

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.15
(145 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
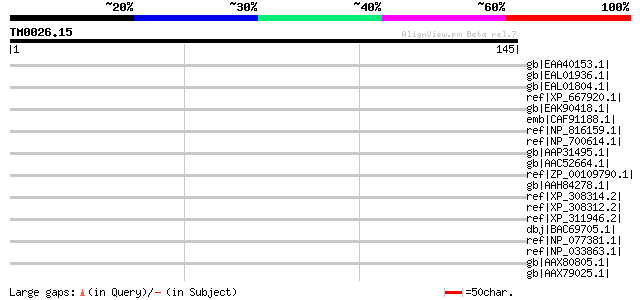
Sequences producing significant alignments: (bits) Value
gb|EAA40153.1| GLP_393_11021_14275 [Giardia lamblia ATCC 50803] 35 0.38
gb|EAL01936.1| hypothetical protein CaO19.11806 [Candida albican... 35 0.49
gb|EAL01804.1| hypothetical protein CaO19.4332 [Candida albicans... 35 0.49
ref|XP_667920.1| hypothetical protein Chro.70489 [Cryptosporidiu... 35 0.49
gb|EAK90418.1| secreted acid phosphatase (calcineurin family),si... 35 0.49
emb|CAF91188.1| unnamed protein product [Tetraodon nigroviridis] 34 0.64
ref|NP_816159.1| membrane protein, putative [Enterococcus faecal... 33 1.1
ref|NP_700614.1| hypothetical protein PF10_0140 [Plasmodium falc... 33 1.4
gb|AAP31495.1| putative cation-transporting ATPase [Western X ph... 33 1.9
gb|AAC52664.1| autoantigen Ku86 32 3.2
ref|ZP_00109790.1| COG2801: Transposase and inactivated derivati... 32 3.2
gb|AAH84278.1| LOC495105 protein [Xenopus laevis] 32 3.2
ref|XP_308314.2| ENSANGP00000009384 [Anopheles gambiae str. PEST... 32 4.2
ref|XP_308312.2| ENSANGP00000024414 [Anopheles gambiae str. PEST... 32 4.2
ref|XP_311946.2| ENSANGP00000011051 [Anopheles gambiae str. PEST... 32 4.2
dbj|BAC69705.1| putative cation transporter [Streptomyces avermi... 32 4.2
ref|NP_077381.1| axin 1 [Rattus norvegicus] gi|2982198|gb|AAC400... 31 5.5
ref|NP_033863.1| axin 1 [Mus musculus] gi|20532379|sp|O35625|AXN... 31 5.5
gb|AAX80805.1| hypothetical protein Tb04.4J6.140 [Trypanosoma br... 31 5.5
gb|AAX79025.1| hypothetical protein, conserved [Trypanosoma brucei] 31 5.5
>gb|EAA40153.1| GLP_393_11021_14275 [Giardia lamblia ATCC 50803]
Length = 1084
Score = 35.0 bits (79), Expect = 0.38
Identities = 20/69 (28%), Positives = 33/69 (46%), Gaps = 1/69 (1%)
Query: 77 SHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDEL 136
SH L + + + EL D + + Y+++K +E E+ LEG+ + WE PPD +
Sbjct: 925 SHSLTGLPPQQQLSIGELPDDRDMKKYINNK-SGLFDDESEISILEGKYFRWEGPPDSSI 983
Query: 137 RARSFVGMA 145
G A
Sbjct: 984 AGEPMHGGA 992
>gb|EAL01936.1| hypothetical protein CaO19.11806 [Candida albicans SC5314]
Length = 1486
Score = 34.7 bits (78), Expect = 0.49
Identities = 17/55 (30%), Positives = 29/55 (51%)
Query: 75 VGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWE 129
VGS L++ S+ ++D L + EN+ +++L E EEL G+ W W+
Sbjct: 1013 VGSFNLVSQGSDKQQDELTIAKLKENKVVKENELFGMNDEIEELEKRAGKFWKWK 1067
>gb|EAL01804.1| hypothetical protein CaO19.4332 [Candida albicans SC5314]
Length = 530
Score = 34.7 bits (78), Expect = 0.49
Identities = 17/55 (30%), Positives = 29/55 (51%)
Query: 75 VGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWE 129
VGS L++ S+ ++D L + EN+ +++L E EEL G+ W W+
Sbjct: 57 VGSFNLVSQGSDKQQDELTIAKLKENKVVKENELFGMNDEIEELEKRAGKFWKWK 111
>ref|XP_667920.1| hypothetical protein Chro.70489 [Cryptosporidium hominis]
gi|54659092|gb|EAL37687.1| hypothetical protein
Chro.70489 [Cryptosporidium hominis]
Length = 826
Score = 34.7 bits (78), Expect = 0.49
Identities = 23/64 (35%), Positives = 30/64 (45%), Gaps = 5/64 (7%)
Query: 64 YAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEG 123
Y FE P V H LN+ S P ++ E+V Y DS ++S PM + P
Sbjct: 135 YTIEEFESP-VNSPHPYLNVSSSPTEETEEMVKTKAIFIYTDSWIISSPMGTDITP---- 189
Query: 124 ELWN 127
ELWN
Sbjct: 190 ELWN 193
>gb|EAK90418.1| secreted acid phosphatase (calcineurin family),signal peptide
[Cryptosporidium parvum] gi|66363254|ref|XP_628593.1|
secreted acid phosphatase (calcineurin family)
[Cryptosporidium parvum]
Length = 826
Score = 34.7 bits (78), Expect = 0.49
Identities = 23/64 (35%), Positives = 30/64 (45%), Gaps = 5/64 (7%)
Query: 64 YAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEG 123
Y FE P V H LN+ S P ++ E+V Y DS ++S PM + P
Sbjct: 135 YTIEEFESP-VNSPHPYLNVSSSPTEETEEMVKTKAIFIYTDSWIISSPMGTDITP---- 189
Query: 124 ELWN 127
ELWN
Sbjct: 190 ELWN 193
>emb|CAF91188.1| unnamed protein product [Tetraodon nigroviridis]
Length = 335
Score = 34.3 bits (77), Expect = 0.64
Identities = 21/71 (29%), Positives = 34/71 (47%), Gaps = 7/71 (9%)
Query: 60 VCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELP 119
VC P + F LP+ G+ +++N EK + V D + D + ++ +EEE P
Sbjct: 212 VCKPRI-KHFALPVWAGTWKIMNFAFRQEKPEWDCVWPDYKQPISDKRDKTEELEEENEP 270
Query: 120 PLEGELWNWED 130
P +WED
Sbjct: 271 P------SWED 275
>ref|NP_816159.1| membrane protein, putative [Enterococcus faecalis V583]
gi|29344471|gb|AAO82229.1| membrane protein, putative
[Enterococcus faecalis V583]
Length = 681
Score = 33.5 bits (75), Expect = 1.1
Identities = 21/67 (31%), Positives = 39/67 (57%), Gaps = 3/67 (4%)
Query: 73 IVVGSHQLLNMKSEPEKDPL--ELVPDDENEHYVD-SKLMSKPMEEEELPPLEGELWNWE 129
+ G + LLN ++ E+ PL +L+P++E + + + ++ +EE E L E+ N+E
Sbjct: 471 VKTGLNSLLNRSTQEEQQPLKDDLLPEEEIVEVEEPTNVDNEELEEIEDKALTDEVDNFE 530
Query: 130 DPPDDEL 136
D DDE+
Sbjct: 531 DLTDDEM 537
>ref|NP_700614.1| hypothetical protein PF10_0140 [Plasmodium falciparum 3D7]
gi|23495005|gb|AAN35338.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 2379
Score = 33.1 bits (74), Expect = 1.4
Identities = 16/48 (33%), Positives = 28/48 (58%)
Query: 83 MKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWED 130
M +EPEK P+ + PD+E E D+K M K ++ + ++G + E+
Sbjct: 1561 MLNEPEKIPIVIPPDNEKEKCGDNKTMGKEGDKRKEDVIDGNEKHMEE 1608
>gb|AAP31495.1| putative cation-transporting ATPase [Western X phytoplasma]
Length = 926
Score = 32.7 bits (73), Expect = 1.9
Identities = 14/26 (53%), Positives = 18/26 (68%)
Query: 86 EPEKDPLELVPDDENEHYVDSKLMSK 111
EPEKD LE P EN+H +D KL+ +
Sbjct: 757 EPEKDLLEHKPRTENDHLLDKKLLKR 782
>gb|AAC52664.1| autoantigen Ku86
Length = 732
Score = 32.0 bits (71), Expect = 3.2
Identities = 22/85 (25%), Positives = 37/85 (42%), Gaps = 6/85 (7%)
Query: 55 LMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKD-----PLELVPDDENEHYVDSKLM 109
L N + C P + + ++ S L+ KSE E P +P+ E + + L
Sbjct: 438 LKNNKKCTPTEAQLSAIDDLIESMSLVK-KSEEEDTIEDLFPTSKIPNPEFQRFFQCLLH 496
Query: 110 SKPMEEEELPPLEGELWNWEDPPDD 134
+E LPP++ + N DPP +
Sbjct: 497 RVLHPQERLPPIQQHILNMLDPPTE 521
>ref|ZP_00109790.1| COG2801: Transposase and inactivated derivatives [Nostoc
punctiforme PCC 73102]
Length = 522
Score = 32.0 bits (71), Expect = 3.2
Identities = 12/29 (41%), Positives = 17/29 (58%)
Query: 100 NEHYVDSKLMSKPMEEEELPPLEGELWNW 128
N HY+ M+K M E+ + P+ ELW W
Sbjct: 258 NYHYLSEYPMTKHMIEDHVQPVPSELWEW 286
>gb|AAH84278.1| LOC495105 protein [Xenopus laevis]
Length = 795
Score = 32.0 bits (71), Expect = 3.2
Identities = 20/57 (35%), Positives = 34/57 (59%), Gaps = 4/57 (7%)
Query: 84 KSEPEKDPLELVPDDENEHYVDSKLMSKPM---EEEELPPLEGELWNWEDPPDDELR 137
KSEPEK+ + L+P ++ H + + M KP+ EE+E E + + E+ P++ LR
Sbjct: 139 KSEPEKEVIHLLP-IKDRHGIIPQTMEKPVIPSEEQEENEEEMDTEHTEEVPEEPLR 194
>ref|XP_308314.2| ENSANGP00000009384 [Anopheles gambiae str. PEST]
gi|55245362|gb|EAA04742.3| ENSANGP00000009384 [Anopheles
gambiae str. PEST]
Length = 5724
Score = 31.6 bits (70), Expect = 4.2
Identities = 25/79 (31%), Positives = 40/79 (49%), Gaps = 7/79 (8%)
Query: 63 PYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDE----NEHYVDSKLMSKPMEEEEL 118
P R+ ELP S + +++K P+K +E VP D + K + KP +EEL
Sbjct: 5369 PTPKRAKELP--QESIEEVSLKPVPQKSEVEEVPADTVSIAEVKDISPKKVKKPKRKEEL 5426
Query: 119 PPLEGELWNWEDPPDDELR 137
PLE ++ E P +E++
Sbjct: 5427 EPLEKPVYP-EIPEQEEVQ 5444
>ref|XP_308312.2| ENSANGP00000024414 [Anopheles gambiae str. PEST]
gi|55245361|gb|EAA45411.2| ENSANGP00000024414 [Anopheles
gambiae str. PEST]
Length = 1400
Score = 31.6 bits (70), Expect = 4.2
Identities = 25/79 (31%), Positives = 40/79 (49%), Gaps = 7/79 (8%)
Query: 63 PYAPRSFELPIVVGSHQLLNMKSEPEKDPLELVPDDE----NEHYVDSKLMSKPMEEEEL 118
P R+ ELP S + +++K P+K +E VP D + K + KP +EEL
Sbjct: 476 PTPKRAKELP--QESIEEVSLKPVPQKSEVEEVPADTVSIAEVKDISPKKVKKPKRKEEL 533
Query: 119 PPLEGELWNWEDPPDDELR 137
PLE ++ E P +E++
Sbjct: 534 EPLEKPVYP-EIPEQEEVQ 551
>ref|XP_311946.2| ENSANGP00000011051 [Anopheles gambiae str. PEST]
gi|55241807|gb|EAA07598.2| ENSANGP00000011051
[Anopheles gambiae str. PEST]
Length = 368
Score = 31.6 bits (70), Expect = 4.2
Identities = 15/64 (23%), Positives = 29/64 (44%)
Query: 15 LHDEDRDRAWVNDEHQVRRIYVGSAAVRSFDQRGRWCETYLMNGRVCLPYAPRSFELPIV 74
+ + +R ++W +D + VG A + + GR CE Y + CL + + L +
Sbjct: 5 MQNNNRTKSWTDDLKVCNQTGVGEAINQIYKDDGRRCEGYESRDKKCLCISDNNTSLYAI 64
Query: 75 VGSH 78
+ H
Sbjct: 65 LSGH 68
>dbj|BAC69705.1| putative cation transporter [Streptomyces avermitilis MA-4680]
gi|29828536|ref|NP_823170.1| putative cation transporter
[Streptomyces avermitilis MA-4680]
Length = 335
Score = 31.6 bits (70), Expect = 4.2
Identities = 16/56 (28%), Positives = 24/56 (42%), Gaps = 11/56 (19%)
Query: 101 EHYVDSKLMSKPMEEEELPPLEGE-----------LWNWEDPPDDELRARSFVGMA 145
EH S+ + +++EE PPL+ E W W PD+ +R G A
Sbjct: 23 EHLASSRFVIVDVDQEEEPPLDDEPLSRRLGLDDAAWKWLGRPDEHVRVEYHAGTA 78
>ref|NP_077381.1| axin 1 [Rattus norvegicus] gi|2982198|gb|AAC40066.1| rAxin [Rattus
norvegicus] gi|7513916|pir||T08422 negative regualtor
axin [imported] - rat
Length = 832
Score = 31.2 bits (69), Expect = 5.5
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELV-------PDDENEHYV 104
E+ +NGRV LP+ PR++ +P ++ EP+K EL+ E E +
Sbjct: 363 ESVQVNGRVPLPHIPRTYRMP--------KEIRVEPQKFAEELIHRLEAVQRTREAEEKL 414
Query: 105 DSKLMSKPMEEE 116
+ +L MEEE
Sbjct: 415 EERLKRVRMEEE 426
>ref|NP_033863.1| axin 1 [Mus musculus] gi|20532379|sp|O35625|AXN1_MOUSE Axin-1 (Axis
inhibition protein 1) (Fused protein)
Length = 863
Score = 31.2 bits (69), Expect = 5.5
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 15/72 (20%)
Query: 52 ETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLELV-------PDDENEHYV 104
E+ +NGRV LP+ PR++ +P ++ EP+K EL+ E E +
Sbjct: 358 ESIQVNGRVPLPHIPRTYRMP--------KEIRVEPQKFAEELIHRLEAVQRTREAEEKL 409
Query: 105 DSKLMSKPMEEE 116
+ +L MEEE
Sbjct: 410 EERLKRVRMEEE 421
>gb|AAX80805.1| hypothetical protein Tb04.4J6.140 [Trypanosoma brucei]
Length = 688
Score = 31.2 bits (69), Expect = 5.5
Identities = 17/50 (34%), Positives = 25/50 (50%)
Query: 91 PLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDELRARS 140
P+ LV D E DS + P++EEE+ P + PP+ +L A S
Sbjct: 322 PVRLVSSDAEESSDDSPCAASPLQEEEISPSVKHKKHRVYPPEGDLSAPS 371
>gb|AAX79025.1| hypothetical protein, conserved [Trypanosoma brucei]
Length = 711
Score = 31.2 bits (69), Expect = 5.5
Identities = 17/50 (34%), Positives = 25/50 (50%)
Query: 91 PLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDELRARS 140
P+ LV D E DS + P++EEE+ P + PP+ +L A S
Sbjct: 345 PVRLVSSDAEESSDDSPCAASPLQEEEISPSVKHKKHRVYPPEGDLSAPS 394
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.137 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 289,414,083
Number of Sequences: 2540612
Number of extensions: 12124548
Number of successful extensions: 29517
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 29498
Number of HSP's gapped (non-prelim): 52
length of query: 145
length of database: 863,360,394
effective HSP length: 121
effective length of query: 24
effective length of database: 555,946,342
effective search space: 13342712208
effective search space used: 13342712208
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0026.15


