

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.15
(313 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
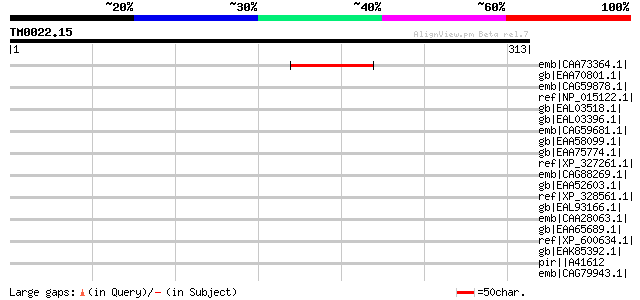
Score E
Sequences producing significant alignments: (bits) Value
emb|CAA73364.1| Pge1 protein [Lotus corniculatus var. japonicus] 60 1e-07
gb|EAA70801.1| hypothetical protein FG04147.1 [Gibberella zeae P... 43 0.014
emb|CAG59878.1| unnamed protein product [Candida glabrata CBS138... 43 0.014
ref|NP_015122.1| Iron-regulated transcriptional activator, requi... 40 0.067
gb|EAL03518.1| hypothetical protein CaO19.12424 [Candida albican... 40 0.11
gb|EAL03396.1| hypothetical protein CaO19.4959 [Candida albicans... 40 0.11
emb|CAG59681.1| unnamed protein product [Candida glabrata CBS138... 39 0.20
gb|EAA58099.1| hypothetical protein AN6124.2 [Aspergillus nidula... 39 0.20
gb|EAA75774.1| hypothetical protein FG05699.1 [Gibberella zeae P... 38 0.33
ref|XP_327261.1| predicted protein [Neurospora crassa] gi|289242... 38 0.33
emb|CAG88269.1| unnamed protein product [Debaryomyces hansenii C... 37 0.57
gb|EAA52603.1| hypothetical protein MG05295.4 [Magnaporthe grise... 37 0.57
ref|XP_328561.1| hypothetical protein [Neurospora crassa] gi|289... 37 0.97
gb|EAL93166.1| isochorismatase family hydrolase, putative [Aspe... 35 2.2
emb|CAA28063.1| rpoC2 [Marchantia polymorpha] gi|66985|pir||RNLV... 35 2.2
gb|EAA65689.1| hypothetical protein AN0859.2 [Aspergillus nidula... 35 2.8
ref|XP_600634.1| PREDICTED: similar to PC-LKC gene product, part... 35 3.7
gb|EAK85392.1| hypothetical protein UM04510.1 [Ustilago maydis 5... 34 4.8
pir||A41612 vitellogenic carboxypeptidase (EC 3.4.16.-) precurso... 33 8.2
emb|CAG79943.1| Mutyl [Yarrowia lipolytica CLIB99] gi|49649200|e... 33 8.2
>emb|CAA73364.1| Pge1 protein [Lotus corniculatus var. japonicus]
Length = 210
Score = 59.7 bits (143), Expect = 1e-07
Identities = 23/50 (46%), Positives = 37/50 (74%)
Query: 170 KVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKY 219
+ FAS ++++WAR +GK++G+++++ RSD G KRK + LGCER KY
Sbjct: 155 EAFASHTDLIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGGKY 204
>gb|EAA70801.1| hypothetical protein FG04147.1 [Gibberella zeae PH-1]
gi|46116610|ref|XP_384323.1| hypothetical protein
FG04147.1 [Gibberella zeae PH-1]
Length = 207
Score = 42.7 bits (99), Expect = 0.014
Identities = 27/96 (28%), Positives = 48/96 (49%), Gaps = 5/96 (5%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQS 230
+F + D+++ + + K G+ IV R+ N + T L C+R + Y K ++
Sbjct: 16 IFRTFDDLMASVQRVAKDQGYGIVKLRASNYRDGKPTRYDLVCDRGG--VKYNSTAKKRN 73
Query: 231 TGTKKCYCPFRLRA-RGTKVG--WKVSVMNGYHIHE 263
T+K CPFR +A ++G W+ ++ G H HE
Sbjct: 74 PSTRKIDCPFRAKAVCEVQLGNQWRFAIQEGRHNHE 109
>emb|CAG59878.1| unnamed protein product [Candida glabrata CBS138]
gi|50289031|ref|XP_446945.1| unnamed protein product
[Candida glabrata]
Length = 535
Score = 42.7 bits (99), Expect = 0.014
Identities = 40/146 (27%), Positives = 60/146 (40%), Gaps = 22/146 (15%)
Query: 160 VDYTQSFTTDKV--FASRDEILEWARNLGKQHGFIIVITRSDN-----------GGLKRK 206
+D Q D V F ++EI W + + G IVI RSD+ G K +
Sbjct: 1 MDSNQLIHLDSVPNFKDKNEIKPWLQKIFYPQGIEIVIERSDSLKVVFKCKAAKRGRKSQ 60
Query: 207 TFMILGCERCDKYIPYKEVLKHQSTGTKKC-----YCPFRLRARGT--KVGWKVSVMNGY 259
+ + + + +K +S G K+C +CPFR+RA + + W V V+N
Sbjct: 61 EELTEEERLRQEALAREHEMKRKSMGNKRCVSRYNHCPFRVRATYSLKRKKWSVVVLNNT 120
Query: 260 HIHERSETLLGHPYVGRPFTNKRLAD 285
H H L Y + F K AD
Sbjct: 121 HTHPLKFNPLSEEY--KKFKEKLRAD 144
>ref|NP_015122.1| Iron-regulated transcriptional activator, required for iron
homeostasis and resistance to oxidative stress; similar
to Aft1p; Aft2p [Saccharomyces cerevisiae]
gi|1370420|emb|CAA97916.1| unnamed protein product
[Saccharomyces cerevisiae] gi|2132231|pir||S65221
hypothetical protein YPL202c - yeast (Saccharomyces
cerevisiae)
Length = 416
Score = 40.4 bits (93), Expect = 0.067
Identities = 35/122 (28%), Positives = 50/122 (40%), Gaps = 12/122 (9%)
Query: 146 KLVRAEDVQDEPLAVDYTQSFTTDKV--FASRDEILEWARNLGKQHGFIIVITRSDNGGL 203
KL + D ++ D + D V F R EI W + + G IVI RSD+ +
Sbjct: 23 KLTASPDNLASMMSKDQNKLIHLDPVPSFEDRHEIKPWLQKIFYPQGIDIVIERSDSSKV 82
Query: 204 KRKTFMILGCERCDKYIPYKEVLKHQSTGTKKCYCPFRLRARGT--KVGWKVSVMNGYHI 261
K C + K + + ++ CPFR+RA + W V VMN H
Sbjct: 83 TFK------CRSVRSKVGLNP--KSKGSSSRSHACPFRIRAAYSVRLQKWNVVVMNNIHS 134
Query: 262 HE 263
HE
Sbjct: 135 HE 136
>gb|EAL03518.1| hypothetical protein CaO19.12424 [Candida albicans SC5314]
Length = 573
Score = 39.7 bits (91), Expect = 0.11
Identities = 34/120 (28%), Positives = 49/120 (40%), Gaps = 22/120 (18%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYK----E 224
++VF SRDE+ E+ + +GF +VI S+ K + CE +Y K +
Sbjct: 208 EQVFNSRDELNEFIAEFARDNGFGVVIAHSN------KKAIYYTCELGGRYRHKKNKKID 261
Query: 225 VLKHQSTG----------TKKCYCPFRLRARGTKV--GWKVSVMNGYHIHERSETLLGHP 272
V K G TKK CPF + A K W + H H + + L HP
Sbjct: 262 VTKQIDVGDGYMLDPDTKTKKLKCPFAMTASYKKSANAWTLRTTCNEHNHPQLDPLSNHP 321
>gb|EAL03396.1| hypothetical protein CaO19.4959 [Candida albicans SC5314]
Length = 573
Score = 39.7 bits (91), Expect = 0.11
Identities = 34/120 (28%), Positives = 49/120 (40%), Gaps = 22/120 (18%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYK----E 224
++VF SRDE+ E+ + +GF +VI S+ K + CE +Y K +
Sbjct: 208 EQVFNSRDELNEFIAEFARDNGFGVVIAHSN------KKAIYYTCELGGRYRHKKNKKID 261
Query: 225 VLKHQSTG----------TKKCYCPFRLRARGTKV--GWKVSVMNGYHIHERSETLLGHP 272
V K G TKK CPF + A K W + H H + + L HP
Sbjct: 262 VTKQIDVGDGYMLDPDTKTKKLKCPFAMTASYKKSANAWTLRTTCNEHNHPQLDPLSNHP 321
>emb|CAG59681.1| unnamed protein product [Candida glabrata CBS138]
gi|50288649|ref|XP_446754.1| unnamed protein product
[Candida glabrata]
Length = 437
Score = 38.9 bits (89), Expect = 0.20
Identities = 34/117 (29%), Positives = 55/117 (46%), Gaps = 13/117 (11%)
Query: 152 DVQDEPLAVDYTQSFTTDKV--FASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFM 209
D + L V+ Q D V F + ++ W + + G IVI RSD KT +
Sbjct: 20 DNLENALMVNENQLIHIDPVPDFQEKIDVKVWLQEMFFPMGIDIVIERSD------KTKI 73
Query: 210 ILGCERCD--KYIPYKEVLKHQSTGTKKCYCPFRLRAR-GTKV-GWKVSVMNGYHIH 262
I C+ D + P K++ + + K+ CPFR+R T++ WK+ ++N H H
Sbjct: 74 IFKCKPSDYRSHEP-KDLRQKEPVDDKRHICPFRVRCTFSTQLKKWKIVIINNSHSH 129
>gb|EAA58099.1| hypothetical protein AN6124.2 [Aspergillus nidulans FGSC A4]
gi|67539908|ref|XP_663728.1| hypothetical protein
AN6124_2 [Aspergillus nidulans FGSC A4]
gi|49097602|ref|XP_410261.1| hypothetical protein
AN6124.2 [Aspergillus nidulans FGSC A4]
Length = 339
Score = 38.9 bits (89), Expect = 0.20
Identities = 29/98 (29%), Positives = 45/98 (45%), Gaps = 8/98 (8%)
Query: 172 FASRDEILEWARNLGKQHGFIIVITRSDNGGLK---RKTFMILGCERCDKYIPY----KE 224
+ + +L + K HG+ +V+ S K R + L C+R Y P +E
Sbjct: 33 YPDKTSLLASVQAHAKAHGYNVVVKSSSTPTEKKPGRTAKVWLRCDRGGHYRPRNGLTEE 92
Query: 225 VLKHQSTGTKKCYCPFRLRARGTKVGWKVSVMNGYHIH 262
K + T ++ CPF L A GT W ++V+NG H H
Sbjct: 93 TRKRRRT-SRLMDCPFMLVAAGTPGIWTLTVLNGTHNH 129
>gb|EAA75774.1| hypothetical protein FG05699.1 [Gibberella zeae PH-1]
gi|46122643|ref|XP_385875.1| hypothetical protein
FG05699.1 [Gibberella zeae PH-1]
Length = 362
Score = 38.1 bits (87), Expect = 0.33
Identities = 28/100 (28%), Positives = 45/100 (45%), Gaps = 7/100 (7%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYI-----PYKEV 225
+F S +E+ + + ++ G+ IV RS G + RCD PY+ +
Sbjct: 15 IFPSYEELFKDLSDRMEKDGYKIVKARSHRGKVGGADIPGNDIVRCDLVCDRGGRPYRCM 74
Query: 226 LKHQSTGTKKCYCPFRLRA--RGTKVGWKVSVMNGYHIHE 263
T TKK CP++ +A R T GW +++ H HE
Sbjct: 75 ATKHKTTTKKTDCPWKAKAVHRKTMGGWVLTITCDQHNHE 114
>ref|XP_327261.1| predicted protein [Neurospora crassa] gi|28924225|gb|EAA33379.1|
predicted protein [Neurospora crassa]
Length = 569
Score = 38.1 bits (87), Expect = 0.33
Identities = 36/121 (29%), Positives = 57/121 (46%), Gaps = 19/121 (15%)
Query: 173 ASRDEILEWARNLGKQHGFIIVITRSDNG--GLKRKTFMILGCERCDKYIPYKEV---LK 227
A +LE+ ++G + +VI RS GLK+ F+ C+R K P K V ++
Sbjct: 316 AIHKHVLEYCTSVG----YAVVIGRSKKTVPGLKKVLFV---CDRAGK--PPKRVSPEMR 366
Query: 228 HQSTGTKKCYCPFRLRARGTKVGWKV----SVMNGYHIHERSETLLGHPYVGRPFTNKRL 283
+ T ++KC CPF A + W + ++ H H SE+ L HP R +K +
Sbjct: 367 KRKTTSRKCDCPFGFFAIEQRTQWTIRYRPDAIHLQHNHGPSESPLLHP-AARKLDSKMV 425
Query: 284 A 284
A
Sbjct: 426 A 426
>emb|CAG88269.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50422877|ref|XP_460016.1| unnamed protein product
[Debaryomyces hansenii]
Length = 481
Score = 37.4 bits (85), Expect = 0.57
Identities = 29/115 (25%), Positives = 47/115 (40%), Gaps = 11/115 (9%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDN---------GGLKRKTFMILGCERCDKY 219
++V +RD++ E+ + + +GF +VI S+ GG R+ G E
Sbjct: 159 EQVLHNRDDLNEFIQEFARDNGFGVVIAHSNKKAIYYTCELGGRYRQKKSKKGMEDARHL 218
Query: 220 IPYKEVLKHQSTGTKKCYCPFRLRARGTKVG--WKVSVMNGYHIHERSETLLGHP 272
+ +T TKK CPF + A K W + H H + + L HP
Sbjct: 219 EVDNGYILDPNTKTKKLRCPFSMTATYKKSTGMWTLKTTCNEHNHPQLDPLSNHP 273
>gb|EAA52603.1| hypothetical protein MG05295.4 [Magnaporthe grisea 70-15]
gi|39939890|ref|XP_359482.1| hypothetical protein
MG05295.4 [Magnaporthe grisea 70-15]
Length = 466
Score = 37.4 bits (85), Expect = 0.57
Identities = 24/96 (25%), Positives = 44/96 (45%), Gaps = 5/96 (5%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQS 230
++ S +++L + K+ G+ +V R+ N + T L C+R + Y K ++
Sbjct: 248 IYRSFEDLLSAVQQFSKEQGYGVVKLRASNYRDGKPTRYDLVCDRGG--VKYSSTAKKRN 305
Query: 231 TGTKKCYCPFRLRA---RGTKVGWKVSVMNGYHIHE 263
T+K CP+R +A W+ +V H HE
Sbjct: 306 PSTRKVDCPWRAKAVCEVQLSNQWRFAVQEARHNHE 341
>ref|XP_328561.1| hypothetical protein [Neurospora crassa] gi|28924768|gb|EAA33880.1|
hypothetical protein [Neurospora crassa]
Length = 704
Score = 36.6 bits (83), Expect = 0.97
Identities = 24/96 (25%), Positives = 46/96 (47%), Gaps = 5/96 (5%)
Query: 171 VFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQS 230
++ + +++L + + K G+ +V R+ N + T L C+R + Y K ++
Sbjct: 460 IYRTFEDLLASVQKVAKDQGYGVVKLRASNYREGKPTRYDLVCDRGG--VKYSSTAKKRN 517
Query: 231 TGTKKCYCPFRLRA-RGTKVG--WKVSVMNGYHIHE 263
T+K CP+R +A +G W+ +V H HE
Sbjct: 518 PSTRKVDCPWRAKAVCEVNLGNQWRFAVQEARHNHE 553
>gb|EAL93166.1| isochorismatase family hydrolase, putative [Aspergillus fumigatus
Af293]
Length = 368
Score = 35.4 bits (80), Expect = 2.2
Identities = 27/83 (32%), Positives = 39/83 (46%), Gaps = 8/83 (9%)
Query: 187 KQHGFIIVITRSDNGGLK---RKTFMILGCERCDKYIPY----KEVLKHQSTGTKKCYCP 239
K HG+ +V+ S K R + L C+R Y P +E K + T ++ CP
Sbjct: 28 KAHGYNVVVKSSSTPTEKKPGRTAKVWLRCDRGGHYRPRNGLTEETRKRRRT-SRLMDCP 86
Query: 240 FRLRARGTKVGWKVSVMNGYHIH 262
F L A GT W ++V+N H H
Sbjct: 87 FMLVAAGTPGIWTLTVLNPTHNH 109
>emb|CAA28063.1| rpoC2 [Marchantia polymorpha] gi|66985|pir||RNLVC2 DNA-directed RNA
polymerase (EC 2.7.7.6) beta'-2 chain - liverwort
(Marchantia polymorpha) chloroplast
gi|11466681|ref|NP_039277.1| RNA polymerase beta'' chain
[Marchantia polymorpha] gi|133444|sp|P06274|RPOC2_MARPO
DNA-directed RNA polymerase beta'' chain (PEP)
(Plastid-encoded RNA polymerase beta'' subunit) (RNA
polymerase beta'' subunit)
Length = 1386
Score = 35.4 bits (80), Expect = 2.2
Identities = 25/76 (32%), Positives = 39/76 (50%), Gaps = 6/76 (7%)
Query: 14 QSETQHSILTKVTRSSIYSEQLSEQRSSKKRR------KSSSPLLLLKLTGDGKMRENEN 67
+S+ ++IL K+ S Y+ L E KK+ K+ + L LK+ +G ++ NE
Sbjct: 535 KSQLTNNILNKINNSKNYNFILQEYNIKKKKNFYFLKNKNLTCPLFLKIKKNGVLKNNEI 594
Query: 68 FAILIFPAPSSVNSKI 83
FAIL P+ NS I
Sbjct: 595 FAILDDPSYKVKNSGI 610
>gb|EAA65689.1| hypothetical protein AN0859.2 [Aspergillus nidulans FGSC A4]
gi|67517161|ref|XP_658463.1| hypothetical protein
AN0859_2 [Aspergillus nidulans FGSC A4]
gi|49085806|ref|XP_404996.1| hypothetical protein
AN0859.2 [Aspergillus nidulans FGSC A4]
Length = 404
Score = 35.0 bits (79), Expect = 2.8
Identities = 24/110 (21%), Positives = 46/110 (41%), Gaps = 5/110 (4%)
Query: 169 DKVFASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKH 228
D + + E ++ + + HG+ ++ R++ G + + L C++ Y +
Sbjct: 172 DATYRTETEAVQAIKTFAQDHGYAVITKRTNKGREGKIEAIYLTCDQGQVY--QSTAKER 229
Query: 229 QSTGTKKCYCPFRLRARGTKVG--WKVSVMNGYHIHERSETLLGHPYVGR 276
Q +++ CPF +R K W V V + H H S HP + R
Sbjct: 230 QREPSRRTGCPFSIRISHHKTDNLWHVKVRDPSHNHGPSRP-GSHPSIRR 278
>ref|XP_600634.1| PREDICTED: similar to PC-LKC gene product, partial [Bos taurus]
Length = 173
Score = 34.7 bits (78), Expect = 3.7
Identities = 28/109 (25%), Positives = 47/109 (42%), Gaps = 15/109 (13%)
Query: 16 ETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFPA 75
ET + ++ S Y+ Q+S+QR + +R+ +L + D N+N P
Sbjct: 46 ETLYWFAITISVSDSYNNQVSQQREGRVQRE------MLVIVED----RNDNA-----PV 90
Query: 76 PSSVNSKIGFGDRIPKLKRGFLELSGCRKRKLPSEERLETSREMGVRVW 124
S++ + K + FL + C KR P EE+ T +G R W
Sbjct: 91 FQSISFSANVSEGEEKWELCFLPMYRCPKRPSPGEEKGATCGTLGFRFW 139
>gb|EAK85392.1| hypothetical protein UM04510.1 [Ustilago maydis 521]
gi|49076254|ref|XP_402125.1| hypothetical protein
UM04510.1 [Ustilago maydis 521]
Length = 1004
Score = 34.3 bits (77), Expect = 4.8
Identities = 25/64 (39%), Positives = 32/64 (49%)
Query: 15 SETQHSILTKVTRSSIYSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFP 74
S T S + T SS S LS+ S+K RKSS PLL + + D + N N + P
Sbjct: 245 SPTLSSPTSASTPSSSVSSSLSKAASNKPSRKSSFPLLFGRKSLDSQKCANVNNGLHQEP 304
Query: 75 APSS 78
PSS
Sbjct: 305 LPSS 308
>pir||A41612 vitellogenic carboxypeptidase (EC 3.4.16.-) precursor - yellow
fever mosquito gi|473361|gb|AAA17682.1| vitellogenic
carboxypeptidase
Length = 441
Score = 33.5 bits (75), Expect = 8.2
Identities = 20/55 (36%), Positives = 27/55 (48%)
Query: 31 YSEQLSEQRSSKKRRKSSSPLLLLKLTGDGKMRENENFAILIFPAPSSVNSKIGF 85
Y + + S ++ +S PL L L DGK+ E N A + P SSV S GF
Sbjct: 26 YKKLMRGSASPRRPGESGEPLFLTPLLQDGKIEEARNKARVNHPMLSSVESYSGF 80
>emb|CAG79943.1| Mutyl [Yarrowia lipolytica CLIB99] gi|49649200|emb|CAG81538.1|
Mutyl [Yarrowia lipolytica CLIB99]
gi|49647723|emb|CAG82169.1| Mutyl [Yarrowia lipolytica
CLIB99] gi|49646428|emb|CAG82793.1| Mutyl [Yarrowia
lipolytica CLIB99] gi|49645984|emb|CAG84050.1| Mutyl
[Yarrowia lipolytica CLIB99] gi|49523824|emb|CAF21314.1|
putative MutA transposase [Yarrowia lipolytica]
gi|50553866|ref|XP_504344.1| Mutyl [Yarrowia lipolytica]
gi|50551715|ref|XP_503332.1| Mutyl [Yarrowia lipolytica]
gi|50548773|ref|XP_501856.1| Mutyl [Yarrowia lipolytica]
gi|50546070|ref|XP_500562.1| Mutyl [Yarrowia lipolytica]
gi|50545163|ref|XP_500119.1| Mutyl [Yarrowia lipolytica]
Length = 1178
Score = 33.5 bits (75), Expect = 8.2
Identities = 33/132 (25%), Positives = 58/132 (43%), Gaps = 14/132 (10%)
Query: 172 FASRDEILEWARNLGKQHGFIIVITRSDNGGLKRKTFMILGCERCDKYIPYKEVLKHQST 231
F + E + +A+ + HGF++ I GG K + IL C + KY + +
Sbjct: 320 FKEKAEAIAYAQTTARLHGFVLHI-----GGGKARN--ILQCHK-SKYDDESGEIGKPTN 371
Query: 232 GTKK--CYCPFRLRARGTKVGWKVSVMNGYHIHERSET--LLGHPYVGRPFTNKRLADGP 287
+++ CYCPF++R K+G + H ++ E L + +P +
Sbjct: 372 ASRRVACYCPFQIRV--NKLGPNDFGLVYCHEQKKDENGDALCAKHNHKPSASVSTNGAY 429
Query: 288 RPVRLANLMVGC 299
R + LAN+ GC
Sbjct: 430 RQLDLANIRKGC 441
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 516,037,680
Number of Sequences: 2540612
Number of extensions: 21233520
Number of successful extensions: 46083
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 46074
Number of HSP's gapped (non-prelim): 22
length of query: 313
length of database: 863,360,394
effective HSP length: 128
effective length of query: 185
effective length of database: 538,162,058
effective search space: 99559980730
effective search space used: 99559980730
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0022.15


