

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0024.11
(148 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
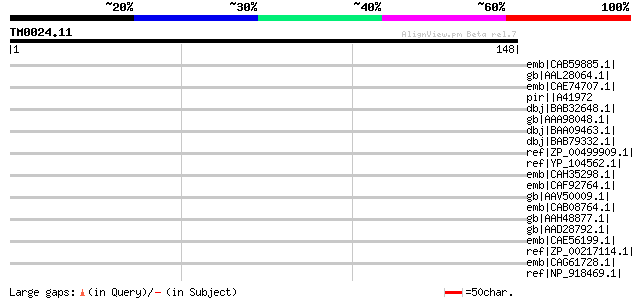
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB59885.1| SPAC1071.09c [Schizosaccharomyces pombe] gi|1911... 35 0.37
gb|AAL28064.1| replication protein [Aneurinibacillus thermoaerop... 35 0.37
emb|CAE74707.1| Hypothetical protein CBG22522 [Caenorhabditis br... 35 0.37
pir||A41972 replication-associated protein homolog RepE - Bacill... 35 0.49
dbj|BAB32648.1| replication protein [Bacillus sp. B-3] gi|131289... 35 0.49
gb|AAA98048.1| replicative protein 35 0.49
dbj|BAA09463.1| troponin T [Halocynthia roretzi] 34 0.64
dbj|BAB79332.1| ORF3 [TT virus] 34 0.64
ref|ZP_00499909.1| COG1840: ABC-type Fe3+ transport system, peri... 34 0.83
ref|YP_104562.1| ABC transporter, periplasmic substrate-binding ... 34 0.83
emb|CAH35298.1| putative exported protein [Burkholderia pseudoma... 34 0.83
emb|CAF92764.1| unnamed protein product [Tetraodon nigroviridis] 34 0.83
gb|AAV50009.1| anthranilate N-hydroxycinnamoyl/benzoyltransferas... 33 1.1
emb|CAB08764.1| SPAC57A7.06 [Schizosaccharomyces pombe] gi|19114... 33 1.9
gb|AAH48877.1| RNA polymerase III 80 kDa subunit RPC5 [Danio rer... 32 2.4
gb|AAD28792.1| U2 small nuclear ribonucleoprotein auxiliary fact... 32 2.4
emb|CAE56199.1| Hypothetical protein CBG23823 [Caenorhabditis br... 32 2.4
ref|ZP_00217114.1| COG1840: ABC-type Fe3+ transport system, peri... 32 2.4
emb|CAG61728.1| unnamed protein product [Candida glabrata CBS138... 32 3.2
ref|NP_918469.1| P0002B05.14 [Oryza sativa (japonica cultivar-gr... 32 3.2
>emb|CAB59885.1| SPAC1071.09c [Schizosaccharomyces pombe]
gi|19115271|ref|NP_594359.1| hypothetical protein
SPAC1071.09c [Schizosaccharomyces pombe 972h-]
gi|7490498|pir||T37491 dnaj protein - fission yeast
(Schizosaccharomyces pombe)
Length = 282
Score = 35.0 bits (79), Expect = 0.37
Identities = 24/110 (21%), Positives = 55/110 (49%), Gaps = 9/110 (8%)
Query: 14 TKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRS-----KE 68
T +D DF ++ + ++QG++ +EF+ S Q S+ + LK+ ++ S +E
Sbjct: 104 TDADIDFDWKEWLDELYQGVVSGETLNEFKASYQYSEEEKCDVLKAYEKGKGSMDVILEE 163
Query: 69 ISVARLPESEYYRAYSPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSE 118
+ + + + +R + N + +GKI + +F+ + K +R+ +E
Sbjct: 164 VMCCEISDEDRFR----QVINNAIKDGKISKYKRFAPNEKKRKRRAKAAE 209
>gb|AAL28064.1| replication protein [Aneurinibacillus thermoaerophilus]
Length = 258
Score = 35.0 bits (79), Expect = 0.37
Identities = 25/81 (30%), Positives = 42/81 (50%), Gaps = 3/81 (3%)
Query: 7 VTTIPKGTKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRS 66
V TI + K + F T+ RN + L E + + F+ ++++ K + S K +FR+
Sbjct: 18 VETINEREKVRWVFLTLTVRNTDAES-LPETIKAMFKGFQRLTNY--KAFKTSVKGYFRA 74
Query: 67 KEISVARLPESEYYRAYSPEF 87
E++ R PESE + Y P F
Sbjct: 75 LEVTKNRDPESESFGTYHPHF 95
>emb|CAE74707.1| Hypothetical protein CBG22522 [Caenorhabditis briggsae]
Length = 1187
Score = 35.0 bits (79), Expect = 0.37
Identities = 36/131 (27%), Positives = 69/131 (52%), Gaps = 19/131 (14%)
Query: 15 KSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRY--LKSQKRWFRSKEIS-- 70
+SDYD STI F + G+ R++ E + S+ IS + K++ LK ++ F+ E+
Sbjct: 200 RSDYDLSTIYFTKFI--GLPRQLKTVEIKFSEVISAMDMKKFNKLKESEKIFKPVELMIN 257
Query: 71 -VARLPESEYYRAYSPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSEIAY--SGHARR 127
+ R E E +++ +P ++ + + +++F+F + SSK+++A+ S + R
Sbjct: 258 LIMRRREIEKFQS-TPNYK---IAKKNFEQVFEFVT-----ILDSSKNDVAFLKSLISSR 308
Query: 128 KAANSCHAYTD 138
N H YTD
Sbjct: 309 TQKNILH-YTD 318
>pir||A41972 replication-associated protein homolog RepE - Bacillus coagulans
Length = 259
Score = 34.7 bits (78), Expect = 0.49
Identities = 21/68 (30%), Positives = 37/68 (53%), Gaps = 3/68 (4%)
Query: 20 FSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEY 79
F T+ RN + L E +++ F+ +++K K + S K +FR+ E++ R P SE+
Sbjct: 31 FLTLTVRNTSPES-LPETISAMFEGFNRLTKY--KAFKTSVKGYFRALEVTKNRDPHSEW 87
Query: 80 YRAYSPEF 87
+ Y P F
Sbjct: 88 FGTYHPHF 95
>dbj|BAB32648.1| replication protein [Bacillus sp. B-3] gi|13128940|ref|NP_077030.1|
replication protein [Bacillus sp. B-3]
Length = 290
Score = 34.7 bits (78), Expect = 0.49
Identities = 24/81 (29%), Positives = 42/81 (51%), Gaps = 3/81 (3%)
Query: 7 VTTIPKGTKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRS 66
V T+ + + F T+ RN + L E +++ F+ +++ I K + S K +FR+
Sbjct: 49 VETVNQRENVKWLFLTLTVRNTSPES-LPETISAMFEGFRRL--IQYKAFKTSVKGYFRA 105
Query: 67 KEISVARLPESEYYRAYSPEF 87
E++ R PESE + Y P F
Sbjct: 106 LEVTKNRDPESESFGTYHPHF 126
>gb|AAA98048.1| replicative protein
Length = 259
Score = 34.7 bits (78), Expect = 0.49
Identities = 21/68 (30%), Positives = 37/68 (53%), Gaps = 3/68 (4%)
Query: 20 FSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEY 79
F T+ RN + L E +++ F+ +++K K + S K +FR+ E++ R P SE+
Sbjct: 31 FLTLTVRNTSPES-LPETISAMFEGFNRLTKY--KAFKTSVKGYFRALEVTKNRDPHSEW 87
Query: 80 YRAYSPEF 87
+ Y P F
Sbjct: 88 FGTYHPHF 95
>dbj|BAA09463.1| troponin T [Halocynthia roretzi]
Length = 248
Score = 34.3 bits (77), Expect = 0.64
Identities = 36/135 (26%), Positives = 55/135 (40%), Gaps = 19/135 (14%)
Query: 4 TTSVTTIPKGTKSDYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRW 63
TTS+ +P G K D D T K R+ L+ ++ + F+ KQ + E+ L+ +KR
Sbjct: 24 TTSIPRLPDGEKVDLDVITQK-RHQKDLEELQALITAHFEQRKQEEEQLEELRLRLEKR- 81
Query: 64 FRSKEISVARLPESEYYRAYSPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSEIA--- 120
KE RL E E E + + +R K +D K + K + A
Sbjct: 82 ---KEERAERLRIRE-------EKTKERVAREREERKHKEEEDEKKRKEEEEKKKAAIAN 131
Query: 121 ----YSGHARRKAAN 131
Y G+ R N
Sbjct: 132 MSLHYGGYLARAEKN 146
>dbj|BAB79332.1| ORF3 [TT virus]
Length = 277
Score = 34.3 bits (77), Expect = 0.64
Identities = 32/143 (22%), Positives = 58/143 (40%), Gaps = 13/143 (9%)
Query: 11 PKGTKSDYDFSTIKFRNHVHQG---------ILREILASEFQNSKQISKIWEKRYLKSQK 61
P T +Y + T ++ +H + R +L E QN+ ++ ++++
Sbjct: 116 PINTLEEYKYQTRRYSDHPQSSTHGTSDVDCLARRVL-KECQNTNRLMTLFQQASSSKDP 174
Query: 62 RWFRSKEISVARLPESEYYRAYSPEFRNETLVEGKIKRIFKFSKDSKSH-LKRSSKSEIA 120
W S + + P+ + YS + + K K SK SK H SS S +
Sbjct: 175 DWKPSTK--EPKKPQKKTPTLYSKHSKKSKRAAARKKNSHKKSKRSKKHNSSSSSNSSSS 232
Query: 121 YSGHARRKAANSCHAYTDSTRES 143
S +R ++ S +DS +ES
Sbjct: 233 NSESSRGESDTSSETSSDSEKES 255
>ref|ZP_00499909.1| COG1840: ABC-type Fe3+ transport system, periplasmic component
[Burkholderia pseudomallei S13]
gi|67755337|ref|ZP_00494233.1| COG1840: ABC-type Fe3+
transport system, periplasmic component [Burkholderia
pseudomallei Pasteur] gi|67736573|ref|ZP_00487431.1|
COG1840: ABC-type Fe3+ transport system, periplasmic
component [Burkholderia pseudomallei 668]
gi|67714276|ref|ZP_00483637.1| COG1840: ABC-type Fe3+
transport system, periplasmic component [Burkholderia
pseudomallei 1710b] gi|67682037|ref|ZP_00476321.1|
COG1840: ABC-type Fe3+ transport system, periplasmic
component [Burkholderia pseudomallei 1710a]
gi|67670350|ref|ZP_00467157.1| COG1840: ABC-type Fe3+
transport system, periplasmic component [Burkholderia
pseudomallei 1655] gi|67651968|ref|ZP_00449389.1|
COG1840: ABC-type Fe3+ transport system, periplasmic
component [Burkholderia mallei SAVP1]
gi|67646222|ref|ZP_00444486.1| COG1840: ABC-type Fe3+
transport system, periplasmic component [Burkholderia
mallei NCTC 10247] gi|67643396|ref|ZP_00442142.1|
COG1840: ABC-type Fe3+ transport system, periplasmic
component [Burkholderia mallei GB8 horse 4]
gi|67636302|ref|ZP_00435249.1| COG1840: ABC-type Fe3+
transport system, periplasmic component [Burkholderia
mallei 10399] gi|67630391|ref|ZP_00430248.1| COG1840:
ABC-type Fe3+ transport system, periplasmic component
[Burkholderia mallei 10229]
Length = 331
Score = 33.9 bits (76), Expect = 0.83
Identities = 28/80 (35%), Positives = 40/80 (50%), Gaps = 6/80 (7%)
Query: 10 IPK-GTKS-DYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSK 67
IPK GT S Y S +K H G R++L +F S + K+W YL+ +
Sbjct: 237 IPKEGTISVPYVMSLVKGAPHEANG--RKVL--DFVLSDEGQKLWANAYLRPVRAQALGA 292
Query: 68 EISVARLPESEYYRAYSPEF 87
+++ LP SEY RA S +F
Sbjct: 293 DVAAQFLPASEYARAKSVDF 312
>ref|YP_104562.1| ABC transporter, periplasmic substrate-binding protein
[Burkholderia mallei ATCC 23344]
gi|52427467|gb|AAU48060.1| ABC transporter, periplasmic
substrate-binding protein [Burkholderia mallei ATCC
23344]
Length = 353
Score = 33.9 bits (76), Expect = 0.83
Identities = 28/80 (35%), Positives = 40/80 (50%), Gaps = 6/80 (7%)
Query: 10 IPK-GTKS-DYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSK 67
IPK GT S Y S +K H G R++L +F S + K+W YL+ +
Sbjct: 259 IPKEGTISVPYVMSLVKGAPHEANG--RKVL--DFVLSDEGQKLWANAYLRPVRAQALGA 314
Query: 68 EISVARLPESEYYRAYSPEF 87
+++ LP SEY RA S +F
Sbjct: 315 DVAAQFLPASEYARAKSVDF 334
>emb|CAH35298.1| putative exported protein [Burkholderia pseudomallei K96243]
gi|53718939|ref|YP_107925.1| hypothetical protein
BPSL1303 [Burkholderia pseudomallei K96243]
Length = 353
Score = 33.9 bits (76), Expect = 0.83
Identities = 28/80 (35%), Positives = 40/80 (50%), Gaps = 6/80 (7%)
Query: 10 IPK-GTKS-DYDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSK 67
IPK GT S Y S +K H G R++L +F S + K+W YL+ +
Sbjct: 259 IPKEGTISVPYVMSLVKGAPHEANG--RKVL--DFVLSDEGQKLWASAYLRPVRAQALGA 314
Query: 68 EISVARLPESEYYRAYSPEF 87
+++ LP SEY RA S +F
Sbjct: 315 DVAAQFLPASEYARAKSVDF 334
>emb|CAF92764.1| unnamed protein product [Tetraodon nigroviridis]
Length = 229
Score = 33.9 bits (76), Expect = 0.83
Identities = 19/53 (35%), Positives = 28/53 (51%), Gaps = 5/53 (9%)
Query: 41 EFQNSKQISKIWEKRYLKSQ-----KRWFRSKEISVARLPESEYYRAYSPEFR 88
E Q ++ + WEK+ +++ KR+FR +E S PE E YSP FR
Sbjct: 84 ELQELREKQEGWEKQRRETENKNKWKRFFRRREASNENSPEKEQQTRYSPLFR 136
>gb|AAV50009.1| anthranilate N-hydroxycinnamoyl/benzoyltransferase [Malus x
domestica]
Length = 398
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/67 (29%), Positives = 31/67 (45%)
Query: 61 KRWFRSKEISVARLPESEYYRAYSPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSEIA 120
KRW IS R+P S+ S ++ V +R+F FSK+ + LK + +EI
Sbjct: 146 KRWLLDGTISPIRIPFSKIEDQVSDKYIGNIRVPLLKERVFHFSKEKIAQLKAKANAEIQ 205
Query: 121 YSGHARR 127
R+
Sbjct: 206 LENTKRK 212
>emb|CAB08764.1| SPAC57A7.06 [Schizosaccharomyces pombe]
gi|19114287|ref|NP_593375.1| hypothetical protein
SPAC57A7.06 [Schizosaccharomyces pombe 972h-]
gi|7490688|pir||T38948 hypothetical coiled-coil protein
- fission yeast (Schizosaccharomyces pombe)
gi|3183291|sp|P87137|YDM6_SCHPO Hypothetical protein
C57A7.06 in chromosome I
Length = 929
Score = 32.7 bits (73), Expect = 1.9
Identities = 26/99 (26%), Positives = 49/99 (49%), Gaps = 10/99 (10%)
Query: 34 LREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRAYSPEFRNETLV 93
+RE++ E + +K+++KI K Y K +K KE +A +P+SE + NE +
Sbjct: 436 MRELMFREERKAKRVAKIKSKTYRKIRK---NRKEKEMALIPKSE------EDLENERIK 486
Query: 94 EGKIKRIFKFSKDSKSHLKRSSKS-EIAYSGHARRKAAN 131
+ + + + ++ K+ + K E A G R+A N
Sbjct: 487 SEEARALERMTQRHKNTSSWTRKMLERASHGEGTREAVN 525
>gb|AAH48877.1| RNA polymerase III 80 kDa subunit RPC5 [Danio rerio]
gi|47086541|ref|NP_997919.1| RNA polymerase III 80 kDa
subunit RPC5 [Danio rerio]
Length = 696
Score = 32.3 bits (72), Expect = 2.4
Identities = 15/40 (37%), Positives = 23/40 (57%)
Query: 91 TLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSGHARRKAA 130
T ++ K++++F FSKD H K + +SG R KAA
Sbjct: 425 TGIQSKLEKVFNFSKDDFVHKKDVQIEPVHFSGDQRLKAA 464
>gb|AAD28792.1| U2 small nuclear ribonucleoprotein auxiliary factor subunit-related
protein [Takifugu rubripes]
Length = 605
Score = 32.3 bits (72), Expect = 2.4
Identities = 29/83 (34%), Positives = 36/83 (42%), Gaps = 13/83 (15%)
Query: 53 EKRYLKSQKRWFRSKEISVARLPESEYYRAY-----SPEFRNETLVEGKI--------KR 99
E+RY + +KR RSKE +R E R SP+ R+E G KR
Sbjct: 454 EERYSRGEKRETRSKERGKSRSSSREGKRTKRSRERSPKKRSEHENNGSADCKKRRHHKR 513
Query: 100 IFKFSKDSKSHLKRSSKSEIAYS 122
K K SK H KRS S+ S
Sbjct: 514 TKKSKKKSKKHKKRSRVSDAGTS 536
>emb|CAE56199.1| Hypothetical protein CBG23823 [Caenorhabditis briggsae]
Length = 278
Score = 32.3 bits (72), Expect = 2.4
Identities = 36/114 (31%), Positives = 52/114 (45%), Gaps = 15/114 (13%)
Query: 24 KFRNHVHQGILREILASEFQNSK-------QISKIWEKRYLK---SQKRWFRSKEISVAR 73
KF N Q + R EF+NS+ +I K LK S+ FR+ EI R
Sbjct: 163 KFGNSEIQKV-RNSEIPEFRNSEIQKFGNSEIQKFKNSEILKFRNSEMPEFRNSEIQKFR 221
Query: 74 LPESEYYR-AYSPEFRNETLVEGKIKRIFKFSKDSKSHLKRSSKSEIAYSGHAR 126
E + +R A PEFRN + E + I KF S +++ SEI G+++
Sbjct: 222 NSEIQKFRNAGIPEFRNSEIPEFRNSEIQKF---WNSEIQKFRNSEIREFGNSK 272
>ref|ZP_00217114.1| COG1840: ABC-type Fe3+ transport system, periplasmic component
[Burkholderia cepacia R18194]
Length = 342
Score = 32.3 bits (72), Expect = 2.4
Identities = 23/70 (32%), Positives = 35/70 (49%), Gaps = 4/70 (5%)
Query: 18 YDFSTIKFRNHVHQGILREILASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPES 77
Y S +K H G +++L +F S + K+W YL+ + S +I+ LP S
Sbjct: 258 YVMSLVKGAPHDANG--KKVL--DFVLSDEGQKLWANAYLRPVRAQTLSADIASKFLPAS 313
Query: 78 EYYRAYSPEF 87
EY RA S +F
Sbjct: 314 EYARAKSVDF 323
>emb|CAG61728.1| unnamed protein product [Candida glabrata CBS138]
gi|50292665|ref|XP_448765.1| unnamed protein product
[Candida glabrata]
Length = 1326
Score = 32.0 bits (71), Expect = 3.2
Identities = 21/49 (42%), Positives = 29/49 (58%), Gaps = 2/49 (4%)
Query: 36 EILASEFQNSKQISKIWE--KRYLKSQKRWFRSKEISVARLPESEYYRA 82
++L SE Q K I+K KRY++S KR FRS EI +R+ + RA
Sbjct: 526 QLLNSENQTFKDINKKSHEFKRYVQSAKRTFRSMEIEKSRIIWPQTVRA 574
>ref|NP_918469.1| P0002B05.14 [Oryza sativa (japonica cultivar-group)]
gi|15289795|dbj|BAB63494.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 195
Score = 32.0 bits (71), Expect = 3.2
Identities = 21/74 (28%), Positives = 36/74 (48%), Gaps = 4/74 (5%)
Query: 38 LASEFQNSKQISKIWEKRYLKSQKRWFRSKEISVARLPESEYYRAYSPEFRNETLVEGKI 97
+A E + + Q + + KS++RW + + R Y+AYS E + V+G
Sbjct: 121 VAEEERGTGQAAAEAAAKRSKSKRRWLALGDPDMERKRRVASYKAYSVEGK----VKGSF 176
Query: 98 KRIFKFSKDSKSHL 111
++ FK+ KD HL
Sbjct: 177 RKSFKWIKDRYLHL 190
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 219,703,255
Number of Sequences: 2540612
Number of extensions: 7800365
Number of successful extensions: 24272
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 24251
Number of HSP's gapped (non-prelim): 67
length of query: 148
length of database: 863,360,394
effective HSP length: 124
effective length of query: 24
effective length of database: 548,324,506
effective search space: 13159788144
effective search space used: 13159788144
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0024.11


