RGIE NP
Sbjct: 619 VDDIVVKSKKAFTHGKDLQETFENLRKFSVKLNPEKCMFGVRVGKLLGFLVSKRGIEANP 678
Query: 601 EKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEE 660
+K AIQQM+ P N +E+QRLTGR+A+LSRFL KS +R PFFK LR F+W EC++
Sbjct: 679 DKIAAIQQMEPPKNTREMQRLTGRMASLSRFLSKSAERGLPFFKTLRAVNNFKWNEECQK 738
Query: 661 AFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGA 720
AF LK+ L + P LS P +G PL LY A + +S+V++QE + VYFVS LQG
Sbjct: 739 AFDDLKDYLHNMPTLSSPQKGEPLLLYVAATPVTVSAVLVQEQGNKQMPVYFVSEALQGP 798
Query: 721 EVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSE 780
+ RY ++EK A+++ +R+LR YF S + + + P+ +VL +++GR+ W++EL
Sbjct: 799 KTRYIEVEKMIYAIVMASRKLRHYFLSHDITIPSTYPIGEVLSNKEIAGRIAKWAMELLP 858
Query: 781 YGLQYDKRGTVGAQSLADFVVELTPDRFERVDTQ---WTLFVDGSSNSSGSGAGVALEGP 837
+ L+Y R + +Q LA FVVE TP E+ + + WT+F DG+ N++G+GA ++ P
Sbjct: 859 FDLKYTSRTAIKSQVLAKFVVEWTPTELEKKEEEEKPWTVFSDGACNATGAGAAAVVKTP 918
Query: 838 GELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVK 897
+ L+ S + F +TNN AEYE ++ ++ +R + R L+I+TDS+LV TF+ +
Sbjct: 919 MKQTLKYSARLNFPSTNNTAEYEGVLLAMRKSRALGARRLIIKTDSKLVAGHFSKTFEAR 978
Query: 898 DPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYP 957
+ + KYLE R F + V+ + R EN AD L K A+ +P N E L +P
Sbjct: 979 EEIMTKYLEEARLNERHFLGITVKAITREENGEADELTKAAAAGQPLENS--FFEILEHP 1036
Query: 958 SIEGELMSCV-NRGRTWMDPIISILAGDP-AEVEQCTKEQQREASHYTLIDGHLYRRGFS 1015
S + + + C+ G W DPI+ L + E E+ + Q A Y +IDG LY+ G +
Sbjct: 1037 SYDKKEVICIQEEGLDWRDPILKFLVSNKLPEKEEEARRVQLMARKYKVIDGQLYKSGVT 1096
Query: 1016 TPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKE 1075
PLLKCV+ E+ ++ E+HEG+C +H RS+A KV+R G YWPT+ KD ++K C+
Sbjct: 1097 APLLKCVTKEEGMQMVVEIHEGICGAHQAPRSVASKVIRQGIYWPTIMKDTEQYIKTCRA 1156
Query: 1076 CQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEP 1135
CQ SKAPPKEL + WPF WG+D+VGP P A+ ++F++VA++YF++WIEAE
Sbjct: 1157 CQKVGSSSKAPPKELQPIPPVWPFYRWGIDIVGPLPRAKGDLRFVIVAIEYFSRWIEAEA 1216
Query: 1136 LAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQ 1195
+A+ITSA + F WK I+CRFGIP+ IV DNG QF S + ++ C+ + +++ FASV HPQ
Sbjct: 1217 VARITSAAMQKFVWKNIICRFGIPKEIVCDNGKQFESEKFKDLCQGLHLKINFASVGHPQ 1276
Query: 1196 TNGQVESANRVILRGLRRRL-AEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDA 1254
TNG VE AN ++ +++RL +KG W ++L +VLW+ TT T TPFR+ YG +
Sbjct: 1277 TNGAVERANGKVVEAIKKRLEGSSKGKWPEDLLSVLWALRTTVVWATGMTPFRLVYGDEP 1336
Query: 1255 MLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRV 1314
M P E+ R F +E++ V L++L E R EA + A + ++N +VR
Sbjct: 1337 MTPSEVG--VNSPRVIFYQEDEQGRKVSLEMLDEIRVEALQKMEAYTEGTRKQYNKKVRP 1394
Query: 1315 RDMQVGDLVL-KWRSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSL 1373
R+++ GDLVL K + KL WEGP+ + K + GAY L LDG L ++N +SL
Sbjct: 1395 RNIEEGDLVLKKVLNEVAVGKLESKWEGPFIVRKKMEIGAYKLAHLDGEELNHTWNAISL 1454
Query: 1374 RYFY 1377
+ FY
Sbjct: 1455 KKFY 1458
>gb|AAL76007.1| prpol [Zea mays]
Length = 1317
Score = 929 bits (2402), Expect = 0.0
Identities = 508/1306 (38%), Positives = 770/1306 (58%), Gaps = 35/1306 (2%)
Query: 96 GGGGDTHAARKRHVRAVNSVHEVAFG-------FVHPDITISMADFEGIKPHKDDPIVVQ 148
GG A +K+ A V V + H IT S D + +D +V+
Sbjct: 22 GGSSSEPANKKQKKEAQRRVQHVGVQGPFIKSRWSHIPITFSQEDLQLKDYPHNDAMVIS 81
Query: 149 LRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMVRGYIDL 208
+ F + VL+D GS+ADII+ AF ++ ++ + L GF G Q++ G I +
Sbjct: 82 CVIKGFLVHNVLVDTGSAADIIFAKAFRQMQEPEDKIHDATHPLCGFGGRQIVALGKITM 141
Query: 209 DTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGR 268
FG R +V + ++ + YN IIGR TLN A++ A+L +K P G +
Sbjct: 142 SVTFGFINNTRTEQVVFDIVDMEYPYNAIIGRGTLNAFEAILHPAYLCMKIPSDQGPIA- 200
Query: 269 LKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKAL 328
+ Q+ AR N + + C + + E ++P P+ + +
Sbjct: 201 IHGSQEAARRAEGNWTDSKAIHNIDGAEACEQYKFRREKA-----ASADQPKPMLLCEDI 255
Query: 329 KFGDRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPV 388
++ + +G++L+EEQE L + L N D+FAWS D+ G++ + I H L ++PS +P
Sbjct: 256 --AEQKVLLGSQLSEEQEKTLIRFLFNNKDVFAWSANDLCGVNRDVIEHSLNVDPSFRPR 313
Query: 389 SQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLN 448
Q R++ DK + + EV +LL+A IREVKYP WLAN VMVKKANGKWRMC+D+TDLN
Sbjct: 314 KQRLRKMSDDKAEGARNEVKRLLSAGVIREVKYPEWLANTVMVKKANGKWRMCIDFTDLN 373
Query: 449 KACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYR 508
KACPKD +PLP IDSLVD A+ +EL+SL+D YSGYHQI M DE KT+F+T YCY
Sbjct: 374 KACPKDEFPLPRIDSLVDAAASSELMSLLDCYSGYHQIWMKKEDEPKTSFITPSGTYCYL 433
Query: 509 TMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKH 568
MP GLKNAG ++ R+ +V Q+GRN+ YVDD+IVKS + +H DL+E F R+
Sbjct: 434 RMPEGLKNAGGSFSRMTAKVLQSQIGRNVLTYVDDIIVKSTKQENHIADLQETFASFRQA 493
Query: 569 SMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAAL 628
++LNPEKC FGV+ GKFLG +++++GIE NP K +AI +M+ P+ K QRLTGR+A+L
Sbjct: 494 GLKLNPEKCVFGVKKGKFLGCLVSTKGIEANPSKIEAILRMEPPTTKKGAQRLTGRLASL 553
Query: 629 SRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYF 688
+RF+ +S +R+ PFF+ L+ VF+W ++AF LK+ L L+ P+ G PL LY
Sbjct: 554 NRFISRSAERNLPFFEVLKSAEVFQWGPIQQKAFEELKQYLIDLTTLTPPMSGAPLLLYV 613
Query: 689 AVSDSALSSVMLQE-IDGE---HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPY 744
A S SA+S+ ++QE +DG+ +YFVS L ++ Y ++EK AVL+ +R+LR Y
Sbjct: 614 AASHSAVSAALVQEKLDGQVKRQAPIYFVSEVLSLSKKNYTELEKVLYAVLMASRKLRHY 673
Query: 745 FQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT 804
FQ++ + V + PL+ +++ + +GR+ W+ EL+E+ ++Y R ++ +Q+LADF+ + T
Sbjct: 674 FQAYNIIVPSSQPLKDIMRNREATGRIGKWAAELNEFCIEYVHRSSIQSQALADFIADWT 733
Query: 805 P----DRFERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYE 860
P + + + WT+F DGS + G+GA L P ++ + + K +F TNN AEYE
Sbjct: 734 PGAQEEETNKDNEAWTVFCDGSWGTFGAGAAAVLVSPSKVKICYAAKLDFNCTNNIAEYE 793
Query: 861 ALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVV 920
AL+ GL+ + + IR +++TDSQ+V + + + KDP L KYL+ VR + F+ V
Sbjct: 794 ALVLGLRKLKAMGIRRAILKTDSQVVSGHIDKSCKAKDPKLEKYLDMVRRIEASFEGFSV 853
Query: 921 EYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSCVN----RGRTWMDP 976
+ +PR +N+ AD LAK A+ P V ET+ PS+E + +N W
Sbjct: 854 KNIPRGQNEHADLLAKSAAQGLP-LPSDVFFETIKAPSVELLERAVLNISPVFSEDWRTE 912
Query: 977 IISILAGD-PAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVH 1035
IIS L G ++ E K + A Y +I+G LY+ G PLLKC+S + +M E+H
Sbjct: 913 IISYLQGKFLSDDEAYNKRIEARARPYVMIEGELYKHGVCAPLLKCLSRTEGIELMKEIH 972
Query: 1036 EGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSA 1095
G+C SHIG R L KV R GFYWP D + V++C+ CQ A K P +
Sbjct: 973 AGLCGSHIGSRPLLGKVFRQGFYWPKAASDAAELVQKCEGCQKCARDQKQPSSLTQLIQP 1032
Query: 1096 PWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCR 1155
WP WG+DL+GP P A+ +++++VAV+YF+KWIEA+PLA ITSA + F+W+ IVCR
Sbjct: 1033 TWPLQRWGLDLLGPLPPAQGNLRYVVVAVEYFSKWIEAKPLATITSATVQKFFWQNIVCR 1092
Query: 1156 FGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRL 1215
FG+P+AI DNGTQF S R+FC ++G ++ FASV HP++NG VE AN +I+ G+ + +
Sbjct: 1093 FGVPKAITVDNGTQFDSEAFRDFCDQIGTKIHFASVRHPESNGLVERANGIIMTGIMKLI 1152
Query: 1216 -AEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEE 1274
+ +G W D+L V+WS+NTT +T TPF++ +G +A+ P + + R E +
Sbjct: 1153 FNQPRGKWPDQLIKVVWSHNTTTSRSTGFTPFKLLFGDEAITPEKAKAGSIRIVASAESD 1212
Query: 1275 NQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSR-VRVRDMQVGDLVLKWRSGAPG- 1332
++A ++E D L R +A + Q K+ R VR+++++ G LVL+ R P
Sbjct: 1213 SEAAYSIEKDALEGIRLQA-VENINKYQAETVKWRDRKVRLKNIEPGHLVLR-RVANPET 1270
Query: 1333 -NKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
KL W+GP+ + G+Y L++++G +PRS+N LR +Y
Sbjct: 1271 VGKLQLKWDGPFLVASSSRPGSYRLKDMNGSDIPRSWNADELRRYY 1316
>gb|AAD11615.1| prpol [Zea mays] gi|7489805|pir||T14595 polyprotein - maize
retrotransposon Cinful-1
Length = 1317
Score = 917 bits (2371), Expect = 0.0
Identities = 511/1306 (39%), Positives = 761/1306 (58%), Gaps = 35/1306 (2%)
Query: 96 GGGGDTHAARKRHVRAVNSVHEVAFG-------FVHPDITISMADFEGIKPHKDDPIVVQ 148
GG A +K+ A V V + H IT S D + +D +V+
Sbjct: 22 GGSSSEPANKKQKKEAQRRVQHVGVQGPFIKSRWSHIPITFSQEDLQLKDYPHNDAMVIS 81
Query: 149 LRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMVRGYIDL 208
+ F + VL+D GS+ADII+ AF ++ ++ + L GF G Q++ G I +
Sbjct: 82 CVIKGFLVHNVLVDTGSAADIIFAKAFRQMQELEDKIHDATHPLCGFGGRQIVALGKITM 141
Query: 209 DTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGR 268
FG R +V + ++ + YN IIGR TLN A++ A+L +K P G +
Sbjct: 142 SVTFGFINNTRTEQVVFDIVDMEYPYNAIIGRGTLNAFEAILHPAYLCMKMPSDQGPIA- 200
Query: 269 LKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKAL 328
+ Q+ AR N + A+ H EA E R E + P
Sbjct: 201 IHGSQEAARRAEGN----WTDSKAI--HNIDGAEAC-EXYKFRREKAASADQPKPMLLCE 253
Query: 329 KFGDRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPV 388
++ + +G++L+EEQE L + L N D+FAWS D+ G++ + I H L ++PS +P
Sbjct: 254 DIAEQKVLLGSQLSEEQEKTLIRFLFNNKDVFAWSANDLCGVNRDVIEHSLNVDPSFRPR 313
Query: 389 SQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLN 448
Q R++ DK + + EV +LL+A IREVKYP WLAN VMVKKANGKWRMC+D+TDLN
Sbjct: 314 KQRLRKMSDDKAEGARNEVKRLLSAGVIREVKYPEWLANTVMVKKANGKWRMCIDFTDLN 373
Query: 449 KACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYR 508
KACPKD +PLP IDSLVD + +EL+SL+D YSGYHQI M DE KT+F+T YCY
Sbjct: 374 KACPKDEFPLPRIDSLVDATASSELMSLLDCYSGYHQIWMKREDEPKTSFITPSGTYCYL 433
Query: 509 TMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKH 568
MP GLKNAG ++ R+ +V Q+GRN+ YVDD+IVKS + +H DL+E F R+
Sbjct: 434 RMPEGLKNAGGSFSRMTAKVLQSQIGRNVLTYVDDIIVKSTKQENHIADLQETFASFRQA 493
Query: 569 SMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAAL 628
++LNPEKC FGV+ GKFLG +++++GIE NP K +AI +M+ P+ K QRLTGR+A+L
Sbjct: 494 GLKLNPEKCVFGVKKGKFLGCLVSTKGIEANPSKIEAILRMEPPTTKKGAQRLTGRLASL 553
Query: 629 SRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYF 688
+RF+ +S +R+ PFF+ L+ VF+W ++AF LK+ L L+ P+ G PL LY
Sbjct: 554 NRFISRSAERNLPFFEVLKSAEVFQWGPIQQKAFEELKQYLIDLTALTPPMPGAPLLLYV 613
Query: 689 AVSDSALSSVMLQE-IDGEHR---IVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPY 744
A S SA+S+ ++QE +DG+ R VYFVS L ++ Y ++EK VL+ +R+LR Y
Sbjct: 614 AASHSAVSAALVQEKLDGQTRKQVPVYFVSEVLSISKKNYTELEKVLYVVLMASRKLRHY 673
Query: 745 FQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT 804
FQ++ + V + PL+ +++ + +GR+ W+ EL+E+ ++Y R ++ +Q+LADF+ + T
Sbjct: 674 FQAYNIIVPSSQPLKDIMRNREATGRIGKWAAELNEFCIEYVHRSSIQSQALADFIADWT 733
Query: 805 PDRFE---RVDTQ-WTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYE 860
P E DT+ WT+F DGS + G+GA L P ++ K +F TNN AEYE
Sbjct: 734 PGAQEEEANKDTEAWTVFCDGSWGTFGAGAAAVLVSPSKVKTCYVAKLDFSCTNNIAEYE 793
Query: 861 ALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVV 920
AL+ GL+ + + IR +++TDSQ+V + + + KDP L KYL+ VR + F+ V
Sbjct: 794 ALLLGLRKLKAMGIRRAVLKTDSQVVSGHIDKSCKAKDPKLEKYLDMVRRVEASFEGFSV 853
Query: 921 EYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSCVN----RGRTWMDP 976
+ +PR +N+ AD LAK A+ P V ET+ PS+E + +N W
Sbjct: 854 KNIPRGQNEHADLLAKSAAQGLP-LPSDVFFETIKAPSVELLERAVLNISPVYSEDWRTE 912
Query: 977 IISILAGD-PAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVH 1035
IIS L G ++ E + + A Y +I+G LY+ G PLLKC+S + +M E+H
Sbjct: 913 IISYLQGKFLSDDEAYNRRIEARARPYVMIEGELYKHGVCAPLLKCLSRTEGIELMKEIH 972
Query: 1036 EGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSA 1095
G+C SHIG R L K+ R GFYWP D + V++C+ CQ A K P +
Sbjct: 973 AGLCGSHIGSRPLLGKIFRQGFYWPKAASDAAELVQKCEGCQKCARDQKQPSSLTQLIQP 1032
Query: 1096 PWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCR 1155
WP WG+DL+GP P A+ +++++VAV+YF+KWIEA+PLA ITSA + F+W+ IVCR
Sbjct: 1033 IWPLQRWGLDLLGPLPPAQGNLRYVVVAVEYFSKWIEAKPLATITSATVQKFFWQNIVCR 1092
Query: 1156 FGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRL 1215
FG+P+AI DNGTQF S R+FC ++G ++ FASV HP++NG VE AN +I+ G+ + +
Sbjct: 1093 FGVPKAITVDNGTQFDSEAFRDFCDQIGTKIHFASVRHPESNGLVERANGIIMTGIMKSI 1152
Query: 1216 -AEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEE 1274
+ +G W D+L V+WS+NTT +T TPF++ +G +A+ P E + R
Sbjct: 1153 FNQPRGKWPDQLTKVVWSHNTTTSRSTGFTPFKLLFGDEAITPEEAKTGSIRVVASAASG 1212
Query: 1275 NQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSR-VRVRDMQVGDLVLKWRSGAPG- 1332
++A+ +VE D L R +A + Q K+ R VR+++++ + R P
Sbjct: 1213 SEADYSVEKDALEGIRLQA-VENINKYQAETIKWRDRKVRLKNIEPDTWCFR-RVANPET 1270
Query: 1333 -NKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
KL W+GP+ + G+Y L+++DG +PRS+N LR +Y
Sbjct: 1271 VGKLQLKWDGPFLVASSSRPGSYRLKDMDGNDIPRSWNADELRRYY 1316
>dbj|BAD18986.1| GAG-POL precursor [Vitis vinifera]
Length = 1027
Score = 912 bits (2356), Expect = 0.0
Identities = 460/1031 (44%), Positives = 656/1031 (63%), Gaps = 25/1031 (2%)
Query: 367 MPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLA 426
M GI P+ HRL + + +PV Q RR D+ + ++ E+DKLL A FIREV YP WLA
Sbjct: 1 MKGIHPSITSHRLNVVSTARPVRQRIRRFHPDRQRVIRNEIDKLLEAGFIREVSYPDWLA 60
Query: 427 NVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQI 486
NVV+V K GKWR+CVDYT+LN ACPKDS+PLP ID +VD SG +LS +DA+SGYHQI
Sbjct: 61 NVVVVPKKEGKWRVCVDYTNLNNACPKDSFPLPRIDQIVDSTSGQGMLSFLDAFSGYHQI 120
Query: 487 RMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIV 546
M P DE+K AF+T YCY+ MPFGLKNAGATYQRLM ++F +G ++EVY+DD++V
Sbjct: 121 PMSPDDEEKIAFITPHDLYCYKVMPFGLKNAGATYQRLMTKIFKPLIGHSVEVYIDDIVV 180
Query: 547 KSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAI 606
KS H L+E F +R++ M+LNP KC+FGV KFLGFM++ RGIE++P++ KA+
Sbjct: 181 KSKTREQHILHLQEVFYLLRRYGMKLNPSKCAFGVSARKFLGFMVSQRGIEVSPDQVKAV 240
Query: 607 QQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLK 666
+ P N KE+QRLTG++ AL RF+ + D PFF +RK WT C+ A R+K
Sbjct: 241 METPPPRNKKELQRLTGKLVALGRFIARFIDELRPFFLAIRKAGTHGWTDNCQNALERIK 300
Query: 667 ELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQ-EIDGEHRIVYFVSHTLQGAEVRYQ 725
L PPILS PI L++Y AVS+ A+S+V+ + E + +Y+VS L E RY
Sbjct: 301 HYLMQPPILSSPIPKEKLYMYLAVSEWAISAVLFRCPSPKEQKPIYYVSRALADVETRYS 360
Query: 726 KIEKAALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQY 785
K+E +LA+ A++LRPYFQ+ PV V TD PLR +L KPDL+GR++ W++ELSE+G+++
Sbjct: 361 KMELISLALRSAAQKLRPYFQAHPVIVLTDQPLRNILHKPDLTGRMLQWAIELSEFGIEF 420
Query: 786 DKRGTVGAQSLADFVVELT--PDRFE--RVDTQWTLFVDGSSNSSGSGAGVALEGPGELV 841
R ++ Q +ADFV+E + P + E R WTL VDG+S SSGSG G+ L+ P
Sbjct: 421 QPRLSMKGQVMADFVLEYSRKPGQHEGSRKKEWWTLRVDGASRSSGSGVGLLLQSPTGEH 480
Query: 842 LEQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNL 901
LEQ+++ F A+NN+AEYEA+++GL LA + + L I +DSQLV V+ ++ KD +
Sbjct: 481 LEQAIRLGFSASNNEAEYEAILSGLDLALALSVSKLRIFSDSQLVVKHVQEEYEAKDARM 540
Query: 902 IKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEG 961
+YL +VR + F E +E + RA+N+RADALA +A++ + PS+
Sbjct: 541 ARYLAKVRNTLQQFTEWTIEKIKRADNRRADALAGIAASLSIKEAILLPIHVQTNPSV-S 599
Query: 962 ELMSCVNR------GRTWMDPI-----ISILAGDPAEVEQCTKEQQREASHYTLIDGHLY 1010
E+ C + WM+ I L GDP + + + +A+ +TLI GHLY
Sbjct: 600 EISICSTTEAPQADDQEWMNDITEYIRTGTLPGDPKQAHKV----RVQAARFTLIGGHLY 655
Query: 1011 RRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFV 1070
+R F+ P L+C+ + + +++E+HEG+ +H GGRSLA + G+YWPT++K+ +V
Sbjct: 656 KRSFTGPYLRCLGHSEAQYVLAELHEGIYGNHSGGRSLAHRAHSQGYYWPTMKKEAAAYV 715
Query: 1071 KQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKW 1130
K+C +CQ +A + P L ++S PWPFA WG+D+V P PTA AQ KF+LVA DYF+KW
Sbjct: 716 KRCDKCQRYAPIPHMPSTTLKSISGPWPFAQWGMDIVRPLPTAPAQKKFLLVATDYFSKW 775
Query: 1131 IEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFAS 1190
+EAE A + F WK I+CRFGIP+ I++DNG QF S R FC E+ I+ +++
Sbjct: 776 VEAEAYASTKDKDVTKFVWKNIICRFGIPQTIIADNGPQFDSIAFRNFCSELNIRNSYST 835
Query: 1191 VEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTY 1250
+PQ+NGQ E+ N+ ++ L++RL +AKG W++ELP VLW+Y TT T TPF + Y
Sbjct: 836 PRYPQSNGQAEATNKTLITALKKRLEQAKGKWVEELPGVLWAYRTTPGRPTGNTPFALAY 895
Query: 1251 GVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNS 1310
G+DA++P+EI T T + + + LD E R+ A IR +QR +A +N
Sbjct: 896 GMDAVIPIEIGLPTIWTNAAKQSDANMQLGRNLDWTDEVRESASIRMADYQQRASAHYNR 955
Query: 1311 RVRVRDMQVGDLVLKW----RSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPR 1366
+VR R ++ G LVL+ + K NWEGPY + K NGAYHL++LDG L R
Sbjct: 956 KVRPRSLKNGTLVLRKFFENTTEVGAGKFQANWEGPYIVSKASDNGAYHLQKLDGTPLLR 1015
Query: 1367 SFNGLSLRYFY 1377
+N +L+ +Y
Sbjct: 1016 PWNVSNLKQYY 1026
>ref|XP_474675.1| OSJNBa0027O01.4 [Oryza sativa (japonica cultivar-group)]
gi|38346188|emb|CAD39529.2| OSJNBa0027O01.4 [Oryza sativa
(japonica cultivar-group)]
Length = 2013
Score = 857 bits (2215), Expect = 0.0
Identities = 523/1420 (36%), Positives = 764/1420 (52%), Gaps = 108/1420 (7%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + R D+ ++ R+++ DT
Sbjct: 657 CPHHPNSNHVAKDCFVYKQFADQYTKTA---------RKNSDEEQSTSRKKDDGDTPAG- 706
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
F H +N I FGG + RK+ + R +N+V
Sbjct: 707 ---------------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 744
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 745 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 795
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 796 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 852
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 853 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTCDKE 911
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGD---RTLKIGT 339
++ + + A+ + EG V TK K G+ +T KI
Sbjct: 912 SCDMAQTREMASAREDIRLAAATAS-----EGEV------PATKISKSGESEAKTKKIPL 960
Query: 340 RLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDK 399
++ +T N D+FAW DMPGI I H L + KP+ Q RR D+
Sbjct: 961 DPSDPTKT------ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDR 1014
Query: 400 GKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLP 459
A+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD + LP
Sbjct: 1015 KDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLP 1074
Query: 460 SIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGA 519
ID +VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGA
Sbjct: 1075 RIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGA 1134
Query: 520 TYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSF 579
TYQR++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+F
Sbjct: 1135 TYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTF 1194
Query: 580 GVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRS 639
GV GK LGFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G+R
Sbjct: 1195 GVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERG 1254
Query: 640 FPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVM 699
PFFK L+K F+W E ++AF K+LL+ PP+L+ P PL LY + + +S+V+
Sbjct: 1255 MPFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVL 1314
Query: 700 LQEIDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVR 753
+ E + E R +YFVS L ++ RY +++K +L+T R+L YFQ V V
Sbjct: 1315 VVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1374
Query: 754 TDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFER 810
T PL +L +++GR+ W++EL + + R ++ +Q+LADFV E T D
Sbjct: 1375 TSFPLGDILHNREVNGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAE 1434
Query: 811 VDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAR 870
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A
Sbjct: 1435 NMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1494
Query: 871 EVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQR 930
+ I+ L++R DSQLV NQV + D N++ Y + VR L F + + +V R N+
Sbjct: 1495 SLGIKRLIVRGDSQLVVNQVMKEWSYLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEA 1554
Query: 931 ADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSCVNRGR-------TWMDPIISILAG 983
AD LA S R+ + V E L P++ + + V W +P+I L
Sbjct: 1555 ADRLANFGSKREAAPS-DVFVEHLYTPTVPHKDTTQVAGTHDAAMVEVDWREPLIRFLTS 1613
Query: 984 DPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASH 1042
++ E+ R + Y L + LY++ S L +CVS E+ ++ ++H G+C +H
Sbjct: 1614 QELPQDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNH 1673
Query: 1043 IGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW 1102
R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+W
Sbjct: 1674 AAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVW 1733
Query: 1103 GVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAI 1162
G+D+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I
Sbjct: 1734 GLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRI 1792
Query: 1163 VSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEA 1218
++DNGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL
Sbjct: 1793 ITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPY 1852
Query: 1219 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQAN 1278
G W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + +
Sbjct: 1853 AGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEED 1912
Query: 1279 MAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTP 1337
+L L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P
Sbjct: 1913 RVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSP 1972
Query: 1338 NWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
WEGP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1973 LWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2012
>ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultivar-group)]
gi|38347303|emb|CAE02298.2| OSJNBa0042F21.5 [Oryza sativa
(japonica cultivar-group)]
Length = 1950
Score = 856 bits (2212), Expect = 0.0
Identities = 522/1420 (36%), Positives = 761/1420 (52%), Gaps = 108/1420 (7%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + R D+ ++ R+++ DT
Sbjct: 594 CPHHPNSNHVAKDCFVYKQFADQYTKTA---------RKNSDEEQSTSRKKDDGDTPAG- 643
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
F H +N I FGG + RK+ + R +N+V
Sbjct: 644 ---------------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 681
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 682 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 732
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 733 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 789
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 790 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTCDKE 848
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGD---RTLKIGT 339
++ + + A+ + EG I TK K G+ +T KI
Sbjct: 849 SCDMAQTREMASAREDIRLAAATAS-----EGE------IPATKTSKSGESEAKTKKIPL 897
Query: 340 RLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDK 399
++ +T N D+FAW DMPGI I H L + KP+ Q RR D+
Sbjct: 898 DPSDPTKT------ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDR 951
Query: 400 GKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLP 459
A+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD + LP
Sbjct: 952 KDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLP 1011
Query: 460 SIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGA 519
ID +VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGA
Sbjct: 1012 RIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGA 1071
Query: 520 TYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSF 579
TYQR++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+F
Sbjct: 1072 TYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTF 1131
Query: 580 GVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRS 639
GV GK LGFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G+R
Sbjct: 1132 GVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERG 1191
Query: 640 FPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVM 699
PFFK L+K F+W E ++AF K+LL+ PPIL+ P PL LY + + +S+V+
Sbjct: 1192 MPFFKLLKKTDNFQWGPEAQKAFEDFKKLLTEPPILASPHPQEPLLLYVSATSQVVSTVL 1251
Query: 700 LQEIDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVR 753
+ E + E R +YFVS L ++ RY +++K +L+T R+L YFQ V V
Sbjct: 1252 VVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1311
Query: 754 TDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFER 810
T PL +L + +GR+ W++EL + + R ++ +Q+LADFV E T D
Sbjct: 1312 TSFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAE 1371
Query: 811 VDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAR 870
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A
Sbjct: 1372 NMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1431
Query: 871 EVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQR 930
+ I+ L++R DSQLV NQV + D N++ Y + VR L F + + +V R N+
Sbjct: 1432 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEA 1491
Query: 931 ADALAKLASTRKPGNNKSVIQETLAYPSI-------EGELMSCVNRGRTWMDPIISILAG 983
AD LA S R+ + V E L P++ + + W +P I L
Sbjct: 1492 ADRLANFGSKREMAPS-DVFVEHLYTPTVPHKDTTQDADTHDVALVEADWREPFIRFLTS 1550
Query: 984 DPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASH 1042
++ E+ R + Y + + LY++ S L +CVS E+ ++ ++H G+C +H
Sbjct: 1551 QELPQDKDEAERISRRSKLYAMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNH 1610
Query: 1043 IGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW 1102
R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+W
Sbjct: 1611 AAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVW 1670
Query: 1103 GVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAI 1162
G+D+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I
Sbjct: 1671 GLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRI 1729
Query: 1163 VSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEA 1218
++DNGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL
Sbjct: 1730 ITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPY 1789
Query: 1219 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQAN 1278
G W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + +
Sbjct: 1790 AGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEED 1849
Query: 1279 MAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTP 1337
+L L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P
Sbjct: 1850 RVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSP 1909
Query: 1338 NWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
WEGP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1910 LWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 1949
>ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultivar-group)]
gi|38344177|emb|CAE03508.2| OSJNBa0053K19.16 [Oryza
sativa (japonica cultivar-group)]
Length = 2010
Score = 856 bits (2211), Expect = 0.0
Identities = 518/1417 (36%), Positives = 763/1417 (53%), Gaps = 102/1417 (7%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + R D+ ++ R+++ DT
Sbjct: 654 CSHHPNSNHVAKDCFVYKQFADQYTKTA---------RKNSDEEQSTSRKKDDGDTPAG- 703
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
F H +N I FGG + RK+ + R +N+V
Sbjct: 704 ---------------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 741
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 742 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 792
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 793 LNILFAKTLDDMQIPHSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 849
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 850 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTCDKE 908
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGDRTLKIGTRLT 342
++ + + A+ + GE + + E++A +T KI +
Sbjct: 909 SCDMAQTREMASAREDIRLAAATAS---EGEEPATKISKSGESEA-----KTKKIPLDPS 960
Query: 343 EEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKA 402
+ +T N D+FAW DMPGI I H L + KP+ Q RR D+ A
Sbjct: 961 DPAKT------ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDA 1014
Query: 403 VQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSID 462
+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD + LP ID
Sbjct: 1015 IKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRID 1074
Query: 463 SLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQ 522
+VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGATYQ
Sbjct: 1075 QVVDSTAGRELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQ 1134
Query: 523 RLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQ 582
R++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+FGV
Sbjct: 1135 RMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVP 1194
Query: 583 GGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPF 642
GK +GFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G+R PF
Sbjct: 1195 SGKLVGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPF 1254
Query: 643 FKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQE 702
FK L+K F+W E ++AF K+LL+ PP+L+ P PL LY + + +S+V++ E
Sbjct: 1255 FKLLKKTDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVE 1314
Query: 703 IDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDL 756
+ E R +YFVS L ++ RY +++K +L+T R+L YFQ V V T
Sbjct: 1315 REEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF 1374
Query: 757 PLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFERVDT 813
PL +L + +GR+ W++EL + + R ++ +Q+LADFV E T D
Sbjct: 1375 PLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAENME 1434
Query: 814 QWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVK 873
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A +
Sbjct: 1435 HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLG 1494
Query: 874 IRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADA 933
I+ L++R DSQLV NQV + D N++ Y + VR L F + + +V R N+ AD
Sbjct: 1495 IKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADR 1554
Query: 934 LAKLASTRKPGNNKSVIQETLAYPSIEGELMSCVNRGR-------TWMDPIISILAGDPA 986
LA S R+ + V E L P++ + + V W +P+I L
Sbjct: 1555 LANFGSKREVAPS-DVFVEHLYTPTVPHKDTTQVAGTHDVAMVETDWREPLIRFLTSQEL 1613
Query: 987 EVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGG 1045
++ E+ R + Y + + LY++ S L +CVS E+ ++ ++H G+C +H
Sbjct: 1614 PQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLQDIHSGICGNHAAA 1673
Query: 1046 RSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVD 1105
R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+WG+D
Sbjct: 1674 RTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLD 1733
Query: 1106 LVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSD 1165
+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I++D
Sbjct: 1734 MVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITD 1792
Query: 1166 NGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEAKGA 1221
NGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL G
Sbjct: 1793 NGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGK 1852
Query: 1222 WLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAV 1281
W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + +
Sbjct: 1853 WVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVD 1912
Query: 1282 ELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTPNWE 1340
+L L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P WE
Sbjct: 1913 DLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWE 1972
Query: 1341 GPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
GP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1973 GPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2009
>gb|AAP52499.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531820|ref|NP_920212.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128689|gb|AAM92802.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 2017
Score = 855 bits (2210), Expect = 0.0
Identities = 522/1420 (36%), Positives = 763/1420 (52%), Gaps = 108/1420 (7%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + R D+ ++ R+++ DT
Sbjct: 661 CPHHPNSNHVAKDCFVYKQFADQYTKTA---------RKNSDEEQSTSRKKDDGDTLAG- 710
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
F H +N I FGG + RK+ + R +N+V
Sbjct: 711 ---------------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 748
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 749 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 799
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 800 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 856
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 857 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTCDKE 915
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGD---RTLKIGT 339
++ + + A+ + EG V TK K G+ +T KI
Sbjct: 916 SCDMAQTREMASAREDIRLAAATAS-----EGEV------PATKISKSGESEAKTKKIPL 964
Query: 340 RLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDK 399
++ +T N D+FAW DMPGI I H L + KP+ Q RR D+
Sbjct: 965 DPSDPTKT------ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDR 1018
Query: 400 GKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLP 459
A+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD + LP
Sbjct: 1019 KDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLP 1078
Query: 460 SIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGA 519
ID +VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGA
Sbjct: 1079 RIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGA 1138
Query: 520 TYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSF 579
TYQR++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+F
Sbjct: 1139 TYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMKLNPEKCTF 1198
Query: 580 GVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRS 639
GV GK LGFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G+R
Sbjct: 1199 GVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERG 1258
Query: 640 FPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVM 699
PFFK L+K F+W E ++AF K+LL+ PP+L+ P PL LY + + +S+V+
Sbjct: 1259 MPFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVL 1318
Query: 700 LQEIDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVR 753
+ E + E R +YFVS L ++ RY +++K +L+T R+L YFQ V V
Sbjct: 1319 VVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1378
Query: 754 TDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFER 810
T PL +L + +GR+ W++EL L + R ++ +Q+LADFV E T D +
Sbjct: 1379 TSFPLGDILHNREANGRIAKWALELMSLDLSFKPRISIKSQALADFVAEWTECQEDTTVK 1438
Query: 811 VDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAR 870
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A
Sbjct: 1439 KMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1498
Query: 871 EVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQR 930
+ I+ L++R DSQLV NQV + D N+ Y + VR L F + + +V R +N+
Sbjct: 1499 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMKAYRQEVRKLEDKFDGLELSHVLRHDNEA 1558
Query: 931 ADALAKLASTRKPGNNKSVIQETLAYPSIEGE-------LMSCVNRGRTWMDPIISILAG 983
AD LA S R+ + V E L P++ + + W +P+I L
Sbjct: 1559 ADRLANFGSKREVAPS-DVFVEHLYTPTVPHKDTTQAAGIHDVAMVETDWREPLIRFLTS 1617
Query: 984 DPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASH 1042
++ E+ R + Y + + LY++ S L +CVS E+ ++ ++H G+C +H
Sbjct: 1618 QELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNH 1677
Query: 1043 IGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW 1102
R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+W
Sbjct: 1678 AAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVW 1737
Query: 1103 GVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAI 1162
G+D+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I
Sbjct: 1738 GLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRI 1796
Query: 1163 VSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEA 1218
++DNGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL
Sbjct: 1797 ITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPY 1856
Query: 1219 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQAN 1278
G W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + +
Sbjct: 1857 AGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEED 1916
Query: 1279 MAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTP 1337
+L L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P
Sbjct: 1917 RVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSP 1976
Query: 1338 NWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
WEGP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1977 LWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEYLRRFY 2016
>ref|NP_918169.1| putative gag-pol polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2012
Score = 855 bits (2209), Expect = 0.0
Identities = 522/1420 (36%), Positives = 761/1420 (52%), Gaps = 108/1420 (7%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + R D+ ++ R+++ DT
Sbjct: 656 CPHHPNSNHVAKDCFVYKQFADQYTKTA---------RKNTDEEQSTSRKKDDGDTPAG- 705
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
F H +N I FGG + RK+ + R +N+V
Sbjct: 706 ---------------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 743
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 744 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 794
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 795 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 851
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 852 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDVKQAVTCDKE 910
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGD---RTLKIGT 339
++ + + A+ + EG I TK K G+ +T KI
Sbjct: 911 SCDMAQTREMASAREDIRLAAATAS-----EGE------IPATKTSKSGESEAKTKKIPL 959
Query: 340 RLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDK 399
++ +T N D+FAW DMPGI I H L + KP+ Q RR D+
Sbjct: 960 DPSDPTKT------ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDR 1013
Query: 400 GKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLP 459
A+++E+ KLLAA FI+EV +P WLAN V+V+K G+W MCVDYTDLNK+CPKD + LP
Sbjct: 1014 KDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWLMCVDYTDLNKSCPKDPFGLP 1073
Query: 460 SIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGA 519
ID +VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGA
Sbjct: 1074 RIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGA 1133
Query: 520 TYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSF 579
TYQR++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+F
Sbjct: 1134 TYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTF 1193
Query: 580 GVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRS 639
GV GK LGFM++ RGI+ NPEK AI MKSPS K+VQ+LTG +AALSRF+ + G+R
Sbjct: 1194 GVPSGKLLGFMVSHRGIQANPEKVTAILNMKSPSTQKDVQKLTGCMAALSRFVSRLGERG 1253
Query: 640 FPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVM 699
PFFK L+K F W E ++AF K+LL+ PP+L+ P PL LY + + +S+V+
Sbjct: 1254 MPFFKLLKKTDSFRWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVL 1313
Query: 700 LQEIDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVR 753
+ E + E R +YFVS L ++ RY +++K +L+T R+L YFQ V V
Sbjct: 1314 VVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1373
Query: 754 TDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFER 810
T PL +L + +GR+ W++EL + + R ++ +Q+LADFV E T D
Sbjct: 1374 TSFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAE 1433
Query: 811 VDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAR 870
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A
Sbjct: 1434 NMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1493
Query: 871 EVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQR 930
+ I+ L++R DSQLV NQV + D N++ Y + VR L F + + +V R N+
Sbjct: 1494 SLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEA 1553
Query: 931 ADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSCVNRGR-------TWMDPIISILAG 983
AD LA S R+ + V E L P++ + + W +P+I L
Sbjct: 1554 ADRLANFGSKREAAPS-DVFVEHLYSPTVPHKDATQAAGAHDVAMVEADWREPLIRFLTS 1612
Query: 984 DPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASH 1042
++ E+ R + Y L + LY++ S L +CVS E+ ++ ++H G+C +H
Sbjct: 1613 QELPQDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNH 1672
Query: 1043 IGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW 1102
R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+W
Sbjct: 1673 AAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVW 1732
Query: 1103 GVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAI 1162
G+D+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I
Sbjct: 1733 GLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRI 1791
Query: 1163 VSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEA 1218
++DNGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL
Sbjct: 1792 ITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPY 1851
Query: 1219 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQAN 1278
G W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + +
Sbjct: 1852 AGKWVQQLPSVLWSLRTTSSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEED 1911
Query: 1279 MAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTP 1337
+L L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P
Sbjct: 1912 RVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSP 1971
Query: 1338 NWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
WEGP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1972 LWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2011
>gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533012|ref|NP_920808.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|19881590|gb|AAM00991.1| Putative retroelement [Oryza
sativa]
Length = 2017
Score = 849 bits (2193), Expect = 0.0
Identities = 505/1325 (38%), Positives = 735/1325 (55%), Gaps = 72/1325 (5%)
Query: 95 FGGGGDTHAARKRHV--RAVNSVH----------EVAFGFV---HPDITISMADFEGIKP 139
FGG + RK+ + R +N+V E+A F HPD + +
Sbjct: 722 FGGPLAYESKRKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRY----- 776
Query: 140 HKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDL----TPYAGTLVGF 195
P+V+ + + +RR L+D GS+ +I++ D + + ++L P+ G + G
Sbjct: 777 ----PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGL 832
Query: 196 AGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHL 255
+ + G I L FG E R + + V +Y+ I+GR L + AV ++
Sbjct: 833 SATPL---GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYM 889
Query: 256 AVKYPLSSGKVGRLKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGR 315
+K P G + L+ D K A C ++ + + A+ + EG
Sbjct: 890 MMKLPGPRGVLS-LRSDIKQAVTCDKESCDMAQTREMASAREDIRLAAATAS-----EGE 943
Query: 316 VNRPTPIEETKALKFGDRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFI 375
V TK K G+ K + ++ T N D+FAW DMPGI I
Sbjct: 944 V------PATKISKSGESEAKTKKIPLDPSDSTKT---ANNKDIFAWKPSDMPGIPREVI 994
Query: 376 CHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKAN 435
H L + KP+ Q RR D+ A+++E+ KLLAA FI+EV +P WLAN V+V+K
Sbjct: 995 EHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKT 1054
Query: 436 GKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDK 495
G+WRMCVDYTDLNK CPKD + LP ID +VD +G ELLS +D YSGYHQIR+ +D K
Sbjct: 1055 GQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLK 1114
Query: 496 TAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHH 555
T+F+T YCY TMPFGLKNAGATYQR++ R F+ Q+GRN+E YVDD++VK+ + D
Sbjct: 1115 TSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLI 1174
Query: 556 QDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNV 615
DLEE F IR M+LNPEKC+FGV GK LGFM++ RGI+ NPEK AI MK PS
Sbjct: 1175 SDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQ 1234
Query: 616 KEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPIL 675
K+VQ+LTG +AALSRF+ + G+R PFFK L+K F W E ++AF K+LL+ PP+L
Sbjct: 1235 KDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFHWGPEAQKAFEDFKKLLTEPPVL 1294
Query: 676 SKPIQGHPLHLYFAVSDSALSSVML--QEIDGE----HRIVYFVSHTLQGAEVRYQKIEK 729
+ P PL LY + + +S+V++ +E DG R +YFVS L ++ RY +++K
Sbjct: 1295 ASPHPQEPLLLYVSATSQVVSTVLVVEREEDGHVQKVQRPIYFVSEVLADSKTRYPQVQK 1354
Query: 730 AALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRG 789
+L+T R+L YFQ V V T PL +L + +GR+ W++EL + + R
Sbjct: 1355 LLYGILITTRKLSHYFQGHLVTVVTSFPLGDILHNREANGRIAKWALELMSLDISFKPRI 1414
Query: 790 TVGAQSLADFVVELTPDR----FERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQS 845
++ +Q+LADFV E T + E+V+ WT+ DGS SG+GAGV L P L
Sbjct: 1415 SIKSQALADFVAEWTECQEDTPAEKVE-HWTMHFDGSKRLSGTGAGVVLISPTGERLSYV 1473
Query: 846 LKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYL 905
L F A++N AEYEAL+ GL++A + I+ L++R DSQLV NQV + D N++ Y
Sbjct: 1474 LWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYR 1533
Query: 906 ERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMS 965
+ VR L F + + +V R N+ AD LA S R+ + V E L P++ + +
Sbjct: 1534 QEVRKLEDKFDGLELSHVLRHNNKAADRLANFGSKREVAPS-DVFVEHLYAPTVPHKDTT 1592
Query: 966 CVNRGR-------TWMDPIISILAGDPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTP 1017
V W +P+I L ++ E+ R + Y + + LY++ S
Sbjct: 1593 QVAGTHDVAMVEADWREPLIRFLTSQELPQDKDEAERISRRSRLYVMHEAELYKKSPSGI 1652
Query: 1018 LLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQ 1077
L +CVS E+ ++ ++H G+C +H R++ K R GF+WPT D V+ C+ CQ
Sbjct: 1653 LQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQ 1712
Query: 1078 VFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLA 1137
FA P +EL T+ WPFA+WG+D+VGPF A + VA+D F+KWIEA+P+
Sbjct: 1713 FFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAFGGYTHLFVAIDKFSKWIEAKPVV 1772
Query: 1138 KITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTN 1197
IT+ +F+ IV RFG+P I++DNG QF+ ++FC++ GI++ +ASV HP +N
Sbjct: 1773 TITADNARDFF-INIVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSN 1831
Query: 1198 GQVESANRVILRGLRR----RLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVD 1253
GQVE AN +IL+G++ RL G W+++LP+VLWS TT T ++PF + YG +
Sbjct: 1832 GQVERANGMILQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAE 1891
Query: 1254 AMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVR 1313
AMLP E++ + R R EE + + +L L E R+ A I+ Q + N VR
Sbjct: 1892 AMLPSEVEFESLRFRNFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVR 1951
Query: 1314 VRDMQVGDLVL-KWRSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLS 1372
R VGDLVL K ++ +KL+P WEGP+ I +V G+Y L+ DG + S+N
Sbjct: 1952 SRAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLIDNSWNIEH 2011
Query: 1373 LRYFY 1377
LR FY
Sbjct: 2012 LRRFY 2016
>ref|XP_470757.1| putative gag-pol precursor [Oryza sativa] gi|18071370|gb|AAL58229.1|
putative gag-pol precursor [Oryza sativa]
Length = 2026
Score = 848 bits (2191), Expect = 0.0
Identities = 504/1324 (38%), Positives = 731/1324 (55%), Gaps = 70/1324 (5%)
Query: 95 FGGGGDTHAARKRHV--RAVNSVH----------EVAFGFV---HPDITISMADFEGIKP 139
FGG + RK+ + R +N+V E+A F HPD + +
Sbjct: 722 FGGPLAYESKRKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRY----- 776
Query: 140 HKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDL----TPYAGTLVGF 195
P+V+ + + +RR L+D GS+ +I++ D + + ++L P+ G + G
Sbjct: 777 ----PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGL 832
Query: 196 AGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHL 255
+ + G I L FG E R + + V +Y+ I+GR L + AV ++
Sbjct: 833 SATPL---GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYM 889
Query: 256 AVKYPLSSGKVGRLKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGR 315
+K P G + L+ D K A C ++ + + A+ + EG
Sbjct: 890 MMKMPGPRGVLS-LRSDIKQAVTCDKESCDMAQTREMASAREDIRLAAATAS-----EGE 943
Query: 316 VNRPTPIEETKALKFGDRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFI 375
V TK K G+ K T+ + TK N D+FAW DMPGI I
Sbjct: 944 V------PATKTSKSGESEAK--TKKIPLDPSNPTKT-ANNKDIFAWKPSDMPGIPREVI 994
Query: 376 CHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKAN 435
H L + KP+ Q RR D+ A+++E+ KLLAA FI+EV +P WLAN V+V+K
Sbjct: 995 EHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVHHPDWLANPVLVRKKT 1054
Query: 436 GKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDK 495
G+WRMCVDYTDLNK+CPKD + LP ID +VD +G ELLS +D YSGYHQIR+ +D K
Sbjct: 1055 GQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLK 1114
Query: 496 TAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHH 555
T+F+T YCY TMPFGLKNAGATYQR++ R F+ Q+GRN+E YVDD++VK+ + D
Sbjct: 1115 TSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLI 1174
Query: 556 QDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNV 615
DLEE F IR M+LNPEKC+FGV GK LGFM++ RGI+ NPEK AI MK PS
Sbjct: 1175 SDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQ 1234
Query: 616 KEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPIL 675
K+VQ+LTG +AALSRF+ + G+R PFFK L+K F+W E ++AF K+LL+ PP+L
Sbjct: 1235 KDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPEAQKAFEDFKKLLTEPPVL 1294
Query: 676 SKPIQGHPLHLYFAVSDSALSSVMLQEIDGE------HRIVYFVSHTLQGAEVRYQKIEK 729
+ P PL LY + + +S+V++ E + E R +YFVS L ++ RY +++K
Sbjct: 1295 ASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQK 1354
Query: 730 AALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRG 789
+L+T R+L YFQ V V T PL +L + +GR+ W++EL + + R
Sbjct: 1355 LLYGILITTRKLSHYFQGHSVTVVTSFPLGDILHNREANGRIAKWALELMSLDISFKPRI 1414
Query: 790 TVGAQSLADFVVELT---PDRFERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSL 846
++ +Q+LADFV E T D WT+ DGS SG+GAGV L P L L
Sbjct: 1415 SIKSQALADFVAEWTECQEDTPAEKMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVL 1474
Query: 847 KFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLE 906
F A++N AEYEAL+ GL++A + I+ L++R DSQLV NQV + D N++ Y +
Sbjct: 1475 WIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQ 1534
Query: 907 RVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSC 966
VR L F + + +V R N+ AD LA S R+ + V E L P++ + +
Sbjct: 1535 EVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKREVAPS-DVFVEHLYTPTVPHKDTTQ 1593
Query: 967 VNRGR-------TWMDPIISILAGDPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPL 1018
V W +P+I L ++ E+ R + Y + + LY++ S L
Sbjct: 1594 VAGTHDVAMVEADWREPLIRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGIL 1653
Query: 1019 LKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 1078
+CVS E+ ++ ++H G+C +H R++ K R GF+WPT D V+ C+ CQ
Sbjct: 1654 QRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQF 1713
Query: 1079 FADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAK 1138
FA P +EL T+ WPFA+WG+D+VGPF A + VA+D F+KWIEA+P+
Sbjct: 1714 FARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVT 1773
Query: 1139 ITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNG 1198
IT+ +F+ IV RFG+P I++DNG QF+ ++ C++ GI++ +ASV HP +NG
Sbjct: 1774 ITADNAPDFF-INIVHRFGVPNRIITDNGRQFTGGVFKDCCEDFGIKICYASVAHPMSNG 1832
Query: 1199 QVESANRVILRGLRR----RLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDA 1254
QVE AN +IL+G++ RL G W+++LP+VLWS TT T ++PF + YG +A
Sbjct: 1833 QVERANGMILQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEA 1892
Query: 1255 MLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRV 1314
MLP E++ + R R EE + + +L L E R+ A I+ Q + N VR
Sbjct: 1893 MLPSEVEFESLRFRNFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRS 1952
Query: 1315 RDMQVGDLVL-KWRSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSL 1373
R VGDLVL K ++ +KL+P WEGP+ I V G+Y L+ DG + S+N L
Sbjct: 1953 RAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIISAVTRPGSYRLKREDGTLVDNSWNIEHL 2012
Query: 1374 RYFY 1377
R FY
Sbjct: 2013 RRFY 2016
>ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46063437|gb|AAS79740.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1756
Score = 843 bits (2178), Expect = 0.0
Identities = 499/1324 (37%), Positives = 727/1324 (54%), Gaps = 70/1324 (5%)
Query: 95 FGGGGDTHAARKRHV--RAVNSVH----------EVAFGFV---HPDITISMADFEGIKP 139
FGG + RK+ + R +N+V E+A F HPD + +
Sbjct: 461 FGGPLAYESKRKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRY----- 515
Query: 140 HKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDL----TPYAGTLVGF 195
P+V+ + + +RR L+D GS+ +I++ D + + ++L P+ G + G
Sbjct: 516 ----PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGL 571
Query: 196 AGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHL 255
+ + G I L FG E R + + V +Y+ I+GR L + AV ++
Sbjct: 572 SATPL---GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYM 628
Query: 256 AVKYPLSSGKVGRLKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGR 315
+K P +V L+ D K A C ++ + + + + EG
Sbjct: 629 MMKMP-GPRRVLSLRSDIKQAVTCDKESCDMAQTREMASAREDIRLAVATAS-----EGE 682
Query: 316 VNRPTPIEETKALKFGDRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFI 375
V TK K G+ K + ++ T N D+FAW DMPGI I
Sbjct: 683 V------PATKISKSGESEAKTKKIPLDPSDSTKT---ANNKDIFAWKPSDMPGIPREVI 733
Query: 376 CHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKAN 435
H L + KP+ Q RR D+ A+++E+ KLLAA FI+EV +P WLAN V+V+K
Sbjct: 734 EHSLHVKEDAKPIKQRLRRFVQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKT 793
Query: 436 GKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDK 495
G+WRMCVDYTDLNK CPKD + LP ID +VD +G ELLS +D YSGYHQIR+ +D K
Sbjct: 794 GQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLK 853
Query: 496 TAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHH 555
T+F+T YCY TMPFGLKNAGATYQR++ R F+ Q+GRN+E YVDD+++K+ + D
Sbjct: 854 TSFITPIGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVIKTKQKDDLI 913
Query: 556 QDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNV 615
DLEE F IR M+LNPEKC+FGV GK LGFM++ RGI+ NPEK AI MK PS
Sbjct: 914 SDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQ 973
Query: 616 KEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPIL 675
K+VQ+LTG +AALSRF+ + G+R PFFK L+K F+W E ++AF K+ L+ PP+L
Sbjct: 974 KDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPEAQKAFEDFKKFLTEPPVL 1033
Query: 676 SKPIQGHPLHLYFAVSDSALSSVMLQEIDGE------HRIVYFVSHTLQGAEVRYQKIEK 729
+ P PL LY + + +S+V++ E + E R +YFVS L ++ RY +++K
Sbjct: 1034 ASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQK 1093
Query: 730 AALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRG 789
+L+T R+L YFQ V V T PL +L + +GR+ W++EL + + R
Sbjct: 1094 LLYGILITTRKLSHYFQGHSVTVVTSFPLGDILHNREANGRIAKWALELMSLDISFKPRI 1153
Query: 790 TVGAQSLADFVVELT---PDRFERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSL 846
++ +Q+LADFV E T D WT+ DGS SG+GAGV L P L L
Sbjct: 1154 SIKSQALADFVAEWTECQEDTPAEKMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVL 1213
Query: 847 KFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLE 906
F A++N AEYEAL+ GL++A + I+ L++R DSQLV NQV + D N++ Y +
Sbjct: 1214 WIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQ 1273
Query: 907 RVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSC 966
VR L F + + +V R N+ AD LA S R+ + V E L P++ + +
Sbjct: 1274 EVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKREVAPS-DVFVEHLYTPTVPHKDTTQ 1332
Query: 967 VNRGR-------TWMDPIISILAGDPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPL 1018
+ W +P I L ++ E+ R + Y + + LY++ S L
Sbjct: 1333 IAGTHDVALVEADWREPFIRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGIL 1392
Query: 1019 LKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 1078
+CVS E+ ++ ++H G+C +H R++ K R GF+WPT D V+ C+ CQ
Sbjct: 1393 QRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQF 1452
Query: 1079 FADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAK 1138
FA P +EL T+ WPFA+WG+D+VGPF A + VA+D F+KWIEA+P+
Sbjct: 1453 FARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVT 1512
Query: 1139 ITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNG 1198
IT+ +F+ IV RFG+P I++DNG QF+ ++FC++ GI++ +ASV HP +NG
Sbjct: 1513 ITADNARDFF-INIVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNG 1571
Query: 1199 QVESANRVILRGLRR----RLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDA 1254
QVE AN +IL+G++ RL G W+ +LP+VLWS TT T ++PF + YG +A
Sbjct: 1572 QVERANGMILQGIKARVFDRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSPFFLVYGAEA 1631
Query: 1255 MLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRV 1314
MLP E++ + R R EE + + +L L E R+ A I+ Q + N VR
Sbjct: 1632 MLPSEVEFESLRFRNFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRS 1691
Query: 1315 RDMQVGDLVL-KWRSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSL 1373
R VGDLVL K ++ +KL+P WEGP+ I +V G+Y L+ DG + S+N L
Sbjct: 1692 RAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHL 1751
Query: 1374 RYFY 1377
R FY
Sbjct: 1752 RRFY 1755
>gb|AAP54912.1| gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37536646|ref|NP_922625.1| gag-pol precursor [Oryza
sativa (japonica cultivar-group)]
gi|13876521|gb|AAK43497.1| gag-pol precursor [Oryza
sativa (japonica cultivar-group)]
Length = 2017
Score = 841 bits (2172), Expect = 0.0
Identities = 500/1324 (37%), Positives = 728/1324 (54%), Gaps = 70/1324 (5%)
Query: 95 FGGGGDTHAARKRHV--RAVNSVH----------EVAFGFV---HPDITISMADFEGIKP 139
FGG + RK+ + R +N+V E+A F HPD + +
Sbjct: 722 FGGPLAYESKRKQKLTEREINAVQPDTPQYLRWSEIAIKFDRSDHPDRVVHPGRY----- 776
Query: 140 HKDDPIVVQLRMNSFNIRRVLLDQGSSADIIYGDAFDKLGLTDNDL----TPYAGTLVGF 195
P+V+ + + +RR L+D GS+ +I++ D + + ++L P+ G + G
Sbjct: 777 ----PLVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGL 832
Query: 196 AGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHL 255
+ + G I L FG E R + + V +Y+ I+GR L + AV ++
Sbjct: 833 SATPL---GQITLPVTFGTRENFRTENISFEVDDFETAYHAILGRPALAKFMAVPHYTYM 889
Query: 256 AVKYPLSSGKVGRLKVDQKMARECYNNCLNLYGKKSALVGHRCYEIEASDENLDPRGEGR 315
+K P G + L+ D K A C ++ + + A+ + EG
Sbjct: 890 MMKMPGPRGVLS-LRSDIKQAVTCDKESCDMAQTREMASAREDIRLAAATAS-----EGE 943
Query: 316 VNRPTPIEETKALKFGDRTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFI 375
V TK K G+ K + ++ T N D+FAW DMPGI I
Sbjct: 944 V------PATKISKSGESEAKTKKIPLDPSDSTKT---ANNKDIFAWKPSDMPGIPREVI 994
Query: 376 CHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKAN 435
H L + KP+ Q RR D+ A+++E+ KLLAA FI+EV +P WLAN V+V+K
Sbjct: 995 EHSLYVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKT 1054
Query: 436 GKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDK 495
G+WRMCVDYTDLN +CPKD + LP ID +VD +G ELLS +D Y GYHQIR+ +D K
Sbjct: 1055 GQWRMCVDYTDLNISCPKDPFGLPRIDQVVDSTAGCELLSFLDCYLGYHQIRLKESDCLK 1114
Query: 496 TAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHH 555
T+F+T YCY TMPFGLKNAGATYQR++ R F+ Q+GRN+E YVDD++VK+ + D
Sbjct: 1115 TSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLI 1174
Query: 556 QDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNV 615
DLEE F IR M+LNPEKC+FGV GK LGFM++ RGI+ NPEK AI MK PS
Sbjct: 1175 SDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKITAILNMKPPSTQ 1234
Query: 616 KEVQRLTGRIAALSRFLPKSGDRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPIL 675
K+VQ+LTG +AALSRF+ + G+R PFFK L+K F+W + ++AF K+LL+ PP+L
Sbjct: 1235 KDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPDAQKAFEDFKKLLTEPPVL 1294
Query: 676 SKPIQGHPLHLYFAVSDSALSSVMLQEIDGE------HRIVYFVSHTLQGAEVRYQKIEK 729
+ P PL LY A + +S+V++ E + E R +YFVS L ++ RY +++K
Sbjct: 1295 ASPHPQEPLLLYIAAASQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQK 1354
Query: 730 AALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRG 789
+L+T R+L YFQS V V T L +L + +GR+ W++EL + + R
Sbjct: 1355 LLYGILITTRKLSHYFQSHSVTVVTSFSLGDILHNREANGRIAKWALELMSLDISFKPRI 1414
Query: 790 TVGAQSLADFVVELT---PDRFERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSL 846
++ +Q+LADFV E T D WT+ DGS SG+GAGV L P L L
Sbjct: 1415 SIKSQALADFVAEWTECQEDTPAEKMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVL 1474
Query: 847 KFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLE 906
F A++N AEYEAL+ GL++A + I+ L++R DSQLV NQV + D N++ Y +
Sbjct: 1475 WIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQ 1534
Query: 907 RVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSC 966
VR L F + + +V R N+ AD LA S R+ + V E L P++ + +
Sbjct: 1535 EVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKREVAPS-DVFVEHLYTPTVPHKDTTQ 1593
Query: 967 VNRGR-------TWMDPIISILAGDPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPL 1018
+ W +P+I L ++ E+ R + Y + + LY++ S L
Sbjct: 1594 IAGTHDVALVEADWREPLIRFLTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGIL 1653
Query: 1019 LKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 1078
+CVS E+ ++ ++H G+C +H R++ K R GF+WPT D V+ C+ CQ
Sbjct: 1654 QRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQF 1713
Query: 1079 FADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAK 1138
FA P +EL T+ WPFA+WG+D+VGPF A + VA+D F+KWIEA+P+
Sbjct: 1714 FARQIHLPAQELQTIPLSWPFAVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVT 1773
Query: 1139 ITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNG 1198
IT+ +F+ IV RFG+P I++DNG QF+ ++FC++ GI++ +ASV HP +NG
Sbjct: 1774 ITADNARDFF-INIVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNG 1832
Query: 1199 QVESANRVILRGLRR----RLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDA 1254
QVE AN +IL+G++ RL G W+ +LP+VLWS TT T ++PF + YG +A
Sbjct: 1833 QVERANGMILQGIKARVFDRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSPFFLVYGAEA 1892
Query: 1255 MLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRV 1314
MLP E++ + R R EE + + +L L E R+ A I+ Q + N VR
Sbjct: 1893 MLPSEVEFESLRFRNYREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRS 1952
Query: 1315 RDMQVGDLVL-KWRSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSL 1373
R VGDLVL K ++ +KL+P WEGP+ I V G+Y L+ DG + S+N L
Sbjct: 1953 RAFLVGDLVLRKIQTTRDRHKLSPLWEGPFIISDVTRPGSYRLKREDGTLVDNSWNIEHL 2012
Query: 1374 RYFY 1377
R FY
Sbjct: 2013 RRFY 2016
>gb|AAP52913.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37532648|ref|NP_920626.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|19881548|gb|AAM00949.1| Putative retroelement [Oryza
sativa]
Length = 1945
Score = 838 bits (2164), Expect = 0.0
Identities = 519/1420 (36%), Positives = 751/1420 (52%), Gaps = 126/1420 (8%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + A K +E T KK
Sbjct: 607 CPHHPNSNHVAKDCFVYKQFADQYTKT------------------ARKNSDEEQSTSRKK 648
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
A F H +N I FGG + RK+ + R +N+V
Sbjct: 649 DDGDAPAG-------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 694
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 695 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 745
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 746 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 802
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 803 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTCDKE 861
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGD---RTLKIGT 339
++ + + A+ + EG I TK K G+ +T KI
Sbjct: 862 SCDMAQTREMASAREDIRLAAATAS-----EGE------IPATKTSKSGESEAKTKKIPL 910
Query: 340 RLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDK 399
++ +T N D+FAW DMPGI I H L + KP+ Q RR D+
Sbjct: 911 DPSDPTKT------ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDR 964
Query: 400 GKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLP 459
A+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD + LP
Sbjct: 965 KDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLP 1024
Query: 460 SIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGA 519
ID +VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGA
Sbjct: 1025 RIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGA 1084
Query: 520 TYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSF 579
TYQR++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+F
Sbjct: 1085 TYQRMIQRFFSTQIGRNVEAYVDDVVVKTKQKGDLISDLEETFVSIRAFRMKLNPEKCTF 1144
Query: 580 GVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRS 639
GV GK LGFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G+R
Sbjct: 1145 GVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERG 1204
Query: 640 FPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVM 699
PFFK L+K F+W E ++AF K+LL+ PP+L+ P PL LY + + +S+V+
Sbjct: 1205 MPFFKLLKKTDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVL 1264
Query: 700 LQEIDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVR 753
+ E + E R +YFVS L ++ RY +++K +L+T R+L YFQ V V
Sbjct: 1265 VVEREEECHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVV 1324
Query: 754 TDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFER 810
T PL +L + +GR+ W++EL + + R ++ +Q+LADFV E T D
Sbjct: 1325 TSFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAE 1384
Query: 811 VDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAR 870
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A
Sbjct: 1385 NMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAI 1444
Query: 871 EVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQR 930
+ I+ L++R DSQLV NQ VR L F+ + + +V R N+
Sbjct: 1445 SLGIKRLIVRGDSQLVVNQ------------------VRKLEDKFEGLELSHVLRHNNEA 1486
Query: 931 ADALAKLASTRKPGNNKSVIQETLAYPSI-------EGELMSCVNRGRTWMDPIISILAG 983
AD LA S R+ + V E L P++ + + W +P I L
Sbjct: 1487 ADRLANFGSKRETAPS-DVFVEHLYTPTVPHKDTTQDADTHDVAMVEADWREPFIRFLTS 1545
Query: 984 DPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASH 1042
++ E+ R + Y + + LY++ S L +CVS E+ ++ ++H G+C +H
Sbjct: 1546 QELPQDKDEAERISRRSKLYVMHESELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNH 1605
Query: 1043 IGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMW 1102
R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+W
Sbjct: 1606 AAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVW 1665
Query: 1103 GVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAI 1162
G+D+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I
Sbjct: 1666 GLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRI 1724
Query: 1163 VSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEA 1218
++DNGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL
Sbjct: 1725 ITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPY 1784
Query: 1219 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQAN 1278
G W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + +
Sbjct: 1785 AGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEED 1844
Query: 1279 MAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTP 1337
+L L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P
Sbjct: 1845 RVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSP 1904
Query: 1338 NWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
WEGP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1905 LWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 1944
>ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultivar-group)]
gi|21741410|emb|CAD40114.1| OSJNBa0035O13.3 [Oryza sativa
(japonica cultivar-group)]
Length = 2008
Score = 836 bits (2159), Expect = 0.0
Identities = 523/1423 (36%), Positives = 758/1423 (52%), Gaps = 119/1423 (8%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ + A K +E T KK
Sbjct: 657 CPHHPNSNHVAKDCFVYKQFADQYTKT------------------ARKNSDEEQSTSRKK 698
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
A F H +N I FGG + RK+ + R +N+V
Sbjct: 699 DDGDAPAG-------FQDHRKE----LNHI---FGGPLAYESKRKQKLTEREINAVQPDT 744
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 745 PQYLRWSEIAIKFDRSDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 795
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 796 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 852
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 853 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTC--- 908
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGD---RTLKIG- 338
K+S + H +E+ ++ E++ P+ TK K G+ +T KI
Sbjct: 909 -----DKESCDMAHT-HEMASAREDIRLAAATASEGEIPV--TKTSKSGESEAKTKKIPL 960
Query: 339 --TRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLG 396
+ T+ E+ L L N D+FAW DMPGI I H L + KP+ Q RR
Sbjct: 961 DPSDPTKTAESALITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFA 1020
Query: 397 GDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSY 456
D+ A+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD +
Sbjct: 1021 QDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPF 1080
Query: 457 PLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKN 516
LP ID +VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKN
Sbjct: 1081 GLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKN 1140
Query: 517 AGATYQRLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEK 576
AGATYQR++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEK
Sbjct: 1141 AGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFVSIRAFRMKLNPEK 1200
Query: 577 CSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSG 636
C+FGV GK LGFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G
Sbjct: 1201 CTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLG 1260
Query: 637 DRSFPFFKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALS 696
+R PFFK L+K F+W E ++AF K+LL+ PP+L+ P PL LY + + +S
Sbjct: 1261 ERGMPFFKLLKKTDNFQWGPEAQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVS 1320
Query: 697 SVMLQEIDGE------HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPV 750
+V+ E + E R +YFVS L ++ RY +++K VT
Sbjct: 1321 TVLAVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGHSVT------------- 1367
Query: 751 KVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDR 807
V T PL +L + +GR+ W++EL + + R ++ +Q+LADFV E T D
Sbjct: 1368 -VVTSFPLGDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDT 1426
Query: 808 FERVDTQWTLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLK 867
WT+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL+
Sbjct: 1427 PAENMEHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLR 1486
Query: 868 LAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAE 927
+A + I+ L++R DSQLV NQV + D N++ Y + VR L F + + +V R
Sbjct: 1487 IAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHN 1546
Query: 928 NQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMSCVNRGR-------TWMDPIISI 980
N+ AD LA S R+ + V E L P++ + + V W +P+I
Sbjct: 1547 NEAADRLANFGSKREVAPS-DVFVEHLYTPTVPHKDTTQVAGTHDVAVVETDWREPLIRF 1605
Query: 981 LAGDPAEVEQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVC 1039
L ++ E+ R + Y + + LY++ S L +CVS E+ ++ ++H G+C
Sbjct: 1606 LTSQELPQDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGIC 1665
Query: 1040 ASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPF 1099
+H R++ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPF
Sbjct: 1666 GNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPF 1725
Query: 1100 AMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIP 1159
A+WG+D+VGPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P
Sbjct: 1726 AVWGLDMVGPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVP 1784
Query: 1160 RAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RL 1215
I++DNGTQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL
Sbjct: 1785 NRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRL 1844
Query: 1216 AEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEEN 1275
G W+ +LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE
Sbjct: 1845 KPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERY 1904
Query: 1276 QANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNK 1334
+ + +L L E R+ A IR Q + N VR R VGDLVL K ++ +K
Sbjct: 1905 EEDRVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHK 1964
Query: 1335 LTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
L+P WEGP+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1965 LSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2007
>emb|CAE02238.2| OSJNBb0054B09.2 [Oryza sativa (japonica cultivar-group)]
gi|50922823|ref|XP_471772.1| OSJNBb0054B09.2 [Oryza
sativa (japonica cultivar-group)]
Length = 1992
Score = 832 bits (2149), Expect = 0.0
Identities = 514/1415 (36%), Positives = 747/1415 (52%), Gaps = 118/1415 (8%)
Query: 2 CEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 61
C H ++ H DC ++ D+ K ++ + + KK
Sbjct: 656 CPHHPNSNHVAKDCFIYKQFADQY-------------------TKTTRKNSDEEQSTSKK 696
Query: 62 KQESAAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRHV--RAVNSVH--- 116
K + A A T H FGG + RK+ + R +N+V
Sbjct: 697 KDDGDAPAGFQDHRTELNHI-------------FGGPLAYESKRKQKLTEREINAVQPDT 743
Query: 117 -------EVAFGFV---HPDITISMADFEGIKPHKDDPIVVQLRMNSFNIRRVLLDQGSS 166
E+A F HPD + + P+V+ + + +RR L+D GS+
Sbjct: 744 PQYLRWSEIAIKFDRLDHPDRVVHPGRY---------PLVLDPVVRNVKLRRTLIDGGSA 794
Query: 167 ADIIYGDAFDKLGLTDNDL----TPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLK 222
+I++ D + + ++L P+ G + G + + G I L FG E R
Sbjct: 795 LNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL---GQITLPVTFGTRENFRTEN 851
Query: 223 VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGRLKVDQKMARECYNN 282
+ + V +Y+ I+GR L + AV ++ +K P G + L+ D K A C
Sbjct: 852 ISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPRGVLS-LRSDIKQAVTCDKE 910
Query: 283 CLNLYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGDRTLKIGTRLT 342
++ + + A+ + EG I TK K G+ K T+
Sbjct: 911 SCDMAQIREMASAREDIRLAAATAS-----EGE------IPATKTSKSGESEAK--TKKI 957
Query: 343 EEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKA 402
+ TK N D+FAW DMPGI I H L + KP+ Q RR D+ A
Sbjct: 958 HLDSSDPTKT-ANNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDA 1016
Query: 403 VQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSID 462
+++E+ KLLAA FI+EV +P WLAN V+V+K G+WRMCVDYTDLNK+CPKD + LP ID
Sbjct: 1017 IKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRID 1076
Query: 463 SLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQ 522
+VD +G ELLS +D YSGYHQIR+ +D KT+F+T YCY TMPFGLKNAGATYQ
Sbjct: 1077 QVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQ 1136
Query: 523 RLMDRVFARQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQ 582
R++ R F+ Q+GRN+E YVDD++VK+ + D DLEE F IR M+LNPEKC+FGV
Sbjct: 1137 RMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVP 1196
Query: 583 GGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPKSGDRSFPF 642
GK LGFM++ RGI+ NPEK AI MK PS K+VQ+LTG +AALSRF+ + G+R PF
Sbjct: 1197 SGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPF 1256
Query: 643 FKCLRKNVVFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQE 702
FK L+K F+W E ++AF K+ L+ PP++S + V+ +E
Sbjct: 1257 FKLLKKTDDFQWGPEAQKAFEDFKKFLTEPPVVSTVL------------------VVERE 1298
Query: 703 IDGE----HRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDLPL 758
+G R +YFVS L ++ RY +++K +L+T R+L YFQ V V T PL
Sbjct: 1299 EEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSFPL 1358
Query: 759 RQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELT---PDRFERVDTQW 815
+L + +GR+ W++EL + + R ++ +Q+LADFV E T D W
Sbjct: 1359 GDILHNREANGRIAKWALELMSLDISFKPRISIKSQALADFVAEWTECQEDTPAEKMEHW 1418
Query: 816 TLFVDGSSNSSGSGAGVALEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKIR 875
T+ DGS SG+GAGV L P L L F A++N AEYEAL+ GL++A + I+
Sbjct: 1419 TMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIK 1478
Query: 876 SLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALA 935
L++R DSQLV NQV + D N++ Y + VR L F + + +V R N+ AD LA
Sbjct: 1479 RLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLA 1538
Query: 936 KLASTRKPGNNKSVIQETLAYPSIEGELMSCVNRGR-------TWMDPIISILAGDPAEV 988
S R+ + V E L P++ + + V W +P+I L
Sbjct: 1539 NFGSKREMAPS-DVFVEHLYAPTVPHKDTTQVAGTHDVAMVEADWREPLIRFLTSQELPQ 1597
Query: 989 EQCTKEQ-QREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRS 1047
++ E+ R + Y + + LY++ S L +CVS E+ ++ ++H G+C +H R+
Sbjct: 1598 DKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAART 1657
Query: 1048 LACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLV 1107
+ K R GF+WPT D V+ C+ CQ FA P +EL T+ WPFA+WG+D+V
Sbjct: 1658 IVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMV 1717
Query: 1108 GPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNG 1167
GPF A + VA+D F+KWIEA+P+ IT+ +F+ IV RFG+P I++DNG
Sbjct: 1718 GPFKKAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNG 1776
Query: 1168 TQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRR----RLAEAKGAWL 1223
TQF+ ++FC++ GI++ +ASV HP +NGQVE AN +IL+G++ RL G W+
Sbjct: 1777 TQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWV 1836
Query: 1224 DELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVEL 1283
+LP+VLWS TT T ++PF + YG +AMLP E++ + R R EE + + +L
Sbjct: 1837 QQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRFRNFREERYEEDRVDDL 1896
Query: 1284 DLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL-KWRSGAPGNKLTPNWEGP 1342
L E R+ A IR Q + N VR R VGDLVL K ++ +KL+P WEGP
Sbjct: 1897 HRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTRDRHKLSPLWEGP 1956
Query: 1343 YRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1377
+ I +V G+Y L+ DG + S+N LR FY
Sbjct: 1957 FIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 1991
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,329,851,306
Number of Sequences: 2540612
Number of extensions: 99582116
Number of successful extensions: 316609
Number of sequences better than 10.0: 36763
Number of HSP's better than 10.0 without gapping: 3976
Number of HSP's successfully gapped in prelim test: 32787
Number of HSP's that attempted gapping in prelim test: 281705
Number of HSP's gapped (non-prelim): 58850
length of query: 1377
length of database: 863,360,394
effective HSP length: 141
effective length of query: 1236
effective length of database: 505,134,102
effective search space: 624345750072
effective search space used: 624345750072
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0029b.1


