

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.17
(341 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
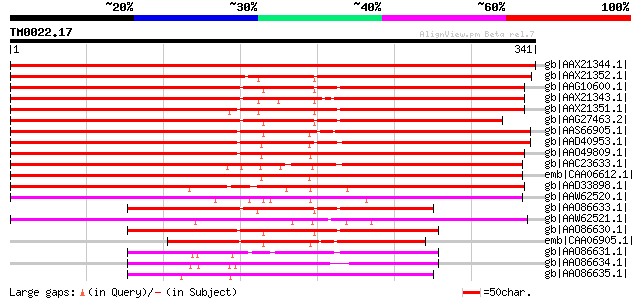
Score E
Sequences producing significant alignments: (bits) Value
gb|AAX21344.1| phantastica transcription factor b [Lotus cornicu... 683 0.0
gb|AAX21352.1| phantastica transcription factor b [Glycine max] 473 e-132
gb|AAG10600.1| MYB-related transcription factor PHAN1 [Pisum sat... 468 e-130
gb|AAX21343.1| phantastica transcription factor a [Lotus cornicu... 462 e-129
gb|AAX21351.1| phantastica transcription factor a [Glycine max] 454 e-126
gb|AAG27463.2| myb-related transcription factor [Medicago trunca... 443 e-123
gb|AAS66905.1| phantastica [Nicotiana tabacum] 433 e-120
gb|AAD40953.1| phantastica [Lycopersicon esculentum] 424 e-117
gb|AAO49809.1| phantastica-like MYB protein [Eschscholzia califo... 409 e-113
gb|AAC23633.1| putative MYB family transcription factor [Arabido... 400 e-110
emb|CAA06612.1| MYB-related transcription factor [Antirrhinum ma... 393 e-108
gb|AAD33898.1| rough sheath 2 [Zea mays] gi|4583431|gb|AAD25082.... 338 1e-91
gb|AAW62520.1| PHANTASTICA-like protein [Selaginella kraussiana] 243 5e-63
gb|AAO86633.1| phantastica transcription factor [Acacia hindsii] 236 6e-61
gb|AAW62521.1| PHANTASTICA-like protein [Selaginella viticulosa] 221 2e-56
gb|AAO86630.1| phantastica transcription factor [Aquilegia formosa] 220 4e-56
emb|CAA06905.1| phantastica protein [Nicotiana tabacum] gi|74891... 158 2e-37
gb|AAO86631.1| phantastica transcription factor [Vitex cannabifo... 137 6e-31
gb|AAO86634.1| phantastica transcription factor [Pachira aquatica] 136 8e-31
gb|AAO86635.1| phantastica transcription factor [Fraxinus americ... 132 1e-29
>gb|AAX21344.1| phantastica transcription factor b [Lotus corniculatus var.
japonicus]
Length = 341
Score = 683 bits (1763), Expect = 0.0
Identities = 341/341 (100%), Positives = 341/341 (100%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP
Sbjct: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPS 180
ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPS
Sbjct: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPS 180
Query: 181 SISVTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGFSKELEEGH 240
SISVTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGFSKELEEGH
Sbjct: 181 SISVTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGFSKELEEGH 240
Query: 241 RALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQTAALNRIENACRE 300
RALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQTAALNRIENACRE
Sbjct: 241 RALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQTAALNRIENACRE 300
Query: 301 QLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQTGAP 341
QLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQTGAP
Sbjct: 301 QLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQTGAP 341
>gb|AAX21352.1| phantastica transcription factor b [Glycine max]
Length = 357
Score = 473 bits (1217), Expect = e-132
Identities = 252/352 (71%), Positives = 284/352 (80%), Gaps = 15/352 (4%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRW +EEDALL +YV+QYGPREWNLVSQRMNT LNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWRAEEDALLRSYVKQYGPREWNLVSQRMNTYLNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQA YGNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQREK EIN I P
Sbjct: 61 SLTEEEQRLVIHLQAKYGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREKKEINRIADP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTN----SSAMLPSWLSNYDSTS 176
I+++KYEH+LE FAEKLVKE PSF MAAS AFL T+ +S++ PSWLSN S +
Sbjct: 121 INNSKYEHILESFAEKLVKERPSPSFVMAASDG-AFLLTDTPAPASSLRPSWLSNSSSAA 179
Query: 177 T-PPSSISVTLSLSPSTVATP--------RGLENNAPFVLRNVTAHNGSVPSFSDHILMS 227
PSS+SV LSLS STVATP RG +NAPFVL NV+A +G++P+ SD + MS
Sbjct: 180 AIGPSSLSVKLSLSSSTVATPPFSWLPPERG-PDNAPFVLGNVSALHGAIPTLSDSMHMS 238
Query: 228 ELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQ 287
++V KELEEGHRALA HKKEA WRL R+ELQLESEKA RRRE +EEFEA IKALQEE+
Sbjct: 239 QMVEHCKELEEGHRALATHKKEAAWRLSRVELQLESEKANRRREKIEEFEAKIKALQEEE 298
Query: 288 TAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQTG 339
AAL RIE REQL LRRDAE+KEQKLAE+W +KHLR TRLLEQ+ + G
Sbjct: 299 KAALGRIEAEYREQLAALRRDAENKEQKLAEQWDAKHLRFTRLLEQLGCRAG 350
>gb|AAG10600.1| MYB-related transcription factor PHAN1 [Pisum sativum]
Length = 359
Score = 468 bits (1203), Expect = e-130
Identities = 246/346 (71%), Positives = 282/346 (81%), Gaps = 15/346 (4%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MK+RQRW +EEDALL AYV+QYGPREWNLVSQRMNTPLNRD KSCLERWKNYLKPGIKKG
Sbjct: 5 MKDRQRWRAEEDALLRAYVKQYGPREWNLVSQRMNTPLNRDAKSCLERWKNYLKPGIKKG 64
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQ LVI LQA +GNKWKKIAA+VPGRTAKRLGKWWEV+KEKQQRE IN V P
Sbjct: 65 SLTEEEQHLVISLQATHGNKWKKIAAQVPGRTAKRLGKWWEVFKEKQQRETKGINKTVDP 124
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSA----MLPSWLSNYDSTS 176
I+D+KYEH+LE FAEKLVKE PSF MAA SN ++LHT++ A +LPSWLSN ++T+
Sbjct: 125 INDSKYEHILESFAEKLVKERPSPSFVMAA-SNSSYLHTDAQAATPGLLPSWLSNSNNTA 183
Query: 177 -TPPSSISVTLSLSPSTVATP-------RGLENNAPFVLRNVTAHNGSVPSFSDHILMSE 228
P+S SVTLSLSPSTVA P RG +NAP VL NV H G+V S+ ++++MSE
Sbjct: 184 PVRPNSPSVTLSLSPSTVAAPPPWMQPVRG-PDNAPLVLGNVAPH-GAVLSYGENMVMSE 241
Query: 229 LVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQT 288
L+ KELEEGH ALAAHKKEA WRL R+ELQLESEKA RRRE +EE EA IKAL+EEQ
Sbjct: 242 LIDCCKELEEGHHALAAHKKEAAWRLSRVELQLESEKASRRREKMEEIEAKIKALREEQA 301
Query: 289 AALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQM 334
AL+RIE REQL GLRRDAE+KEQKLAE+W +KHLRLT+ LEQ+
Sbjct: 302 VALDRIEGEYREQLAGLRRDAEAKEQKLAEQWAAKHLRLTKFLEQV 347
>gb|AAX21343.1| phantastica transcription factor a [Lotus corniculatus var.
japonicus]
Length = 356
Score = 462 bits (1189), Expect = e-129
Identities = 248/347 (71%), Positives = 283/347 (81%), Gaps = 16/347 (4%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRW+SEEDALL AYV+QYGPREW+LVSQRMNT L+RD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWTSEEDALLCAYVKQYGPREWHLVSQRMNTTLHRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQA +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQREK EI+ + P
Sbjct: 61 SLTEEEQRLVIRLQAKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREKQEISKSIGP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTN------SSAMLPSWLSNYD- 173
+ D+KY+H+LE FAEKLVKEH PS+ MAA SN FLHT+ +SA+LP WLSN
Sbjct: 121 VDDSKYDHILETFAEKLVKEHPSPSYLMAA-SNGPFLHTDTPAATPASALLPPWLSNSSN 179
Query: 174 --STSTPPSSISVTLSLSPSTVATP----RGLENNAPFVLRNVTAHNGSVPSFSDHILMS 227
+T+ P S SVTLSLSPSTVA P RGLENNA + N TA +G+VP+FSD++L+S
Sbjct: 180 NPATAGQPPSPSVTLSLSPSTVAGPPPPWRGLENNA-LAMAN-TAPHGTVPAFSDNMLVS 237
Query: 228 ELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQ 287
ELV KELEE H ALAAHKKEA WRLRR+ELQLESEKA RRRE +EE EA IKAL+E+Q
Sbjct: 238 ELVDCCKELEEVHGALAAHKKEATWRLRRVELQLESEKANRRREKIEETEAKIKALREQQ 297
Query: 288 TAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQM 334
AAL RIE REQL GLRRDAE+KEQKLAE+WT KH RL + +EQ+
Sbjct: 298 NAALERIEAEYREQLAGLRRDAETKEQKLAEQWTVKHSRLMKFMEQI 344
>gb|AAX21351.1| phantastica transcription factor a [Glycine max]
Length = 361
Score = 454 bits (1168), Expect = e-126
Identities = 244/351 (69%), Positives = 279/351 (78%), Gaps = 19/351 (5%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MK+RQRW +EEDALL AYV+QYGPREWNLVSQRMNTPLNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKDRQRWRAEEDALLRAYVKQYGPREWNLVSQRMNTPLNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQA +GNKWKKIAA+VPGRTAKRLGKWWEV+KEKQQRE + + P
Sbjct: 61 SLTEEEQRLVINLQATHGNKWKKIAAQVPGRTAKRLGKWWEVFKEKQQRETKGNSCTIDP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEH----TLPSFAMAASSNEAFLHTN----SSAMLPSWLSNY 172
ISD+KYEH+LE FAEKLVKE T SF M A+SN +FLH + + A+LPSWLSN
Sbjct: 121 ISDSKYEHILESFAEKLVKERPSTSTSTSFVM-ATSNSSFLHADAPAPAPALLPSWLSNS 179
Query: 173 DSTS-TPPSSISVTLSLSPSTVATP--------RGLENNAPFVLRNVTAHNGSVPSFSDH 223
+ T+ P S SVTLSLSPSTVA P RG +N +P VL NV H G+V +F ++
Sbjct: 180 NGTAPVRPPSPSVTLSLSPSTVAAPPPWMQPPVRGQDNASPLVLGNVAPH-GAVLAFGEN 238
Query: 224 ILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKAL 283
++MSELV KEL+E H ALA HKKEA WRL R+ELQLESEKA RRRE +EE EA IKAL
Sbjct: 239 MVMSELVECCKELDEVHHALAGHKKEAAWRLSRVELQLESEKAGRRREKMEEIEAKIKAL 298
Query: 284 QEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQM 334
+EEQTAAL+RIE REQL GLRRDAESKEQKLAE+W +KHLRLT+ LEQ+
Sbjct: 299 REEQTAALDRIEAEYREQLAGLRRDAESKEQKLAEQWAAKHLRLTKFLEQV 349
>gb|AAG27463.2| myb-related transcription factor [Medicago truncatula]
Length = 334
Score = 443 bits (1139), Expect = e-123
Identities = 238/333 (71%), Positives = 269/333 (80%), Gaps = 16/333 (4%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MK+RQRW +EEDALL AYV+QYGPREWNLVSQRMNTPLNRD KSCLERWKNYLKPGIKKG
Sbjct: 4 MKDRQRWRAEEDALLRAYVKQYGPREWNLVSQRMNTPLNRDAKSCLERWKNYLKPGIKKG 63
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE-KIEINGIVS 119
SLT+EEQRLVI LQA +GNKWKKIAA+VPGRTAKRLGKWWEV+KEKQQRE K IN V
Sbjct: 64 SLTEEEQRLVISLQATHGNKWKKIAAQVPGRTAKRLGKWWEVFKEKQQRETKGSINRTVD 123
Query: 120 PISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSA----MLPSWLSNYDST 175
PI+D+KYEH+LE FAEKLVKE PSF MAA SN ++LHT++ A +LPSWLSN ++
Sbjct: 124 PINDSKYEHILESFAEKLVKERPSPSFVMAA-SNSSYLHTDAQAPTPGLLPSWLSNSNNA 182
Query: 176 S-TPPSSISVTLSLSPSTVATP-------RGLENNAPFVLRNVTAHNGSVPSFSDHILMS 227
+ P+S SVTLSLSPSTVA P RG +NAP VL NV H G+V S+ + ++MS
Sbjct: 183 APVRPNSPSVTLSLSPSTVAAPPPWMQPVRG-PDNAPLVLGNVAPH-GAVLSYGESMVMS 240
Query: 228 ELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQ 287
ELV KELEE H ALAAHKKEA WRL R+ELQLESEKA RRRE +EE EA IKAL+EEQ
Sbjct: 241 ELVDCCKELEEVHHALAAHKKEAAWRLSRVELQLESEKASRRREKMEEIEAKIKALREEQ 300
Query: 288 TAALNRIENACREQLGGLRRDAESKEQKLAEKW 320
AL+RIE REQL GLRRDAE+KEQKLAE+W
Sbjct: 301 AVALDRIEGEYREQLAGLRRDAETKEQKLAEQW 333
>gb|AAS66905.1| phantastica [Nicotiana tabacum]
Length = 363
Score = 433 bits (1114), Expect = e-120
Identities = 232/357 (64%), Positives = 273/357 (75%), Gaps = 22/357 (6%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
M+ERQRW SEEDALL AYV+QYGP+EW+LVSQRMNT LNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MRERQRWRSEEDALLRAYVKQYGPKEWHLVSQRMNTALNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQA +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQ RE+ E N +V P
Sbjct: 61 SLTQEEQRLVIHLQAKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQHREQKENNKVVDP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSA-----MLPSWLSNYDST 175
+ + KY+H+LE FAEK+VKE ++P M A+SN FLH ++ A +LP WLSN +T
Sbjct: 121 VDEGKYDHILETFAEKIVKERSVPGLLM-ATSNGGFLHADAPAPSPQTLLPPWLSNSTAT 179
Query: 176 STPPS-SISVTLSLSPSTV-------------ATPRGLENNAPFVLRNVTAHNGSVPSFS 221
ST S S SVTLSLSPSTV T RG E NAP +L + H+G P
Sbjct: 180 STVRSPSPSVTLSLSPSTVPPTPTPTPGIPWLQTDRGPE-NAPLILSSF-PHHGVAPPCG 237
Query: 222 DHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIK 281
++ ++ELV KEL+EGHRA AAHKKEA WRLRR+ELQLESEK C+ RE +EE EA +K
Sbjct: 238 ENPFVTELVECCKELDEGHRAWAAHKKEAAWRLRRVELQLESEKICKVREKMEEIEAKMK 297
Query: 282 ALQEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQT 338
AL+EEQ A L+RIE +EQL GLRRDAE+KEQKLAE+W SKHLRL++ LEQM Q+
Sbjct: 298 ALREEQKATLDRIEAEYKEQLAGLRRDAEAKEQKLAEQWASKHLRLSKFLEQMGCQS 354
>gb|AAD40953.1| phantastica [Lycopersicon esculentum]
Length = 360
Score = 424 bits (1090), Expect = e-117
Identities = 226/355 (63%), Positives = 269/355 (75%), Gaps = 21/355 (5%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
M+ERQRW +EEDALL AYV+QYGP+EW LVSQRMNTPLNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MRERQRWRAEEDALLRAYVRQYGPKEWPLVSQRMNTPLNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT++EQRLVI LQA +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE+ E N +V P
Sbjct: 61 SLTEDEQRLVIQLQAKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREQKENNKVVDP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSS-----AMLPSWLSNYDST 175
+ + KY+H+LE FAEK+VKE ++P M A+SN FLH ++S +LP WLSN +
Sbjct: 121 VDEGKYDHILETFAEKIVKERSVPGLLM-ATSNGGFLHADASTPTPQTLLPPWLSNSSAP 179
Query: 176 ST-PPSSISVTLSLSPSTV-----------ATPRGLENNAPFVLRNVTAHNGSVPSFSDH 223
ST SS SVTLSLSPSTV T RG +NAP +L + H SV ++
Sbjct: 180 STVRSSSPSVTLSLSPSTVPPTPTPGIPWLQTDRG-PDNAPLILSSFPHH--SVAPCGEN 236
Query: 224 ILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKAL 283
++EL K+L+EGHR AHKKEA WRLRR+ELQLESEKA + RE +EE EA +KAL
Sbjct: 237 PFITELAECCKDLDEGHRTWTAHKKEAAWRLRRVELQLESEKASKVREKMEEIEAKMKAL 296
Query: 284 QEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQMKIQT 338
+EEQ A L+RIE +EQL GLRRDAE+KEQKLAE+WTSKH+RL + LEQM Q+
Sbjct: 297 REEQKATLDRIEAEYKEQLAGLRRDAEAKEQKLAEQWTSKHMRLAKFLEQMGCQS 351
>gb|AAO49809.1| phantastica-like MYB protein [Eschscholzia californica subsp.
californica]
Length = 360
Score = 409 bits (1051), Expect = e-113
Identities = 215/350 (61%), Positives = 259/350 (73%), Gaps = 17/350 (4%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRW +EEDALL AYV+QYGPREWNLVSQRMNT L+RD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWRAEEDALLRAYVKQYGPREWNLVSQRMNTHLDRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQA +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE+ E + + P
Sbjct: 61 SLTEEEQRLVIRLQAKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREQKETSKTIDP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSS-----AMLPSWLSNYDST 175
I + KY+ +LE FAEKLVKE P M +SN +L +N++ +LP WLS+ +
Sbjct: 121 IEEGKYDQILETFAEKLVKERPNPPLYM-GTSNGGYLQSNAATVPPPTLLPPWLSSSSAP 179
Query: 176 STPPSSISVTLSLSPSTVA----------TPRGLENNAPFVLRNV-TAHNGSVPSFSDHI 224
T S SVTL+LSPST+A G ++N VL N H PS D +
Sbjct: 180 PTTSSPPSVTLTLSPSTIAPCTSMSWLQPDRGGNDSNPSLVLGNFPPTHVPVPPSGGDRL 239
Query: 225 LMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQ 284
++ +LV +ELEE HRAL AHKKEA WRL+R+ELQLESEKACRRRE +EE E ++AL+
Sbjct: 240 MVPDLVECCRELEESHRALVAHKKEAAWRLKRVELQLESEKACRRREKMEEIEMKVRALR 299
Query: 285 EEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQM 334
EEQ L+R+E R+QL GLRRDAE+KEQKLA++W +KHLRL + LEQ+
Sbjct: 300 EEQKVTLDRMEADYRDQLAGLRRDAEAKEQKLADQWAAKHLRLMKFLEQI 349
>gb|AAC23633.1| putative MYB family transcription factor [Arabidopsis thaliana]
gi|5823325|gb|AAD53101.1| putative transcription factor
[Arabidopsis thaliana] gi|29427581|sp|O80931|ASL1_ARATH
ASYMMETRIC LEAVES1 protein (Myb-related protein 91)
(AtMYB91) (PHANTASTICA protein)
gi|15224328|ref|NP_181299.1| myb family transcription
factor (MYB91) [Arabidopsis thaliana]
gi|41619204|gb|AAS10048.1| MYB transcription factor
[Arabidopsis thaliana]
Length = 367
Score = 400 bits (1028), Expect = e-110
Identities = 220/364 (60%), Positives = 264/364 (72%), Gaps = 37/364 (10%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRWS EEDALL AYV+Q+GPREW+LVS+RMN PLNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWSGEEDALLRAYVRQFGPREWHLVSERMNKPLNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQ +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE+ E N V P
Sbjct: 61 SLTEEEQRLVIRLQEKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREEKESNKRVEP 120
Query: 121 ISDTKYEHMLEGFAEKLVKE--HTLPSFAMA-----ASSNEAFLHTNS-----SAMLPSW 168
I ++KY+ +LE FAEKLVKE + +P+ A A A+SN FLH+ + ++P W
Sbjct: 121 IDESKYDRILESFAEKLVKERSNVVPAAAAAATVVMANSNGGFLHSEQQVQPPNPVIPPW 180
Query: 169 LSNYDS----TSTPPSSISVTLSLSPSTVAT--------------PRGLENN-APFVLRN 209
L+ ++ + PP SVTL+LSPSTVA P EN VL +
Sbjct: 181 LATSNNGNNVVARPP---SVTLTLSPSTVAAAAPQPPIPWLQQQQPERAENGPGGLVLGS 237
Query: 210 VTAHNGSVPSFSDHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRR 269
+ S S+ + +SELV +ELEEGHRA A HKKEA WRLRRLELQLESEK CR+
Sbjct: 238 MMP---SCSGSSESVFLSELVECCRELEEGHRAWADHKKEAAWRLRRLELQLESEKTCRQ 294
Query: 270 RETVEEFEANIKALQEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTR 329
RE +EE EA +KAL+EEQ A+ +IE REQL GLRRDAE+K+QKLA++WTS+H+RLT+
Sbjct: 295 REKMEEIEAKMKALREEQKNAMEKIEGEYREQLVGLRRDAEAKDQKLADQWTSRHIRLTK 354
Query: 330 LLEQ 333
LEQ
Sbjct: 355 FLEQ 358
>emb|CAA06612.1| MYB-related transcription factor [Antirrhinum majus]
gi|7489304|pir||T17027 MYB-related transcription factor
- garden snapdragon
Length = 357
Score = 393 bits (1010), Expect = e-108
Identities = 210/344 (61%), Positives = 255/344 (74%), Gaps = 11/344 (3%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRW EEDALL AYV++YGPR+W+LV+QRMN PLNRD KSCLERWKNYLKPGIKK
Sbjct: 1 MKERQRWRPEEDALLRAYVKEYGPRDWHLVTQRMNKPLNRDAKSCLERWKNYLKPGIKKE 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQ LVI LQA +GNKWKKIAAEVPGRTAKRLGKWWEV+KEK+QRE+ + I P
Sbjct: 61 SLTQEEQILVINLQAKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKKQREEKDNKKITEP 120
Query: 121 ISDTKYEHMLEGFAEKLVKEHTLPS-FAMAASSNEAFLHTNSS-----AMLPSWLSNYDS 174
I + KY+ +LE FAEK+VKE + M +SN FL + S ++LP WL++
Sbjct: 121 IEEGKYDRILETFAEKIVKERVVSRIITMPPTSNSGFLQNDPSPHSAQSVLPPWLASSSM 180
Query: 175 TST-PPSSISVTLSLSPSTV----ATPRGLENNAPFVLRNVTAHNGSVPSFSDHILMSEL 229
T+T P S SVTLSLSPS V A P +N N+++ P ++ ++ EL
Sbjct: 181 TTTIRPQSPSVTLSLSPSVVPPAPAIPWLHPDNTTHGPSNLSSLGVVAPFMGENHIVPEL 240
Query: 230 VGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEANIKALQEEQTA 289
+ +ELEEG RA AAH+KEA WRL+R+ELQLESEKACRRRE +EE EA +KAL+EEQ A
Sbjct: 241 LECCRELEEGQRAWAAHRKEAAWRLKRVELQLESEKACRRREKMEEIEAKMKALREEQKA 300
Query: 290 ALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQ 333
+L+RIE REQL GLRR+AE KEQKLAE+W +KHLRLT+ LEQ
Sbjct: 301 SLDRIEAEYREQLAGLRREAEVKEQKLAEQWAAKHLRLTKFLEQ 344
>gb|AAD33898.1| rough sheath 2 [Zea mays] gi|4583431|gb|AAD25082.1| rough sheath2
protein [Zea mays]
Length = 370
Score = 338 bits (868), Expect = 1e-91
Identities = 191/361 (52%), Positives = 238/361 (65%), Gaps = 33/361 (9%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRW EEDA+L AYV+QYGPREW+LVSQRMN L+RD KSCLERWKNYL+PGIKKG
Sbjct: 1 MKERQRWRPEEDAVLRAYVRQYGPREWHLVSQRMNVALDRDAKSCLERWKNYLRPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEIN---GI 117
SLT+EEQRLVI LQA +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE +
Sbjct: 61 SLTEEEQRLVIRLQAKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQRELRDSRRPPPE 120
Query: 118 VSPISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTST 177
SP +YE +LE FAEKLV E P A AA S ++ +LP WLS+ +
Sbjct: 121 PSPDERGRYEWLLENFAEKLVGER--PQQAAAAPSPLLM----AAPVLPPWLSSNAGPAA 174
Query: 178 P-----------PSSISVTLSLSPSTVA----------TPRGLENNAPFVLRNVTAHNGS 216
P S SVTLSL+ + VA R + AP+ + + H G+
Sbjct: 175 AAAAAVAHPPPRPPSPSVTLSLASAAVAPGPPAPAPWMPDRAAADAAPYGFPSPSQHGGA 234
Query: 217 VP---SFSDHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETV 273
P + D ++EL +ELEEG RA AAH++EA WRL+R+E QLE E+ RRRE
Sbjct: 235 APPGMAVVDGQALAELAECCRELEEGRRAWAAHRREAAWRLKRVEQQLEMEREMRRREVW 294
Query: 274 EEFEANIKALQEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTRLLEQ 333
EEFEA ++ ++ EQ AA R+E RE++ LRRDA+ KE+K+AE+W +KH R+ + +EQ
Sbjct: 295 EEFEAKMRTMRLEQAAAAERVERDHREKVAELRRDAQVKEEKMAEQWAAKHARVAKFVEQ 354
Query: 334 M 334
M
Sbjct: 355 M 355
>gb|AAW62520.1| PHANTASTICA-like protein [Selaginella kraussiana]
Length = 404
Score = 243 bits (621), Expect = 5e-63
Identities = 155/379 (40%), Positives = 209/379 (54%), Gaps = 46/379 (12%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MK++QRW EEDALL AYV+QYGP +WNLVS+RM TPL+RD KSC ERWKNYLKPG+K+G
Sbjct: 1 MKDKQRWQPEEDALLCAYVKQYGPNDWNLVSERMATPLDRDPKSCHERWKNYLKPGLKRG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
L++EEQ LVI LQ YGNKWK+IAAEVPGRTAKRLGKWWEV+KE++Q+E I+ + +
Sbjct: 61 PLSEEEQNLVIRLQEKYGNKWKRIAAEVPGRTAKRLGKWWEVHKERRQKEAIQRHQRIQT 120
Query: 121 ISDTKYEHMLEG-----FAEKLVKEHTLPSFAMAASSNE---------------AFLHTN 160
T + M G F T ++S++E AF T+
Sbjct: 121 GVHTSHLSMFYGQTVAPFIPPAQSFSTCAEVVSSSSASEGESQCRNEPRMNLPAAFPPTS 180
Query: 161 SSA----------MLPSW--------LSNYDSTSTPPSSISVTLSLSPSTVA----TPRG 198
S +LP+W S S P + + LSLS + A T G
Sbjct: 181 SEPVLTLGPTVLDLLPAWKPAPRAASTSELPSLMAPEAIMKPNLSLSLDSGAESGDTDTG 240
Query: 199 LE-NNAPFVLRNVTAHNGSVPSFSDHILMSELV---GFSKELEEGHRALAAHKKEAEWRL 254
NN V + + + I EL+ G KELEE + KK A L
Sbjct: 241 THFNNNKKVSTIIPKDDEFCNEINSDISPGELIPLLGLVKELEENKESWNVQKKNAASTL 300
Query: 255 RRLELQLESEKACRRRETVEEFEANIKALQEEQTAALNRIENACREQLGGLRRDAESKEQ 314
R L+ QLE E+ +R++ + E E+ I+AL++E+ L+++E E + L RDAE KE+
Sbjct: 301 RELKQQLECERIEKRKQKMLEVESKIQALRKEEKLYLDKLELDYAELVAKLDRDAELKEE 360
Query: 315 KLAEKWTSKHLRLTRLLEQ 333
KL E W+ K+ +L + EQ
Sbjct: 361 KLVESWSLKYNKLVLMFEQ 379
>gb|AAO86633.1| phantastica transcription factor [Acacia hindsii]
Length = 210
Score = 236 bits (603), Expect = 6e-61
Identities = 131/212 (61%), Positives = 155/212 (72%), Gaps = 16/212 (7%)
Query: 77 YGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKYEHMLEGFAEK 136
+GNKWKKIA+EVPGRTAKRLGKWWEV+KEKQQREK +I+ V PI D KY+H+LE FAEK
Sbjct: 1 HGNKWKKIASEVPGRTAKRLGKWWEVFKEKQQREKKQISKTVDPIYDRKYDHLLETFAEK 60
Query: 137 LVKEHTLPSFAMAASSNEAFLHT----NSSAMLPSWLSNYDSTS-TPPSSISVTLSLSPS 191
LV E S+ M A SN AF+HT ++S LP WLSN +++ P S SVTL LSPS
Sbjct: 61 LVNERPSSSYLMTA-SNGAFMHTDVPSSASTSLPPWLSNSNTSKVVRPPSPSVTLGLSPS 119
Query: 192 TVATP--------RGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGFSKELEEGHRAL 243
TV P RG NAP +L NVT H G VP+F D++L+SEL ++ELEEGH+A
Sbjct: 120 TVVAPPFPWLQPERG-PGNAPLILGNVTPH-GPVPAFGDNMLISELFECARELEEGHQAW 177
Query: 244 AAHKKEAEWRLRRLELQLESEKACRRRETVEE 275
AAHKKEA WRL R+ELQLESEKACRRRE +EE
Sbjct: 178 AAHKKEAAWRLSRVELQLESEKACRRREKMEE 209
>gb|AAW62521.1| PHANTASTICA-like protein [Selaginella viticulosa]
Length = 391
Score = 221 bits (563), Expect = 2e-56
Identities = 137/381 (35%), Positives = 200/381 (51%), Gaps = 46/381 (12%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
+K+RQRW EEDA L AYV +YGP+ W L+S+R L+RD KSC ERWKNYLKPGIK+G
Sbjct: 5 LKDRQRWQPEEDAXLCAYVTEYGPQNWQLISERTGKRLDRDPKSCEERWKNYLKPGIKRG 64
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVS- 119
LT EEQ+LVI LQ YGNKWK+IAA+VPGRTAKRLGKWWEVYKE++ +E ++ GI +
Sbjct: 65 PLTDEEQQLVIKLQTKYGNKWKRIAAQVPGRTAKRLGKWWEVYKERKLKENKKLLGIAAT 124
Query: 120 ---------------PISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAM 164
P+ + L+ + P+ + + + + +T+ ++
Sbjct: 125 GNGDSIITAEEEMHLPVVAPIFSRPFLSSTPDLLHGNGDPALSTSGNDHHFGNNTSGASN 184
Query: 165 LPSWLSNYDSTSTPPSSI--SVTLSLSPSTVAT---------PRGLENNAPFVLRNVTAH 213
+ N + PSS+ + SP VAT P+ LE L N +A
Sbjct: 185 AETLSENEPLLTLGPSSLVQNAAKQCSPEMVATSGVTMPKWIPKELELQIASTL-NESAV 243
Query: 214 NGSV---PSFSDHILMSELVGFS---------------KELEEGHRALAAHKKEAEWRLR 255
S+ SF D + + S K+L++ +K+ + R
Sbjct: 244 ESSLSLSSSFDDGVDCLDPASVSSDNRESSAIQVFELFKQLKDQRNNWILERKKVWSKFR 303
Query: 256 RLELQLESEKACRRRETVEEFEANIKALQEEQTAALNRIENACREQLGGLRRDAESKEQK 315
L+ QL+SE+ ++R+ +EE ++AL++E+ L ++E E + L RDAE KE K
Sbjct: 304 ELKRQLDSERLEKQRQKIEEVGIKVRALKDEEARFLRKVEQDYEELVSNLERDAELKENK 363
Query: 316 LAEKWTSKHLRLTRLLEQMKI 336
+ E WT KH +L E+ I
Sbjct: 364 IMEVWTIKHEKLVHAYERYTI 384
>gb|AAO86630.1| phantastica transcription factor [Aquilegia formosa]
Length = 214
Score = 220 bits (561), Expect = 4e-56
Identities = 122/216 (56%), Positives = 152/216 (69%), Gaps = 16/216 (7%)
Query: 77 YGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKYEHMLEGFAEK 136
+GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE+ E N PI + KY+ +LE FAEK
Sbjct: 1 HGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREQKENNKAPEPIEEGKYDSILETFAEK 60
Query: 137 LVKEHTLPSFAMAASSNEAFLHTNSSA----MLPSWLSNYDSTSTPPSSISVTLSLSPST 192
LVKE P F M A+SN FLH++ A MLP W+++ + T+ PSS SVTL+LSPST
Sbjct: 61 LVKECPNPPFLM-ATSNGGFLHSDPPAPPPTMLPPWMASSNGTTVRPSSPSVTLTLSPST 119
Query: 193 VATP----------RGLENNAPFVLRNVTAHNGSVPSFSDHILMSELVGFSKELEEGHRA 242
V P RG N L ++++H GS + D+ ++++LV +ELEEGHRA
Sbjct: 120 VTPPPSIPWLQSADRGAAENPSLGLGSLSSH-GSGSTGGDNHMVADLVECCRELEEGHRA 178
Query: 243 LAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEA 278
AHKKEA WRL+R+ELQLESEKACRRR+ +EE EA
Sbjct: 179 WVAHKKEAAWRLKRVELQLESEKACRRRDKMEEIEA 214
>emb|CAA06905.1| phantastica protein [Nicotiana tabacum] gi|7489154|pir||T02170
hypothetical protein phan - common tobacco (fragment)
Length = 182
Score = 158 bits (400), Expect = 2e-37
Identities = 97/185 (52%), Positives = 122/185 (65%), Gaps = 20/185 (10%)
Query: 103 YKEKQQREKIEINGIVSPISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSS 162
+KEKQQRE+ E N +V P+ + KY+H+LE FAEK+VKE ++P MA +SN FLHT++
Sbjct: 1 FKEKQQREQKENNKVVDPVDEGKYDHILETFAEKIVKERSVPGLLMA-TSNGGFLHTDAP 59
Query: 163 A-----MLPSWLSNYDSTSTPPS-SISVTLSLSPSTVA-----------TPRGLENNAPF 205
A +LP WLSN +TST S S SVTLSLSPSTV T RG EN AP
Sbjct: 60 APSPQTLLPPWLSNSTATSTVRSQSPSVTLSLSPSTVPPTPTPGIPWLQTDRGPEN-APL 118
Query: 206 VLRNVTAHNGSVPSFSDHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEK 265
+L + H+G P ++ ++EL KEL+EGHRA AAHKKEA WRLRR+ELQLESEK
Sbjct: 119 ILSSFP-HHGVAPPCGENPFVTELAECCKELDEGHRAWAAHKKEAAWRLRRVELQLESEK 177
Query: 266 ACRRR 270
+ R
Sbjct: 178 TSKVR 182
>gb|AAO86631.1| phantastica transcription factor [Vitex cannabifolia]
Length = 231
Score = 137 bits (344), Expect = 6e-31
Identities = 93/239 (38%), Positives = 123/239 (50%), Gaps = 45/239 (18%)
Query: 77 YGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGI------------VSP--IS 122
+GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQ ++ ++ N + SP +
Sbjct: 1 HGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQLKQALQKNHLDYGGGSAVPGSAASPEKAA 60
Query: 123 DTKYEHMLEGFAEKLVKEHTL-----------------------PSFAMAASSNEAFLHT 159
KY+H+L+ FAEK V+ L PS + + EA
Sbjct: 61 QGKYDHILDTFAEKYVQPKLLSFQSPSNLIMPAANLSIPEPPPVPSLGSVSINTEA--GN 118
Query: 160 NSSAMLPSWLSNYDSTSTPPSSISVTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPS 219
+ LP W+ + +TP ++ S+T S S + + L + P VL T G P
Sbjct: 119 SGCTSLPPWM---NMITTPTATSSLTSSSSTPSPSVSLTLSPSEPAVLDPET---GVAPR 172
Query: 220 FSDHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEA 278
F + L+ KE+EEG HKKEA WRL RLE QLESEK+ RR+E +EE EA
Sbjct: 173 FFPVQQVGVLIQHCKEVEEGRENWMRHKKEATWRLNRLEQQLESEKSRRRKEKMEEIEA 231
>gb|AAO86634.1| phantastica transcription factor [Pachira aquatica]
Length = 225
Score = 136 bits (343), Expect = 8e-31
Identities = 92/237 (38%), Positives = 129/237 (53%), Gaps = 47/237 (19%)
Query: 77 YGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEING------------IVSPI--- 121
+GNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ ++ + G V+P+
Sbjct: 1 HGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQLKQLQKNQGGRKEFSPQGSCSTVAPVVSS 60
Query: 122 SDTKYEHMLEGFAEKLVKEH--------------TLPS-----FAMAASSNEAFLHTNSS 162
S +Y+H+LE FAEK V + +LP S + T+SS
Sbjct: 61 SPGQYDHILETFAEKYVLSNYSAMNLPPIMAPTISLPDPDPDPVLSLGSGSSGTATTSSS 120
Query: 163 AMLPSWLSNYDSTSTPPSSISVTLSLSPS-TVATPRGLENNAPFVLRNVTAHNGSVPSFS 221
+LP W+++ +T++ SS + + + SPS +++ G P ++R S
Sbjct: 121 VVLPLWMNHNHTTASSLSSSTSSTTPSPSVSLSLSPGEPGLDPDLIR------------S 168
Query: 222 DHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEEFEA 278
+ + V + KELEEG ++ HKKEA WRL RLE QLE+EKA +R+E +EE EA
Sbjct: 169 CRLKLVRWVQYCKELEEGRQSWMQHKKEATWRLSRLEQQLEAEKARKRKEKMEEIEA 225
>gb|AAO86635.1| phantastica transcription factor [Fraxinus americana]
Length = 233
Score = 132 bits (333), Expect = 1e-29
Identities = 89/232 (38%), Positives = 112/232 (47%), Gaps = 33/232 (14%)
Query: 77 YGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE------------------KIEINGIV 118
+GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQ ++ + G
Sbjct: 1 HGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQLKQLQKSHNSRQEYDNPVAGSAVAGGGSP 60
Query: 119 SPISDTKYEHMLEGFAEKLVKEHT---------------LPSFAMAASSNEAFLHTNSSA 163
KY+H+LE FAEK V+ LP S + S
Sbjct: 61 EKAVQGKYDHILETFAEKYVQPKMFAFQSPNIIIPPNLPLPEPHPVLSIGSSRPVQLSGP 120
Query: 164 MLPSWLSNYDSTSTPPSSISVTLSLSPSTVATPRGLENNAPFVLRNVTAHNGSVPSFSDH 223
+LP W++N T SS + S S + + L + P VL + G F
Sbjct: 121 ILPPWMNNTTHPHTQTSSSLTSSSSSTHSPSVSLTLSPSEPAVLDPIHPEPGLPTRFLPI 180
Query: 224 ILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRRRETVEE 275
M+ L+ + KELEEG + HKKEA WRL RLE QLESEK +RRE +EE
Sbjct: 181 KQMNTLIQYCKELEEGRQIWLHHKKEATWRLTRLEQQLESEKTRKRREKMEE 232
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.127 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 577,653,609
Number of Sequences: 2540612
Number of extensions: 23424502
Number of successful extensions: 81006
Number of sequences better than 10.0: 1941
Number of HSP's better than 10.0 without gapping: 1272
Number of HSP's successfully gapped in prelim test: 681
Number of HSP's that attempted gapping in prelim test: 77110
Number of HSP's gapped (non-prelim): 3165
length of query: 341
length of database: 863,360,394
effective HSP length: 128
effective length of query: 213
effective length of database: 538,162,058
effective search space: 114628518354
effective search space used: 114628518354
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0022.17


