

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
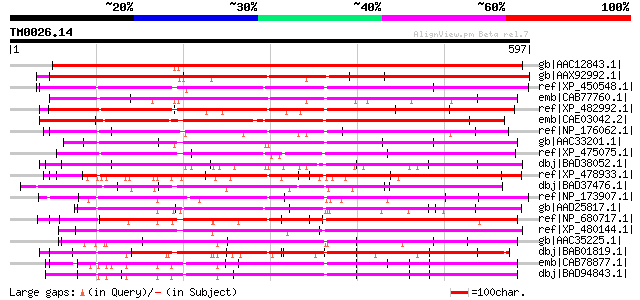
Query= TM0026.14
(597 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAC12843.1| hypothetical protein [Arabidopsis thaliana] gi|74... 623 e-177
gb|AAX92992.1| hypothetical protein LOC_Os11g24530 [Oryza sativa... 584 e-165
ref|XP_450548.1| putative pentatricopeptide (PPR) repeat-contain... 440 e-122
emb|CAB77760.1| hypothetical protein [Arabidopsis thaliana] gi|1... 426 e-117
ref|XP_482992.1| pentatricopeptide (PPR) repeat-containing prote... 417 e-115
emb|CAE03042.2| OSJNBa0084A10.17 [Oryza sativa (japonica cultiva... 411 e-113
ref|NP_176062.1| pentatricopeptide (PPR) repeat-containing prote... 407 e-112
gb|AAC33201.1| Hypothetical protein [Arabidopsis thaliana] gi|15... 405 e-111
ref|XP_475075.1| unknown protein [Oryza sativa (japonica cultiva... 377 e-103
dbj|BAD38052.1| putative pentatricopeptide (PPR) repeat-containi... 374 e-102
ref|XP_478933.1| pentatricopeptide (PPR) repeat-containing prote... 372 e-101
dbj|BAD37476.1| pentatricopeptide (PPR) repeat-containing protei... 365 2e-99
ref|NP_173907.1| pentatricopeptide (PPR) repeat-containing prote... 348 2e-94
gb|AAD25817.1| hypothetical protein [Arabidopsis thaliana] gi|15... 346 1e-93
ref|NP_680717.1| pentatricopeptide (PPR) repeat-containing prote... 346 1e-93
ref|XP_480144.1| putative pentatricopeptide (PPR) repeat-contain... 345 2e-93
gb|AAC35225.1| hypothetical protein [Arabidopsis thaliana] gi|15... 342 3e-92
dbj|BAB01819.1| unnamed protein product [Arabidopsis thaliana] g... 338 3e-91
emb|CAB78877.1| putative protein [Arabidopsis thaliana] gi|45393... 326 1e-87
dbj|BAD94843.1| putative protein [Arabidopsis thaliana] 326 1e-87
>gb|AAC12843.1| hypothetical protein [Arabidopsis thaliana] gi|7485810|pir||T00484
hypothetical protein At2g35030 [imported] - Arabidopsis
thaliana gi|15226879|ref|NP_181048.1| pentatricopeptide
(PPR) repeat-containing protein [Arabidopsis thaliana]
Length = 627
Score = 623 bits (1606), Expect = e-177
Identities = 308/566 (54%), Positives = 411/566 (72%), Gaps = 26/566 (4%)
Query: 50 ISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTW 109
I LCK G+ ARKLFD +P+RD WT +I GYIK G ++EAR+LFD+ D++K+VVTW
Sbjct: 53 IGELCKVGKIAEARKLFDGLPERDVVTWTHVITGYIKLGDMREARELFDRVDSRKNVVTW 112
Query: 110 TTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLV 169
T MV+GY++ Q+ AE LF EMPERN +SWN MI GY Q+G+I+KAL+LF MP+RN+V
Sbjct: 113 TAMVSGYLRSKQLSIAEMLFQEMPERNVVSWNTMIDGYAQSGRIDKALELFDEMPERNIV 172
Query: 170 SWNIIIKALARCGRIEDAQ-------------WHLI-------------RCRKGMMPLRN 203
SWN ++KAL + GRI++A W + R MP RN
Sbjct: 173 SWNSMVKALVQRGRIDEAMNLFERMPRRDVVSWTAMVDGLAKNGKVDEARRLFDCMPERN 232
Query: 204 VVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQ 263
++SWNAMITGYAQN R+DEA +LF+ MPERD ASWN M+TGF +N E+N+A LF +P+
Sbjct: 233 IISWNAMITGYAQNNRIDEADQLFQVMPERDFASWNTMITGFIRNREMNKACGLFDRMPE 292
Query: 264 KDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQ 323
K+VI+WT+M+TGY ++ +EEAL +F+KM +G +KPN GT+V++L ACS LA L EGQQ
Sbjct: 293 KNVISWTTMITGYVENKENEEALNVFSKMLRDGSVKPNVGTYVSILSACSDLAGLVEGQQ 352
Query: 324 IHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHH 383
IHQLISK+ Q+N V SAL+NMYSK GEL ARK+FD+GL+ QRDLISWN MIA YAHH
Sbjct: 353 IHQLISKSVHQKNEIVTSALLNMYSKSGELIAARKMFDNGLVCQRDLISWNSMIAVYAHH 412
Query: 384 GYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDH 443
G+G EAI ++N+M++ GF+ + VTY+ LL ACSHAGLV++G+++F L+++ S+ ++E+H
Sbjct: 413 GHGKEAIEMYNQMRKHGFKPSAVTYLNLLFACSHAGLVEKGMEFFKDLVRDESLPLREEH 472
Query: 444 YACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVE 503
Y CLVDLCGRAGRLK+ I +LS S +G +L+ CNVH I K V KK+L+
Sbjct: 473 YTCLVDLCGRAGRLKDVTNFINCDDARLSRSFYGAILSACNVHNEVSIAKEVVKKVLETG 532
Query: 504 HENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSH 563
++AGTY L+SN+YA+ GK +EAA +RMKMK+KGLKKQPGCSW++VG +FVVGDKSH
Sbjct: 533 SDDAGTYVLMSNIYAANGKREEAAEMRMKMKEKGLKKQPGCSWVKVGKQNHLFVVGDKSH 592
Query: 564 SQSEMLEYLLLGLHTKMKKFGDILDD 589
Q E L+ +L L KM+K ++ D
Sbjct: 593 PQFEALDSILSDLRNKMRKNKNVTSD 618
>gb|AAX92992.1| hypothetical protein LOC_Os11g24530 [Oryza sativa (japonica
cultivar-group)]
Length = 622
Score = 584 bits (1506), Expect = e-165
Identities = 298/578 (51%), Positives = 396/578 (67%), Gaps = 30/578 (5%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSV 106
N ++ L GR AR+LFD MP RD WT M+ Y + G ++EAR LFD+PDA+++V
Sbjct: 45 NWRVAELAAAGRVSDARRLFDGMPDRDVVSWTAMVAAYARRGMLQEARVLFDRPDARRNV 104
Query: 107 VTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKR 166
VTWT +++GY + +++EAE+LF M ERN +SWN M+ Y G++E A LF RMP R
Sbjct: 105 VTWTALLSGYARARRVDEAEALFEGMAERNVVSWNTMLEAYTAVGRVEDASALFNRMPVR 164
Query: 167 NLVSWNIIIKALARCGRIEDAQWHLIR----------------CRKGM----------MP 200
+ SWNI++ L R G +E A+ R R G MP
Sbjct: 165 DAGSWNILLCGLVRSGSLERARKMFERMPVRDVMSWTTMISGLARNGSVDDAWVLFDAMP 224
Query: 201 LRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAE 260
RNVVSWNAMI+GYA+N R++EAL+LF +MP RD+ASWN M+TGF QN +L A +LF E
Sbjct: 225 ERNVVSWNAMISGYARNHRIEEALDLFTKMPIRDVASWNIMITGFIQNKDLKSARQLFDE 284
Query: 261 LPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTE 320
+P+++VITWT+MM GY Q SE ALK+F M G ++PN TF+ L ACS LA+L E
Sbjct: 285 MPKRNVITWTTMMNGYLQCMQSEMALKLFNCMLVQG-IQPNQVTFLGSLDACSNLAALCE 343
Query: 321 GQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAY 380
GQQ+HQ+I KT Q +T V S L+N+Y+KCGE+ +AR +FD + ++DLISWNG+IAAY
Sbjct: 344 GQQVHQMICKTPSQFDTFVESTLMNLYAKCGEIRLARNVFDFSM--EKDLISWNGIIAAY 401
Query: 381 AHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVK 440
AHHG+G EA++L+ MQE G++ ND TYV LL+ACSHAGLVDEG++ F+ ++K+ SI V+
Sbjct: 402 AHHGFGIEAMHLYKNMQENGYKPNDATYVGLLSACSHAGLVDEGLKIFESMVKDNSIVVR 461
Query: 441 EDHYACLVDLCGRAGRLKEAFYIIEGLGVK-LSLSVWGPLLAGCNVHGNADIGKLVAKKI 499
++HY CLVDLC RAGRL++A +I +K S +VW LL GCN HGN IG L AK +
Sbjct: 462 DEHYTCLVDLCSRAGRLEDAKRLISWFKIKPTSSTVWSALLGGCNSHGNESIGDLAAKHL 521
Query: 500 LKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVG 559
L+ E +NAGTY+LL N+YAS GKWKEAA +R +M +GLKKQPGCSWIEV N V VFV
Sbjct: 522 LEAEPDNAGTYTLLCNIYASAGKWKEAAEIRSEMNVRGLKKQPGCSWIEVANKVHVFVSR 581
Query: 560 DKSHSQSEMLEYLLLGLHTKMKKFGDILDDDLSRDVEL 597
DKSHS+S+++ LL +H M+ G + D + DVEL
Sbjct: 582 DKSHSESDLINDLLQDIHRIMRMAGTVPRDHMLIDVEL 619
Score = 178 bits (452), Expect = 4e-43
Identities = 114/370 (30%), Positives = 198/370 (52%), Gaps = 22/370 (5%)
Query: 32 ASTFSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIK 91
++ F+ P + N + L + G + ARK+F++MP RD WTTMI G + G++
Sbjct: 155 SALFNRMPVRDAGSWNILLCGLVRSGSLERARKMFERMPVRDVMSWTTMISGLARNGSVD 214
Query: 92 EARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNG 151
+A LFD +++VV+W M++GY + ++IEEA LF +MP R+ SWNIMI G+ QN
Sbjct: 215 DAWVLFDAM-PERNVVSWNAMISGYARNHRIEEALDLFTKMPIRDVASWNIMITGFIQNK 273
Query: 152 QIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCR--KGMMPLRNVVSWNA 209
++ A LF MPKRN+++W ++ +C + E A L C +G+ P N V++
Sbjct: 274 DLKSARQLFDEMPKRNVITWTTMMNGYLQCMQSEMA-LKLFNCMLVQGIQP--NQVTFLG 330
Query: 210 MITGYAQNRRLDEALELFERM----PERDMASWNAMLTGFFQNGELNRAEKLFAELPQKD 265
+ + L E ++ + + + D + ++ + + GE+ A +F +KD
Sbjct: 331 SLDACSNLAALCEGQQVHQMICKTPSQFDTFVESTLMNLYAKCGEIRLARNVFDFSMEKD 390
Query: 266 VITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIH 325
+I+W ++ YA HG EA+ ++ MQ N G KPN+ T+V +L ACS + EG +I
Sbjct: 391 LISWNGIIAAYAHHGFGIEAMHLYKNMQEN-GYKPNDATYVGLLSACSHAGLVDEGLKIF 449
Query: 326 QLISKTGFQENTRVV-----SALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAY 380
+ + K +N+ VV + L+++ S+ G L A+++ ++ W+ ++
Sbjct: 450 ESMVK----DNSIVVRDEHYTCLVDLCSRAGRLEDAKRLISWFKIKPTSSTVWSALLGGC 505
Query: 381 AHHGYGNEAI 390
H GNE+I
Sbjct: 506 NSH--GNESI 513
Score = 81.3 bits (199), Expect = 8e-14
Identities = 60/232 (25%), Positives = 116/232 (49%), Gaps = 16/232 (6%)
Query: 201 LRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAE 260
+ +V N + A R+ +A LF+ MP+RD+ SW AM+ + + G L A LF
Sbjct: 38 VNHVQDSNWRVAELAAAGRVSDARRLFDGMPDRDVVSWTAMVAAYARRGMLQEARVLFDR 97
Query: 261 L-PQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLT 319
+++V+TWT++++GYA+ +EA +F G + N ++ T+L A + + +
Sbjct: 98 PDARRNVVTWTALLSGYARARRVDEAEALF-----EGMAERNVVSWNTMLEAYTAVGRVE 152
Query: 320 EGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAA 379
+ + + ++ L+ + G L ARK+F+ + RD++SW MI+
Sbjct: 153 DASALFNRMPVRDAGSWNILLCGLV----RSGSLERARKMFE--RMPVRDVMSWTTMISG 206
Query: 380 YAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKL 431
A +G ++A LF+ M E N V++ +++ + ++E + F K+
Sbjct: 207 LARNGSVDDAWVLFDAMPE----RNVVSWNAMISGYARNHRIEEALDLFTKM 254
>ref|XP_450548.1| putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa (japonica cultivar-group)]
gi|48716903|dbj|BAD23598.1| putative pentatricopeptide
(PPR) repeat-containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 755
Score = 440 bits (1132), Expect = e-122
Identities = 227/563 (40%), Positives = 337/563 (59%), Gaps = 10/563 (1%)
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F P + N IS G AR FD P++D W M+ Y++ G ++EAR
Sbjct: 123 FDEMPVRDSVTYNVMISSHANHGLVSLARHYFDLAPEKDAVSWNGMLAAYVRNGRVEEAR 182
Query: 95 KLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIE 154
LF+ + V++W +++GYV+ ++ EA LF MP R+ +SWNIM+ GY + G +
Sbjct: 183 GLFNSR-TEWDVISWNALMSGYVQWGKMSEARELFDRMPGRDVVSWNIMVSGYARRGDMV 241
Query: 155 KALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGY 214
+A LF P R++ +W ++ A+ G +E+A R MP RN VSWNAM+ Y
Sbjct: 242 EARRLFDAAPVRDVFTWTAVVSGYAQNGMLEEA-----RRVFDAMPERNAVSWNAMVAAY 296
Query: 215 AQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMT 274
Q R +DEA ELF MP R++ASWN MLTG+ Q G L A+ +F +PQKD ++W +M+
Sbjct: 297 IQRRMMDEAKELFNMMPCRNVASWNTMLTGYAQAGMLEEAKAVFDTMPQKDAVSWAAMLA 356
Query: 275 GYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQ 334
Y+Q G SEE L++F +M G N F VL C+ +A+L G Q+H + + G+
Sbjct: 357 AYSQGGCSEETLQLFIEM-GRCGEWVNRSAFACVLSTCADIAALECGMQLHGRLIRAGYG 415
Query: 335 ENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFN 394
V +AL+ MY KCG + AR F++ + +RD++SWN MIA YA HG+G EA+ +F+
Sbjct: 416 VGCFVGNALLAMYFKCGNMEDARNAFEE--MEERDVVSWNTMIAGYARHGFGKEALEIFD 473
Query: 395 KMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRA 454
M+ + +D+T V +L ACSH+GLV++GI YF + + + K +HY C++DL GRA
Sbjct: 474 MMRTTSTKPDDITLVGVLAACSHSGLVEKGISYFYSMHHDFGVTAKPEHYTCMIDLLGRA 533
Query: 455 GRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLS 514
GRL EA +++ + + ++WG LL +H N ++G+ A+KI ++E ENAG Y LLS
Sbjct: 534 GRLAEAHDLMKDMPFEPDSTMWGALLGASRIHRNPELGRSAAEKIFELEPENAGMYVLLS 593
Query: 515 NMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLL 574
N+YAS GKW++A +R+ M+++G+KK PG SWIEV N V F GD H + E + L
Sbjct: 594 NIYASSGKWRDARKMRVMMEERGVKKVPGFSWIEVQNKVHTFSAGDCVHPEKEKIYAFLE 653
Query: 575 GLHTKMKKFGDILDDDL-SRDVE 596
L +MKK G + D+ DVE
Sbjct: 654 DLDMRMKKAGYVSATDMVLHDVE 676
Score = 209 bits (532), Expect = 2e-52
Identities = 132/459 (28%), Positives = 238/459 (51%), Gaps = 25/459 (5%)
Query: 32 ASTFSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIK 91
+S+ S E+ R N I+ + GR A +LF MP+R + M+ GY G +
Sbjct: 27 SSSSSGRLEPEVIRSNKAITAHMRAGRVADAERLFAAMPRRSTSTYNAMLAGYSANGRLP 86
Query: 92 EARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNG 151
A LF + + ++ T+++ + + +A LF EMP R+ +++N+MI + +G
Sbjct: 87 LAASLF-RAIPRPDNYSYNTLLHALAVSSSLADARGLFDEMPVRDSVTYNVMISSHANHG 145
Query: 152 QIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLR---NVVSWN 208
+ A F P+++ VSWN ++ A R GR+E+A +G+ R +V+SWN
Sbjct: 146 LVSLARHYFDLAPEKDAVSWNGMLAAYVRNGRVEEA--------RGLFNSRTEWDVISWN 197
Query: 209 AMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVIT 268
A+++GY Q ++ EA ELF+RMP RD+ SWN M++G+ + G++ A +LF P +DV T
Sbjct: 198 ALMSGYVQWGKMSEARELFDRMPGRDVVSWNIMVSGYARRGDMVEARRLFDAAPVRDVFT 257
Query: 269 WTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLI 328
WT++++GYAQ+G+ EEA ++F M + N ++ A + E +++ ++
Sbjct: 258 WTAVVSGYAQNGMLEEARRVFDAMPERNAVSWN-----AMVAAYIQRRMMDEAKELFNMM 312
Query: 329 SKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNE 388
N + ++ Y++ G L A+ +FD + Q+D +SW M+AAY+ G E
Sbjct: 313 P----CRNVASWNTMLTGYAQAGMLEEAKAVFD--TMPQKDAVSWAAMLAAYSQGGCSEE 366
Query: 389 AINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLV 448
+ LF +M G N + +L+ C+ ++ G+Q +L++ V L+
Sbjct: 367 TLQLFIEMGRCGEWVNRSAFACVLSTCADIAALECGMQLHGRLIR-AGYGVGCFVGNALL 425
Query: 449 DLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHG 487
+ + G +++A E + + +S W ++AG HG
Sbjct: 426 AMYFKCGNMEDARNAFEEMEERDVVS-WNTMIAGYARHG 463
>emb|CAB77760.1| hypothetical protein [Arabidopsis thaliana]
gi|15235498|ref|NP_192184.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|4263522|gb|AAD15348.1| hypothetical protein
[Arabidopsis thaliana] gi|25345450|pir||A85035
hypothetical protein AT4g02750 [imported] - Arabidopsis
thaliana
Length = 781
Score = 426 bits (1095), Expect = e-117
Identities = 222/596 (37%), Positives = 348/596 (58%), Gaps = 63/596 (10%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFD-QPDAKKS 105
N IS + G + ARKLFD+MP+RD W MI GY++ + +AR+LF+ P+ +
Sbjct: 99 NGMISGYLRNGEFELARKLFDEMPERDLVSWNVMIKGYVRNRNLGKARELFEIMPE--RD 156
Query: 106 VVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRR--- 162
V +W TM++GY + +++A S+F MPE+N++SWN ++ Y QN ++E+A LF+
Sbjct: 157 VCSWNTMLSGYAQNGCVDDARSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSREN 216
Query: 163 ----------------------------MPKRNLVSWNIIIKALARCGRIEDAQ------ 188
M R++VSWN II A+ G+I++A+
Sbjct: 217 WALVSWNCLLGGFVKKKKIVEARQFFDSMNVRDVVSWNTIITGYAQSGKIDEARQLFDES 276
Query: 189 -------WHLI-------------RCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFE 228
W + R MP RN VSWNAM+ GY Q R++ A ELF+
Sbjct: 277 PVQDVFTWTAMVSGYIQNRMVEEARELFDKMPERNEVSWNAMLAGYVQGERMEMAKELFD 336
Query: 229 RMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKM 288
MP R++++WN M+TG+ Q G+++ A+ LF ++P++D ++W +M+ GY+Q G S EAL++
Sbjct: 337 VMPCRNVSTWNTMITGYAQCGKISEAKNLFDKMPKRDPVSWAAMIAGYSQSGHSFEALRL 396
Query: 289 FTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYS 348
F +M+ GG + N +F + L C+ + +L G+Q+H + K G++ V +AL+ MY
Sbjct: 397 FVQMEREGG-RLNRSSFSSALSTCADVVALELGKQLHGRLVKGGYETGCFVGNALLLMYC 455
Query: 349 KCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTY 408
KCG + A +F + + +D++SWN MIA Y+ HG+G A+ F M+ G + +D T
Sbjct: 456 KCGSIEEANDLFKE--MAGKDIVSWNTMIAGYSRHGFGEVALRFFESMKREGLKPDDATM 513
Query: 409 VELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLG 468
V +L+ACSH GLVD+G QYF + ++ + HYAC+VDL GRAG L++A +++ +
Sbjct: 514 VAVLSACSHTGLVDKGRQYFYTMTQDYGVMPNSQHYACMVDLLGRAGLLEDAHNLMKNMP 573
Query: 469 VKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAAN 528
+ ++WG LL VHGN ++ + A KI +E EN+G Y LLSN+YAS G+W +
Sbjct: 574 FEPDAAIWGTLLGASRVHGNTELAETAADKIFAMEPENSGMYVLLSNLYASSGRWGDVGK 633
Query: 529 VRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFG 584
+R++M+DKG+KK PG SWIE+ N F VGD+ H + + + L L +MKK G
Sbjct: 634 LRVRMRDKGVKKVPGYSWIEIQNKTHTFSVGDEFHPEKDEIFAFLEELDLRMKKAG 689
Score = 166 bits (420), Expect = 2e-39
Identities = 128/435 (29%), Positives = 212/435 (48%), Gaps = 62/435 (14%)
Query: 140 WNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMM 199
WN+ I Y + G+ +AL +F+RMP+ + VS+N +I R G E L R M
Sbjct: 67 WNVAISSYMRTGRCNEALRVFKRMPRWSSVSYNGMISGYLRNGEFE-----LARKLFDEM 121
Query: 200 PLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFA 259
P R++VSWN MI GY +NR L +A ELFE MPERD+ SWN ML+G+ QNG ++ A +F
Sbjct: 122 PERDLVSWNVMIKGYVRNRNLGKARELFEIMPERDVCSWNTMLSGYAQNGCVDDARSVFD 181
Query: 260 ELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLT 319
+P+K+ ++W ++++ Y Q+ EEA +F K + N L N +LG +
Sbjct: 182 RMPEKNDVSWNALLSAYVQNSKMEEACMLF-KSRENWALVSWN----CLLGGFVKKKKIV 236
Query: 320 EGQQIHQLISKTGFQENTRVV---SALINMYSKCGELHIARKIFDDGLLRQRDLISWNGM 376
E +Q + N R V + +I Y++ G++ AR++FD+ + +D+ +W M
Sbjct: 237 EARQFFDSM-------NVRDVVSWNTIITGYAQSGKIDEARQLFDESPV--QDVFTWTAM 287
Query: 377 IAAYAHHGYGNEAINLFNKM---QELGFQANDVTYVE----------------------- 410
++ Y + EA LF+KM E+ + A YV+
Sbjct: 288 VSGYIQNRMVEEARELFDKMPERNEVSWNAMLAGYVQGERMEMAKELFDVMPCRNVSTWN 347
Query: 411 -LLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEA---FYIIEG 466
++T + G + E FDK+ K + +A ++ ++G EA F +E
Sbjct: 348 TMITGYAQCGKISEAKNLFDKMPKRDPVS-----WAAMIAGYSQSGHSFEALRLFVQMER 402
Query: 467 LGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHEN---AGTYSLLSNMYASVGKW 523
G +L+ S + L+ C ++GK + +++K +E G LL MY G
Sbjct: 403 EGGRLNRSSFSSALSTCADVVALELGKQLHGRLVKGGYETGCFVGNALLL--MYCKCGSI 460
Query: 524 KEAANVRMKMKDKGL 538
+EA ++ +M K +
Sbjct: 461 EEANDLFKEMAGKDI 475
>ref|XP_482992.1| pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa (japonica cultivar-group)]
gi|51964972|ref|XP_507270.1| PREDICTED
OSJNBb0092C08.13-1 gene product [Oryza sativa (japonica
cultivar-group)] gi|42409025|dbj|BAD10278.1|
pentatricopeptide (PPR) repeat-containing protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 563
Score = 417 bits (1071), Expect = e-115
Identities = 211/539 (39%), Positives = 329/539 (60%), Gaps = 12/539 (2%)
Query: 45 RCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGA-IKEARKLFDQPDAK 103
R N I+ L + G AR++FD MP+RD W ++ + G + AR LFD ++
Sbjct: 18 RDNQRITALARAGDVAAARRVFDAMPRRDAVSWNALLTALWRAGRDLPAARSLFDDMPSR 77
Query: 104 KSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM 163
+V++W +++ G + + A + F P RN SWN M+ G + G +E A LF +M
Sbjct: 78 -NVISWNSIIAGCLAHGDLAAASAYFARAPRRNVASWNAMLAGLVRLGSMEDARSLFDQM 136
Query: 164 PKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEA 223
P+RN+VS+ ++ LARCG + A R MP RN+VSW AMI+GY N L+EA
Sbjct: 137 PERNVVSYTTMVDGLARCGEVASA-----RELFDAMPTRNLVSWAAMISGYVDNNMLEEA 191
Query: 224 LELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSE 283
+LFE MPE+++ + AM+TG+ + G+L A +LF + KDVI+W ++++GY +GL E
Sbjct: 192 RKLFEAMPEKNVVACTAMITGYCKEGDLQNARRLFDGIRAKDVISWNAIISGYVHNGLGE 251
Query: 284 EALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSAL 343
EA K++ M G +KP+ T + +L ACS LA L +G+ H ++ K + + + +AL
Sbjct: 252 EATKLYIIMLREG-IKPDQATLIALLTACSSLALLRQGRSTHAVVIKAMLESSISICNAL 310
Query: 344 INMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQA 403
+ MYSKCG + + +F L+ +D++SWN +IAAYA HG + I LF++M+ G
Sbjct: 311 MTMYSKCGNVDESELVFMS--LKSQDIVSWNTIIAAYAQHGRYQKVIALFHEMELCGLIP 368
Query: 404 NDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYI 463
ND+T++ +L+AC HAG VDE ++ FD + +I + +HYAC+VD+ RAG+L++A
Sbjct: 369 NDITFLSMLSACGHAGRVDESLKLFDLMFSKYAISPRAEHYACIVDILSRAGQLEKACSY 428
Query: 464 IEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKW 523
I+ + + +VWG LL HGN +G+L AK ++ + E++G Y +LSN+YA+ G W
Sbjct: 429 IKEMPSEAEKNVWGTLLCASQTHGNVQLGELAAKMLVLSDFESSGAYVMLSNIYAAAGMW 488
Query: 524 KEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEML--EYLLLGLHTKM 580
E VR +MK+KG+KKQPG SW E+ + V +FV GD SH + +M+ E + H +M
Sbjct: 489 GEVNRVRSQMKEKGVKKQPGHSWTEIADKVHMFVGGDASHPEMDMILSELRKISFHMQM 547
Score = 177 bits (450), Expect = 7e-43
Identities = 116/415 (27%), Positives = 216/415 (51%), Gaps = 13/415 (3%)
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F+ P + N+ ++ L + G + AR LFDQMP+R+ +TTM+DG +CG + AR
Sbjct: 102 FARAPRRNVASWNAMLAGLVRLGSMEDARSLFDQMPERNVVSYTTMVDGLARCGEVASAR 161
Query: 95 KLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIE 154
+LFD +++V+W M++GYV N +EEA LF MPE+N ++ MI GY + G ++
Sbjct: 162 ELFDAMPT-RNLVSWAAMISGYVDNNMLEEARKLFEAMPEKNVVACTAMITGYCKEGDLQ 220
Query: 155 KALDLFRRMPKRNLVSWNIIIKALARCGRIEDA-QWHLIRCRKGMMPLRNVVSWNAMITG 213
A LF + ++++SWN II G E+A + ++I R+G+ P + + A++T
Sbjct: 221 NARRLFDGIRAKDVISWNAIISGYVHNGLGEEATKLYIIMLREGIKP--DQATLIALLTA 278
Query: 214 YAQNRRLDEALE----LFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITW 269
+ L + + + M E ++ NA++T + + G ++ +E +F L +D+++W
Sbjct: 279 CSSLALLRQGRSTHAVVIKAMLESSISICNALMTMYSKCGNVDESELVFMSLKSQDIVSW 338
Query: 270 TSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLI- 328
+++ YAQHG ++ + +F +M+ GL PN+ TF+++L AC + E ++ L+
Sbjct: 339 NTIIAAYAQHGRYQKVIALFHEMEL-CGLIPNDITFLSMLSACGHAGRVDESLKLFDLMF 397
Query: 329 SKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNE 388
SK + ++++ S+ G+L A + + + + W ++ A HG +
Sbjct: 398 SKYAISPRAEHYACIVDILSRAGQLEKACSYIKE-MPSEAEKNVWGTLLCASQTHG-NVQ 455
Query: 389 AINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDH 443
L KM L + YV L + AG+ E + +K + ++ + H
Sbjct: 456 LGELAAKMLVLSDFESSGAYVMLSNIYAAAGMWGE-VNRVRSQMKEKGVKKQPGH 509
Score = 111 bits (277), Expect = 8e-23
Identities = 94/372 (25%), Positives = 156/372 (41%), Gaps = 69/372 (18%)
Query: 190 HLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLT------ 243
H +R + R +V N IT A+ + A +F+ MP RD SWNA+LT
Sbjct: 2 HRLRHHAALASSRLLVRDNQRITALARAGDVAAARRVFDAMPRRDAVSWNALLTALWRAG 61
Query: 244 --------------------------GFFQNGELNRAEKLFAELPQKDVITWTSMMTGYA 277
G +G+L A FA P+++V +W +M+ G
Sbjct: 62 RDLPAARSLFDDMPSRNVISWNSIIAGCLAHGDLAAASAYFARAPRRNVASWNAMLAGLV 121
Query: 278 QHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLAS---LTEGQQIHQLISKTGF- 333
+ G E+A +F +M + + T V L C +AS L + L+S
Sbjct: 122 RLGSMEDARSLFDQMPERNVV--SYTTMVDGLARCGEVASARELFDAMPTRNLVSWAAMI 179
Query: 334 --------------------QENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISW 373
++N +A+I Y K G+L AR++FD +R +D+ISW
Sbjct: 180 SGYVDNNMLEEARKLFEAMPEKNVVACTAMITGYCKEGDLQNARRLFDG--IRAKDVISW 237
Query: 374 NGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLK 433
N +I+ Y H+G G EA L+ M G + + T + LLTACS L+ +G ++K
Sbjct: 238 NAIISGYVHNGLGEEATKLYIIMLREGIKPDQATLIALLTACSSLALLRQGRSTHAVVIK 297
Query: 434 NRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGK 493
++ L+ + + G + E+ + L + +S W ++A HG
Sbjct: 298 -AMLESSISICNALMTMYSKCGNVDESELVFMSLKSQDIVS-WNTIIAAYAQHGR----- 350
Query: 494 LVAKKILKVEHE 505
+K++ + HE
Sbjct: 351 --YQKVIALFHE 360
>emb|CAE03042.2| OSJNBa0084A10.17 [Oryza sativa (japonica cultivar-group)]
gi|50924392|ref|XP_472556.1| OSJNBa0084A10.17 [Oryza
sativa (japonica cultivar-group)]
Length = 655
Score = 411 bits (1057), Expect = e-113
Identities = 208/535 (38%), Positives = 326/535 (60%), Gaps = 13/535 (2%)
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F+ P+ + N+ +S + G AR+LFD MP RD W TMI GYIK ++EAR
Sbjct: 119 FNRIPDRNVVSWNAMVSGYARNGMVKRARELFDMMPWRDDVSWLTMISGYIKRKHVREAR 178
Query: 95 KLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIE 154
+LFD + + V +++GYV+L + AE LF +M RN +SWN+MI GY + G +
Sbjct: 179 ELFDSMPSPPTSVC-NALLSGYVELGYMRAAEVLFGQMQTRNPVSWNVMITGYARAGSMG 237
Query: 155 KALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGY 214
A LF MP+++++S I++ + G + DA W + + MP R+ V+WN M+ G+
Sbjct: 238 IAQRLFDEMPEKDVLSRTAIMRGYLQNGSV-DAAWKVFKD----MPHRDTVAWNTMMDGF 292
Query: 215 AQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMT 274
+N RLD+AL+LF MP+RD SWNA+L G+ Q G+++ A F P KD I+W ++++
Sbjct: 293 VRNDRLDDALKLFSEMPDRDQISWNAILQGYVQQGDMDSANAWFRRAPNKDAISWNTLIS 352
Query: 275 GYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQ 334
GY G AL + ++M GGLKP+ T V+ C+ L SL G+ +H KTGF+
Sbjct: 353 GYKDEG----ALSLLSEM-IRGGLKPDQATLSVVISICASLVSLGCGKMVHLWAIKTGFE 407
Query: 335 ENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFN 394
+ V+S+LI+MYSKCG + A ++F+ L+ QRD ++WN MIA YA+HG +EA+ +F+
Sbjct: 408 HDALVMSSLISMYSKCGLISEASQVFE--LILQRDTVTWNAMIATYAYHGLADEALKVFD 465
Query: 395 KMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRA 454
M + GF+ + T++ +L+AC+H G + EG +F + ++ ++ + DHY+C+VDL GR+
Sbjct: 466 MMTKAGFRPDHATFLSILSACAHKGYLYEGCYHFRSMQEDWNLVPRSDHYSCMVDLLGRS 525
Query: 455 GRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLS 514
G + +A+ + + W L + CN HG +G+++A+ +LK + G Y+LLS
Sbjct: 526 GFIHQAYDFTRRIPSDHRTTAWETLFSVCNSHGEIQLGEIIARNVLKARPSDGGMYTLLS 585
Query: 515 NMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEML 569
N+YA+ W AA+VR MK++GLKK+ GCSWIE+ V F D +H E +
Sbjct: 586 NIYAAKEMWSSAASVRGFMKERGLKKETGCSWIELKGEVVTFSSNDSNHPLIEQI 640
Score = 159 bits (401), Expect = 3e-37
Identities = 119/433 (27%), Positives = 203/433 (46%), Gaps = 29/433 (6%)
Query: 99 QPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALD 158
Q A V T +N + + ++ A +F EM ERN +WN M+ G +N + +A
Sbjct: 27 QRHAAGDVFQSNTAINEHFRAGRVAAARRVFDEMSERNVFTWNCMVSGLIRNRMLAEARK 86
Query: 159 LFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNR 218
+F MP RN VSW ++ ARCGR+ +A+ R +P RNVVSWNAM++GYA+N
Sbjct: 87 VFDAMPVRNSVSWAALLTGYARCGRVAEARELFNR-----IPDRNVVSWNAMVSGYARNG 141
Query: 219 RLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQ 278
+ A ELF+ MP RD SW M++G+ + + A +LF +P ++++GY +
Sbjct: 142 MVKRARELFDMMPWRDDVSWLTMISGYIKRKHVREARELFDSMPSPPTSVCNALLSGYVE 201
Query: 279 HGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLAS-LTEGQQIHQLISKTGFQENT 337
G A +F +MQ + N G+A L + ++S+T
Sbjct: 202 LGYMRAAEVLFGQMQTRNPVSWNVMITGYARAGSMGIAQRLFDEMPEKDVLSRT------ 255
Query: 338 RVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQ 397
A++ Y + G + A K+F D + RD ++WN M+ + + ++A+ LF++M
Sbjct: 256 ----AIMRGYLQNGSVDAAWKVFKD--MPHRDTVAWNTMMDGFVRNDRLDDALKLFSEMP 309
Query: 398 ELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRL 457
+ + +++ +L G +D +F + +I + L+ G L
Sbjct: 310 D----RDQISWNAILQGYVQQGDMDSANAWFRRAPNKDAIS-----WNTLISGYKDEGAL 360
Query: 458 KEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHE-NAGTYSLLSNM 516
+I G G+K + +++ C + GK+V +K E +A S L +M
Sbjct: 361 SLLSEMIRG-GLKPDQATLSVVISICASLVSLGCGKMVHLWAIKTGFEHDALVMSSLISM 419
Query: 517 YASVGKWKEAANV 529
Y+ G EA+ V
Sbjct: 420 YSKCGLISEASQV 432
>ref|NP_176062.1| pentatricopeptide (PPR) repeat-containing protein [Arabidopsis
thaliana] gi|25345446|pir||G96608 hypothetical protein
F25P12.87 [imported] - Arabidopsis thaliana
gi|9954744|gb|AAG09095.1| Hypothetical protein
[Arabidopsis thaliana]
Length = 704
Score = 407 bits (1047), Expect = e-112
Identities = 217/559 (38%), Positives = 325/559 (57%), Gaps = 41/559 (7%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSV 106
NS +S G AR+LFD+M +R+ W ++ GYIK I EAR +F+ +++V
Sbjct: 52 NSIVSGYFSNGLPKEARQLFDEMSERNVVSWNGLVSGYIKNRMIVEARNVFELMP-ERNV 110
Query: 107 VTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKR 166
V+WT MV GY++ + EAESLF+ MPERNE+SW +M GG +G+I+KA L+ MP +
Sbjct: 111 VSWTAMVKGYMQEGMVGEAESLFWRMPERNEVSWTVMFGGLIDDGRIDKARKLYDMMPVK 170
Query: 167 NLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALEL 226
++V+ +I L R GR+++A+ R+ RNVV+W MITGY QN R+D A +L
Sbjct: 171 DVVASTNMIGGLCREGRVDEARLIFDEMRE-----RNVVTWTTMITGYRQNNRVDVARKL 225
Query: 227 FERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVI------------------- 267
FE MPE+ SW +ML G+ +G + AE+ F +P K VI
Sbjct: 226 FEVMPEKTEVSWTSMLLGYTLSGRIEDAEEFFEVMPMKPVIACNAMIVGFGEVGEISKAR 285
Query: 268 ------------TWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGL 315
TW M+ Y + G EAL +F +MQ G ++P+ + +++L C+ L
Sbjct: 286 RVFDLMEDRDNATWRGMIKAYERKGFELEALDLFAQMQKQG-VRPSFPSLISILSVCATL 344
Query: 316 ASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNG 375
ASL G+Q+H + + F ++ V S L+ MY KCGEL A+ +FD +D+I WN
Sbjct: 345 ASLQYGRQVHAHLVRCQFDDDVYVASVLMTMYVKCGELVKAKLVFDR--FSSKDIIMWNS 402
Query: 376 MIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNR 435
+I+ YA HG G EA+ +F++M G N VT + +LTACS+AG ++EG++ F+ +
Sbjct: 403 IISGYASHGLGEEALKIFHEMPSSGTMPNKVTLIAILTACSYAGKLEEGLEIFESMESKF 462
Query: 436 SIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLV 495
+ +HY+C VD+ GRAG++ +A +IE + +K +VWG LL C H D+ ++
Sbjct: 463 CVTPTVEHYSCTVDMLGRAGQVDKAMELIESMTIKPDATVWGALLGACKTHSRLDLAEVA 522
Query: 496 AKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQV 555
AKK+ + E +NAGTY LLS++ AS KW + A VR M+ + K PGCSWIEVG V +
Sbjct: 523 AKKLFENEPDNAGTYVLLSSINASRSKWGDVAVVRKNMRTNNVSKFPGCSWIEVGKKVHM 582
Query: 556 FVVGD-KSHSQSEMLEYLL 573
F G K+H + M+ +L
Sbjct: 583 FTRGGIKNHPEQAMILMML 601
Score = 174 bits (442), Expect = 6e-42
Identities = 113/351 (32%), Positives = 187/351 (53%), Gaps = 11/351 (3%)
Query: 39 PNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFD 98
P ++ + I LC+EGR D AR +FD+M +R+ WTTMI GY + + ARKLF+
Sbjct: 168 PVKDVVASTNMIGGLCREGRVDEARLIFDEMRERNVVTWTTMITGYRQNNRVDVARKLFE 227
Query: 99 QPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALD 158
+K+ V+WT+M+ GY +IE+AE F MP + ++ N MI G+G+ G+I KA
Sbjct: 228 VM-PEKTEVSWTSMLLGYTLSGRIEDAEEFFEVMPMKPVIACNAMIVGFGEVGEISKARR 286
Query: 159 LFRRMPKRNLVSWNIIIKALARCG-RIEDAQWHLIRCRKGMMP----LRNVVSWNAMITG 213
+F M R+ +W +IKA R G +E ++G+ P L +++S A +
Sbjct: 287 VFDLMEDRDNATWRGMIKAYERKGFELEALDLFAQMQKQGVRPSFPSLISILSVCATLAS 346
Query: 214 YAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMM 273
R++ L + D+ + ++T + + GEL +A+ +F KD+I W S++
Sbjct: 347 LQYGRQVH--AHLVRCQFDDDVYVASVLMTMYVKCGELVKAKLVFDRFSSKDIIMWNSII 404
Query: 274 TGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQ-LISKTG 332
+GYA HGL EEALK+F +M ++G + PN T + +L ACS L EG +I + + SK
Sbjct: 405 SGYASHGLGEEALKIFHEMPSSGTM-PNKVTLIAILTACSYAGKLEEGLEIFESMESKFC 463
Query: 333 FQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHH 383
S ++M + G++ A ++ + ++ D W ++ A H
Sbjct: 464 VTPTVEHYSCTVDMLGRAGQVDKAMELIESMTIKP-DATVWGALLGACKTH 513
Score = 141 bits (356), Expect = 5e-32
Identities = 116/487 (23%), Positives = 203/487 (40%), Gaps = 122/487 (25%)
Query: 118 KLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKA 177
++ +I EA F + + SWN ++ GY NG ++A LF M +RN
Sbjct: 29 RIGKINEARKFFDSLQFKAIGSWNSIVSGYFSNGLPKEARQLFDEMSERN---------- 78
Query: 178 LARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMAS 237
VVSWN +++GY +NR + EA +FE MPER++ S
Sbjct: 79 --------------------------VVSWNGLVSGYIKNRMIVEARNVFELMPERNVVS 112
Query: 238 WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGG 297
W AM+ G+ Q G + AE LF +P+++ ++WT M G G ++A K++ M
Sbjct: 113 WTAMVKGYMQEGMVGEAESLFWRMPERNEVSWTVMFGGLIDDGRIDKARKLYDMMPVKDV 172
Query: 298 LKPNNGTFVTVLGACSGLASLTEGQQIHQLISK----------TGFQENTRV-------- 339
+ N ++G + E + I + + TG+++N RV
Sbjct: 173 VASTN-----MIGGLCREGRVDEARLIFDEMRERNVVTWTTMITGYRQNNRVDVARKLFE 227
Query: 340 ----------------------------------------VSALINMYSKCGELHIARKI 359
+A+I + + GE+ AR++
Sbjct: 228 VMPEKTEVSWTSMLLGYTLSGRIEDAEEFFEVMPMKPVIACNAMIVGFGEVGEISKARRV 287
Query: 360 FDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAG 419
FD L+ RD +W GMI AY G+ EA++LF +MQ+ G + + + + +L+ C+
Sbjct: 288 FD--LMEDRDNATWRGMIKAYERKGFELEALDLFAQMQKQGVRPSFPSLISILSVCATLA 345
Query: 420 LVDEGIQYFDKLLKNRSIQVKEDHY--ACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWG 477
+ G Q L++ Q +D Y + L+ + + G L +A + + K + +W
Sbjct: 346 SLQYGRQVHAHLVR---CQFDDDVYVASVLMTMYVKCGELVKAKLVFDRFSSK-DIIMWN 401
Query: 478 PLLAGCNVHGNADIGKLVAKKILKVEHE--------NAGTYSLLSNMYASVGKWKEAANV 529
+++G HG + ++ LK+ HE N T + + GK +E +
Sbjct: 402 SIISGYASHG-------LGEEALKIFHEMPSSGTMPNKVTLIAILTACSYAGKLEEGLEI 454
Query: 530 RMKMKDK 536
M+ K
Sbjct: 455 FESMESK 461
>gb|AAC33201.1| Hypothetical protein [Arabidopsis thaliana]
gi|15217508|ref|NP_172412.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|25345445|pir||D86227 hypothetical protein [imported]
- Arabidopsis thaliana
Length = 705
Score = 405 bits (1041), Expect = e-111
Identities = 213/555 (38%), Positives = 333/555 (59%), Gaps = 40/555 (7%)
Query: 62 ARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQ 121
ARKLFD+MP R+ W ++ GY+K G I EARK+FD +++VV+WT +V GYV +
Sbjct: 67 ARKLFDEMPDRNIISWNGLVSGYMKNGEIDEARKVFDLMP-ERNVVSWTALVKGYVHNGK 125
Query: 122 IEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARC 181
++ AESLF++MPE+N++SW +M+ G+ Q+G+I+ A L+ +P ++ ++ +I L +
Sbjct: 126 VDVAESLFWKMPEKNKVSWTVMLIGFLQDGRIDDACKLYEMIPDKDNIARTSMIHGLCKE 185
Query: 182 GRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAM 241
GR+++A+ M R+V++W M+TGY QN R+D+A ++F+ MPE+ SW +M
Sbjct: 186 GRVDEAREIFDE-----MSERSVITWTTMVTGYGQNNRVDDARKIFDVMPEKTEVSWTSM 240
Query: 242 LTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKM--------- 292
L G+ QNG + AE+LF +P K VI +M++G Q G +A ++F M
Sbjct: 241 LMGYVQNGRIEDAEELFEVMPVKPVIACNAMISGLGQKGEIAKARRVFDSMKERNDASWQ 300
Query: 293 ------QANG---------------GLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKT 331
+ NG G++P T +++L C+ LASL G+Q+H + +
Sbjct: 301 TVIKIHERNGFELEALDLFILMQKQGVRPTFPTLISILSVCASLASLHHGKQVHAQLVRC 360
Query: 332 GFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAIN 391
F + V S L+ MY KCGEL ++ IFD +D+I WN +I+ YA HG G EA+
Sbjct: 361 QFDVDVYVASVLMTMYIKCGELVKSKLIFDR--FPSKDIIMWNSIISGYASHGLGEEALK 418
Query: 392 LFNKMQELGF-QANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDL 450
+F +M G + N+VT+V L+ACS+AG+V+EG++ ++ + ++ HYAC+VD+
Sbjct: 419 VFCEMPLSGSTKPNEVTFVATLSACSYAGMVEEGLKIYESMESVFGVKPITAHYACMVDM 478
Query: 451 CGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTY 510
GRAGR EA +I+ + V+ +VWG LL C H D+ + AKK++++E EN+GTY
Sbjct: 479 LGRAGRFNEAMEMIDSMTVEPDAAVWGSLLGACRTHSQLDVAEFCAKKLIEIEPENSGTY 538
Query: 511 SLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGD-KSHSQSEML 569
LLSNMYAS G+W + A +R MK + ++K PGCSW EV N V F G SH + E +
Sbjct: 539 ILLSNMYASQGRWADVAELRKLMKTRLVRKSPGCSWTEVENKVHAFTRGGINSHPEQESI 598
Query: 570 EYLLLGLHTKMKKFG 584
+L L +++ G
Sbjct: 599 LKILDELDGLLREAG 613
Score = 202 bits (513), Expect = 3e-50
Identities = 121/407 (29%), Positives = 225/407 (54%), Gaps = 29/407 (7%)
Query: 86 KCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIG 145
+ G I EARKLFD D+K S+ +W +MV GY +A LF EMP+RN +SWN ++
Sbjct: 29 RIGKIHEARKLFDSCDSK-SISSWNSMVAGYFANLMPRDARKLFDEMPDRNIISWNGLVS 87
Query: 146 GYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQ---WHLIRCRKGMMPLR 202
GY +NG+I++A +F MP+RN+VSW ++K G+++ A+ W MP +
Sbjct: 88 GYMKNGEIDEARKVFDLMPERNVVSWTALVKGYVHNGKVDVAESLFW--------KMPEK 139
Query: 203 NVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELP 262
N VSW M+ G+ Q+ R+D+A +L+E +P++D + +M+ G + G ++ A ++F E+
Sbjct: 140 NKVSWTVMLIGFLQDGRIDDACKLYEMIPDKDNIARTSMIHGLCKEGRVDEAREIFDEMS 199
Query: 263 QKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQ 322
++ VITWT+M+TGY Q+ ++A K+F M + ++ ++L + + +
Sbjct: 200 ERSVITWTTMVTGYGQNNRVDDARKIFDVMP-----EKTEVSWTSMLMGYVQNGRIEDAE 254
Query: 323 QIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAH 382
++ +++ + +A+I+ + GE+ AR++FD +++R+ SW +I +
Sbjct: 255 ELFEVMP----VKPVIACNAMISGLGQKGEIAKARRVFDS--MKERNDASWQTVIKIHER 308
Query: 383 HGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKED 442
+G+ EA++LF MQ+ G + T + +L+ C+ + G Q +L++ Q D
Sbjct: 309 NGFELEALDLFILMQKQGVRPTFPTLISILSVCASLASLHHGKQVHAQLVR---CQFDVD 365
Query: 443 HY--ACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHG 487
Y + L+ + + G L ++ I + K + +W +++G HG
Sbjct: 366 VYVASVLMTMYIKCGELVKSKLIFDRFPSK-DIIMWNSIISGYASHG 411
Score = 117 bits (292), Expect = 1e-24
Identities = 66/258 (25%), Positives = 141/258 (54%), Gaps = 20/258 (7%)
Query: 172 NIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP 231
N+ I L+R G+I +A+ C +++ SWN+M+ GY N +A +LF+ MP
Sbjct: 21 NVRITHLSRIGKIHEARKLFDSCDS-----KSISSWNSMVAGYFANLMPRDARKLFDEMP 75
Query: 232 ERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTK 291
+R++ SWN +++G+ +NGE++ A K+F +P+++V++WT+++ GY +G + A +F K
Sbjct: 76 DRNIISWNGLVSGYMKNGEIDEARKVFDLMPERNVVSWTALVKGYVHNGKVDVAESLFWK 135
Query: 292 MQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCG 351
M + N ++ +L + + +++++I ++ +++I+ K G
Sbjct: 136 MP-----EKNKVSWTVMLIGFLQDGRIDDACKLYEMIP----DKDNIARTSMIHGLCKEG 186
Query: 352 ELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVEL 411
+ AR+IFD+ + +R +I+W M+ Y + ++A +F+ M E +V++ +
Sbjct: 187 RVDEAREIFDE--MSERSVITWTTMVTGYGQNNRVDDARKIFDVMPE----KTEVSWTSM 240
Query: 412 LTACSHAGLVDEGIQYFD 429
L G +++ + F+
Sbjct: 241 LMGYVQNGRIEDAEELFE 258
>ref|XP_475075.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|45680436|gb|AAS75237.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 661
Score = 377 bits (969), Expect = e-103
Identities = 201/529 (37%), Positives = 317/529 (58%), Gaps = 14/529 (2%)
Query: 55 KEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVN 114
K G+ D AR+LFD M ++ WT M+ GY + G + EAR+LFD + +V +WTTMV
Sbjct: 135 KAGQVDRARRLFDGMAVKNVVAWTCMVSGYCRAGLVDEARRLFDLMPYR-NVFSWTTMVQ 193
Query: 115 GYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNII 174
GY + EA +F +MPERN ++W +M+ Y NG I++AL+LF RMP+ N SWN +
Sbjct: 194 GYAHNGMLREAREMFDKMPERNVVAWTVMVKAYVDNGCIQEALELFNRMPQMNSYSWNAM 253
Query: 175 IKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERD 234
L G+++DA + MP +NVVSW M+TG AQN + A E F+RMP++D
Sbjct: 254 ATGLMSAGKVDDAVQLFDK-----MPHKNVVSWTIMVTGLAQNGFVSRAREFFDRMPKKD 308
Query: 235 MASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQA 294
+ +WN+M+T + +G++N A++LF +P K+++TW ++ GY+ + L +EAL++F M
Sbjct: 309 IPAWNSMITAYTNDGQVNDAQRLFDIMPSKNLVTWNIIIDGYSMNNLKDEALRLFLLM-L 367
Query: 295 NGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELH 354
+ P++ T ++VL S E +QIH L +K G+Q T + + L++MYS+ G+L
Sbjct: 368 RSAVSPDSTTLISVLVVSE---STMEVRQIHGLSTKLGYQPETNLGNTLVSMYSRSGDLS 424
Query: 355 IARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTA 414
A F L ++D I+W MI A A+HG A+ F +M G++ + T+ +L+A
Sbjct: 425 SAWLAF--RRLNEKDAITWTSMIQALANHGCAPCALQGFAQMLRCGYKPSSTTFTAVLSA 482
Query: 415 CSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKL-SL 473
C+H GLV++G + F + + +HY+CLVDL GRAG ++EA +++G+ +
Sbjct: 483 CNHVGLVEKGRKIFKSISHVYGLGPTIEHYSCLVDLLGRAGYVREAKEVVDGMQRDMCDE 542
Query: 474 SVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKM 533
++ G LL C H ++G+ V + ++K++ +G Y+LL+N++AS G W E A+V M
Sbjct: 543 AILGTLLGSCMTHNEVEVGRAVGEDLVKIDPSGSGGYTLLANVFASGGMWNEVASVWKIM 602
Query: 534 KDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQ-SEMLEYLLLGLHTKMK 581
K +KK PG S IEV VF D+ HSQ +E+ E L L +MK
Sbjct: 603 KGSKVKKTPGFSQIEVNARNHVFYSRDQMHSQRTEIYEMLNSRLVPQMK 651
Score = 92.4 bits (228), Expect = 4e-17
Identities = 67/248 (27%), Positives = 120/248 (48%), Gaps = 30/248 (12%)
Query: 210 MITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITW 269
M+ GY + ++D A LF+ M +++ +W M++G+ + G ++ A +LF +P ++V +W
Sbjct: 129 MLDGYVKAGQVDRARRLFDGMAVKNVVAWTCMVSGYCRAGLVDEARRLFDLMPYRNVFSW 188
Query: 270 TSMMTGYAQHGLSEEALKMFTKMQANGGLK--------PNNGTFVTVL------------ 309
T+M+ GYA +G+ EA +MF KM + +NG L
Sbjct: 189 TTMVQGYAHNGMLREAREMFDKMPERNVVAWTVMVKAYVDNGCIQEALELFNRMPQMNSY 248
Query: 310 ---GACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLR 366
+GL S + QL K +N + ++ ++ G + AR+ FD +
Sbjct: 249 SWNAMATGLMSAGKVDDAVQLFDKMP-HKNVVSWTIMVTGLAQNGFVSRAREFFD--RMP 305
Query: 367 QRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQ 426
++D+ +WN MI AY + G N+A LF+ M N VT+ ++ S L DE ++
Sbjct: 306 KKDIPAWNSMITAYTNDGQVNDAQRLFDIMP----SKNLVTWNIIIDGYSMNNLKDEALR 361
Query: 427 YFDKLLKN 434
F +L++
Sbjct: 362 LFLLMLRS 369
>dbj|BAD38052.1| putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa (japonica cultivar-group)]
Length = 922
Score = 374 bits (961), Expect = e-102
Identities = 209/565 (36%), Positives = 314/565 (54%), Gaps = 39/565 (6%)
Query: 60 DHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKL 119
D A K F+ M +R+++ W+TMI G I A ++++ D KS+ T ++ G +
Sbjct: 277 DTAIKFFESMIERNEYTWSTMIAALSHGGRIDAAIAVYER-DPVKSIACRTALITGLAQC 335
Query: 120 NQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALA 179
+I++A LF ++PE +SWN +I GY QNG + +A +LF +MP RN +SW +I A
Sbjct: 336 GRIDDARILFEQIPEPIVVSWNALITGYMQNGMVNEAKELFDKMPFRNTISWAGMIAGYA 395
Query: 180 RCGRIEDAQWHLIRC-RKGMMP-----------LRNVVSW-------------------- 207
+ GR E+A L R GM+P N+V+
Sbjct: 396 QNGRSEEALGLLQELHRSGMLPSLSSLTSIFFACSNIVALETGTQVHSLAVKVGCQFNSF 455
Query: 208 --NAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKD 265
NA+IT Y + R ++ A ++F RM +D+ SWN+ L QN L+ A F + +D
Sbjct: 456 ACNALITMYGKCRNMEYARQVFSRMVTKDIVSWNSFLAALVQNDLLDEARNTFDNMLSRD 515
Query: 266 VITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIH 325
++WT++++ YA S EA+ F M L PN+ +LG C L + GQQIH
Sbjct: 516 DVSWTTIISAYAHAEQSNEAMGAFKTMFCEHEL-PNSPILTILLGVCGSLGASKIGQQIH 574
Query: 326 QLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGY 385
+ K G V +ALI+MY KCG +R+IFD L+ +RD+ +WN +I YA HG
Sbjct: 575 TVAIKLGMDSELIVANALISMYFKCGCAD-SRRIFD--LMEERDIFTWNTIITGYAQHGL 631
Query: 386 GNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYA 445
G EAI ++ M+ G N+VT+V LL ACSHAGLVDEG ++F + ++ + +HYA
Sbjct: 632 GREAIKMYQHMESAGVLPNEVTFVGLLNACSHAGLVDEGWKFFKSMSQDYGLTPLPEHYA 691
Query: 446 CLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHE 505
C+VDL GR G ++ A I + ++ +W LL C +H NA+IGK A+K+ ++E
Sbjct: 692 CMVDLLGRTGDVQGAEQFIYDMPIEPDTVIWSALLGACKIHKNAEIGKRAAEKLFRIEPS 751
Query: 506 NAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQ 565
NAG Y +LSN+Y+S+G W E A VR MK +G+ K+PGCSW ++ + + FV GDK H Q
Sbjct: 752 NAGNYVMLSNIYSSLGMWGEVAEVRKIMKQQGVIKEPGCSWTQIKDKMHSFVTGDKQHEQ 811
Query: 566 SEMLEYLLLGLHTKMKKFGDILDDD 590
E + L L+T +K G + D +
Sbjct: 812 IEEIVATLEELYTLLKATGYVPDTE 836
Score = 202 bits (513), Expect = 3e-50
Identities = 161/606 (26%), Positives = 269/606 (43%), Gaps = 117/606 (19%)
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F P ++ NS IS C G D AR L+D + + ++ GY + G + EAR
Sbjct: 57 FDAMPRRDIIAWNSMISAYCHNGMPDAARDLYDAISGGNMRTGAILLSGYGRLGRVLEAR 116
Query: 95 KLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIE 154
++FD +++ V W M++ YV+ I A LF MP R+ SWN M+ GY + Q+
Sbjct: 117 RVFDGM-LERNTVAWNAMISCYVQNGDITMARRLFDAMPSRDVSSWNSMLTGYCHSLQMV 175
Query: 155 KALDLFRRMPKRNLVSWNIIIKALARCGRIED--AQWHLI--RCRKGMMP---------- 200
A +LF +MP+RNLVSW ++I GRIE+ W + R+G++P
Sbjct: 176 DARNLFEKMPERNLVSWTVMISGY---GRIENHGKAWDIFCKMHREGLLPDQSNFASALS 232
Query: 201 -----------------------LRNVVSWNAMITGYAQNRR-LDEALELFERMPERDMA 236
R+VV A++ Y+++ LD A++ FE M ER+
Sbjct: 233 AVKGLGNLDVLESLRVLALKTGFERDVVIGTAILNVYSRDTSVLDTAIKFFESMIERNEY 292
Query: 237 SWN-------------------------------AMLTGFFQNGELNRAEKLFAELPQKD 265
+W+ A++TG Q G ++ A LF ++P+
Sbjct: 293 TWSTMIAALSHGGRIDAAIAVYERDPVKSIACRTALITGLAQCGRIDDARILFEQIPEPI 352
Query: 266 VITWTSMMTGYAQHGLSEEALKMFTKM---------------QANG-------------- 296
V++W +++TGY Q+G+ EA ++F KM NG
Sbjct: 353 VVSWNALITGYMQNGMVNEAKELFDKMPFRNTISWAGMIAGYAQNGRSEEALGLLQELHR 412
Query: 297 -GLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHI 355
G+ P+ + ++ ACS + +L G Q+H L K G Q N+ +ALI MY KC +
Sbjct: 413 SGMLPSLSSLTSIFFACSNIVALETGTQVHSLAVKVGCQFNSFACNALITMYGKCRNMEY 472
Query: 356 ARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTAC 415
AR++F + +D++SWN +AA + +EA N F+ M +DV++ +++A
Sbjct: 473 ARQVF--SRMVTKDIVSWNSFLAALVQNDLLDEARNTFDNM----LSRDDVSWTTIISAY 526
Query: 416 SHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLK---EAFYIIEGLGVKLS 472
+HA +E + F + + L+ +CG G K + + LG+
Sbjct: 527 AHAEQSNEAMGAFKTMFCEHELP-NSPILTILLGVCGSLGASKIGQQIHTVAIKLGMDSE 585
Query: 473 LSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMK 532
L V L++ G AD ++ +E + T++ + YA G +EA +
Sbjct: 586 LIVANALISMYFKCGCADSRRIFD----LMEERDIFTWNTIITGYAQHGLGREAIKMYQH 641
Query: 533 MKDKGL 538
M+ G+
Sbjct: 642 MESAGV 647
Score = 174 bits (441), Expect = 7e-42
Identities = 131/454 (28%), Positives = 228/454 (49%), Gaps = 47/454 (10%)
Query: 118 KLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKA 177
+L ++ EA +F MP R+ ++WN MI Y NG + A DL+ + N+ + I++
Sbjct: 46 RLGRVGEAREVFDAMPRRDIIAWNSMISAYCHNGMPDAARDLYDAISGGNMRTGAILLSG 105
Query: 178 LARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMAS 237
R GR+ +A+ R GM+ RN V+WNAMI+ Y QN + A LF+ MP RD++S
Sbjct: 106 YGRLGRVLEAR----RVFDGMLE-RNTVAWNAMISCYVQNGDITMARRLFDAMPSRDVSS 160
Query: 238 WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGG 297
WN+MLTG+ + ++ A LF ++P++++++WT M++GY + +A +F KM G
Sbjct: 161 WNSMLTGYCHSLQMVDARNLFEKMPERNLVSWTVMISGYGRIENHGKAWDIFCKMHRE-G 219
Query: 298 LKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSK-CGELHIA 356
L P+ F + L A GL +L + + L KTGF+ + + +A++N+YS+ L A
Sbjct: 220 LLPDQSNFASALSAVKGLGNLDVLESLRVLALKTGFERDVVIGTAILNVYSRDTSVLDTA 279
Query: 357 RKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNK---------------MQELGF 401
K F+ + +R+ +W+ MIAA +H G + AI ++ + + + G
Sbjct: 280 IKFFES--MIERNEYTWSTMIAALSHGGRIDAAIAVYERDPVKSIACRTALITGLAQCG- 336
Query: 402 QAND-------------VTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLV 448
+ +D V++ L+T G+V+E + FDK+ +I +A ++
Sbjct: 337 RIDDARILFEQIPEPIVVSWNALITGYMQNGMVNEAKELFDKMPFRNTIS-----WAGMI 391
Query: 449 DLCGRAGRLKEAFYIIEGL---GVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHE 505
+ GR +EA +++ L G+ SLS + C+ + G V +KV +
Sbjct: 392 AGYAQNGRSEEALGLLQELHRSGMLPSLSSLTSIFFACSNIVALETGTQVHSLAVKVGCQ 451
Query: 506 -NAGTYSLLSNMYASVGKWKEAANVRMKMKDKGL 538
N+ + L MY + A V +M K +
Sbjct: 452 FNSFACNALITMYGKCRNMEYARQVFSRMVTKDI 485
Score = 156 bits (395), Expect = 2e-36
Identities = 131/478 (27%), Positives = 227/478 (47%), Gaps = 55/478 (11%)
Query: 42 EMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPD 101
E+ C++ I L + GR AR++FD MP+RD W +MI Y G AR L+D
Sbjct: 33 EVSGCSARIRDLGRLGRVGEAREVFDAMPRRDIIAWNSMISAYCHNGMPDAARDLYDAIS 92
Query: 102 AKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFR 161
++ T +++GY +L ++ EA +F M ERN ++WN MI Y QNG I A LF
Sbjct: 93 G-GNMRTGAILLSGYGRLGRVLEARRVFDGMLERNTVAWNAMISCYVQNGDITMARRLFD 151
Query: 162 RMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLD 221
MP R++ SWN ++ ++ DA R MP RN+VSW MI+GY +
Sbjct: 152 AMPSRDVSSWNSMLTGYCHSLQMVDA-----RNLFEKMPERNLVSWTVMISGYGRIENHG 206
Query: 222 EALELFERMPER----DMASWNAMLTGFFQNGELNRAEKL----FAELPQKDVITWTSMM 273
+A ++F +M D +++ + L+ G L+ E L ++DV+ T+++
Sbjct: 207 KAWDIFCKMHREGLLPDQSNFASALSAVKGLGNLDVLESLRVLALKTGFERDVVIGTAIL 266
Query: 274 TGYAQH-GLSEEALKMFTKMQANGGLKPNNGTFVTVLGACS------GLASLTEGQQIHQ 326
Y++ + + A+K F M ++ N T+ T++ A S ++ E +
Sbjct: 267 NVYSRDTSVLDTAIKFFESM-----IERNEYTWSTMIAALSHGGRIDAAIAVYERDPVKS 321
Query: 327 LISKT---------GFQENTRVV------------SALINMYSKCGELHIARKIFDDGLL 365
+ +T G ++ R++ +ALI Y + G ++ A+++FD +
Sbjct: 322 IACRTALITGLAQCGRIDDARILFEQIPEPIVVSWNALITGYMQNGMVNEAKELFDK--M 379
Query: 366 RQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGI 425
R+ ISW GMIA YA +G EA+ L ++ G + + + ACS+ ++ G
Sbjct: 380 PFRNTISWAGMIAGYAQNGRSEEALGLLQELHRSGMLPSLSSLTSIFFACSNIVALETGT 439
Query: 426 QYFDKLLKNRSIQVKEDHYAC--LVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLA 481
Q +K + + + +AC L+ + G+ ++ A + + K +S W LA
Sbjct: 440 QVHSLAVK---VGCQFNSFACNALITMYGKCRNMEYARQVFSRMVTKDIVS-WNSFLA 493
Score = 76.6 bits (187), Expect = 2e-12
Identities = 46/167 (27%), Positives = 88/167 (52%), Gaps = 11/167 (6%)
Query: 232 ERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTK 291
E +++ +A + + G + A ++F +P++D+I W SM++ Y +G+ + A ++
Sbjct: 31 ELEVSGCSARIRDLGRLGRVGEAREVFDAMPRRDIIAWNSMISAYCHNGMPDAARDLYDA 90
Query: 292 MQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCG 351
+ +GG N T +L L + E +++ + + NT +A+I+ Y + G
Sbjct: 91 I--SGG---NMRTGAILLSGYGRLGRVLEARRVFDGM----LERNTVAWNAMISCYVQNG 141
Query: 352 ELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQE 398
++ +AR++FD + RD+ SWN M+ Y H +A NLF KM E
Sbjct: 142 DITMARRLFD--AMPSRDVSSWNSMLTGYCHSLQMVDARNLFEKMPE 186
>ref|XP_478933.1| pentatricopeptide (PPR) repeat-containing protein-like protein
[Oryza sativa (japonica cultivar-group)]
gi|50509373|dbj|BAD30928.1| pentatricopeptide (PPR)
repeat-containing protein-like protein [Oryza sativa
(japonica cultivar-group)] gi|34393605|dbj|BAC83258.1|
pentatricopeptide (PPR) repeat-containing protein-like
protein [Oryza sativa (japonica cultivar-group)]
Length = 808
Score = 372 bits (955), Expect = e-101
Identities = 192/516 (37%), Positives = 319/516 (61%), Gaps = 16/516 (3%)
Query: 84 YIKCG---AIKEARKLFDQ-PDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELS 139
Y+KC A +ARK+ D+ PD K +TWTTMV GYV+ + A S+F E+ + ++
Sbjct: 210 YMKCDTPEASWDARKVLDEMPD--KDDLTWTTMVVGYVRRGDVNAARSVFEEVDGKFDVV 267
Query: 140 WNIMIGGYGQNGQIEKALDLFRRMPKRNL----VSWNIIIKALARCGRI---EDAQWHLI 192
WN MI GY Q+G A +LFRRM + ++ ++ A A G + +I
Sbjct: 268 WNAMISGYVQSGMCADAFELFRRMVSEKVPLDEFTFTSVLSACANAGFFVHGKSVHGQII 327
Query: 193 RCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELN 252
R + +P + NA++T Y++ ++ A +F+ M +D+ SWN +L+G+ +G L+
Sbjct: 328 RLQPNFVPEAALPVNNALVTLYSKGGKIVIAKRIFDTMNLKDVVSWNTILSGYIDSGCLD 387
Query: 253 RAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGAC 312
+A ++F +P K+ ++W M++GY GLSE+ALK+F +M+A +KP + T+ + AC
Sbjct: 388 KAVEVFKVMPYKNDLSWMVMVSGYVHGGLSEDALKLFNQMRAED-VKPCDYTYAGAIAAC 446
Query: 313 SGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLIS 372
L +L G+Q+H + + GF+ + +AL+ MY+KCG ++ AR +F ++ D +S
Sbjct: 447 GELGALKHGRQLHAHLVQCGFEASNSAGNALLTMYAKCGAVNDARLVFL--VMPNLDSVS 504
Query: 373 WNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLL 432
WN MI+A HG+G EA+ LF++M G + ++++ +LTAC+HAGLVDEG YF+ +
Sbjct: 505 WNAMISALGQHGHGREALELFDQMVAEGIDPDRISFLTILTACNHAGLVDEGFHYFESMK 564
Query: 433 KNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIG 492
++ I EDHYA L+DL GR+GR+ EA +I+ + + + S+W +L+GC +G+ + G
Sbjct: 565 RDFGISPGEDHYARLIDLLGRSGRIGEARDLIKTMPFEPTPSIWEAILSGCRTNGDMEFG 624
Query: 493 KLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNT 552
A ++ ++ ++ GTY LLSN Y++ G+W +AA VR M+D+G+KK+PGCSWIEVG+
Sbjct: 625 AYAADQLFRMIPQHDGTYILLSNTYSAAGRWVDAARVRKLMRDRGVKKEPGCSWIEVGSK 684
Query: 553 VQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILD 588
+ VF+VGD H +++ + L + +M+K G + D
Sbjct: 685 IHVFLVGDTKHPEAQEVYQFLEVIGARMRKLGYVPD 720
Score = 111 bits (277), Expect = 8e-23
Identities = 100/438 (22%), Positives = 194/438 (43%), Gaps = 66/438 (15%)
Query: 55 KEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKK---SVVTWTT 111
+ G + AR +F+++ + +W MI GY++ G +A +LF + ++K T+T+
Sbjct: 246 RRGDVNAARSVFEEVDGKFDVVWNAMISGYVQSGMCADAFELFRRMVSEKVPLDEFTFTS 305
Query: 112 MVNGYVKLNQIEEAESLFYE--------MPERNELSWNIMIGGYGQNGQIEKALDLFRRM 163
+++ +S+ + +PE N ++ Y + G+I A +F M
Sbjct: 306 VLSACANAGFFVHGKSVHGQIIRLQPNFVPEAALPVNNALVTLYSKGGKIVIAKRIFDTM 365
Query: 164 PKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEA 223
+++VSWN I+ G ++ A + K +MP +N +SW M++GY ++A
Sbjct: 366 NLKDVVSWNTILSGYIDSGCLDKA----VEVFK-VMPYKNDLSWMVMVSGYVHGGLSEDA 420
Query: 224 LELFERMPERDM---------------------------------------ASWNAMLTG 244
L+LF +M D+ ++ NA+LT
Sbjct: 421 LKLFNQMRAEDVKPCDYTYAGAIAACGELGALKHGRQLHAHLVQCGFEASNSAGNALLTM 480
Query: 245 FFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGT 304
+ + G +N A +F +P D ++W +M++ QHG EAL++F +M A G+ P+ +
Sbjct: 481 YAKCGAVNDARLVFLVMPNLDSVSWNAMISALGQHGHGREALELFDQMVAE-GIDPDRIS 539
Query: 305 FVTVLGACSGLASLTEGQQIHQLISKT-GFQENTRVVSALINMYSKCGELHIARKIFDDG 363
F+T+L AC+ + EG + + + G + LI++ + G + AR +
Sbjct: 540 FLTILTACNHAGLVDEGFHYFESMKRDFGISPGEDHYARLIDLLGRSGRIGEARDLIKTM 599
Query: 364 LLRQRDLISWNGMIAAYAHHG---YGNEAINLFNKMQELGFQANDVTYVELLTACSHAGL 420
I W +++ +G +G A + +M +D TY+ L S AG
Sbjct: 600 PFEPTPSI-WEAILSGCRTNGDMEFGAYAADQLFRM----IPQHDGTYILLSNTYSAAGR 654
Query: 421 VDEGIQYFDKLLKNRSIQ 438
+ + KL+++R ++
Sbjct: 655 WVDAAR-VRKLMRDRGVK 671
Score = 84.7 bits (208), Expect = 8e-15
Identities = 73/315 (23%), Positives = 136/315 (43%), Gaps = 44/315 (13%)
Query: 39 PNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFD 98
P + + N+ ++ K G+ A+++FD M +D W T++ GYI G + +A ++F
Sbjct: 335 PEAALPVNNALVTLYSKGGKIVIAKRIFDTMNLKDVVSWNTILSGYIDSGCLDKAVEVFK 394
Query: 99 QPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERN----ELSWNIMIGGYGQNGQIE 154
K ++W MV+GYV E+A LF +M + + ++ I G+ G ++
Sbjct: 395 VMPYKND-LSWMVMVSGYVHGGLSEDALKLFNQMRAEDVKPCDYTYAGAIAACGELGALK 453
Query: 155 KALDLFRRMP----KRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAM 210
L + + + + N ++ A+CG + DA+ + +MP + VSWNAM
Sbjct: 454 HGRQLHAHLVQCGFEASNSAGNALLTMYAKCGAVNDARLVFL-----VMPNLDSVSWNAM 508
Query: 211 ITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWT 270
I+ Q+ EALELF++M AE D I++
Sbjct: 509 ISALGQHGHGREALELFDQM---------------------------VAEGIDPDRISFL 541
Query: 271 SMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISK 330
+++T GL +E F M+ + G+ P + ++ + E + LI
Sbjct: 542 TILTACNHAGLVDEGFHYFESMKRDFGISPGEDHYARLIDLLGRSGRIGEAR---DLIKT 598
Query: 331 TGFQENTRVVSALIN 345
F+ + A+++
Sbjct: 599 MPFEPTPSIWEAILS 613
Score = 84.0 bits (206), Expect = 1e-14
Identities = 84/380 (22%), Positives = 164/380 (43%), Gaps = 48/380 (12%)
Query: 226 LFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELP--QKDVITWTSMMTGYAQHGLSE 283
LF P+ + +++ G L A F +P ++D + +MM+ +A+ L+
Sbjct: 85 LFRSDPDPGPVAATSLVAAHAAAGRLRDAAAFFDAVPPARRDTVLHNAMMSAFARASLAA 144
Query: 284 EALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQ--QIHQLISKTGFQENTRVVS 341
A+ +F + +G L+P++ +F ++ A + +L Q+H + K+G V +
Sbjct: 145 PAVSVFHALLGSGSLRPDDYSFTALISAVGQMHNLAAPHCTQLHCSVLKSGAAAVLSVSN 204
Query: 342 ALINMYSKCGELHI---ARKIFDD--------------GLLRQRDL-------------- 370
ALI +Y KC ARK+ D+ G +R+ D+
Sbjct: 205 ALIALYMKCDTPEASWDARKVLDEMPDKDDLTWTTMVVGYVRRGDVNAARSVFEEVDGKF 264
Query: 371 -ISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFD 429
+ WN MI+ Y G +A LF +M ++ T+ +L+AC++AG G
Sbjct: 265 DVVWNAMISGYVQSGMCADAFELFRRMVSEKVPLDEFTFTSVLSACANAGFFVHGKSVHG 324
Query: 430 KLLKNRSIQVKEDHYA---CLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVH 486
++++ + V E LV L + G++ A I + + +K +S W +L+G
Sbjct: 325 QIIRLQPNFVPEAALPVNNALVTLYSKGGKIVIAKRIFDTMNLKDVVS-WNTILSGYIDS 383
Query: 487 GNADIGKLVAKKILKV-EHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCS 545
G D A ++ KV ++N ++ ++ + Y G ++A + +M+ + +K C
Sbjct: 384 GCLD----KAVEVFKVMPYKNDLSWMVMVSGYVHGGLSEDALKLFNQMRAEDVKP---CD 436
Query: 546 WIEVGNTVQVFVVGDKSHSQ 565
+ G +G H +
Sbjct: 437 YTYAGAIAACGELGALKHGR 456
Score = 83.6 bits (205), Expect = 2e-14
Identities = 82/341 (24%), Positives = 145/341 (42%), Gaps = 60/341 (17%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQ---PDAK 103
N+ +S G D A ++F MP ++ W M+ GY+ G ++A KLF+Q D K
Sbjct: 374 NTILSGYIDSGCLDKAVEVFKVMPYKNDLSWMVMVSGYVHGGLSEDALKLFNQMRAEDVK 433
Query: 104 KSVVTW-----------------------------------TTMVNGYVKLNQIEEAESL 128
T+ ++ Y K + +A +
Sbjct: 434 PCDYTYAGAIAACGELGALKHGRQLHAHLVQCGFEASNSAGNALLTMYAKCGAVNDARLV 493
Query: 129 FYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNL----VSWNIIIKALARCGRI 184
F MP + +SWN MI GQ+G +AL+LF +M + +S+ I+ A G +
Sbjct: 494 FLVMPNLDSVSWNAMISALGQHGHGREALELFDQMVAEGIDPDRISFLTILTACNHAGLV 553
Query: 185 EDAQWHLIRCRK---GMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP-ERDMASWNA 240
++ +H K G+ P + + +I ++ R+ EA +L + MP E + W A
Sbjct: 554 DEG-FHYFESMKRDFGISPGED--HYARLIDLLGRSGRIGEARDLIKTMPFEPTPSIWEA 610
Query: 241 MLTGFFQNGELN----RAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANG 296
+L+G NG++ A++LF +PQ D T+ + Y+ G +A ++ K+ +
Sbjct: 611 ILSGCRTNGDMEFGAYAADQLFRMIPQHDG-TYILLSNTYSAAGRWVDAARV-RKLMRDR 668
Query: 297 GLKPNNGTFVTVLGA-----CSGLASLTEGQQIHQLISKTG 332
G+K G +G+ G E Q+++Q + G
Sbjct: 669 GVKKEPGCSWIEVGSKIHVFLVGDTKHPEAQEVYQFLEVIG 709
>dbj|BAD37476.1| pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa (japonica cultivar-group)]
gi|51535392|dbj|BAD37262.1| pentatricopeptide (PPR)
repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 908
Score = 365 bits (937), Expect = 2e-99
Identities = 202/574 (35%), Positives = 317/574 (55%), Gaps = 30/574 (5%)
Query: 13 LMHAPKLNTHPTYILRTRLASTFSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQR 72
L+ A + HP ++ R F P ++ N+ + + G D AR+LFD+MPQR
Sbjct: 104 LLAAYSASPHPDHLAAAR--RLFDEMPQRDVVTWNTLLGAYARRGLMDEARRLFDEMPQR 161
Query: 73 DKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEM 132
+ W TM+ G+ G + +A +FD AK S + +TMV+G+ K + EAE L +
Sbjct: 162 NAASWNTMVTGFFAAGQVVKALDVFDAMPAKDSA-SLSTMVSGFTKNGMLHEAEELLTKR 220
Query: 133 PERNEL-----SWNIMIGGYGQNGQIEKALDLFRRMPK--------------RNLVSWNI 173
++ ++N +I YGQ G+ A LF +PK RN+VSWN
Sbjct: 221 LSVTDMDKAVDAYNTLIVAYGQAGRFSDAKRLFDMIPKGQYQHNMLKRKGFERNVVSWNS 280
Query: 174 IIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPER 233
++ + G + A R MP +++VSWN MI+GY Q + E+ +LF MP+
Sbjct: 281 MMICYIKAGDVCSA-----RALFNEMPDKDLVSWNTMISGYTQASDMKESEKLFWEMPDP 335
Query: 234 DMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQ 293
D SWN ++ GF Q GE A F +P++ I+W +M++GY ++G ++K+F+KM
Sbjct: 336 DTVSWNLIIQGFMQKGEAEHARGFFDRMPERGTISWNTMISGYEKNGNYISSVKLFSKML 395
Query: 294 ANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGEL 353
G + P+ TF +VL AC+ + L G QIHQL+ K+ F +T + +ALI MYS+CG L
Sbjct: 396 EVGEI-PDRHTFSSVLAACASIPMLGLGAQIHQLVEKS-FVPDTAISNALITMYSRCGAL 453
Query: 354 HIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLT 413
+ A IF + ++DL+SWN +I Y HHG +A+ LF +M+ +T+V LL+
Sbjct: 454 NDAEAIFKQ-MHTKKDLVSWNALIGCYEHHGRATKALQLFKEMRRAKVMPTHITFVSLLS 512
Query: 414 ACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSL 473
AC +AGLV EG FD ++ I + +HYA LV+L GR G+L +A +I + +
Sbjct: 513 ACVNAGLVSEGRMVFDTMVHEYGIVARIEHYAALVNLIGRHGQLDDALEVINSMPMAPDR 572
Query: 474 SVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKM 533
SVWG L C N + ++ AK++ + +++ Y L+ N++A GKW AA VR +M
Sbjct: 573 SVWGAFLGACTAKKNEPLAQMAAKELSTINPDSSAPYVLIHNLHAHEGKWGSAAVVREEM 632
Query: 534 KDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSE 567
+ +G+ KQPG SWI++ + VF+ GD H ++
Sbjct: 633 ERQGIYKQPGYSWIDLEGKMHVFISGDTWHPNAQ 666
Score = 153 bits (386), Expect = 2e-35
Identities = 91/325 (28%), Positives = 169/325 (52%), Gaps = 22/325 (6%)
Query: 125 AESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRI 184
A LF +P R+ ++WN ++ G + A + F MP R+ VSWN ++ A +
Sbjct: 55 ARRLFDALPARSVVTWNSLLAGLARRPDARAAREFFDAMPVRDAVSWNTLLAAYSASPHP 114
Query: 185 EDAQWHLIRCRK--GMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAML 242
+ HL R+ MP R+VV+WN ++ YA+ +DEA LF+ MP+R+ ASWN M+
Sbjct: 115 D----HLAAARRLFDEMPQRDVVTWNTLLGAYARRGLMDEARRLFDEMPQRNAASWNTMV 170
Query: 243 TGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNN 302
TGFF G++ +A +F +P KD + ++M++G+ ++G+ EA ++ TK + +
Sbjct: 171 TGFFAAGQVVKALDVFDAMPAKDSASLSTMVSGFTKNGMLHEAEELLTKRLSVTDMDKAV 230
Query: 303 GTFVTVLGACSGLASLTE----------GQQIHQLISKTGFQENTRVVSALINMYSKCGE 352
+ T++ A ++ GQ H ++ + GF+ N ++++ Y K G+
Sbjct: 231 DAYNTLIVAYGQAGRFSDAKRLFDMIPKGQYQHNMLKRKGFERNVVSWNSMMICYIKAGD 290
Query: 353 LHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELL 412
+ AR +F++ + +DL+SWN MI+ Y E+ LF +M + + V++ ++
Sbjct: 291 VCSARALFNE--MPDKDLVSWNTMISGYTQASDMKESEKLFWEMPD----PDTVSWNLII 344
Query: 413 TACSHAGLVDEGIQYFDKLLKNRSI 437
G + +FD++ + +I
Sbjct: 345 QGFMQKGEAEHARGFFDRMPERGTI 369
Score = 100 bits (250), Expect = 1e-19
Identities = 78/277 (28%), Positives = 132/277 (47%), Gaps = 32/277 (11%)
Query: 172 NIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP 231
N + AL R GR A R +P R+VV+WN+++ G A+ A E F+ MP
Sbjct: 40 NRSLAALLRAGRYGAA-----RRLFDALPARSVVTWNSLLAGLARRPDARAAREFFDAMP 94
Query: 232 ERDMASWNAMLTGFFQN---GELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKM 288
RD SWN +L + + L A +LF E+PQ+DV+TW +++ YA+ GL +EA ++
Sbjct: 95 VRDAVSWNTLLAAYSASPHPDHLAAARRLFDEMPQRDVVTWNTLLGAYARRGLMDEARRL 154
Query: 289 FTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQL-ISKTGFQENTRVVSALINMY 347
F +M N T VT A GQ + L + +++ +S +++ +
Sbjct: 155 FDEMPQRNAASWN--TMVTGFFAA--------GQVVKALDVFDAMPAKDSASLSTMVSGF 204
Query: 348 SKCGELHIARKIFDDGLL---RQRDLISWNGMIAAYAHHGYGNEAINLF----------N 394
+K G LH A ++ L + + ++N +I AY G ++A LF N
Sbjct: 205 TKNGMLHEAEELLTKRLSVTDMDKAVDAYNTLIVAYGQAGRFSDAKRLFDMIPKGQYQHN 264
Query: 395 KMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKL 431
++ GF+ N V++ ++ AG V F+++
Sbjct: 265 MLKRKGFERNVVSWNSMMICYIKAGDVCSARALFNEM 301
>ref|NP_173907.1| pentatricopeptide (PPR) repeat-containing protein [Arabidopsis
thaliana] gi|11067273|gb|AAG28801.1| hypothetical
protein [Arabidopsis thaliana] gi|25345466|pir||E86383
hypothetical protein [imported] - Arabidopsis thaliana
Length = 790
Score = 348 bits (894), Expect = 2e-94
Identities = 192/528 (36%), Positives = 311/528 (58%), Gaps = 18/528 (3%)
Query: 80 MIDGYIKCGA----IKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPER 135
++ Y KC + + ARK+FD+ +K +WTTM+ GYVK + E L M +
Sbjct: 190 LVSVYSKCASSPSLLHSARKVFDEI-LEKDERSWTTMMTGYVKNGYFDLGEELLEGMDDN 248
Query: 136 NEL-SWNIMIGGYGQNGQIEKALDLFRRMPKRNL----VSWNIIIKALARCGRIE-DAQW 189
+L ++N MI GY G ++AL++ RRM + ++ +I+A A G ++ Q
Sbjct: 249 MKLVAYNAMISGYVNRGFYQEALEMVRRMVSSGIELDEFTYPSVIRACATAGLLQLGKQV 308
Query: 190 HLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNG 249
H R+ N++++ Y + + DEA +FE+MP +D+ SWNA+L+G+ +G
Sbjct: 309 HAYVLRREDFSFHFD---NSLVSLYYKCGKFDEARAIFEKMPAKDLVSWNALLSGYVSSG 365
Query: 250 ELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVL 309
+ A+ +F E+ +K++++W M++G A++G EE LK+F+ M+ G +P + F +
Sbjct: 366 HIGEAKLIFKEMKEKNILSWMIMISGLAENGFGEEGLKLFSCMKREG-FEPCDYAFSGAI 424
Query: 310 GACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRD 369
+C+ L + GQQ H + K GF + +ALI MY+KCG + AR++F + D
Sbjct: 425 KSCAVLGAYCNGQQYHAQLLKIGFDSSLSAGNALITMYAKCGVVEEARQVFRT--MPCLD 482
Query: 370 LISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFD 429
+SWN +IAA HG+G EA++++ +M + G + + +T + +LTACSHAGLVD+G +YFD
Sbjct: 483 SVSWNALIAALGQHGHGAEAVDVYEEMLKKGIRPDRITLLTVLTACSHAGLVDQGRKYFD 542
Query: 430 KLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNA 489
+ I DHYA L+DL R+G+ +A +IE L K + +W LL+GC VHGN
Sbjct: 543 SMETVYRIPPGADHYARLIDLLCRSGKFSDAESVIESLPFKPTAEIWEALLSGCRVHGNM 602
Query: 490 DIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEV 549
++G + A K+ + E+ GTY LLSNM+A+ G+W+E A VR M+D+G+KK+ CSWIE+
Sbjct: 603 ELGIIAADKLFGLIPEHDGTYMLLSNMHAATGQWEEVARVRKLMRDRGVKKEVACSWIEM 662
Query: 550 GNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILDDD-LSRDVE 596
V F+V D SH ++E + L L +M++ G + D + DVE
Sbjct: 663 ETQVHTFLVDDTSHPEAEAVYIYLQDLGKEMRRLGYVPDTSFVLHDVE 710
Score = 145 bits (366), Expect = 4e-33
Identities = 123/508 (24%), Positives = 225/508 (44%), Gaps = 78/508 (15%)
Query: 34 TFSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEA 93
TF P + + N I CK ++AR+LFD++ + DK TTM+ GY G I A
Sbjct: 42 TFGFQPRAHI--LNRLIDVYCKSSELNYARQLFDEISEPDKIARTTMVSGYCASGDITLA 99
Query: 94 RKLFDQ-PDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQ 152
R +F++ P + V + M+ G+ N A +LF +M N
Sbjct: 100 RGVFEKAPVCMRDTVMYNAMITGFSHNNDGYSAINLFCKMKHEGFKPDNFTFASV----- 154
Query: 153 IEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMIT 212
L ++ +C Q+H + G + +V NA+++
Sbjct: 155 ---------------LAGLALVADDEKQC-----VQFHAAALKSGAGYITSVS--NALVS 192
Query: 213 GYAQ----NRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQK-DVI 267
Y++ L A ++F+ + E+D SW M+TG+ +NG + E+L + ++
Sbjct: 193 VYSKCASSPSLLHSARKVFDEILEKDERSWTTMMTGYVKNGYFDLGEELLEGMDDNMKLV 252
Query: 268 TWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQL 327
+ +M++GY G +EAL+M +M + G++ + T+ +V+ AC+ L G+Q+H
Sbjct: 253 AYNAMISGYVNRGFYQEALEMVRRM-VSSGIELDEFTYPSVIRACATAGLLQLGKQVHAY 311
Query: 328 ISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAY------- 380
+ + + ++L+++Y KCG+ AR IF+ + +DL+SWN +++ Y
Sbjct: 312 VLRRE-DFSFHFDNSLVSLYYKCGKFDEARAIFEK--MPAKDLVSWNALLSGYVSSGHIG 368
Query: 381 ------------------------AHHGYGNEAINLFNKMQELGFQANDVTYVELLTACS 416
A +G+G E + LF+ M+ GF+ D + + +C+
Sbjct: 369 EAKLIFKEMKEKNILSWMIMISGLAENGFGEEGLKLFSCMKREGFEPCDYAFSGAIKSCA 428
Query: 417 HAGLVDEGIQYFDKLLK---NRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSL 473
G G QY +LLK + S+ L+ + + G ++EA + + S+
Sbjct: 429 VLGAYCNGQQYHAQLLKIGFDSSLSAGN----ALITMYAKCGVVEEARQVFRTMPCLDSV 484
Query: 474 SVWGPLLAGCNVHGNADIGKLVAKKILK 501
S W L+A HG+ V +++LK
Sbjct: 485 S-WNALIAALGQHGHGAEAVDVYEEMLK 511
Score = 60.1 bits (144), Expect = 2e-07
Identities = 57/259 (22%), Positives = 113/259 (43%), Gaps = 40/259 (15%)
Query: 317 SLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDD-------------- 362
SL + +H I GFQ +++ LI++Y K EL+ AR++FD+
Sbjct: 29 SLQLARAVHGNIITFGFQPRAHILNRLIDVYCKSSELNYARQLFDEISEPDKIARTTMVS 88
Query: 363 ------------GLLRQ-----RDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQAND 405
G+ + RD + +N MI ++H+ G AINLF KM+ GF+ ++
Sbjct: 89 GYCASGDITLARGVFEKAPVCMRDTVMYNAMITGFSHNNDGYSAINLFCKMKHEGFKPDN 148
Query: 406 VTYVELLTACS-HAGLVDEGIQYFDKLLKNRS--IQVKEDHYACLVDLCGRAGRLKEAFY 462
T+ +L + A + +Q+ LK+ + I + + C + L +
Sbjct: 149 FTFASVLAGLALVADDEKQCVQFHAAALKSGAGYITSVSNALVSVYSKCASSPSLLHSAR 208
Query: 463 IIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAG--TYSLLSNMYASV 520
+ ++ W ++ G +G D+G +++L+ +N Y+ + + Y +
Sbjct: 209 KVFDEILEKDERSWTTMMTGYVKNGYFDLG----EELLEGMDDNMKLVAYNAMISGYVNR 264
Query: 521 GKWKEAANVRMKMKDKGLK 539
G ++EA + +M G++
Sbjct: 265 GFYQEALEMVRRMVSSGIE 283
>gb|AAD25817.1| hypothetical protein [Arabidopsis thaliana]
gi|15227144|ref|NP_179798.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|25345463|pir||F84608 hypothetical protein At2g22070
[imported] - Arabidopsis thaliana
Length = 786
Score = 346 bits (887), Expect = 1e-93
Identities = 194/519 (37%), Positives = 309/519 (59%), Gaps = 15/519 (2%)
Query: 79 TMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNEL 138
++++ Y KCG A+ +FD+ + + +W M+ ++++ Q++ A + F +M ER+ +
Sbjct: 186 SLLNMYAKCGDPMMAKFVFDRM-VVRDISSWNAMIALHMQVGQMDLAMAQFEQMAERDIV 244
Query: 139 SWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWN--IIIKALARCGRIEDA----QWHLI 192
+WN MI G+ Q G +ALD+F +M + +L+S + + L+ C +E Q H
Sbjct: 245 TWNSMISGFNQRGYDLRALDIFSKMLRDSLLSPDRFTLASVLSACANLEKLCIGKQIHSH 304
Query: 193 RCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMA--SWNAMLTGFFQNGE 250
G + +V NA+I+ Y++ ++ A L E+ +D+ + A+L G+ + G+
Sbjct: 305 IVTTGF-DISGIVL-NALISMYSRCGGVETARRLIEQRGTKDLKIEGFTALLDGYIKLGD 362
Query: 251 LNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLG 310
+N+A+ +F L +DV+ WT+M+ GY QHG EA+ +F M GG +PN+ T +L
Sbjct: 363 MNQAKNIFVSLKDRDVVAWTAMIVGYEQHGSYGEAINLFRSM-VGGGQRPNSYTLAAMLS 421
Query: 311 ACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLR-QRD 369
S LASL+ G+QIH K+G + V +ALI MY+K G + A + FD L+R +RD
Sbjct: 422 VASSLASLSHGKQIHGSAVKSGEIYSVSVSNALITMYAKAGNITSASRAFD--LIRCERD 479
Query: 370 LISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFD 429
+SW MI A A HG+ EA+ LF M G + + +TYV + +AC+HAGLV++G QYFD
Sbjct: 480 TVSWTSMIIALAQHGHAEEALELFETMLMEGLRPDHITYVGVFSACTHAGLVNQGRQYFD 539
Query: 430 KLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNA 489
+ I HYAC+VDL GRAG L+EA IE + ++ + WG LL+ C VH N
Sbjct: 540 MMKDVDKIIPTLSHYACMVDLFGRAGLLQEAQEFIEKMPIEPDVVTWGSLLSACRVHKNI 599
Query: 490 DIGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEV 549
D+GK+ A+++L +E EN+G YS L+N+Y++ GKW+EAA +R MKD +KK+ G SWIEV
Sbjct: 600 DLGKVAAERLLLLEPENSGAYSALANLYSACGKWEEAAKIRKSMKDGRVKKEQGFSWIEV 659
Query: 550 GNTVQVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILD 588
+ V VF V D +H + + + + ++KK G + D
Sbjct: 660 KHKVHVFGVEDGTHPEKNEIYMTMKKIWDEIKKMGYVPD 698
Score = 194 bits (492), Expect = 9e-48
Identities = 130/452 (28%), Positives = 214/452 (46%), Gaps = 40/452 (8%)
Query: 75 HLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMPE 134
+L +++ Y K G ARKLFD+ + + +W T+++ Y K ++ F ++P+
Sbjct: 50 YLMNNLMNVYSKTGYALHARKLFDEMPLR-TAFSWNTVLSAYSKRGDMDSTCEFFDQLPQ 108
Query: 135 RNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNL-----VSWNIIIKALARCGRIEDAQW 189
R+ +SW MI GY GQ KA+ + M K + N++ A +
Sbjct: 109 RDSVSWTTMIVGYKNIGQYHKAIRVMGDMVKEGIEPTQFTLTNVLASVAATRCMETGKKV 168
Query: 190 HLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNG 249
H + G+ NV N+++ YA+ A +F+RM RD++SWNAM+ Q G
Sbjct: 169 HSFIVKLGLR--GNVSVSNSLLNMYAKCGDPMMAKFVFDRMVVRDISSWNAMIALHMQVG 226
Query: 250 ELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVL 309
+++ A F ++ ++D++TW SM++G+ Q G AL +F+KM + L P+ T +VL
Sbjct: 227 QMDLAMAQFEQMAERDIVTWNSMISGFNQRGYDLRALDIFSKMLRDSLLSPDRFTLASVL 286
Query: 310 GACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFD-------- 361
AC+ L L G+QIH I TGF + V++ALI+MYS+CG + AR++ +
Sbjct: 287 SACANLEKLCIGKQIHSHIVTTGFDISGIVLNALISMYSRCGGVETARRLIEQRGTKDLK 346
Query: 362 --------DGL---------------LRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQE 398
DG L+ RD+++W MI Y HG EAINLF M
Sbjct: 347 IEGFTALLDGYIKLGDMNQAKNIFVSLKDRDVVAWTAMIVGYEQHGSYGEAINLFRSMVG 406
Query: 399 LGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLK 458
G + N T +L+ S + G Q +K+ I A L+ + +AG +
Sbjct: 407 GGQRPNSYTLAAMLSVASSLASLSHGKQIHGSAVKSGEIYSVSVSNA-LITMYAKAGNIT 465
Query: 459 EAFYIIEGLGVKLSLSVWGPLLAGCNVHGNAD 490
A + + + W ++ HG+A+
Sbjct: 466 SASRAFDLIRCERDTVSWTSMIIALAQHGHAE 497
Score = 135 bits (341), Expect = 3e-30
Identities = 94/356 (26%), Positives = 178/356 (49%), Gaps = 24/356 (6%)
Query: 191 LIRCR---KGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQ 247
L+ CR G+M +V N ++ Y++ A +LF+ MP R SWN +L+ + +
Sbjct: 35 LVHCRVIKSGLM--FSVYLMNNLMNVYSKTGYALHARKLFDEMPLRTAFSWNTVLSAYSK 92
Query: 248 NGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVT 307
G+++ + F +LPQ+D ++WT+M+ GY G +A+++ M G++P T
Sbjct: 93 RGDMDSTCEFFDQLPQRDSVSWTTMIVGYKNIGQYHKAIRVMGDM-VKEGIEPTQFTLTN 151
Query: 308 VLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQ 367
VL + + + G+++H I K G + N V ++L+NMY+KCG+ +A+ +FD ++
Sbjct: 152 VLASVAATRCMETGKKVHSFIVKLGLRGNVSVSNSLLNMYAKCGDPMMAKFVFDRMVV-- 209
Query: 368 RDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQY 427
RD+ SWN MIA + G + A+ F +M E + VT+ +++ + G +
Sbjct: 210 RDISSWNAMIALHMQVGQMDLAMAQFEQMAE----RDIVTWNSMISGFNQRGYDLRALDI 265
Query: 428 FDKLLKNRSIQVKEDHYACLVDLCGRAGRL---KEAFYIIEGLGVKLSLSVWGPLLAGCN 484
F K+L++ + A ++ C +L K+ I G +S V L++ +
Sbjct: 266 FSKMLRDSLLSPDRFTLASVLSACANLEKLCIGKQIHSHIVTTGFDISGIVLNALISMYS 325
Query: 485 VHGNADIGKLVAK----KILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDK 536
G + + + + K LK+E ++ L + Y +G +A N+ + +KD+
Sbjct: 326 RCGGVETARRLIEQRGTKDLKIE-----GFTALLDGYIKLGDMNQAKNIFVSLKDR 376
Score = 49.7 bits (117), Expect = 3e-04
Identities = 43/192 (22%), Positives = 82/192 (42%), Gaps = 37/192 (19%)
Query: 322 QQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLR--------------- 366
Q +H + K+G + +++ L+N+YSK G ARK+FD+ LR
Sbjct: 34 QLVHCRVIKSGLMFSVYLMNNLMNVYSKTGYALHARKLFDEMPLRTAFSWNTVLSAYSKR 93
Query: 367 --------------QRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELL 412
QRD +SW MI Y + G ++AI + M + G + T +L
Sbjct: 94 GDMDSTCEFFDQLPQRDSVSWTTMIVGYKNIGQYHKAIRVMGDMVKEGIEPTQFTLTNVL 153
Query: 413 TACSHAGLVDEGIQ---YFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGV 469
+ + ++ G + + KL ++ V L+++ + G A ++ + + V
Sbjct: 154 ASVAATRCMETGKKVHSFIVKLGLRGNVSVSNS----LLNMYAKCGDPMMAKFVFDRMVV 209
Query: 470 KLSLSVWGPLLA 481
+ +S W ++A
Sbjct: 210 R-DISSWNAMIA 220
Score = 39.7 bits (91), Expect = 0.28
Identities = 30/141 (21%), Positives = 66/141 (46%), Gaps = 10/141 (7%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMP-QRDKHLWTTMIDGYIKCGAIKEARKLFD---QPDA 102
N+ I+ K G A + FD + +RD WT+MI + G +EA +LF+
Sbjct: 452 NALITMYAKAGNITSASRAFDLIRCERDTVSWTSMIIALAQHGHAEEALELFETMLMEGL 511
Query: 103 KKSVVTWTTMVNGYVKLNQIEEAESLFYEMPERNEL-----SWNIMIGGYGQNGQIEKAL 157
+ +T+ + + + + F M + +++ + M+ +G+ G +++A
Sbjct: 512 RPDHITYVGVFSACTHAGLVNQGRQYFDMMKDVDKIIPTLSHYACMVDLFGRAGLLQEAQ 571
Query: 158 DLFRRMP-KRNLVSWNIIIKA 177
+ +MP + ++V+W ++ A
Sbjct: 572 EFIEKMPIEPDVVTWGSLLSA 592
Score = 36.6 bits (83), Expect = 2.4
Identities = 36/145 (24%), Positives = 67/145 (45%), Gaps = 18/145 (12%)
Query: 57 GRTDHARKLFDQMPQRDKHL-----WTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTT 111
G + R+ FD M DK + + M+D + + G ++EA++ ++ + VVTW +
Sbjct: 529 GLVNQGRQYFDMMKDVDKIIPTLSHYACMVDLFGRAGLLQEAQEFIEKMPIEPDVVTWGS 588
Query: 112 M-----VNGYVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFR----- 161
+ V+ + L ++ AE L PE N +++ + Y G+ E+A + +
Sbjct: 589 LLSACRVHKNIDLGKV-AAERLLLLEPE-NSGAYSALANLYSACGKWEEAAKIRKSMKDG 646
Query: 162 RMPKRNLVSWNIIIKALARCGRIED 186
R+ K SW I +K +ED
Sbjct: 647 RVKKEQGFSW-IEVKHKVHVFGVED 670
>ref|NP_680717.1| pentatricopeptide (PPR) repeat-containing protein [Arabidopsis
thaliana]
Length = 575
Score = 346 bits (887), Expect = 1e-93
Identities = 180/488 (36%), Positives = 300/488 (60%), Gaps = 15/488 (3%)
Query: 104 KSVVTWTTMVNGYVK-LNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRR 162
K+ +TW +++ G K +++ EA LF E+PE + S+NIM+ Y +N EKA F R
Sbjct: 4 KNTITWNSLLIGISKDPSRMMEAHQLFDEIPEPDTFSYNIMLSCYVRNVNFEKAQSFFDR 63
Query: 163 MPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDE 222
MP ++ SWN +I AR G +E A+ M +N VSWNAMI+GY + L++
Sbjct: 64 MPFKDAASWNTMITGYARRGEMEKARELFYS-----MMEKNEVSWNAMISGYIECGDLEK 118
Query: 223 ALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELP-QKDVITWTSMMTGYAQHGL 281
A F+ P R + +W AM+TG+ + ++ AE +F ++ K+++TW +M++GY ++
Sbjct: 119 ASHFFKVAPVRGVVAWTAMITGYMKAKKVELAEAMFKDMTVNKNLVTWNAMISGYVENSR 178
Query: 282 SEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVS 341
E+ LK+F M G ++PN+ + L CS L++L G+QIHQ++SK+ + ++
Sbjct: 179 PEDGLKLFRAMLEEG-IRPNSSGLSSALLGCSELSALQLGRQIHQIVSKSTLCNDVTALT 237
Query: 342 ALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGF 401
+LI+MY KCGEL A K+F+ +++++D+++WN MI+ YA HG ++A+ LF +M +
Sbjct: 238 SLISMYCKCGELGDAWKLFE--VMKKKDVVAWNAMISGYAQHGNADKALCLFREMIDNKI 295
Query: 402 QANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRAGRLKEAF 461
+ + +T+V +L AC+HAGLV+ G+ YF+ ++++ ++ + DHY C+VDL GRAG+L+EA
Sbjct: 296 RPDWITFVAVLLACNHAGLVNIGMAYFESMVRDYKVEPQPDHYTCMVDLLGRAGKLEEAL 355
Query: 462 YIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLSNMYASVG 521
+I + + +V+G LL C VH N ++ + A+K+L++ +NA Y L+N+YAS
Sbjct: 356 KLIRSMPFRPHAAVFGTLLGACRVHKNVELAEFAAEKLLQLNSQNAAGYVQLANIYASKN 415
Query: 522 KWKEAANVRMKMKDKGL-----KKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLLGL 576
+W++ A VR +MK+ + K PG SWIE+ N V F D+ H + + + L L
Sbjct: 416 RWEDVARVRKRMKESNVVKVERVKVPGYSWIEIRNKVHHFRSSDRIHPELDSIHKKLKEL 475
Query: 577 HTKMKKFG 584
KMK G
Sbjct: 476 EKKMKLAG 483
Score = 167 bits (423), Expect = 9e-40
Identities = 101/342 (29%), Positives = 174/342 (50%), Gaps = 46/342 (13%)
Query: 58 RTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYV 117
R A +LFD++P+ D + M+ Y++ ++A+ FD+ K +W TM+ GY
Sbjct: 22 RMMEAHQLFDEIPEPDTFSYNIMLSCYVRNVNFEKAQSFFDRMPF-KDAASWNTMITGYA 80
Query: 118 KLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKA 177
+ ++E+A LFY M E+NE+SWN MI GY + G +EKA F+ P R +V+W +I
Sbjct: 81 RRGEMEKARELFYSMMEKNEVSWNAMISGYIECGDLEKASHFFKVAPVRGVVAWTAMITG 140
Query: 178 LARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPE----- 232
+ ++E A+ K M +N+V+WNAMI+GY +N R ++ L+LF M E
Sbjct: 141 YMKAKKVELAE----AMFKDMTVNKNLVTWNAMISGYVENSRPEDGLKLFRAMLEEGIRP 196
Query: 233 ----------------------------------RDMASWNAMLTGFFQNGELNRAEKLF 258
D+ + ++++ + + GEL A KLF
Sbjct: 197 NSSGLSSALLGCSELSALQLGRQIHQIVSKSTLCNDVTALTSLISMYCKCGELGDAWKLF 256
Query: 259 AELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASL 318
+ +KDV+ W +M++GYAQHG +++AL +F +M N ++P+ TFV VL AC+ +
Sbjct: 257 EVMKKKDVVAWNAMISGYAQHGNADKALCLFREMIDN-KIRPDWITFVAVLLACNHAGLV 315
Query: 319 TEGQ-QIHQLISKTGFQENTRVVSALINMYSKCGELHIARKI 359
G ++ + + ++++ + G+L A K+
Sbjct: 316 NIGMAYFESMVRDYKVEPQPDHYTCMVDLLGRAGKLEEALKL 357
Score = 131 bits (329), Expect = 7e-29
Identities = 90/322 (27%), Positives = 160/322 (48%), Gaps = 16/322 (4%)
Query: 33 STFSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKE 92
S F P + N+ I+ + G + AR+LF M ++++ W MI GYI+CG +++
Sbjct: 59 SFFDRMPFKDAASWNTMITGYARRGEMEKARELFYSMMEKNEVSWNAMISGYIECGDLEK 118
Query: 93 ARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMP-ERNELSWNIMIGGYGQNG 151
A F + VV WT M+ GY+K ++E AE++F +M +N ++WN MI GY +N
Sbjct: 119 ASHFFKVAPV-RGVVAWTAMITGYMKAKKVELAEAMFKDMTVNKNLVTWNAMISGYVENS 177
Query: 152 QIEKALDLFRRMPKRNL-VSWNIIIKALARCGRIE----DAQWHLIRCRKGMMPLRNVVS 206
+ E L LFR M + + + + + AL C + Q H I + + +V +
Sbjct: 178 RPEDGLKLFRAMLEEGIRPNSSGLSSALLGCSELSALQLGRQIHQIVSKSTL--CNDVTA 235
Query: 207 WNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAEL----P 262
++I+ Y + L +A +LFE M ++D+ +WNAM++G+ Q+G ++A LF E+
Sbjct: 236 LTSLISMYCKCGELGDAWKLFEVMKKKDVVAWNAMISGYAQHGNADKALCLFREMIDNKI 295
Query: 263 QKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQ 322
+ D IT+ +++ GL + F M + ++P + ++ L E
Sbjct: 296 RPDWITFVAVLLACNHAGLVNIGMAYFESMVRDYKVEPQPDHYTCMVDLLGRAGKLEEAL 355
Query: 323 QIHQLISKTGFQENTRVVSALI 344
+LI F+ + V L+
Sbjct: 356 ---KLIRSMPFRPHAAVFGTLL 374
Score = 103 bits (258), Expect = 1e-20
Identities = 84/285 (29%), Positives = 131/285 (45%), Gaps = 30/285 (10%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSV 106
N+ IS + G + A F P R WT MI GY+K ++ A +F K++
Sbjct: 104 NAMISGYIECGDLEKASHFFKVAPVRGVVAWTAMITGYMKAKKVELAEAMFKDMTVNKNL 163
Query: 107 VTWTTMVNGYVKLNQIEEAESLFYEMPER----NELSWNIMIGGYGQNGQIEKALDLFRR 162
VTW M++GYV+ ++ E+ LF M E N + + G + ++ + +
Sbjct: 164 VTWNAMISGYVENSRPEDGLKLFRAMLEEGIRPNSSGLSSALLGCSELSALQLGRQIHQI 223
Query: 163 MPKRNL----VSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNR 218
+ K L + +I +CG + DA W L +M ++VV+WNAMI+GYAQ+
Sbjct: 224 VSKSTLCNDVTALTSLISMYCKCGELGDA-WKLFE----VMKKKDVVAWNAMISGYAQHG 278
Query: 219 RLDEALELFERMPER----DMASWNAMLTGFFQNGELNRAEKLFAEL-------PQKDVI 267
D+AL LF M + D ++ A+L G +N F + PQ D
Sbjct: 279 NADKALCLFREMIDNKIRPDWITFVAVLLACNHAGLVNIGMAYFESMVRDYKVEPQPD-- 336
Query: 268 TWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGAC 312
+T M+ + G EEALK+ M +P+ F T+LGAC
Sbjct: 337 HYTCMVDLLGRAGKLEEALKLIRSMP----FRPHAAVFGTLLGAC 377
>ref|XP_480144.1| putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa (japonica cultivar-group)]
gi|37806252|dbj|BAC99769.1| putative pentatricopeptide
(PPR) repeat-containing protein [Oryza sativa (japonica
cultivar-group)] gi|27817913|dbj|BAC55678.1| putative
pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa (japonica cultivar-group)]
Length = 613
Score = 345 bits (886), Expect = 2e-93
Identities = 182/517 (35%), Positives = 304/517 (58%), Gaps = 10/517 (1%)
Query: 76 LWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVK-LNQIEEAESLFYEMPE 134
L T + ++ G + A + F +K+ T+ ++ GY + L ++ +A LF +P
Sbjct: 19 LSTVAVAAAVRRGDLTGAEEAFASTP-RKTTATYNCLLAGYARALGRLADARHLFDRIPT 77
Query: 135 RNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRC 194
+ +S+N ++ + +G + A LF MP R++VSWN ++ L++ G +E+A+ +
Sbjct: 78 PDAVSYNTLLSCHFASGDADGARRLFASMPVRDVVSWNTMVSGLSKSGAVEEAKAVFLA- 136
Query: 195 RKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPER-DMASWNAMLTGFFQNGELNR 253
MP+RN VSWNAM++G+A +R + A E F PE+ D W AM++G+ G + +
Sbjct: 137 ----MPVRNSVSWNAMVSGFACSRDMSAAEEWFRNAPEKGDAVLWTAMVSGYMDIGNVVK 192
Query: 254 AEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACS 313
A + F +P +++++W +++ GY ++ +++AL++F M ++PN T +VL CS
Sbjct: 193 AIEYFEAMPVRNLVSWNAVVAGYVKNSHADDALRLFRTMVREANVQPNASTLSSVLLGCS 252
Query: 314 GLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISW 373
L++L G+QIHQ K N V ++L++MY KCG+L A K+F G + RD+++W
Sbjct: 253 NLSALGFGKQIHQWCMKLPLSRNLTVGTSLVSMYCKCGDLSSACKLF--GEMHTRDVVAW 310
Query: 374 NGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLK 433
N MI+ YA HG G EAINLF +M++ G + N +T+V +LTAC H GL D GI+ F+ + +
Sbjct: 311 NAMISGYAQHGDGKEAINLFERMKDEGVEPNWITFVAVLTACIHTGLCDFGIRCFEGMQE 370
Query: 434 NRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGK 493
I+ + DHY+C+VDL RAG+L+ A +I + + S +G LLA C V+ N + +
Sbjct: 371 LYGIEPRVDHYSCMVDLLCRAGKLERAVDLIRSMPFEPHPSAYGTLLAACRVYKNLEFAE 430
Query: 494 LVAKKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTV 553
L A K+++ + ++AG Y L+N+YA +W + + VR MKD + K PG SWIE+ +
Sbjct: 431 LAAGKLIEKDPQSAGAYVQLANIYAGANQWDDVSRVRRWMKDNAVVKTPGYSWIEIKGVM 490
Query: 554 QVFVVGDKSHSQSEMLEYLLLGLHTKMKKFGDILDDD 590
F D+ H Q ++ L L +MK G + D D
Sbjct: 491 HEFRSNDRLHPQLYLIHEKLGQLAERMKAMGYVPDLD 527
Score = 156 bits (395), Expect = 2e-36
Identities = 105/347 (30%), Positives = 168/347 (48%), Gaps = 59/347 (17%)
Query: 57 GRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGY 116
GR AR LFD++P D + T++ + G AR+LF + VV+W TMV+G
Sbjct: 63 GRLADARHLFDRIPTPDAVSYNTLLSCHFASGDADGARRLFASMPV-RDVVSWNTMVSGL 121
Query: 117 VKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMP-KRNLVSWNIII 175
K +EEA+++F MP RN +SWN M+ G+ + + A + FR P K + V W ++
Sbjct: 122 SKSGAVEEAKAVFLAMPVRNSVSWNAMVSGFACSRDMSAAEEWFRNAPEKGDAVLWTAMV 181
Query: 176 KALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMP---- 231
G + A + MP+RN+VSWNA++ GY +N D+AL LF M
Sbjct: 182 SGYMDIGNVVKAIEYF-----EAMPVRNLVSWNAVVAGYVKNSHADDALRLFRTMVREAN 236
Query: 232 ------------------------------------ERDMASWNAMLTGFFQNGELNRAE 255
R++ ++++ + + G+L+ A
Sbjct: 237 VQPNASTLSSVLLGCSNLSALGFGKQIHQWCMKLPLSRNLTVGTSLVSMYCKCGDLSSAC 296
Query: 256 KLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGAC--S 313
KLF E+ +DV+ W +M++GYAQHG +EA+ +F +M+ + G++PN TFV VL AC +
Sbjct: 297 KLFGEMHTRDVVAWNAMISGYAQHGDGKEAINLFERMK-DEGVEPNWITFVAVLTACIHT 355
Query: 314 GLASL----TEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIA 356
GL EG Q G + S ++++ + G+L A
Sbjct: 356 GLCDFGIRCFEGMQ-----ELYGIEPRVDHYSCMVDLLCRAGKLERA 397
>gb|AAC35225.1| hypothetical protein [Arabidopsis thaliana]
gi|15227619|ref|NP_180537.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|25345458|pir||C84700 hypothetical protein At2g29760
[imported] - Arabidopsis thaliana
Length = 738
Score = 342 bits (876), Expect = 3e-92
Identities = 189/567 (33%), Positives = 309/567 (54%), Gaps = 44/567 (7%)
Query: 60 DHARKLFDQMPQRDKHLWTTMIDGY--------------------------------IKC 87
++ARK+FD++P+ + W T+I Y IK
Sbjct: 81 EYARKVFDEIPKPNSFAWNTLIRAYASGPDPVLSIWAFLDMVSESQCYPNKYTFPFLIKA 140
Query: 88 GAIKEARKLFDQPD--AKKSVV-----TWTTMVNGYVKLNQIEEAESLFYEMPERNELSW 140
A + L A KS V ++++ Y ++ A +F + E++ +SW
Sbjct: 141 AAEVSSLSLGQSLHGMAVKSAVGSDVFVANSLIHCYFSCGDLDSACKVFTTIKEKDVVSW 200
Query: 141 NIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNI-IIKALARCGRIEDAQWHLIRCR--KG 197
N MI G+ Q G +KAL+LF++M ++ + ++ ++ L+ C +I + ++ C +
Sbjct: 201 NSMINGFVQKGSPDKALELFKKMESEDVKASHVTMVGVLSACAKIRNLEFGRQVCSYIEE 260
Query: 198 MMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKL 257
N+ NAM+ Y + +++A LF+ M E+D +W ML G+ + + A ++
Sbjct: 261 NRVNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEKDNVTWTTMLDGYAISEDYEAAREV 320
Query: 258 FAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLAS 317
+PQKD++ W ++++ Y Q+G EAL +F ++Q +K N T V+ L AC+ + +
Sbjct: 321 LNSMPQKDIVAWNALISAYEQNGKPNEALIVFHELQLQKNMKLNQITLVSTLSACAQVGA 380
Query: 318 LTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMI 377
L G+ IH I K G + N V SALI+MYSKCG+L +R++F+ + +RD+ W+ MI
Sbjct: 381 LELGRWIHSYIKKHGIRMNFHVTSALIHMYSKCGDLEKSREVFNS--VEKRDVFVWSAMI 438
Query: 378 AAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSI 437
A HG GNEA+++F KMQE + N VT+ + ACSH GLVDE F ++ N I
Sbjct: 439 GGLAMHGCGNEAVDMFYKMQEANVKPNGVTFTNVFCACSHTGLVDEAESLFHQMESNYGI 498
Query: 438 QVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAK 497
+E HYAC+VD+ GR+G L++A IE + + S SVWG LL C +H N ++ ++
Sbjct: 499 VPEEKHYACIVDVLGRSGYLEKAVKFIEAMPIPPSTSVWGALLGACKIHANLNLAEMACT 558
Query: 498 KILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFV 557
++L++E N G + LLSN+YA +GKW+ + +R M+ GLKK+PGCS IE+ + F+
Sbjct: 559 RLLELEPRNDGAHVLLSNIYAKLGKWENVSELRKHMRVTGLKKEPGCSSIEIDGMIHEFL 618
Query: 558 VGDKSHSQSEMLEYLLLGLHTKMKKFG 584
GD +H SE + L + K+K G
Sbjct: 619 SGDNAHPMSEKVYGKLHEVMEKLKSNG 645
Score = 144 bits (364), Expect = 6e-33
Identities = 97/372 (26%), Positives = 184/372 (49%), Gaps = 43/372 (11%)
Query: 153 IEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGMMPLRNVVSWNAMIT 212
+E A +F +PK N +WN +I+A A + W + N ++ +I
Sbjct: 80 LEYARKVFDEIPKPNSFAWNTLIRAYASGPDPVLSIWAFLDMVSESQCYPNKYTFPFLIK 139
Query: 213 GYAQNRRLDEALELFERMPERDMAS----WNAMLTGFFQNGELNRAEKLFAELPQKDVIT 268
A+ L L + + S N+++ +F G+L+ A K+F + +KDV++
Sbjct: 140 AAAEVSSLSLGQSLHGMAVKSAVGSDVFVANSLIHCYFSCGDLDSACKVFTTIKEKDVVS 199
Query: 269 WTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLI 328
W SM+ G+ Q G ++AL++F KM++ +K ++ T V VL AC+ + +L G+Q+ I
Sbjct: 200 WNSMINGFVQKGSPDKALELFKKMESE-DVKASHVTMVGVLSACAKIRNLEFGRQVCSYI 258
Query: 329 SKTGFQENTRVVSALINMYSKCGELHIARKIFD--------------DGL---------- 364
+ N + +A+++MY+KCG + A+++FD DG
Sbjct: 259 EENRVNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEKDNVTWTTMLDGYAISEDYEAAR 318
Query: 365 -----LRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQ-ELGFQANDVTYVELLTACSHA 418
+ Q+D+++WN +I+AY +G NEA+ +F+++Q + + N +T V L+AC+
Sbjct: 319 EVLNSMPQKDIVAWNALISAYEQNGKPNEALIVFHELQLQKNMKLNQITLVSTLSACAQV 378
Query: 419 GLVDEGIQYFDKLLKNRSIQVKEDHYACLVDL---CGRAGRLKEAFYIIEGLGVKLSLSV 475
G ++ G ++ +K I++ + L+ + CG + +E F +E K + V
Sbjct: 379 GALELG-RWIHSYIKKHGIRMNFHVTSALIHMYSKCGDLEKSREVFNSVE----KRDVFV 433
Query: 476 WGPLLAGCNVHG 487
W ++ G +HG
Sbjct: 434 WSAMIGGLAMHG 445
Score = 126 bits (316), Expect = 2e-27
Identities = 96/388 (24%), Positives = 179/388 (45%), Gaps = 49/388 (12%)
Query: 57 GRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLF---DQPDAKKSVVTW---- 109
G D A K+F + ++D W +MI+G+++ G+ +A +LF + D K S VT
Sbjct: 180 GDLDSACKVFTTIKEKDVVSWNSMINGFVQKGSPDKALELFKKMESEDVKASHVTMVGVL 239
Query: 110 -------------------------------TTMVNGYVKLNQIEEAESLFYEMPERNEL 138
M++ Y K IE+A+ LF M E++ +
Sbjct: 240 SACAKIRNLEFGRQVCSYIEENRVNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEKDNV 299
Query: 139 SWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDA--QWHLIRCRK 196
+W M+ GY + E A ++ MP++++V+WN +I A + G+ +A +H ++ +K
Sbjct: 300 TWTTMLDGYAISEDYEAAREVLNSMPQKDIVAWNALISAYEQNGKPNEALIVFHELQLQK 359
Query: 197 GMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMAS----WNAMLTGFFQNGELN 252
M N ++ + ++ AQ L+ + + + + +A++ + + G+L
Sbjct: 360 NMK--LNQITLVSTLSACAQVGALELGRWIHSYIKKHGIRMNFHVTSALIHMYSKCGDLE 417
Query: 253 RAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGAC 312
++ ++F + ++DV W++M+ G A HG EA+ MF KMQ +KPN TF V AC
Sbjct: 418 KSREVFNSVEKRDVFVWSAMIGGLAMHGCGNEAVDMFYKMQ-EANVKPNGVTFTNVFCAC 476
Query: 313 SGLASLTEGQQI-HQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLI 371
S + E + + HQ+ S G + + ++++ + G L A K F + +
Sbjct: 477 SHTGLVDEAESLFHQMESNYGIVPEEKHYACIVDVLGRSGYLEKAVK-FIEAMPIPPSTS 535
Query: 372 SWNGMIAAYAHHGYGNEAINLFNKMQEL 399
W ++ A H N A ++ EL
Sbjct: 536 VWGALLGACKIHANLNLAEMACTRLLEL 563
Score = 105 bits (263), Expect = 3e-21
Identities = 70/276 (25%), Positives = 141/276 (50%), Gaps = 7/276 (2%)
Query: 251 LNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLG 310
L A K+F E+P+ + W +++ YA ++ F M + PN TF ++
Sbjct: 80 LEYARKVFDEIPKPNSFAWNTLIRAYASGPDPVLSIWAFLDMVSESQCYPNKYTFPFLIK 139
Query: 311 ACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDL 370
A + ++SL+ GQ +H + K+ + V ++LI+ Y CG+L A K+F ++++D+
Sbjct: 140 AAAEVSSLSLGQSLHGMAVKSAVGSDVFVANSLIHCYFSCGDLDSACKVFT--TIKEKDV 197
Query: 371 ISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDK 430
+SWN MI + G ++A+ LF KM+ +A+ VT V +L+AC+ ++ G Q
Sbjct: 198 VSWNSMINGFVQKGSPDKALELFKKMESEDVKASHVTMVGVLSACAKIRNLEFGRQVCSY 257
Query: 431 LLKNRSIQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNAD 490
+ +NR + V ++D+ + G +++A + + + K +++ W +L G + + +
Sbjct: 258 IEENR-VNVNLTLANAMLDMYTKCGSIEDAKRLFDAMEEKDNVT-WTTMLDGYAISEDYE 315
Query: 491 IGKLVAKKILKVEHENAGTYSLLSNMYASVGKWKEA 526
+ V + + ++ ++ L + Y GK EA
Sbjct: 316 AAREV---LNSMPQKDIVAWNALISAYEQNGKPNEA 348
>dbj|BAB01819.1| unnamed protein product [Arabidopsis thaliana]
gi|15228653|ref|NP_189568.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
Length = 600
Score = 338 bits (867), Expect = 3e-91
Identities = 171/439 (38%), Positives = 278/439 (62%), Gaps = 13/439 (2%)
Query: 141 NIMIGGYGQNGQ--IEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRKGM 198
N +I Y + G + A+ LF +M +R+ VSWN ++ L + G + DA R
Sbjct: 156 NALIDCYSRCGGLGVRDAMKLFEKMSERDTVSWNSMLGGLVKAGELRDA-----RRLFDE 210
Query: 199 MPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLF 258
MP R+++SWN M+ GYA+ R + +A ELFE+MPER+ SW+ M+ G+ + G++ A +F
Sbjct: 211 MPQRDLISWNTMLDGYARCREMSKAFELFEKMPERNTVSWSTMVMGYSKAGDMEMARVMF 270
Query: 259 AE--LPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLA 316
+ LP K+V+TWT ++ GYA+ GL +EA ++ +M A+G LK + +++L AC+
Sbjct: 271 DKMPLPAKNVVTWTIIIAGYAEKGLLKEADRLVDQMVASG-LKFDAAAVISILAACTESG 329
Query: 317 SLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGM 376
L+ G +IH ++ ++ N V++AL++MY+KCG L A +F+D + ++DL+SWN M
Sbjct: 330 LLSLGMRIHSILKRSNLGSNAYVLNALLDMYAKCGNLKKAFDVFND--IPKKDLVSWNTM 387
Query: 377 IAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRS 436
+ HG+G EAI LF++M+ G + + VT++ +L +C+HAGL+DEGI YF + K
Sbjct: 388 LHGLGVHGHGKEAIELFSRMRREGIRPDKVTFIAVLCSCNHAGLIDEGIDYFYSMEKVYD 447
Query: 437 IQVKEDHYACLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVA 496
+ + +HY CLVDL GR GRLKEA +++ + ++ ++ +WG LL C +H DI K V
Sbjct: 448 LVPQVEHYGCLVDLLGRVGRLKEAIKVVQTMPMEPNVVIWGALLGACRMHNEVDIAKEVL 507
Query: 497 KKILKVEHENAGTYSLLSNMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVF 556
++K++ + G YSLLSN+YA+ W+ A++R KMK G++K G S +E+ + + F
Sbjct: 508 DNLVKLDPCDPGNYSLLSNIYAAAEDWEGVADIRSKMKSMGVEKPSGASSVELEDGIHEF 567
Query: 557 VVGDKSHSQSEMLEYLLLG 575
V DKSH +S+ + Y +LG
Sbjct: 568 TVFDKSHPKSDQI-YQMLG 585
Score = 160 bits (405), Expect = 1e-37
Identities = 99/369 (26%), Positives = 188/369 (50%), Gaps = 48/369 (13%)
Query: 73 DKHLWTTMIDGYIKCGA--IKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFY 130
D ++ +ID Y +CG +++A KLF++ +++ V+W +M+ G VK ++ +A LF
Sbjct: 151 DIYVPNALIDCYSRCGGLGVRDAMKLFEKM-SERDTVSWNSMLGGLVKAGELRDARRLFD 209
Query: 131 EMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNLVSWNIIIKALARCGRIEDAQWH 190
EMP+R+ +SWN M+ GY + ++ KA +LF +MP+RN VSW+ ++ ++ G +E A+
Sbjct: 210 EMPQRDLISWNTMLDGYARCREMSKAFELFEKMPERNTVSWSTMVMGYSKAGDMEMAR-- 267
Query: 191 LIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERM-------------------P 231
+ K +P +NVV+W +I GYA+ L EA L ++M
Sbjct: 268 -VMFDKMPLPAKNVVTWTIIIAGYAEKGLLKEADRLVDQMVASGLKFDAAAVISILAACT 326
Query: 232 ERDMAS--------------------WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTS 271
E + S NA+L + + G L +A +F ++P+KD+++W +
Sbjct: 327 ESGLLSLGMRIHSILKRSNLGSNAYVLNALLDMYAKCGNLKKAFDVFNDIPKKDLVSWNT 386
Query: 272 MMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKT 331
M+ G HG +EA+++F++M+ G++P+ TF+ VL +C+ + EG + K
Sbjct: 387 MLHGLGVHGHGKEAIELFSRMRRE-GIRPDKVTFIAVLCSCNHAGLIDEGIDYFYSMEKV 445
Query: 332 -GFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAI 390
L+++ + G L A K+ + + +++ W ++ A H + A
Sbjct: 446 YDLVPQVEHYGCLVDLLGRVGRLKEAIKVVQT-MPMEPNVVIWGALLGACRMHNEVDIAK 504
Query: 391 NLFNKMQEL 399
+ + + +L
Sbjct: 505 EVLDNLVKL 513
Score = 137 bits (344), Expect = 1e-30
Identities = 98/330 (29%), Positives = 168/330 (50%), Gaps = 23/330 (6%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQPDAKKSV 106
NS + L K G AR+LFD+MPQRD W TM+DGY +C + +A +LF++ +++
Sbjct: 189 NSMLGGLVKAGELRDARRLFDEMPQRDLISWNTMLDGYARCREMSKAFELFEKM-PERNT 247
Query: 107 VTWTTMVNGYVKLNQIEEAESLFYEM--PERNELSWNIMIGGYGQNGQIEKALDLFRRMP 164
V+W+TMV GY K +E A +F +M P +N ++W I+I GY + G +++A L +M
Sbjct: 248 VSWSTMVMGYSKAGDMEMARVMFDKMPLPAKNVVTWTIIIAGYAEKGLLKEADRLVDQMV 307
Query: 165 KRNL-VSWNIIIKALARC---GRIE-DAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRR 219
L +I LA C G + + H I R + N NA++ YA+
Sbjct: 308 ASGLKFDAAAVISILAACTESGLLSLGMRIHSILKRSNLG--SNAYVLNALLDMYAKCGN 365
Query: 220 LDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQK----DVITWTSMMTG 275
L +A ++F +P++D+ SWN ML G +G A +LF+ + ++ D +T+ +++
Sbjct: 366 LKKAFDVFNDIPKKDLVSWNTMLHGLGVHGHGKEAIELFSRMRREGIRPDKVTFIAVLCS 425
Query: 276 YAQHGLSEEALKMFTKMQANGGLKP---NNGTFVTVLGACSGLASLTEGQQIHQLISKTG 332
GL +E + F M+ L P + G V +LG L ++ +++
Sbjct: 426 CNHAGLIDEGIDYFYSMEKVYDLVPQVEHYGCLVDLLGRVGRL------KEAIKVVQTMP 479
Query: 333 FQENTRVVSALINMYSKCGELHIARKIFDD 362
+ N + AL+ E+ IA+++ D+
Sbjct: 480 MEPNVVIWGALLGACRMHNEVDIAKEVLDN 509
Score = 99.4 bits (246), Expect = 3e-19
Identities = 92/328 (28%), Positives = 152/328 (46%), Gaps = 51/328 (15%)
Query: 250 ELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVL 309
+ N A ++F ++ + +V S++ +AQ+ +A +F++MQ GL +N T+ +L
Sbjct: 66 QTNLAVRVFNQVQEPNVHLCNSLIRAHAQNSQPYQAFFVFSEMQ-RFGLFADNFTYPFLL 124
Query: 310 GACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKC------------------- 350
ACSG + L + +H I K G + V +ALI+ YS+C
Sbjct: 125 KACSGQSWLPVVKMMHNHIEKLGLSSDIYVPNALIDCYSRCGGLGVRDAMKLFEKMSERD 184
Query: 351 --------------GELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKM 396
GEL AR++FD+ + QRDLISWN M+ YA ++A LF KM
Sbjct: 185 TVSWNSMLGGLVKAGELRDARRLFDE--MPQRDLISWNTMLDGYARCREMSKAFELFEKM 242
Query: 397 QELGFQANDVTYVELLTACSHAGLVDEGIQYFDKL-LKNRSIQVKEDHYACLVDLCGRAG 455
E N V++ ++ S AG ++ FDK+ L +++ + ++ G
Sbjct: 243 PE----RNTVSWSTMVMGYSKAGDMEMARVMFDKMPLPAKNVVT----WTIIIAGYAEKG 294
Query: 456 RLKEAFYIIEGL---GVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSL 512
LKEA +++ + G+K + +LA C G +G + ILK + + Y L
Sbjct: 295 LLKEADRLVDQMVASGLKFDAAAVISILAACTESGLLSLGMRI-HSILKRSNLGSNAYVL 353
Query: 513 --LSNMYASVGKWKEAANVRMKMKDKGL 538
L +MYA G K+A +V + K L
Sbjct: 354 NALLDMYAKCGNLKKAFDVFNDIPKKDL 381
Score = 88.6 bits (218), Expect = 5e-16
Identities = 77/298 (25%), Positives = 144/298 (47%), Gaps = 31/298 (10%)
Query: 35 FSPNPNSEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEAR 94
F P ++ N+ + + A +LF++MP+R+ W+TM+ GY K G ++ AR
Sbjct: 208 FDEMPQRDLISWNTMLDGYARCREMSKAFELFEKMPERNTVSWSTMVMGYSKAGDMEMAR 267
Query: 95 KLFDQ-PDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMP----ERNELSWNIMIGGYGQ 149
+FD+ P K+VVTWT ++ GY + ++EA+ L +M + + + ++ +
Sbjct: 268 VMFDKMPLPAKNVVTWTIIIAGYAEKGLLKEADRLVDQMVASGLKFDAAAVISILAACTE 327
Query: 150 NGQIEKALDLFRRMPKRNLVSWNIIIKAL----ARCGRIEDAQWHLIRCRKGMMPLRNVV 205
+G + + + + + NL S ++ AL A+CG ++ A +P +++V
Sbjct: 328 SGLLSLGMRIHSILKRSNLGSNAYVLNALLDMYAKCGNLKKAFDVF-----NDIPKKDLV 382
Query: 206 SWNAMITGYAQNRRLDEALELFERMPER----DMASWNAMLTGFFQNGELNRA------- 254
SWN M+ G + EA+ELF RM D ++ A+L G ++
Sbjct: 383 SWNTMLHGLGVHGHGKEAIELFSRMRREGIRPDKVTFIAVLCSCNHAGLIDEGIDYFYSM 442
Query: 255 EKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGAC 312
EK++ +PQ V + ++ + G +EA+K+ M ++PN + +LGAC
Sbjct: 443 EKVYDLVPQ--VEHYGCLVDLLGRVGRLKEAIKVVQTMP----MEPNVVIWGALLGAC 494
Score = 57.4 bits (137), Expect = 1e-06
Identities = 45/172 (26%), Positives = 86/172 (49%), Gaps = 12/172 (6%)
Query: 316 ASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNG 375
A+L + +Q+H I + E+ + LI+ S C + ++A ++F+ +++ ++ N
Sbjct: 30 ANLNQVKQLHAQIIRRNLHEDLHIAPKLISALSLCRQTNLAVRVFNQ--VQEPNVHLCNS 87
Query: 376 MIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAG---LVDEGIQYFDKLL 432
+I A+A + +A +F++MQ G A++ TY LL ACS +V + +KL
Sbjct: 88 LIRAHAQNSQPYQAFFVFSEMQRFGLFADNFTYPFLLKACSGQSWLPVVKMMHNHIEKLG 147
Query: 433 KNRSIQVKEDHYACLVDLCGRAGRL--KEAFYIIEGLGVKLSLSVWGPLLAG 482
+ I V L+D R G L ++A + E + + ++S W +L G
Sbjct: 148 LSSDIYVPN----ALIDCYSRCGGLGVRDAMKLFEKMSERDTVS-WNSMLGG 194
>emb|CAB78877.1| putative protein [Arabidopsis thaliana] gi|4539394|emb|CAB37460.1|
putative protein [Arabidopsis thaliana]
gi|15234006|ref|NP_193610.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|7486269|pir||T04867 hypothetical protein F28A21.160 -
Arabidopsis thaliana
Length = 871
Score = 326 bits (835), Expect = 1e-87
Identities = 176/550 (32%), Positives = 306/550 (55%), Gaps = 49/550 (8%)
Query: 79 TMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMP----- 133
+++ Y+K + ARK+FD+ ++ V++W +++NGYV E+ S+F +M
Sbjct: 235 SLVAFYLKNQRVDSARKVFDEM-TERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIE 293
Query: 134 ----------------------------------ERNELSWNIMIGGYGQNGQIEKALDL 159
R + N ++ Y + G ++ A +
Sbjct: 294 IDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAV 353
Query: 160 FRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRK-GMMPLRNVVSWNAMITGYAQNR 218
FR M R++VS+ +I AR G +A + G+ P +V + A++ A+ R
Sbjct: 354 FREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISP--DVYTVTAVLNCCARYR 411
Query: 219 RLDEALELFERMPERDMAS----WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMT 274
LDE + E + E D+ NA++ + + G + AE +F+E+ KD+I+W +++
Sbjct: 412 LLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIG 471
Query: 275 GYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQ 334
GY+++ + EAL +F + P+ T VL AC+ L++ +G++IH I + G+
Sbjct: 472 GYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYF 531
Query: 335 ENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFN 394
+ V ++L++MY+KCG L +A +FDD + +DL+SW MIA Y HG+G EAI LFN
Sbjct: 532 SDRHVANSLVDMYAKCGALLLAHMLFDD--IASKDLVSWTVMIAGYGMHGFGKEAIALFN 589
Query: 395 KMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRA 454
+M++ G +A+++++V LL ACSH+GLVDEG ++F+ + I+ +HYAC+VD+ R
Sbjct: 590 QMRQAGIEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVDMLART 649
Query: 455 GRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLS 514
G L +A+ IE + + ++WG LL GC +H + + + VA+K+ ++E EN G Y L++
Sbjct: 650 GDLIKAYRFIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGYYVLMA 709
Query: 515 NMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLL 574
N+YA KW++ +R ++ +GL+K PGCSWIE+ V +FV GD S+ ++E +E L
Sbjct: 710 NIYAEAEKWEQVKRLRKRIGQRGLRKNPGCSWIEIKGRVNIFVAGDSSNPETENIEAFLR 769
Query: 575 GLHTKMKKFG 584
+ +M + G
Sbjct: 770 KVRARMIEEG 779
Score = 144 bits (362), Expect = 1e-32
Identities = 109/455 (23%), Positives = 209/455 (44%), Gaps = 66/455 (14%)
Query: 47 NSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIK---EARKLFDQPDAK 103
N+ + R C+ G ++A KL + D ID C ++ +++ L D +
Sbjct: 65 NTQLRRFCESGNLENAVKLLCVSGKWD-------IDPRTLCSVLQLCADSKSLKDGKEVD 117
Query: 104 KSVVTWTTMVNG---------YVKLNQIEEAESLFYEMPERNELSWNIMIGGYGQNGQIE 154
+ +++ Y ++EA +F E+ L WNI++ ++G
Sbjct: 118 NFIRGNGFVIDSNLGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKSGDFS 177
Query: 155 KALDLFRRMPKRNL----VSWNIIIKALARCGRIEDA-QWHLIRCRKGMMPLRNVVSWNA 209
++ LF++M + +++ + K+ + + Q H + G RN V N+
Sbjct: 178 GSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGFGE-RNSVG-NS 235
Query: 210 MITGYAQNRRLDEALELFERMPERDMASWNAMLTGFFQNGELNRAEKLFAELPQKDVITW 269
++ Y +N+R+D A ++F+ M ERD+ SWN+++ G
Sbjct: 236 LVAFYLKNQRVDSARKVFDEMTERDVISWNSIING------------------------- 270
Query: 270 TSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLIS 329
Y +GL+E+ L +F +M + G++ + T V+V C+ ++ G+ +H +
Sbjct: 271 ------YVSNGLAEKGLSVFVQMLVS-GIEIDLATIVSVFAGCADSRLISLGRAVHSIGV 323
Query: 330 KTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEA 389
K F R + L++MYSKCG+L A+ +F + + R ++S+ MIA YA G EA
Sbjct: 324 KACFSREDRFCNTLLDMYSKCGDLDSAKAVFRE--MSDRSVVSYTSMIAGYAREGLAGEA 381
Query: 390 INLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYA--CL 447
+ LF +M+E G + T +L C+ L+DEG + + + +N + D + L
Sbjct: 382 VKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWIKEN---DLGFDIFVSNAL 438
Query: 448 VDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLAG 482
+D+ + G ++EA + + VK +S W ++ G
Sbjct: 439 MDMYAKCGSMQEAELVFSEMRVKDIIS-WNTIIGG 472
Score = 95.5 bits (236), Expect = 4e-18
Identities = 60/213 (28%), Positives = 112/213 (52%), Gaps = 6/213 (2%)
Query: 249 GELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTV 308
G+L A ++F E+ + + W +M A+ G ++ +F KM ++G ++ ++ TF V
Sbjct: 143 GDLKEASRVFDEVKIEKALFWNILMNELAKSGDFSGSIGLFKKMMSSG-VEMDSYTFSCV 201
Query: 309 LGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQR 368
+ S L S+ G+Q+H I K+GF E V ++L+ Y K + ARK+FD+ + +R
Sbjct: 202 SKSFSSLRSVHGGEQLHGFILKSGFGERNSVGNSLVAFYLKNQRVDSARKVFDE--MTER 259
Query: 369 DLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYF 428
D+ISWN +I Y +G + +++F +M G + + T V + C+ + L+ G
Sbjct: 260 DVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVH 319
Query: 429 DKLLKNRSIQVKEDHYA-CLVDLCGRAGRLKEA 460
+ ++ +ED + L+D+ + G L A
Sbjct: 320 S--IGVKACFSREDRFCNTLLDMYSKCGDLDSA 350
Score = 94.0 bits (232), Expect = 1e-17
Identities = 94/403 (23%), Positives = 163/403 (40%), Gaps = 80/403 (19%)
Query: 42 EMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQP- 100
E + CN+ + K G D A+ +F +M R +T+MI GY + G EA KLF++
Sbjct: 330 EDRFCNTLLDMYSKCGDLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEME 389
Query: 101 ------------------------DAKKSVVTW-------------TTMVNGYVKLNQIE 123
D K V W +++ Y K ++
Sbjct: 390 EEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQ 449
Query: 124 EAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM--PKRNLVSWNIIIKALARC 181
EAE +F EM ++ +SWN +IGGY +N +AL LF + KR + L C
Sbjct: 450 EAELVFSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPAC 509
Query: 182 GRI----EDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMAS 237
+ + + H R G R+V N+++ YA+ L A LF+ + +D+ S
Sbjct: 510 ASLSAFDKGREIHGYIMRNGYFSDRHVA--NSLVDMYAKCGALLLAHMLFDDIASKDLVS 567
Query: 238 WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGG 297
W M+ G Y HG +EA+ +F +M+ G
Sbjct: 568 WTVMIAG-------------------------------YGMHGFGKEAIALFNQMR-QAG 595
Query: 298 LKPNNGTFVTVLGACSGLASLTEGQQIHQLI-SKTGFQENTRVVSALINMYSKCGELHIA 356
++ + +FV++L ACS + EG + ++ + + + +++M ++ G+L A
Sbjct: 596 IEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVDMLARTGDLIKA 655
Query: 357 RKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQEL 399
+ F + + D W ++ H A + K+ EL
Sbjct: 656 YR-FIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFEL 697
Score = 58.9 bits (141), Expect = 4e-07
Identities = 43/171 (25%), Positives = 80/171 (46%), Gaps = 5/171 (2%)
Query: 264 KDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQ 323
+ V + + + + G E A+K+ +G + T +VL C+ SL +G++
Sbjct: 59 RSVTDANTQLRRFCESGNLENAVKLLC---VSGKWDIDPRTLCSVLQLCADSKSLKDGKE 115
Query: 324 IHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHH 383
+ I GF ++ + S L MY+ CG+L A ++FD+ ++ + WN ++ A
Sbjct: 116 VDNFIRGNGFVIDSNLGSKLSLMYTNCGDLKEASRVFDE--VKIEKALFWNILMNELAKS 173
Query: 384 GYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKN 434
G + +I LF KM G + + T+ + + S V G Q +LK+
Sbjct: 174 GDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKS 224
Score = 35.4 bits (80), Expect = 5.3
Identities = 23/59 (38%), Positives = 28/59 (46%)
Query: 41 SEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQ 99
S+ NS + K G A LFD + +D WT MI GY G KEA LF+Q
Sbjct: 532 SDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAIALFNQ 590
>dbj|BAD94843.1| putative protein [Arabidopsis thaliana]
Length = 720
Score = 326 bits (835), Expect = 1e-87
Identities = 176/550 (32%), Positives = 306/550 (55%), Gaps = 49/550 (8%)
Query: 79 TMIDGYIKCGAIKEARKLFDQPDAKKSVVTWTTMVNGYVKLNQIEEAESLFYEMP----- 133
+++ Y+K + ARK+FD+ ++ V++W +++NGYV E+ S+F +M
Sbjct: 84 SLVAFYLKNQRVDSARKVFDEM-TERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIE 142
Query: 134 ----------------------------------ERNELSWNIMIGGYGQNGQIEKALDL 159
R + N ++ Y + G ++ A +
Sbjct: 143 IDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAV 202
Query: 160 FRRMPKRNLVSWNIIIKALARCGRIEDAQWHLIRCRK-GMMPLRNVVSWNAMITGYAQNR 218
FR M R++VS+ +I AR G +A + G+ P +V + A++ A+ R
Sbjct: 203 FREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISP--DVYTVTAVLNCCARYR 260
Query: 219 RLDEALELFERMPERDMAS----WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMT 274
LDE + E + E D+ NA++ + + G + AE +F+E+ KD+I+W +++
Sbjct: 261 LLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIG 320
Query: 275 GYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLASLTEGQQIHQLISKTGFQ 334
GY+++ + EAL +F + P+ T VL AC+ L++ +G++IH I + G+
Sbjct: 321 GYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYF 380
Query: 335 ENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFN 394
+ V ++L++MY+KCG L +A +FDD + +DL+SW MIA Y HG+G EAI LFN
Sbjct: 381 SDRHVANSLVDMYAKCGALLLAHMLFDD--IASKDLVSWTVMIAGYGMHGFGKEAIALFN 438
Query: 395 KMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSIQVKEDHYACLVDLCGRA 454
+M++ G +A+++++V LL ACSH+GLVDEG ++F+ + I+ +HYAC+VD+ R
Sbjct: 439 QMRQAGIEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVDMLART 498
Query: 455 GRLKEAFYIIEGLGVKLSLSVWGPLLAGCNVHGNADIGKLVAKKILKVEHENAGTYSLLS 514
G L +A+ IE + + ++WG LL GC +H + + + VA+K+ ++E EN G Y L++
Sbjct: 499 GDLIKAYRFIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGYYVLMA 558
Query: 515 NMYASVGKWKEAANVRMKMKDKGLKKQPGCSWIEVGNTVQVFVVGDKSHSQSEMLEYLLL 574
N+YA KW++ +R ++ +GL+K PGCSWIE+ V +FV GD S+ ++E +E L
Sbjct: 559 NIYAEAEKWEQVKRLRKRIGQRGLRKNPGCSWIEIKGRVNIFVAGDSSNPETENIEAFLR 618
Query: 575 GLHTKMKKFG 584
+ +M + G
Sbjct: 619 KVRARMIEEG 628
Score = 135 bits (341), Expect = 3e-30
Identities = 93/361 (25%), Positives = 174/361 (47%), Gaps = 47/361 (13%)
Query: 129 FYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRMPKRNL----VSWNIIIKALARCGRI 184
F E+ L WNI++ ++G ++ LF++M + +++ + K+ + +
Sbjct: 1 FDEVKIEKALFWNILMNELAKSGDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSV 60
Query: 185 EDA-QWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMASWNAMLT 243
Q H + G RN V N+++ Y +N+R+D A ++F+ M ERD+ SWN+++
Sbjct: 61 HGGEQLHGFILKSGFGE-RNSVG-NSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIIN 118
Query: 244 GFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNG 303
G Y +GL+E+ L +F +M + G++ +
Sbjct: 119 G-------------------------------YVSNGLAEKGLSVFVQMLVS-GIEIDLA 146
Query: 304 TFVTVLGACSGLASLTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDG 363
T V+V C+ ++ G+ +H + K F R + L++MYSKCG+L A+ +F +
Sbjct: 147 TIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAVFRE- 205
Query: 364 LLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDE 423
+ R ++S+ MIA YA G EA+ LF +M+E G + T +L C+ L+DE
Sbjct: 206 -MSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDE 264
Query: 424 GIQYFDKLLKNRSIQVKEDHYA--CLVDLCGRAGRLKEAFYIIEGLGVKLSLSVWGPLLA 481
G + + + +N + D + L+D+ + G ++EA + + VK +S W ++
Sbjct: 265 GKRVHEWIKEN---DLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIIS-WNTIIG 320
Query: 482 G 482
G
Sbjct: 321 G 321
Score = 94.0 bits (232), Expect = 1e-17
Identities = 94/403 (23%), Positives = 163/403 (40%), Gaps = 80/403 (19%)
Query: 42 EMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQP- 100
E + CN+ + K G D A+ +F +M R +T+MI GY + G EA KLF++
Sbjct: 179 EDRFCNTLLDMYSKCGDLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEME 238
Query: 101 ------------------------DAKKSVVTW-------------TTMVNGYVKLNQIE 123
D K V W +++ Y K ++
Sbjct: 239 EEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQ 298
Query: 124 EAESLFYEMPERNELSWNIMIGGYGQNGQIEKALDLFRRM--PKRNLVSWNIIIKALARC 181
EAE +F EM ++ +SWN +IGGY +N +AL LF + KR + L C
Sbjct: 299 EAELVFSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPAC 358
Query: 182 GRI----EDAQWHLIRCRKGMMPLRNVVSWNAMITGYAQNRRLDEALELFERMPERDMAS 237
+ + + H R G R+V N+++ YA+ L A LF+ + +D+ S
Sbjct: 359 ASLSAFDKGREIHGYIMRNGYFSDRHVA--NSLVDMYAKCGALLLAHMLFDDIASKDLVS 416
Query: 238 WNAMLTGFFQNGELNRAEKLFAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGG 297
W M+ G Y HG +EA+ +F +M+ G
Sbjct: 417 WTVMIAG-------------------------------YGMHGFGKEAIALFNQMR-QAG 444
Query: 298 LKPNNGTFVTVLGACSGLASLTEGQQIHQLI-SKTGFQENTRVVSALINMYSKCGELHIA 356
++ + +FV++L ACS + EG + ++ + + + +++M ++ G+L A
Sbjct: 445 IEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVDMLARTGDLIKA 504
Query: 357 RKIFDDGLLRQRDLISWNGMIAAYAHHGYGNEAINLFNKMQEL 399
+ F + + D W ++ H A + K+ EL
Sbjct: 505 YR-FIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFEL 546
Score = 88.2 bits (217), Expect = 7e-16
Identities = 57/204 (27%), Positives = 106/204 (51%), Gaps = 6/204 (2%)
Query: 258 FAELPQKDVITWTSMMTGYAQHGLSEEALKMFTKMQANGGLKPNNGTFVTVLGACSGLAS 317
F E+ + + W +M A+ G ++ +F KM ++G ++ ++ TF V + S L S
Sbjct: 1 FDEVKIEKALFWNILMNELAKSGDFSGSIGLFKKMMSSG-VEMDSYTFSCVSKSFSSLRS 59
Query: 318 LTEGQQIHQLISKTGFQENTRVVSALINMYSKCGELHIARKIFDDGLLRQRDLISWNGMI 377
+ G+Q+H I K+GF E V ++L+ Y K + ARK+FD+ + +RD+ISWN +I
Sbjct: 60 VHGGEQLHGFILKSGFGERNSVGNSLVAFYLKNQRVDSARKVFDE--MTERDVISWNSII 117
Query: 378 AAYAHHGYGNEAINLFNKMQELGFQANDVTYVELLTACSHAGLVDEGIQYFDKLLKNRSI 437
Y +G + +++F +M G + + T V + C+ + L+ G + ++
Sbjct: 118 NGYVSNGLAEKGLSVFVQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVHS--IGVKAC 175
Query: 438 QVKEDHYA-CLVDLCGRAGRLKEA 460
+ED + L+D+ + G L A
Sbjct: 176 FSREDRFCNTLLDMYSKCGDLDSA 199
Score = 35.4 bits (80), Expect = 5.3
Identities = 23/59 (38%), Positives = 28/59 (46%)
Query: 41 SEMKRCNSDISRLCKEGRTDHARKLFDQMPQRDKHLWTTMIDGYIKCGAIKEARKLFDQ 99
S+ NS + K G A LFD + +D WT MI GY G KEA LF+Q
Sbjct: 381 SDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAIALFNQ 439
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,020,196,256
Number of Sequences: 2540612
Number of extensions: 42507015
Number of successful extensions: 122590
Number of sequences better than 10.0: 1219
Number of HSP's better than 10.0 without gapping: 835
Number of HSP's successfully gapped in prelim test: 387
Number of HSP's that attempted gapping in prelim test: 92456
Number of HSP's gapped (non-prelim): 7218
length of query: 597
length of database: 863,360,394
effective HSP length: 134
effective length of query: 463
effective length of database: 522,918,386
effective search space: 242111212718
effective search space used: 242111212718
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0026.14


