

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.7
(1703 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
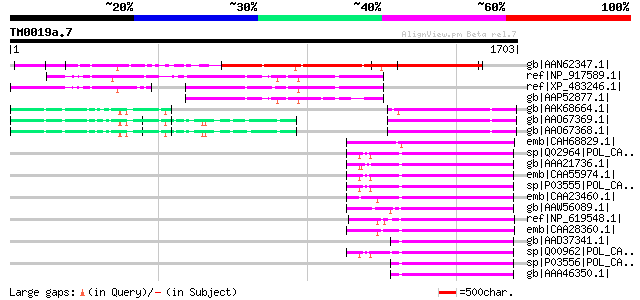
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN62347.1| CTV.20 [Poncirus trifoliata] 805 0.0
ref|NP_917589.1| sumilar to Oryza sativa chromosome 10, OSJNBb00... 413 e-113
ref|XP_483246.1| hypothetical protein [Oryza sativa (japonica cu... 363 3e-98
gb|AAP52877.1| hypothetical protein [Oryza sativa (japonica cult... 318 8e-85
gb|AAK68664.1| ORF I polyprotein [petunia vein clearing virus] g... 315 9e-84
gb|AAO67369.1| polyprotein 1 [Petunia vein clearing virus] 313 3e-83
gb|AAO67368.1| polyprotein 1 [Petunia vein clearing virus] 313 3e-83
emb|CAH68829.1| polyprotein [Carnation etched ring virus] 302 7e-80
sp|Q02964|POL_CAMVE Enzymatic polyprotein [Contains: Aspartic pr... 295 7e-78
gb|AAA21736.1| reverse transcriptase 294 2e-77
emb|CAA55974.1| unnamed protein product [Cauliflower mosaic virus] 293 3e-77
sp|P03555|POL_CAMVC Enzymatic polyprotein [Contains: Aspartic pr... 293 3e-77
emb|CAA23460.1| unnamed protein product [Cauliflower mosaic viru... 293 5e-77
gb|AAW56089.1| polyprotein [Horseradish latent virus] 293 5e-77
ref|NP_619548.1| Enzymatic polyprotein [Contains: Aspartic prote... 291 1e-76
emb|CAA28360.1| unnamed protein product [Carnation etched ring v... 290 2e-76
gb|AAD37341.1| reverse transcriptase [Cauliflower mosaic virus] 289 7e-76
sp|Q00962|POL_CAMVN Enzymatic polyprotein [Contains: Aspartic pr... 288 1e-75
sp|P03556|POL_CAMVD Enzymatic polyprotein [Contains: Aspartic pr... 287 2e-75
gb|AAA46350.1| ORF5; putative 282 8e-74
>gb|AAN62347.1| CTV.20 [Poncirus trifoliata]
Length = 3148
Score = 805 bits (2078), Expect = 0.0
Identities = 432/915 (47%), Positives = 598/915 (65%), Gaps = 58/915 (6%)
Query: 710 NSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTREDSQQAFWK 769
N+ + + Q+ GDPS ++DK+ +LL NL+C+ L DF+ YK TF TR++ R+D+ WK
Sbjct: 1951 NTELQNVTQYSDGDPSHLRDKNAELLHNLRCRKLSDFQLYKTTFFTRLFLRDDANHTTWK 2010
Query: 770 EKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRVALKICQDDKIQQQLT 829
EKFLA LP G+KVR +++ IPY L+YG+L++ + + LKICQD K+Q++L
Sbjct: 2011 EKFLACLPTLLGEKVRNFIKALYDN-RIPYDELTYGELVSFVNKEGLKICQDLKLQKRLK 2069
Query: 830 KEKSQNRRDLGTFCEQFGIQGCPKKPKPRKQ------DPPPKQQWRKRSSRN---DHRK- 879
+E Q++R+LG+FC+QF P K K P K+ ++ +S R HR+
Sbjct: 2070 QEIRQSKRELGSFCKQFNYD--PFKASTSKDCNGKCSSRPYKKHYKSKSHRKLFLGHREN 2127
Query: 880 --PKPRSKPQSSQIP-----KNPPETRPSQGKDVTCYNCGKPGHISRYCRLKRRISELHL 932
KP + S+ P K P+T P K+ C+ CG GH ++YC++ R++ EL L
Sbjct: 2128 FYKKPTRPYKKSRFPRKKDFKATPKT-PFNFKEAICHRCGIKGHTAKYCKMNRKLHELGL 2186
Query: 933 EPEIEDKINNLLIQTSDEEESNPSDSE-------VSEDLNQIQNDDSQSSSSVNTLSINT 985
+ +I KI L+I++S+ E S DS+ + D + + DS+S S + IN
Sbjct: 2187 DDDILSKIAPLMIESSNSESSMSGDSDSLQIDELIDSDTSVSNSSDSESESYLK--KINV 2244
Query: 986 LTNEQDLLFRAINSIPDPEEKKIYLERLRSTLE-DRPPKSPITINKFNLRDTFKRLEKST 1044
LT +Q+ + I DP +K YLE+L T++ ++ S + I K N D + L+K
Sbjct: 2245 LTKDQETFLELVKHISDPNLQKEYLEKLLKTMDFNKAETSKVPIVKKNSYDLTEILDKKK 2304
Query: 1045 VKPVT--IQDLHFEINSLKTEVKSLKQIQKSQ----QLILEKLTKNYEEDDSSIPDSNPA 1098
K IQDL EI +K+E+K LK+ QKS QL+L+K ++ +++S+ +++
Sbjct: 2305 TKKSVPNIQDLQKEIKDIKSEIKDLKEKQKSDSETIQLLLQKHLQDDSDNESTHSENHIE 2364
Query: 1099 PN-DNCE----DFLENINQVTIQKFFIHVKILIG-DFVLEIPALFDTGADSSCISEGLIP 1152
N DN E DFL + QVT +K+ I ++ DF ++ ALFDTGAD +CI + ++P
Sbjct: 2365 QNVDNIESVPHDFLFVLKQVTTRKYLIKATLIFSNDFAIDAIALFDTGADLNCIRQDIVP 2424
Query: 1153 TRYFEKTTEKLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIKK 1212
R+ EKT E+LSAA S+LK+ K+ ++I NG E +T F+L ++ +I+GTPFI
Sbjct: 2425 KRFHEKTKERLSAANNSKLKVDSKVEASIHNNG-FEFKTSFILTNDIHHAVILGTPFINL 2483
Query: 1213 LFPYNTDENGITVQHLGQPILFKFSEPPIDKTLNV--------------ISYKEKQINFL 1258
+ PY + + I+ + + I+F F E P + LN+ I+ K+ + FL
Sbjct: 2484 ITPYTVNYDSISFKVKNKKIIFPFIEKPKTRNLNIVKACSIYQNRINNLINSKQNDLTFL 2543
Query: 1259 KEEISYRTIEEQLQQPSVKSRIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDK 1318
++++S + IE LQ+ +K +I D I+ IC+DLP+AFW RK HMV+LPYE F +K
Sbjct: 2544 QKDLSLQRIENDLQRDFIKRKIFDFKTLIEKEICADLPSAFWNRKQHMVDLPYENSFDEK 2603
Query: 1319 QISTKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLV 1378
QI TKARPIQMN +L C++EINDL++K LI +S+SPWSCAAFYVNK +EIERGTPRLV
Sbjct: 2604 QIPTKARPIQMNMDLERHCKEEINDLVKKGLIVKSRSPWSCAAFYVNKNSEIERGTPRLV 2663
Query: 1379 INYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVP 1438
INYKPLN+AL WIRYPIPNKKDLL + H A +FSKFDMKSGFWQIQ+ KD+YKTAFTVP
Sbjct: 2664 INYKPLNKALKWIRYPIPNKKDLLQKLHSAFIFSKFDMKSGFWQIQIHPKDRYKTAFTVP 2723
Query: 1439 FGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFI 1498
FGQYEW VMPFGLKNAPSEFQRIMN+I+NPYS IVYIDDVLIFS ++DQHFKHL TF
Sbjct: 2724 FGQYEWTVMPFGLKNAPSEFQRIMNDIYNPYSDFCIVYIDDVLIFSNSIDQHFKHLKTFY 2783
Query: 1499 SVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRF 1558
++ GLA+S +KVSLFQTKIRFLGH+I +GTI PI R++ F DKFPD+I+DKTQLQRF
Sbjct: 2784 FATRKAGLAISNSKVSLFQTKIRFLGHHISKGTITPIERSLAFADKFPDKILDKTQLQRF 2843
Query: 1559 LGCLNYVADFCPQLS 1573
LG LNYV DFCP +S
Sbjct: 2844 LGSLNYVLDFCPNIS 2858
Score = 446 bits (1146), Expect = e-123
Identities = 210/285 (73%), Positives = 242/285 (84%)
Query: 1303 KSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAF 1362
K HMV+LPYE F +KQI TKARPIQMN +L C++EINDL++K LI +S+SPWSCAAF
Sbjct: 1123 KQHMVDLPYENSFDEKQIPTKARPIQMNMDLERHCKEEINDLVKKGLIVKSRSPWSCAAF 1182
Query: 1363 YVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQ 1422
YVNK +EIERGTPRLVINYKPLN+AL WIRYPIPNKKDLL + H A +FSKFDMKS FWQ
Sbjct: 1183 YVNKNSEIERGTPRLVINYKPLNKALKWIRYPIPNKKDLLQKLHSAFIFSKFDMKSVFWQ 1242
Query: 1423 IQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVYIDDVLI 1482
IQ+ K++YKTAFTVPFGQYEW VMPFGLKNAPSEFQRIMN+I+NPYS IVYIDDVLI
Sbjct: 1243 IQIHPKERYKTAFTVPFGQYEWTVMPFGLKNAPSEFQRIMNDIYNPYSDFCIVYIDDVLI 1302
Query: 1483 FSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFT 1542
FS ++DQHFKHL TF ++ GLA+S +KVSLFQTKIRFLGH+I +GTI PI R++ F
Sbjct: 1303 FSNSIDQHFKHLKTFYFATRKAGLAISNSKVSLFQTKIRFLGHHISKGTITPIERSLAFA 1362
Query: 1543 DKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDP 1587
DKFPD+I+DKTQLQRFLG LNYV DFCP +S + KPLHDRLKK P
Sbjct: 1363 DKFPDKILDKTQLQRFLGSLNYVLDFCPNISRLSKPLHDRLKKKP 1407
Score = 434 bits (1115), Expect = e-119
Identities = 347/1231 (28%), Positives = 561/1231 (45%), Gaps = 239/1231 (19%)
Query: 15 GLKPLTRKSLDIAVLLCLRDVRHNQFHDSLLGTVETSLSNGPIFFNCFPDLTVSLEDKNI 74
G+KPLT++ LD ++L LRD R F DSLL ++E+SL GPI F+C+P++T+SL+DKN+
Sbjct: 101 GIKPLTKEGLDTSILAVLRDGRFISFDDSLLSSIESSLCKGPIPFDCYPNITISLKDKNV 160
Query: 75 LDVLFLNIKLHGLDMKEDSIPISLIYRVQYKVMNSIKSYFLRTNCEKSGETTLFLTDYDK 134
L + L IK H M + SIP++LI+++ YK M S S R K ET L TD +
Sbjct: 161 LKSMILQIKTHNYKMIKGSIPVALIFKISYKAMVSAFSTQHRFQ-SKRDETLLLQTDLSR 219
Query: 135 ANVIVPKTIQWSDITLPEKWSLEKAT----PAQPAETPEFREIIQHQSGQVDLIFDRRNS 190
AN ++PK IQW DI LPE+W LE A P Q E + + Q+ G+V L+F R S
Sbjct: 220 ANTVIPKPIQWKDINLPEEWILEGAAPPTIPKQLEPNTELQNVTQYSDGKVKLLFRRSMS 279
Query: 191 FSIPR-----SIRTSSNDFQSAMSRRSFS-MTRRNQPESSTRPSLSHDGSTIHLAGYVNT 244
SI T F S + + ++QP ST S S+IH Y
Sbjct: 280 SRFSNKESCSSIPTLERKFTKIPSVINLPYQSTKSQPRFSTSDIPS---SSIHSVDYTTN 336
Query: 245 TP----VKTTYDKGDEDHQSTL--SPTYSSLETPGETSDFIGEITTEFVIDKGSIKADFY 298
P + + + E+ + +L SPT+S++ T + I I EF +DK + DFY
Sbjct: 337 VPHPIYTSSQHVQSQEEKEPSLPTSPTFSAV-----TENVINIIEKEFELDKTLLHNDFY 391
Query: 299 SDRWTQQRKWFFAKYQGEKRKKIQEKFYGFLEKTQQNIFFFEWFQDYANKNWKSYPFQID 358
SD +++ WFF ++ + RK+IQ+ +Y F+ + +I FF+WF+ Y+++N +YPF+
Sbjct: 392 SDLNKEKKIWFFKQFLNQ-RKEIQQIYYEFVNFHKVHILFFDWFEIYSSENNITYPFKES 450
Query: 359 --------MVKWQHKDGKETL-SNLPPKTPFLL-KGAQSTPVLASPFKTRKEEGDVTGKD 408
+ +W+ D +T+ S PP + G + ASP+K K + +
Sbjct: 451 NPITIRKKIPEWKLSDSDKTIESEHPPLRSLTIDHGEPPIQIRASPYKIPKPND--SDSN 508
Query: 409 IKDLMEQSNYTNKYLQILGE--SQVQDSSSRKGKEKVKIEPSSSNLDIPKGCQLEKPLFK 466
+ +++Q+N+ N L +G+ +++++ + P S D K +L++ +FK
Sbjct: 509 LSSIIQQNNFCNTNLNTIGKQLTRIENQFQKSTITVSSTSPIPSKSDYDK--KLKEAIFK 566
Query: 467 PFQISSRSRHSAQTLRAKKESDNELLQKVVEQLQLLKQVIPETPEPPETPEPTPEAVSNP 526
PFQ+S S+ Q ES ++ + + EQL ++ + + SN
Sbjct: 567 PFQVSKTSQKLVQ------ESKSDFAKAIREQLDRIE---------------SASSSSNK 605
Query: 527 TITAPPPPKTRTSTRNTSTSLNNIEDGKVESDSDIQSMKSDVQSIKSIPVNTVIHNSKNQ 586
AP P++ + G +E D SD+++ K +K
Sbjct: 606 VQIAPNTPQSS-------------KIGVLEQDDQTSIASSDLEAFKDEAPTP--RANKIH 650
Query: 587 WRKETKLYYPRA-TAPDLLLEESSNFKSFSANNVYEWNIDGENEYGITKILQNMTMVATA 645
W +L P + PDL ++ ++LQ MTM A A
Sbjct: 651 W----ELAPPTVKSPPDLAIDNR------------------------PRVLQQMTMAANA 682
Query: 646 YVTANNCPESLIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIMEDGKFI 705
Y T + + I E+L+AGF GQLKG
Sbjct: 683 YKTQSGTSDMAIAEILIAGFTGQLKG---------------------------------- 708
Query: 706 SDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTREDSQQ 765
DPS ++DK+ +LL NL+C+ L DF+ YK TF TR++ R+D+
Sbjct: 709 -----------------DPSHLRDKNAELLHNLRCRKLSDFQLYKTTFFTRLFLRDDANH 751
Query: 766 AFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRVALKICQDDKIQ 825
WKEKFLAGLP G+KVR +++ IPY L+YGQL++ + + LKICQD K+Q
Sbjct: 752 TTWKEKFLAGLPTLLGEKVRNSIKALY-DNRIPYDELTYGQLVSFVNKEGLKICQDLKLQ 810
Query: 826 QQLTKEKSQNRRDLGTFCEQFGIQGCPKKPKPRKQDPPPKQQWRKRSSRNDHRKPKPRSK 885
++L +E Q++R+LG+FC+QF P K K + +
Sbjct: 811 KRLKQELRQSKRELGSFCKQFNYD-------------PFKASTSKDCNGDSDSLQIDELI 857
Query: 886 PQSSQIPKNPPETRPSQGKDVTCYNCGKPGHISRYCRLKRRISELHLEPEIEDKINNLLI 945
+ + + S K + + + L + IS+ +L+ E +K+ +
Sbjct: 858 DSDTSVSNSSDSESESYLKKINILTKDQ----ETFLELVKHISDPNLQKEYLEKLLKTMD 913
Query: 946 QTSDEEESNPSDSEVSEDLNQIQNDDSQSSSSVNTLSINTLTNEQDLLFRAINSIPDPEE 1005
E P + S DL +I D ++ SV N QDL +
Sbjct: 914 FNKAETSKVPIVKKNSYDLTEIL-DKKKTKKSV--------PNIQDL------------Q 952
Query: 1006 KKIYLERLRSTLEDRPPKSPITINKFNLRDTFKRLEKSTVKPVTIQDLHFEINSLKTEVK 1065
K+I + ++S ++D K L L+K + H E N ++ V
Sbjct: 953 KEI--KDIKSEIKDLKEKQKSDSETIQL-----LLQKHLQDDSDNESTHSE-NHIEQNVD 1004
Query: 1066 SLKQIQKSQQLILEKLTKNYEEDDSSIPDSNPAPNDNCEDFLENINQVTIQKFFIHVKIL 1125
+++ + +L+++T +K+ I ++
Sbjct: 1005 NIESVPHDFLFVLKQVT--------------------------------TRKYLIKATLI 1032
Query: 1126 I-GDFVLEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSAAEGSRLKIKYKIPSAIIKN 1184
DF ++ ALFDTGAD +CI + ++P R+ EKT E+LSAA S+LK+ K+ ++I N
Sbjct: 1033 FSNDFAIDAIALFDTGADLNCIRQDIVPKRFHEKTKERLSAANNSKLKVDSKVEASIHNN 1092
Query: 1185 GSLEIETPFLLVRNLSQKIIIGTPFIKKLFP 1215
G E +T F+L ++ +I+GTPFI + P
Sbjct: 1093 G-FEFKTSFILTNDIHHAVILGTPFINLITP 1122
Score = 55.8 bits (133), Expect = 1e-05
Identities = 28/71 (39%), Positives = 37/71 (51%), Gaps = 4/71 (5%)
Query: 121 KSGETTLFLTDYDKANVIVPKTIQWSDITLPEKWSLEKAT----PAQPAETPEFREIIQH 176
K ET L TD +AN ++PK IQW DI LPE+W LE A P Q E + + Q+
Sbjct: 1901 KRDETLLLQTDLSRANTVIPKPIQWKDINLPEEWILEGAAPPTFPKQLEPNTELQNVTQY 1960
Query: 177 QSGQVDLIFDR 187
G + D+
Sbjct: 1961 SDGDPSHLRDK 1971
>ref|NP_917589.1| sumilar to Oryza sativa chromosome 10, OSJNBb0036B06.4 [Oryza sativa
(japonica cultivar-group)] gi|20804792|dbj|BAB92476.1|
hypothetical protein [Oryza sativa (japonica
cultivar-group)] gi|18677105|dbj|BAB84832.1| hypothetical
protein [Oryza sativa (japonica cultivar-group)]
Length = 1634
Score = 413 bits (1062), Expect = e-113
Identities = 348/1200 (29%), Positives = 543/1200 (45%), Gaps = 207/1200 (17%)
Query: 123 GETTLFLTDYDKANVIVPKTIQWSDITLPEKWSLEKATPAQPAETPEFREIIQHQSGQVD 182
G TTL T+ + + I PKT++W +ITLPEKW L +A + + E +I+ G V+
Sbjct: 15 GLTTLLQTNPNNRHTI-PKTLKWDEITLPEKWVLSQAVEPKSMDQSEVESLIETLDGDVE 73
Query: 183 LIFDRRNSFSIPRSIRTSSNDFQSAMSRRSFSMTRRNQPESSTRPSLSHDGSTIHLAGYV 242
+ F A +++F +R PS+S D
Sbjct: 74 ITF---------------------ASKQKAFLQSR---------PSVSLDSRP------- 96
Query: 243 NTTPVKTTY----DKGDEDHQSTLSPTYSSLETPGETSDFIGEITTEFVIDKGSIKADFY 298
T P Y D DE S L+ I EF IDK ++ +
Sbjct: 97 RTKPQNVVYATYEDNSDEPSISDFDINVIELDV----GFVIALEEEEFEIDKELLRREIR 152
Query: 299 SDRWTQQRKWFFAKYQGEKRKKIQEKFYGFLEKTQQNIFFFEWFQD-------------- 344
+ + K + + R KI+E ++ + + ++NIFFF+W+++
Sbjct: 153 LPKNRAKTKRYLEEVDESFRMKIREVWHNEMREQRRNIFFFDWYENSQIIHFEEFFKRKN 212
Query: 345 YANKNWKSYPFQIDMVK-----WQHKDGKETLSNLPPKTPFLLK--GAQSTPVLASPFKT 397
K KS + ++K W+ G + S PP L G ++ P+ + T
Sbjct: 213 MIKKEQKSEAEDLTVIKKVSTEWETASGNKVDSVHPPFESIQLSHNGGKACPLKSISKNT 272
Query: 398 RKEEGDVTGKDIKDLMEQSNYTNKYLQILGESQ-----VQDSSSRKGKEKVKIE-PSSSN 451
E V + I L+EQ NY N L+ LG+ + + G+ +VKI PS+S
Sbjct: 273 YGETAKV--EHIGHLVEQQNYANISLRSLGQQTDRIETILMEGYKTGRPEVKINIPSNSQ 330
Query: 452 LDIPKGCQLEKPLFKPFQISSRSRHSAQTLRAKKESDNELLQKVVEQLQLLKQVIPETPE 511
Q P+F P I + Q S+ EL+ ++ +L LK
Sbjct: 331 ---SSSSQSVSPMFVP-TIDPNIKFGKQKAFGPAISE-ELVNELALKLNNLK-------- 377
Query: 512 PPETPEPTPEAVSNPTITAPPPPKTRTSTRNTSTSLNNIEDGKVESDSDIQSMKSDVQSI 571
+ ++N I D + K D+ +
Sbjct: 378 -------------------------------VNKNINEISDNE----------KYDIVN- 395
Query: 572 KSIPVNTVIHNSKNQWRKETKLYYPRATAPDLLLEES---SNFKSFSANNVYEWNIDGEN 628
K +T+ ++N YYPR T DL EE N ++ + EWN+DG
Sbjct: 396 KIFKPSTLTSTTRN--------YYPRPTYADLQFEEMPQIQNMTYYNGKEIVEWNLDGFT 447
Query: 629 EYGITKILQNMTMVATAYVTANNCPESLIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTA 688
EY I + M M A A + AN E ++V GF GQLKGWW+NYL + ++ +IL A
Sbjct: 448 EYQIFTLCHQMIMYANACI-ANGNKEREAANMIVIGFSGQLKGWWNNYLNETQRQEILCA 506
Query: 689 VKTDEEGNPIME-DGKF-----ISDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKS 742
VK D++G P+ + DG ISDA+ +LI+ I HF G+ I +K+ + L NLKCK+
Sbjct: 507 VKRDDQGRPLPDRDGNGNPTGNISDALATLIYNIIYHFAGNYHDIYEKNREQLINLKCKT 566
Query: 743 LGDFRWYKDTFLTRVYTREDSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTL 802
+ DFRWYKDTFL+++YT + Q FWKEK+++GLP F +KVR LR + GG I YH L
Sbjct: 567 MSDFRWYKDTFLSKLYTLPEPNQDFWKEKYISGLPPLFAEKVRNSLRKEG-GGSINYHYL 625
Query: 803 SYGQLIAIIQRVALKICQDDKIQQQLTKEKSQNRRDLGTFCEQFGIQGCPKKPKPRKQDP 862
G++ IQ V ++C D KI+ QL K++ +R++G FC QFG QDP
Sbjct: 626 DIGKITQKIQLVGAELCNDLKIKDQLKKQRILGKREMGDFCYQFGF-----------QDP 674
Query: 863 PPKQQWRKRSSRNDHRKP--KPRSKPQSS-QIPKNPPE--------TRPSQGKDVTCYNC 911
+RKR + H KP KP K + S Q K P+ T+ ++ K+ CY C
Sbjct: 675 ---YVYRKRKT---HSKPMTKPNDKSKMSFQATKRKPKRIYNKNIRTQDTESKETICYKC 728
Query: 912 GKPGHISRYC---RLKRRISELHLEPEIEDKINNLLIQTSDEEESNPSDSEVSEDLNQIQ 968
G GHI+ C ++K+ I L L+ E ED L ++ + SD E + ++N Q
Sbjct: 729 GLKGHIANRCFKSKVKKEIQAL-LDSESEDVKEKLEAILNNIDNDLSSDEEKNAEINCCQ 787
Query: 969 N--------DDSQSSSSVNTLSINTLTNEQDLLFRAINSIPDPEEKKIYLERLRSTLEDR 1020
+ D+S+ S N L LT+ ++ + +I DPEEK+ LE+ S ++
Sbjct: 788 DSGCSCCEPDNSEEESDENIL---VLTSLEEFVLDTFETIQDPEEKRRVLEKFLSRVKTD 844
Query: 1021 PPKSPITINK---FNLRDTFKRLEKSTVKPVTIQDLHFEINSLKTEVKSLKQIQKSQQLI 1077
K I K ++ + FKRL++ K DL K + L++I+K ++
Sbjct: 845 KDKLKKDIQKSKFLSIDEVFKRLDEQKKKNEQ-PDLISLFEDQKIMKQDLEEIRKRLYML 903
Query: 1078 LEKLTKNYEEDDSSIPDSNPAPNDNCEDFLENINQVTIQKFFIHV--KILIGDFVLEIPA 1135
K + EE D I + + + I + QK++ V + + G + I
Sbjct: 904 ESKEGFHMEEKDEPIQED--------DHVVGTIQKYMKQKWYTEVMYRFIDGSYFQHI-T 954
Query: 1136 LFDTGADSSCISEGLIPTRYFEKTTEKLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLL 1195
L D+GAD SCI EG+IP +YF K ++ A+G L ++Y+IP I + I+T FLL
Sbjct: 955 LIDSGADVSCIREGIIPHKYFYKAAHRIRGADGRLLTVEYQIPEIYICISEVCIKTSFLL 1014
Query: 1196 VRNLSQKIIIGTPFIKKLFPYNTDENGITVQHLGQPILFKFSEPPIDKTLNVISYKEKQI 1255
V+NL Q +I+GTPF+ + P+ I + + + + +FS + +I K +Q+
Sbjct: 1015 VKNLKQDVILGTPFLSLIRPFLVTNEDIQFKIMDKQVSLRFSSNTDEILDQLIQTKREQV 1074
>ref|XP_483246.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|42408922|dbj|BAD10179.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1275
Score = 363 bits (932), Expect = 3e-98
Identities = 233/692 (33%), Positives = 362/692 (51%), Gaps = 51/692 (7%)
Query: 591 TKLYYPRATAPDLLLEESSNFKS---FSANNVYEWNIDGENEYGITKILQNMTMVATAYV 647
T+ YYPR T DL EE ++ ++ + EWN+DG EY I + M M A A +
Sbjct: 487 TRNYYPRPTYADLQFEEMPQIQNMIYYNGKEIVEWNLDGFTEYQIFTLCHQMIMYANACI 546
Query: 648 TANNCPESLIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIME-DGKF-- 704
AN E ++V GF GQLKGWW+NYL + ++ +IL AVK D++G P+ + DG
Sbjct: 547 -ANGNKEREAANMIVIGFSGQLKGWWNNYLNETQRQEILCAVKRDDQGRPLPDRDGNGNP 605
Query: 705 ---ISDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTRE 761
ISDA+ +LI+ I HF G+ I +K+ + L NLKCK++ DFRWYKDTFL+++YT
Sbjct: 606 TGNISDALATLIYNIIYHFAGNYHDIYEKNREQLINLKCKTMSDFRWYKDTFLSKLYTLP 665
Query: 762 DSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRVALKICQD 821
+ Q FWKEK+++GLP F +KVR LR + GG I YH L G++ IQ V ++C D
Sbjct: 666 EPNQDFWKEKYISGLPPLFAEKVRNSLRKEG-GGSINYHYLDIGKITQKIQLVGAELCND 724
Query: 822 DKIQQQLTKEKSQNRRDLGTFCEQFGIQG--CPKKPKPRKQDPPPKQQWRKRSSRNDHRK 879
KI+ QL K++ +R++G FC QFG Q +K K + K S + RK
Sbjct: 725 LKIKDQLKKQRILGKREMGDFCYQFGFQDPYVHRKRKTHSKPMTKPNDKSKMSFQATKRK 784
Query: 880 PKPRSKPQSSQIPKNPPETRPSQGKDVTCYNCGKPGHISRYC---RLKRRISELHLEPEI 936
PK +I T+ ++ K+ CY CG GHI+ C ++K+ I L L+ E
Sbjct: 785 PK--------RIYNKNIRTQDTESKETICYKCGLKGHIANRCFKSKVKKEIQAL-LDSES 835
Query: 937 EDKINNLLIQTSDEEESNPSDSEVSEDLNQIQN--------DDSQSSSSVNTLSINTLTN 988
ED L ++ + + SD E + ++N Q+ D+S+ S N L LT+
Sbjct: 836 EDVKEKLEAILNNIDNDSSSDEEKNAEINCCQDSGCSCCEPDNSEEESDENIL---VLTS 892
Query: 989 EQDLLFRAINSIPDPEEKKIYLERLRSTLEDRPPKSPITINK---FNLRDTFKRLEKSTV 1045
++ + +I DPEEK+ LE+ S ++ K I K ++ + FKRL++
Sbjct: 893 LEEFVLDTFETIQDPEEKRRVLEKFLSRVKTDKDKLKKDIQKSKFLSIDEVFKRLDEQKK 952
Query: 1046 KPVTIQDLHFEINSLKTEVKSLKQIQKSQQLILEKLTKNYEEDDSSIPDSNPAPNDNCED 1105
K DL K + L++I+K ++ K + EE D I + +
Sbjct: 953 KNEQ-PDLISLFEDQKIMKQDLEEIRKRLYMLESKEGFHMEEKDEPIQED--------DH 1003
Query: 1106 FLENINQVTIQKFFIHV--KILIGDFVLEIPALFDTGADSSCISEGLIPTRYFEKTTEKL 1163
+ I + QK++ V + + G + I L D+GAD +CI EG+IP +YF K ++
Sbjct: 1004 VVGTIQKYMKQKWYTEVMYRFIDGSYFQHI-TLIDSGADVNCIREGIIPHKYFYKAAHRI 1062
Query: 1164 SAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDENGI 1223
A+G L ++Y+IP I + I+T FLLV+NL Q +I+GTPF+ + P+ I
Sbjct: 1063 RGADGGLLTVEYQIPEIYICITEVCIKTSFLLVKNLKQDVILGTPFLSLIRPFLVTNEDI 1122
Query: 1224 TVQHLGQPILFKFSEPPIDKTLNVISYKEKQI 1255
+ +G+ + +FS + ++ K +Q+
Sbjct: 1123 QFKIMGKQVSLRFSSNTDEILDQLVQTKREQV 1154
Score = 191 bits (484), Expect = 2e-46
Identities = 147/498 (29%), Positives = 231/498 (45%), Gaps = 80/498 (16%)
Query: 4 FNYLHIGSVQVGLKPLTRKSLDIAVLLCLRDVRHNQFHDSLLGTVETSLSNGPIFFNCFP 63
++YLH G +QV ++ LTRK L+++VL CLRD R+ +F DSLLG VE SLSNGP++FN FP
Sbjct: 47 YHYLHFGLIQVAIRSLTRKGLNVSVLACLRDCRNKRFKDSLLGMVEASLSNGPVYFNTFP 106
Query: 64 DLTVSLEDKNILDVLFLNIKLHGLDMKEDSIPISLIYRVQYKVMNSIKSYFLRTNCEKSG 123
D +VSL D NI VL LN++ G D++ S IS+ YRV YK M ++ + G
Sbjct: 107 DFSVSLSDTNIHKVLTLNLQTSGYDLEPGSENISVTYRVYYKAMTTLAP--CAKHYTPKG 164
Query: 124 ETTLFLTDYDKANVIVPKTIQWSDITLPEKWSLEKATPAQPAETPEFREIIQHQSGQVDL 183
TTL T+ + + I PKT++W +ITLPEKW L +A + + E +I+ G V++
Sbjct: 165 LTTLLQTNPNNRHTI-PKTLKWDEITLPEKWVLSQAVEPKSMDQSEVESLIETPDGDVEI 223
Query: 184 IFDRRNSFSIPRSIRTSSNDFQSAMSRRSFSMTRRNQPESSTRPSLSHDGSTIHLAGYVN 243
F A +++F +RPS+S D
Sbjct: 224 TF---------------------ASKQKAF---------LQSRPSVSLDSRP-------R 246
Query: 244 TTPVKTTY----DKGDEDHQSTLSPTYSSLETPGETSDFIGEITTEFVIDKGSIKADFYS 299
T P Y D DE S L + I EF IDK +K +
Sbjct: 247 TKPQNVVYATYEDNSDEPSISDFDINVIEL----DVGFVIAIEEDEFEIDKDLLKRELRL 302
Query: 300 DRWTQQRKWFFAKYQGEKRKKIQEKFYGFLEKTQQNIFFFEWFQDYANKNWKSYPFQIDM 359
+ + K +F + R KI+E ++ + + ++NIFFF+W++ ++++ + + +M
Sbjct: 303 QKNRPKMKRYFERVDEPFRLKIRELWHKEMREQRKNIFFFDWYESSQVRHFEEFFKRKNM 362
Query: 360 VK-------------------WQHKDGKETLSNLPPKTPFLL--KGAQSTPVLASPFKTR 398
+K W+ G + S PP L G ++ P+ + T
Sbjct: 363 IKKEQKSEAEDLTVIKKVSTEWETASGNKVDSVHPPFESVQLSHNGGKACPLKSISKNTY 422
Query: 399 KEEGDVTGKDIKDLMEQSNYTNKYLQILGESQVQDSSSRKGKEKVKIEPSSSNLDIPKGC 458
E V + I L+EQ NY N L+ LG+ E + +E + P+
Sbjct: 423 GETAKV--EHIGHLVEQQNYANISLRSLGQ-------QTNRIETILMEGYKTGR--PEKY 471
Query: 459 QLEKPLFKPFQISSRSRH 476
+ +FKP ++S +R+
Sbjct: 472 DMVNKIFKPSTLTSTTRN 489
>gb|AAP52877.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37532576|ref|NP_920590.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|15217202|gb|AAK92546.1| Hypothetical protein [Oryza
sativa]
Length = 1204
Score = 318 bits (816), Expect = 8e-85
Identities = 228/704 (32%), Positives = 347/704 (48%), Gaps = 113/704 (16%)
Query: 591 TKLYYPRATAPDLLLEES---SNFKSFSANNVYEWNIDGENEYGITKILQNMTMVATAYV 647
T+ YYPR T DL EE N ++ + EWN+DG EY I + M M A A +
Sbjct: 15 TRNYYPRPTYADLQFEEMPQIQNMTYYNGKEIVEWNLDGFTEYQIFTLCHQMIMYANACI 74
Query: 648 TANNCPESLIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIME-DGKF-- 704
AN E ++V GF GQLKGWW+NYL + ++ +IL AVK D++G P+ + DG
Sbjct: 75 -ANGNKEREAANMIVIGFSGQLKGWWNNYLNETQRQEILCAVKRDDQGRPLPDRDGNGNP 133
Query: 705 ---ISDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTRE 761
ISDA+ +LI+ I HF G+ I +K+ + L NLKCK++ DFRWYKDTFL+++YT
Sbjct: 134 TGNISDALATLIYNIIYHFAGNYHDIYEKNREQLINLKCKTMSDFRWYKDTFLSKLYTLP 193
Query: 762 DSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRVALKICQD 821
+ Q FWKEK+++GLP F +KVR LR + GG I YH L G++ IQ V +++C D
Sbjct: 194 EPNQDFWKEKYISGLPPLFAEKVRNSLRKEG-GGSINYHYLDIGKITQKIQLVGVELCND 252
Query: 822 DKIQQQLTKEKSQNRRDLGTFCEQFGIQGCPKKPKPRKQDPPPKQQWRKRSSRNDHRKP- 880
KI+ QL K++ +R++G FC QFG QDP +RKR + H KP
Sbjct: 253 LKIKDQLKKQRILGKREMGDFCYQFGF-----------QDP---YVYRKRKT---HSKPM 295
Query: 881 -KPRSKPQSS-QIPKNPPE--------TRPSQGKDVTCYNCGKPGHISRYC---RLKRRI 927
KP K + S Q K P+ T+ ++ K+ CY CG GHI+ C ++K+ I
Sbjct: 296 TKPNDKSKMSFQATKRKPKRIYNKNIRTQDTESKETICYKCGLKGHIANRCFKSKVKKEI 355
Query: 928 SELHLEPEIEDKINNLLIQTSDEEESNPSDSEVSEDLNQIQN--------DDSQSSSSVN 979
L L+ E ED L ++ + + SD E + ++N Q+ D+S+ S N
Sbjct: 356 QAL-LDSESEDVKEKLEAILNNIDNDSSSDEEKNAEINCCQDSGCSCYEPDNSEEESDEN 414
Query: 980 TLSINTLTNEQDLLFRAINSIPDPEEKKIYLERLRSTLEDRPPKSPITINK---FNLRDT 1036
L LT+ ++ + +I DPEEK+ LE+ S ++ K I K ++ +
Sbjct: 415 IL---VLTSLEEFVLDTFETIQDPEEKRRVLEKFLSRVKTDKDKLKKDIQKSKFLSIDEV 471
Query: 1037 FKRLE---KSTVKPVTIQDLHFEINSLKTEVKSLKQIQKSQQLILEKLTKNYEEDDSSIP 1093
FKRL+ K KP DL + + L++I+K + K + EE D I
Sbjct: 472 FKRLDEQKKKNEKP----DLISLFEDQRIMKQDLEEIRKRLYRLESKEGFHMEEKDEPIQ 527
Query: 1094 DSNPAPNDNCEDFLENINQVTIQKFFIHV--KILIGDFVLEIPALFDTGADSSCISEGLI 1151
+ + + I + QK++ V + + G + I L D+GAD +CI E
Sbjct: 528 ED--------DQVVGTIQKYMKQKWYTEVMYRFIDGSYFQHI-TLIDSGADVNCIRE--- 575
Query: 1152 PTRYFEKTTEKLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIK 1211
+ I+T FLLV+NL Q +I+GTPF+
Sbjct: 576 -----------------------------------VCIKTSFLLVKNLKQDVILGTPFLS 600
Query: 1212 KLFPYNTDENGITVQHLGQPILFKFSEPPIDKTLNVISYKEKQI 1255
+ P+ I + +G+ + +FS + ++ K +Q+
Sbjct: 601 LIRPFLVTNEDIQFKIMGKQVSLRFSSNTDEILDQLVQSKREQV 644
>gb|AAK68664.1| ORF I polyprotein [petunia vein clearing virus]
gi|14575753|ref|NP_127504.1| ORF I polyprotein [petunia
vein clearing virus]
Length = 2179
Score = 315 bits (807), Expect = 9e-84
Identities = 176/439 (40%), Positives = 262/439 (59%), Gaps = 14/439 (3%)
Query: 1270 QLQQPSVKSRIEDILENIQSSICSDLPNAFWERKSH--------MVELPYEKDFSDKQIS 1321
QL + ++ED+++N +C+D F + SH ++LP++K+ +
Sbjct: 1331 QLTTSNELQQLEDVIKN---QLCADSHVDFLSKCSHPLWLNQDFFIQLPFKKN--ENINP 1385
Query: 1322 TKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINY 1381
TKA MN E L KE ++L + LI S S W+C AFYVNK++E RG RLVINY
Sbjct: 1386 TKASHSGMNPEHLQLAIKECDELQQFDLIEPSDSQWACEAFYVNKRSEQVRGKLRLVINY 1445
Query: 1382 KPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQ 1441
+PLN L ++PIPNK L + AK+FSKFD+KSGFWQ+ + ++ KT F +P
Sbjct: 1446 QPLNHFLQDDKFPIPNKLTLFSHLSKAKLFSKFDLKSGFWQLGIHPNERPKTGFCIPDRH 1505
Query: 1442 YEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVI 1501
++W VMPFGLK APS FQ+ M +IF P +VYIDD+L+FS+TL+ H K LN FIS++
Sbjct: 1506 FQWKVMPFGLKTAPSLFQKAMIKIFQPILFSALVYIDDILLFSETLEDHIKLLNQFISLV 1565
Query: 1502 KRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGC 1561
K+ G+ +S K+ L Q KI+FLG + GT P KFPD + Q+Q+FLG
Sbjct: 1566 KKFGVMLSAKKMILAQNKIQFLGMDFADGTFSPAGHISLELQKFPDTNLSVKQIQQFLGI 1625
Query: 1562 LNYVADFCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKI 1621
+NY+ DF P+++ I PL D LKK PP N VKQ+K + + L++P+ + KI
Sbjct: 1626 VNYIRDFIPEVTEHISPLSDMLKKKPPAWGKCQDNAVKQLKQLAQQVKSLHIPS-EGKKI 1684
Query: 1622 IETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDL 1681
++TDASD + +L ++ K +I F S + ++Q+Y + KE+LA+ I KF L
Sbjct: 1685 LQTDASDQYWSAVLLEEHNGKRKICGFASGKFKVSEQHYHSTFKEILAVKNGIKKFNFFL 1744
Query: 1682 INQKFLVCVDCKSAKEILQ 1700
I+ FLV +D ++ ++++
Sbjct: 1745 IHTNFLVEMDMRAFPKMIR 1763
Score = 84.3 bits (207), Expect = 3e-14
Identities = 143/597 (23%), Positives = 234/597 (38%), Gaps = 105/597 (17%)
Query: 1 SKNFNYLHIGSVQVGLKPLTRKSLDIAVLLCLRDVRHNQFHDSLLGTVETSLSNGPIFFN 60
S F ++H G+V++ L RK + L L D R+ ++ + LGT E +L+ G +F
Sbjct: 119 SHQFTHIHFGAVKIALTYHGRKGQPVVARLALLDTRYLEYQHANLGTAEITLNAGTVFIT 178
Query: 61 CFPDLTVSLEDKNILDVLFLNIKLHGLDMKEDSIPISLIYRVQYKVMNSIKSYFLRTNCE 120
FP+ T+SL D N+ L + +++ G + +DSI +L Y++ ++V N L
Sbjct: 179 LFPNFTMSLSDANLSTALKIQVQIQGAPLTKDSIQATLHYQIAWRVQNHAMDLTL----- 233
Query: 121 KSGETTLFLTDYDKANVI----VPKTIQWSDI--TLPEKWSLEKATPAQPAETPEFREII 174
GE LFL + VP+ + D+ LP+ W +P E E+
Sbjct: 234 PGGEEALFLKIDAGSGATQCTQVPRQLSKEDLIKILPDSWVTNYEKLREPEEPLRSTEVS 293
Query: 175 QHQSGQVDLIFDRRNSFSIPRSIRTSSNDFQSAMSRRSFSMTRRNQPE--SSTRPSLSHD 232
+ D+ + S S + M S + + PE S T PSL
Sbjct: 294 MSKR------HDKSVAISFDHSHYKKLRNTHHFMGMISDDVIVLDDPETFSKTLPSLMQT 347
Query: 233 GSTIH---LAGYVNTTPVKTTYD---KGDEDHQSTLSPTYSSLETPGETSDFIGEITTEF 286
IH L G + K +D D D Q YS L + E DF T++
Sbjct: 348 HDWIHHFQLDGRA-VSWYKDPFDGHCPWDIDCQ-----CYSCLYSEDE-EDFEDGFPTKY 400
Query: 287 VIDKGSIKADFYSDRWTQQRKWFFAKYQGEKRKKIQEK--FYGFLEKTQQNIFFFEWFQD 344
KG + ++R Q+ + +K +EK F G L + + +E+
Sbjct: 401 ---KGIPRPGSIAERKMQE--------EANLKKLYEEKDPFVGSLSRPGK----YEYLVR 445
Query: 345 YANKNWKSYP-FQIDMVKWQHKD---------GKETL------------SNLPPKTPFLL 382
Y +W P ++ W + + TL +N PP + F
Sbjct: 446 YDAPSWAKDPHLTVEPTGWDSDEPIPPKQPFTTRNTLPKIYMFNPLNYENNFPPLSSFSK 505
Query: 383 KGAQSTP-------VLASPFKTRKEEGDVTGKDIKDLMEQSNYTNKYLQILGESQVQDSS 435
GA TP VL S K D TG DL N+ + L ++++ +
Sbjct: 506 DGADHTPKIPKRNVVLPSGAK------DPTG----DLEATVNWQTE--NALAQNRMLTTI 553
Query: 436 SRKGKEKV-KIEPSSSNLDIPKGCQLEKPLFKPFQ-ISSRSRHSAQTLRAKKESDNELLQ 493
R KE V K++ + +G L K L + Q ++ R +L + +
Sbjct: 554 DRTLKETVTKVDRVTDQSSKNQG--LIKVLEQQLQDLNKRICPPGTSLFHFFDQQKSEMA 611
Query: 494 KVVEQLQLLKQVIPETPEPPETPEPTPE-------AVSNPTITAPPPPKTRTSTRNT 543
+ EQ++LLK E P+ ET P+ + + S+P + + PP T+ NT
Sbjct: 612 SLKEQIRLLK----EQPQKNETDTPSYQSSYQPFHSFSSPYMPSNPPNSPFTTFANT 664
Score = 49.3 bits (116), Expect = 0.001
Identities = 111/571 (19%), Positives = 210/571 (36%), Gaps = 129/571 (22%)
Query: 459 QLEKPLFKPFQISSRSRHSAQTLRAKKESDNELLQKVVEQLQLLK-QVIPETPEPPETPE 517
Q + LF + I +S ++ + E + + + + +L K +V +T PPE+
Sbjct: 666 QPQPSLFSQYPIQPKSPNTFDLAKLVWEKKDAIAAEKRAKKKLQKDEVKQKTSLPPESKR 725
Query: 518 PTPEA---------VSNP------TITAPPPPKTRTSTRNTSTSLNNIEDGKVESDS-DI 561
P P++ +S+P + P P TS+++ ++ ++ED +DS +
Sbjct: 726 PDPQSSSHLGDQFMISDPALPKVYALNEPSVPSEDTSSQSYISTEESVED----TDSFSV 781
Query: 562 QSMKSDVQSIKSIPVNTVIHNSKNQWRKETKLYYPRATAPDLLLEESSNFKSFSANNVYE 621
S +S S S N N++N + + R T P++ + E
Sbjct: 782 VSEESTQLSQLSSSSNDSPENNENTL---PQTFMVRPTEPEI--------------SEVE 824
Query: 622 WNIDGENEYGITKILQNMT---MVATAY-------VTANNCPES---------------- 655
+DG E I + +T MV T + ++ PE
Sbjct: 825 DEVDGMTEEPIPERRPEITPPKMVGTGFHTFSLDDISITKWPERIQDFHTWMLTKQLVER 884
Query: 656 ---LIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIMEDGKFISDAVNSL 712
LI+ A G L+ WW N + D+K++ LT S
Sbjct: 885 EPFLILSEFTARLSGTLREWW-NSVGPDDKNRFLT------------------SQDFTWN 925
Query: 713 IFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTR------EDSQQA 766
I + +F GD S K++ + +KC S Y + R + R
Sbjct: 926 IRILYSYFCGDQSQNKEELRRQIFEMKCLS------YDRKKIDRHFQRMIKLFYHIGGDI 979
Query: 767 FWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRVALKICQDDKIQQ 826
K+ F++ LP +++ ++ + T + I++ A + D ++
Sbjct: 980 SLKQAFISSLPPILSERISALIKERG--------TSVTQMHVGDIRQTAFYVLDDLCSKR 1031
Query: 827 QLTKEKSQNRRDLGTFCEQF-----GIQGCPKKPKP---RKQDPPPKQQWRKRSSRNDHR 878
+ + + RDL C + G +GC P RK +++R R +
Sbjct: 1032 KFFNQMKKMSRDLEKACTKSDLIIKGDKGCSGYCNPSRRRKYKRFKLPSFKERDGRQYRK 1091
Query: 879 KPKPRSKPQSSQIPKNPPETRPSQGKDVTCYNCGKPGHISRYCRLKRR----ISEL--HL 932
+ + + ++S+ + P +C+ CGK GH SR C ++ ISE+ +
Sbjct: 1092 RRRFFRRSKTSKAMRQKPR---------SCFTCGKIGHFSRNCPQNKKSIKLISEIQKYT 1142
Query: 933 EPEIEDKINNLLIQTSDEEESNPSDSEVSED 963
+IED + ++ + E E E+
Sbjct: 1143 GIDIEDDLESVFSIEDEPSEDTLFSLEFYEE 1173
>gb|AAO67369.1| polyprotein 1 [Petunia vein clearing virus]
Length = 1886
Score = 313 bits (802), Expect = 3e-83
Identities = 172/436 (39%), Positives = 258/436 (58%), Gaps = 8/436 (1%)
Query: 1270 QLQQPSVKSRIEDILEN-----IQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKA 1324
QL + ++ED+++N S + W + ++LP++++ + TKA
Sbjct: 1332 QLTTSNELQQLEDVIKNQLCAESHVDFLSKCSHPLWLNQDFFIQLPFKRN--ENINPTKA 1389
Query: 1325 RPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPL 1384
MN E L KE ++L + LI S S W+C AFYVNK++E RG RLVINY+PL
Sbjct: 1390 SHSGMNPEHLQLALKECDELQQFDLIEPSDSQWACEAFYVNKRSEQVRGKLRLVINYQPL 1449
Query: 1385 NQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEW 1444
N L ++PIPNK L + AK+FSKFD+KSGFWQ+ + ++ KT F +P ++W
Sbjct: 1450 NHFLQDDKFPIPNKLTLFSHLSKAKLFSKFDLKSGFWQLGIHPNERPKTGFCIPDRHFQW 1509
Query: 1445 NVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRN 1504
VMPFGLK APS FQ+ M +IF P +VYIDD+L+FS+TL+ H K LN FIS++K+
Sbjct: 1510 KVMPFGLKTAPSLFQKAMIKIFQPILFSALVYIDDILLFSETLEDHIKLLNQFISLVKKF 1569
Query: 1505 GLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNY 1564
G+ +S K+ L Q KI+FLG + GT P KFPD + Q+Q+FLG +NY
Sbjct: 1570 GVMLSAKKMILAQNKIQFLGMDFADGTFSPAGHISLELQKFPDTNLSVKQIQQFLGIVNY 1629
Query: 1565 VADFCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIET 1624
+ DF P+++ I PL D LKK PP N VKQ+K + + L++P+ + KI++T
Sbjct: 1630 IRDFIPEVTEHISPLSDMLKKKPPAWGKCQDNAVKQLKQLAQQVKSLHIPS-EGKKILQT 1688
Query: 1625 DASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQ 1684
DASD + +L ++ K +I F S + ++Q+Y + KE+LA+ I KF LI+
Sbjct: 1689 DASDQYWSAVLLEEHNGKRKICGFASGKFKVSEQHYHSTFKEILAVKNGIKKFNFFLIHT 1748
Query: 1685 KFLVCVDCKSAKEILQ 1700
FLV +D ++ ++++
Sbjct: 1749 NFLVEMDMRAFPKMIR 1764
Score = 85.1 bits (209), Expect = 2e-14
Identities = 143/596 (23%), Positives = 233/596 (38%), Gaps = 103/596 (17%)
Query: 1 SKNFNYLHIGSVQVGLKPLTRKSLDIAVLLCLRDVRHNQFHDSLLGTVETSLSNGPIFFN 60
S F ++H G+V++ L RK + L L D R+ ++ + LGT E +L+ G +F
Sbjct: 120 SHQFTHIHFGAVKIALTYHGRKGQPVVARLALLDTRYLEYQHANLGTAEITLNAGTVFIT 179
Query: 61 CFPDLTVSLEDKNILDVLFLNIKLHGLDMKEDSIPISLIYRVQYKVMNSIKSYFLRTNCE 120
FP+ T+SL D N+ L + +++ G + +DSI +L Y++ ++V N L
Sbjct: 180 LFPNFTMSLSDANLSTALKIQVQIQGAPLTKDSIQATLHYQIAWRVQNHAMDLTL----- 234
Query: 121 KSGETTLFLTDYDKAN-----VIVPKTIQWSDI--TLPEKWSLEKATPAQPAETPEFREI 173
GE LFL D N VP+ + D+ LP+ W +P E E+
Sbjct: 235 PGGEEALFL-KIDAGNGATQCTQVPRQLSKEDLIKILPDSWVTNYEKLKEPEEPLRSTEV 293
Query: 174 IQHQSGQVDLIFDRRNSFSIPRSIRTSSNDFQSAMSRRSFSMTRRNQPE--SSTRPSLSH 231
+ D+ + S S + M S + + PE S T PSL
Sbjct: 294 SMSKR------HDKSVAISFDHSHYKKLRNTHHFMGMISDDVIVLDDPETFSKTLPSLMQ 347
Query: 232 DGSTIH---LAGYVNTTPVKTTYD---KGDEDHQSTLSPTYSSLETPGETSDFIGEITTE 285
IH L G + K +D D D Q YS L + E DF T+
Sbjct: 348 THDWIHHFQLDGRA-VSWYKDPFDGHCPWDIDCQ-----CYSCLYSEDE-EDFEDGFPTK 400
Query: 286 FVIDKGSIKADFYSDRWTQQRKWFFAKYQGEKRKKIQEKFYGFLEKTQQNIFFFEWFQDY 345
+ KG + ++R Q+ Y+ ++ F G L + + +E+ Y
Sbjct: 401 Y---KGIPRPGSIAERKMQEEANLKKLYED------KDPFVGSLSRPGK----YEYLVRY 447
Query: 346 ANKNWKSYP-FQIDMVKWQHKD---------GKETL------------SNLPPKTPFLLK 383
+W P ++ W + + TL +N PP + F
Sbjct: 448 DAPSWAKDPHLTVEPTGWDSDEPILPNQPFITRNTLPKIYMFNPLNYENNFPPLSSFSKD 507
Query: 384 GAQSTP-------VLASPFKTRKEEGDVTGKDIKDLMEQSNYTNKYLQILGESQVQDSSS 436
GA TP VL S K D TG DL N+ + L ++++ +
Sbjct: 508 GADHTPKIPKRNVVLPSGAK------DPTG----DLEATVNWQTE--NALAQNRMLTTID 555
Query: 437 RKGKEKV-KIEPSSSNLDIPKGCQLEKPLFKPFQ-ISSRSRHSAQTLRAKKESDNELLQK 494
R KE V K++ S +G L + L + Q ++ R +L + +
Sbjct: 556 RTLKETVTKVDRVSDQSSKNQG--LIRVLEQQLQDLNKRICPPGTSLFHFFDQQKSEMAS 613
Query: 495 VVEQLQLLKQVIPETPEPPETPEPTPEA-------VSNPTITAPPPPKTRTSTRNT 543
+ EQ++LLK E P+ ET P+ ++ S+P + + PP T+ NT
Sbjct: 614 LKEQIRLLK----EQPQKNETDTPSYQSSYQPFHNFSSPYMPSNPPNSPFTNFANT 665
Score = 54.3 bits (129), Expect = 4e-05
Identities = 115/583 (19%), Positives = 216/583 (36%), Gaps = 129/583 (22%)
Query: 447 PSSSNLDIPKGCQLEKPLFKPFQISSRSRHSAQTLRAKKESDNELLQKVVEQLQLLK-QV 505
P+S + Q + LF + I +S ++ + E + + ++ + +L K +V
Sbjct: 655 PNSPFTNFANTPQPQPSLFSQYPIQPKSPNTFDLAKLVWEKKDAIAEEKRAKKKLQKDEV 714
Query: 506 IPETPEPPETPEPTPEA---------VSNPT------ITAPPPPKTRTSTRNTSTSLNNI 550
+T PPE+ P P++ +S+PT + P P TS+++ ++ ++
Sbjct: 715 KQKTSLPPESKRPDPQSSSHLGDQFMISDPTLPKVYELNEPSVPSEDTSSQSYISTEESV 774
Query: 551 EDGKVESDS-DIQSMKSDVQSIKSIPVNTVIHNSKNQWRKETKLYYPRATAPDLLLEESS 609
ED +DS + S +S S S N N++N + + R T P++
Sbjct: 775 ED----TDSFSVVSEESTQLSQLSSSSNDSPENNENTL---PQTFMVRPTEPEI------ 821
Query: 610 NFKSFSANNVYEWNIDGENEYGITKILQNMT---MVATAY-------VTANNCPES---- 655
+ E +DG E I + +T MV T + ++ PE
Sbjct: 822 --------SEVEDEVDGMTEEPIPERRPEITPPKMVGTGFHTFSLDDISITKWPERIQDF 873
Query: 656 ---------------LIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIME 700
LI+ A G L+ WW N + D+K++ LT
Sbjct: 874 HTWMLTKQLVEREPFLILSEFTARLSGTLREWW-NSVGPDDKNRFLT------------- 919
Query: 701 DGKFISDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTR 760
S I + +F GD S K++ + +KC S Y + R + R
Sbjct: 920 -----SQDFTWNIRILYSYFCGDQSQNKEELRRQIFEMKCLS------YDRKKIDRHFQR 968
Query: 761 ------EDSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRV 814
K+ F++ LP +++ ++ + T + I++
Sbjct: 969 MIKLFYHIGGDISLKQAFISSLPPILSERISALIKERG--------TSVTQMHVGDIRQT 1020
Query: 815 ALKICQDDKIQQQLTKEKSQNRRDLGTFCEQF-----GIQGCPKKPKP---RKQDPPPKQ 866
A + D +++ + + RDL C + G +GC P RK
Sbjct: 1021 AFYVLDDLCSKRKFFNQMKKMSRDLEKACTKSDLIIKGDKGCSGYCNPSRRRKYKRFKLP 1080
Query: 867 QWRKRSSRNDHRKPKPRSKPQSSQIPKNPPETRPSQGKDVTCYNCGKPGHISRYCRLKRR 926
+++R R ++ + K ++S+ + P +C+ CGK GH SR C ++
Sbjct: 1081 SFKERDGRQYRKRRRFFRKSKTSKAMRQKPR---------SCFTCGKIGHFSRNCPQNKK 1131
Query: 927 ----ISEL--HLEPEIEDKINNLLIQTSDEEESNPSDSEVSED 963
ISE+ + +IED + ++ + E E E+
Sbjct: 1132 SIKLISEIQKYTGIDIEDDLESVFSIEDEPSEDTLFSLEFYEE 1174
>gb|AAO67368.1| polyprotein 1 [Petunia vein clearing virus]
Length = 2180
Score = 313 bits (802), Expect = 3e-83
Identities = 172/436 (39%), Positives = 258/436 (58%), Gaps = 8/436 (1%)
Query: 1270 QLQQPSVKSRIEDILEN-----IQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKA 1324
QL + ++ED+++N S + W + ++LP++++ + TKA
Sbjct: 1332 QLTTSNELQQLEDVIKNQLCAESHVDFLSKCSHPLWLNQDFFIQLPFKRN--ENINPTKA 1389
Query: 1325 RPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPL 1384
MN E L KE ++L + LI S S W+C AFYVNK++E RG RLVINY+PL
Sbjct: 1390 SHSGMNPEHLQLALKECDELQQFDLIEPSDSQWACEAFYVNKRSEQVRGKLRLVINYQPL 1449
Query: 1385 NQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEW 1444
N L ++PIPNK L + AK+FSKFD+KSGFWQ+ + ++ KT F +P ++W
Sbjct: 1450 NHFLQDDKFPIPNKLTLFSHLSKAKLFSKFDLKSGFWQLGIHPNERPKTGFCIPDRHFQW 1509
Query: 1445 NVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRN 1504
VMPFGLK APS FQ+ M +IF P +VYIDD+L+FS+TL+ H K LN FIS++K+
Sbjct: 1510 KVMPFGLKTAPSLFQKAMIKIFQPILFSALVYIDDILLFSETLEDHIKLLNQFISLVKKF 1569
Query: 1505 GLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNY 1564
G+ +S K+ L Q KI+FLG + GT P KFPD + Q+Q+FLG +NY
Sbjct: 1570 GVMLSAKKMILAQNKIQFLGMDFADGTFSPAGHISLELQKFPDTNLSVKQIQQFLGIVNY 1629
Query: 1565 VADFCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIET 1624
+ DF P+++ I PL D LKK PP N VKQ+K + + L++P+ + KI++T
Sbjct: 1630 IRDFIPEVTEHISPLSDMLKKKPPAWGKCQDNAVKQLKQLAQQVKSLHIPS-EGKKILQT 1688
Query: 1625 DASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQ 1684
DASD + +L ++ K +I F S + ++Q+Y + KE+LA+ I KF LI+
Sbjct: 1689 DASDQYWSAVLLEEHNGKRKICGFASGKFKVSEQHYHSTFKEILAVKNGIKKFNFFLIHT 1748
Query: 1685 KFLVCVDCKSAKEILQ 1700
FLV +D ++ ++++
Sbjct: 1749 NFLVEMDMRAFPKMIR 1764
Score = 85.1 bits (209), Expect = 2e-14
Identities = 143/596 (23%), Positives = 233/596 (38%), Gaps = 103/596 (17%)
Query: 1 SKNFNYLHIGSVQVGLKPLTRKSLDIAVLLCLRDVRHNQFHDSLLGTVETSLSNGPIFFN 60
S F ++H G+V++ L RK + L L D R+ ++ + LGT E +L+ G +F
Sbjct: 120 SHQFTHIHFGAVKIALTYHGRKGQPVVARLALLDTRYLEYQHANLGTAEITLNAGTVFIT 179
Query: 61 CFPDLTVSLEDKNILDVLFLNIKLHGLDMKEDSIPISLIYRVQYKVMNSIKSYFLRTNCE 120
FP+ T+SL D N+ L + +++ G + +DSI +L Y++ ++V N L
Sbjct: 180 LFPNFTMSLSDANLSTALKIQVQIQGAPLTKDSIQATLHYQIAWRVQNHAMDLTL----- 234
Query: 121 KSGETTLFLTDYDKAN-----VIVPKTIQWSDI--TLPEKWSLEKATPAQPAETPEFREI 173
GE LFL D N VP+ + D+ LP+ W +P E E+
Sbjct: 235 PGGEEALFL-KIDAGNGATQCTQVPRQLSKEDLIKILPDSWVTNYEKLKEPEEPLRSTEV 293
Query: 174 IQHQSGQVDLIFDRRNSFSIPRSIRTSSNDFQSAMSRRSFSMTRRNQPE--SSTRPSLSH 231
+ D+ + S S + M S + + PE S T PSL
Sbjct: 294 SMSKR------HDKSVAISFDHSHYKKLRNTHHFMGMISDDVIVLDDPETFSKTLPSLMQ 347
Query: 232 DGSTIH---LAGYVNTTPVKTTYD---KGDEDHQSTLSPTYSSLETPGETSDFIGEITTE 285
IH L G + K +D D D Q YS L + E DF T+
Sbjct: 348 THDWIHHFQLDGRA-VSWYKDPFDGHCPWDIDCQ-----CYSCLYSEDE-EDFEDGFPTK 400
Query: 286 FVIDKGSIKADFYSDRWTQQRKWFFAKYQGEKRKKIQEKFYGFLEKTQQNIFFFEWFQDY 345
+ KG + ++R Q+ Y+ ++ F G L + + +E+ Y
Sbjct: 401 Y---KGIPRPGSIAERKMQEEANLKKLYED------KDPFVGSLSRPGK----YEYLVRY 447
Query: 346 ANKNWKSYP-FQIDMVKWQHKD---------GKETL------------SNLPPKTPFLLK 383
+W P ++ W + + TL +N PP + F
Sbjct: 448 DAPSWAKDPHLTVEPTGWDSDEPILPNQPFITRNTLPKIYMFNPLNYENNFPPLSSFSKD 507
Query: 384 GAQSTP-------VLASPFKTRKEEGDVTGKDIKDLMEQSNYTNKYLQILGESQVQDSSS 436
GA TP VL S K D TG DL N+ + L ++++ +
Sbjct: 508 GADHTPKIPKRNVVLPSGAK------DPTG----DLEATVNWQTE--NALAQNRMLTTID 555
Query: 437 RKGKEKV-KIEPSSSNLDIPKGCQLEKPLFKPFQ-ISSRSRHSAQTLRAKKESDNELLQK 494
R KE V K++ S +G L + L + Q ++ R +L + +
Sbjct: 556 RTLKETVTKVDRVSDQSSKNQG--LIRVLEQQLQDLNKRICPPGTSLFHFFDQQKSEMAS 613
Query: 495 VVEQLQLLKQVIPETPEPPETPEPTPEA-------VSNPTITAPPPPKTRTSTRNT 543
+ EQ++LLK E P+ ET P+ ++ S+P + + PP T+ NT
Sbjct: 614 LKEQIRLLK----EQPQKNETDTPSYQSSYQPFHNFSSPYMPSNPPNSPFTNFANT 665
Score = 55.5 bits (132), Expect = 2e-05
Identities = 114/583 (19%), Positives = 216/583 (36%), Gaps = 129/583 (22%)
Query: 447 PSSSNLDIPKGCQLEKPLFKPFQISSRSRHSAQTLRAKKESDNELLQKVVEQLQLLK-QV 505
P+S + Q + LF + I +S ++ + E + + ++ + +L K +V
Sbjct: 655 PNSPFTNFANTPQPQPSLFSQYPIQPKSPNTFDLAKLVWEKKDAIAEEKRAKKKLQKDEV 714
Query: 506 IPETPEPPETPEPTPEA---------VSNPT------ITAPPPPKTRTSTRNTSTSLNNI 550
+T PPE+ P P++ +S+PT + P P TS+++ ++ ++
Sbjct: 715 KQKTSLPPESKRPDPQSSSHLGDQFMISDPTLPKVYELNEPSVPSEDTSSQSYISTEESV 774
Query: 551 EDGKVESDS-DIQSMKSDVQSIKSIPVNTVIHNSKNQWRKETKLYYPRATAPDLLLEESS 609
ED +DS + S +S S S N N++N + + R T P++
Sbjct: 775 ED----TDSFSVVSEESTQLSQLSSSSNDSPENNENTL---PQTFMVRPTEPEI------ 821
Query: 610 NFKSFSANNVYEWNIDGENEYGITKILQNMT---MVATAY-------VTANNCPES---- 655
+ E +DG E I + +T MV T + ++ PE
Sbjct: 822 --------SEVEDEVDGMTEEPIPERRPEITPPKMVGTGFHTFSLDDISITKWPERIQDF 873
Query: 656 ---------------LIVEVLVAGFCGQLKGWWDNYLTQDEKDQILTAVKTDEEGNPIME 700
LI+ A G L+ WW N + D+K++ LT
Sbjct: 874 HTWMLTKQLVEREPFLILSEFTARLSGTLREWW-NSVGPDDKNRFLT------------- 919
Query: 701 DGKFISDAVNSLIFTIAQHFVGDPSLIKDKSGDLLSNLKCKSLGDFRWYKDTFLTRVYTR 760
S I + +F GD S K++ + +KC S Y + R + R
Sbjct: 920 -----SQDFTWNIRILYSYFCGDQSQNKEELRRQIFEMKCLS------YDRKKIDRHFQR 968
Query: 761 ------EDSQQAFWKEKFLAGLPKSFGDKVREKLRSQNPGGEIPYHTLSYGQLIAIIQRV 814
K+ F++ LP +++ ++ + G + + I++
Sbjct: 969 MIKLFYHIGGDISLKQAFISSLPPILSERISALIKERGTSGTQMH--------VGDIRQT 1020
Query: 815 ALKICQDDKIQQQLTKEKSQNRRDLGTFCEQF-----GIQGCPKKPKP---RKQDPPPKQ 866
+ D +++ + + RDL C + G +GC P RK
Sbjct: 1021 GFYVLDDLCSKRKFFNQMKKMSRDLEKACTKSDLIIKGDKGCSGYCNPSRRRKYKRFKLP 1080
Query: 867 QWRKRSSRNDHRKPKPRSKPQSSQIPKNPPETRPSQGKDVTCYNCGKPGHISRYCRLKRR 926
+++R R ++ + K ++S+ + P +C+ CGK GH SR C ++
Sbjct: 1081 SFKERDGRQYRKRRRFFRKSKTSKAMRQKPR---------SCFTCGKIGHFSRNCPQNKK 1131
Query: 927 ----ISEL--HLEPEIEDKINNLLIQTSDEEESNPSDSEVSED 963
ISE+ + +IED + ++ + E E E+
Sbjct: 1132 SIKLISEIQKYTGIDIEDDLESVFSIEDEPSEDTLFSLEFYEE 1174
>emb|CAH68829.1| polyprotein [Carnation etched ring virus]
Length = 656
Score = 302 bits (773), Expect = 7e-80
Identities = 201/579 (34%), Positives = 302/579 (51%), Gaps = 18/579 (3%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEK---LSAAEGSRLKIKYKIPSAIIKNGSL 1187
LE+ DTG+ S+ +IP Y++ T EK + A G +++ I+ G
Sbjct: 26 LELHCYVDTGSSLCMASKYVIPEEYWQ-TAEKPLNIKIANGKIIQLTKVCSKLPIRLGGE 84
Query: 1188 EIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDENGITVQHLGQP--ILFKFSEPPIDKTL 1245
P L + +++G F + P+ + I HL + I+ K ++
Sbjct: 85 RFLIPTLFQQESGIDLLLGNNFCQLYSPFIQYTDRIYF-HLNKQSVIIGKITKAYQYGVK 143
Query: 1246 NVISYKEKQINFLKEEISYRTIEEQLQQPSVKSRIEDILENIQSSICSDLPNAFWERKSH 1305
+ +K+ + E T + L S IE++ IQ S S + ER S
Sbjct: 144 GFLESMKKKYKVNRPEPINITSNQHLFLEEGGSNIEELCYEIQISKFSAIEEML-ERVSS 202
Query: 1306 MVELPYEK---------DFSDKQISTKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSP 1356
+ EK + D + + +P+ + K+I + L+ K+I+ SKSP
Sbjct: 203 ENPIDPEKSKQWMTATIELIDPKTIVEVKPMSYSPSDREEFDKQIKEPLDLKVIKPSKSP 262
Query: 1357 WSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDM 1416
AF V +AE RG R+V+NYK +N+A + +PNK +LLA K++S FD
Sbjct: 263 HMSPAFLVENEAERRRGKKRMVVNYKAMNKATKGDAHNLPNKDELLALVRGKKIYSSFDC 322
Query: 1417 KSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVY 1476
KSGFWQ+ L ++ + TAFT P G Y+WNV+PFGLK APS FQR M FN +SK VY
Sbjct: 323 KSGFWQVLLDKESQLLTAFTCPQGHYQWNVVPFGLKQAPSIFQRHMQTAFNQHSKYCCVY 382
Query: 1477 IDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPIN 1536
+DD+L+FS T ++H+ H+ + ++ G+ +SK K LF+ KI FLG I QGT P N
Sbjct: 383 VDDILVFSNTEEEHYIHVLNILRRCEKLGIILSKKKAQLFKEKINFLGLEIDQGTHCPQN 442
Query: 1537 RAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPP-P*SDIHT 1595
+E KFPD+I DK QLQR LG L Y +D+ P+L++I KPL +LK+D +D +
Sbjct: 443 HILEHIHKFPDRIEDKKQLQRSLGILTYASDYIPKLASIRKPLQSKLKEDSTWTWNDTDS 502
Query: 1596 NVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNP 1655
+ +IK +K+ P LY P P +IETDAS+ +GGILK E I + S +
Sbjct: 503 QYMAKIKKNLKSFPKLYHPEPNDKLVIETDASEEFWGGILKAIHNSHEYICRYASGSFKA 562
Query: 1656 AQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVDCKS 1694
A++NY + +KE+LA+ I KF L +FL+ D K+
Sbjct: 563 AERNYHSNEKELLAVFRVIYKFSIYLTPCRFLIRSDNKN 601
>sp|Q02964|POL_CAMVE Enzymatic polyprotein [Contains: Aspartic protease ; Endonuclease;
Reverse transcriptase ] gi|293185|gb|AAA62375.1| reverse
transcriptase
Length = 679
Score = 295 bits (756), Expect = 7e-78
Identities = 205/600 (34%), Positives = 311/600 (51%), Gaps = 55/600 (9%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSA--AEGSRLKIK-------------- 1174
+E+ DTGA S+ +IP ++ + A+GS + I
Sbjct: 38 IELHCFVDTGASLCIASKFVIPEEHWVNAERPIMVKIADGSSITISKVCKDIDLIIAREI 97
Query: 1175 YKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIK----------KLFPYNTDENGIT 1224
+KIP+ + ++ F++ N Q + PFI+ K +P + +
Sbjct: 98 FKIPTVYQQESGID----FIIGNNFCQ---LYEPFIQFTDRVIFTKNKSYPVHIAKLTRA 150
Query: 1225 VQHLGQPIL------FKFSEP-PIDKTLNVISYKEKQINFLKEEISYRTIEEQLQQPSVK 1277
V+ + L K +P P++ + N I K+I L E R EE+L +
Sbjct: 151 VRVGTEGFLESMKKRSKTQQPEPVNISTNKIENPLKEIAILSE--GRRLSEEKLFITQQR 208
Query: 1278 -SRIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHF 1336
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 209 MQKIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREE 261
Query: 1337 CQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIP 1396
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y +P
Sbjct: 262 FDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIGDAYNLP 321
Query: 1397 NKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPS 1456
NK +LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS
Sbjct: 322 NKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPS 381
Query: 1457 EFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLF 1516
FQR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF
Sbjct: 382 IFQRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLF 441
Query: 1517 QTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTII 1576
+ KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I
Sbjct: 442 KKKINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIR 501
Query: 1577 KPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGIL 1635
KPL +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+L
Sbjct: 502 KPLQAKLKENVPWKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGML 561
Query: 1636 K----QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
K + + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 562 KAIKINEGTNTELICRYASGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 621
>gb|AAA21736.1| reverse transcriptase
Length = 680
Score = 294 bits (753), Expect = 2e-77
Identities = 204/599 (34%), Positives = 308/599 (51%), Gaps = 53/599 (8%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSA--AEGSRLKIK-------------- 1174
+E+ DTGA S+ +IP ++ + A+GS + I
Sbjct: 39 IELHCFVDTGASLCIASKFVIPEEHWVNAERPIMVKIADGSSITISKVCKDIDLIIAGEI 98
Query: 1175 YKIPSA---------IIKNGSLEIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDENGITV 1225
+KIP+ II N ++ PF+ + ++I K +P + V
Sbjct: 99 FKIPTVYQQESGIDFIIGNNFCQLYEPFI---QFTDRVIFAK---NKSYPVHIANVTRAV 152
Query: 1226 QHLGQPIL------FKFSEP-PIDKTLNVISYKEKQINFLKEEISYRTIEEQLQQPSVK- 1277
+ + L K +P P++ + N I ++I L E R EE+L +
Sbjct: 153 RVGTEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAILSE--GRRLSEEKLFITQQRM 210
Query: 1278 SRIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFC 1337
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 211 QKIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPNKAIKVKPMKYSPMDREEF 263
Query: 1338 QKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPN 1397
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y +PN
Sbjct: 264 DKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNLPN 323
Query: 1398 KKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSE 1457
K +LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS
Sbjct: 324 KDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPSI 383
Query: 1458 FQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQ 1517
FQR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF+
Sbjct: 384 FQRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFK 443
Query: 1518 TKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIK 1577
KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I K
Sbjct: 444 KKINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRK 503
Query: 1578 PLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILK 1636
PL +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+LK
Sbjct: 504 PLQAKLKENVPWKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLK 563
Query: 1637 ----QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
+ + E I + S + A++NY + KE LA++ +I KF L FL D
Sbjct: 564 AIKINEGINTELICRYASGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPAHFLTRTD 622
>emb|CAA55974.1| unnamed protein product [Cauliflower mosaic virus]
Length = 680
Score = 293 bits (751), Expect = 3e-77
Identities = 204/600 (34%), Positives = 311/600 (51%), Gaps = 55/600 (9%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEKL--SAAEGSRLKIK-------------- 1174
+E+ DTGA S+ +IP ++ + A+GS + I
Sbjct: 39 IELHCFVDTGASLCIASKFVIPEEHWVNAERPILVKIADGSSITISKVCKDIDLIIAGEI 98
Query: 1175 YKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIK----------KLFPYNTDENGIT 1224
+KIP+ + ++ F++ N Q + PFI+ K +P + +
Sbjct: 99 FKIPTVYQQESGID----FIIGNNFCQ---LYEPFIQFTDRVIFTKNKSYPVHIAKLTRA 151
Query: 1225 VQHLGQPIL------FKFSEP-PIDKTLNVISYKEKQINFLKEEISYRTIEEQLQQPSVK 1277
V+ + L K +P P++ + N I ++I L E R EE+L +
Sbjct: 152 VRVGTEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAILSE--GRRLSEEKLFITQQR 209
Query: 1278 -SRIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHF 1336
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 210 MQKIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREE 262
Query: 1337 CQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIP 1396
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y +P
Sbjct: 263 FDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIGDAYNLP 322
Query: 1397 NKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPS 1456
NK +LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS
Sbjct: 323 NKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPS 382
Query: 1457 EFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLF 1516
FQR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF
Sbjct: 383 IFQRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLF 442
Query: 1517 QTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTII 1576
+ KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I
Sbjct: 443 KKKINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIR 502
Query: 1577 KPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGIL 1635
KPL +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+L
Sbjct: 503 KPLQAKLKENVPWRWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGML 562
Query: 1636 K----QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
K + + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 563 KAIKINEGTNTELICRYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 622
>sp|P03555|POL_CAMVC Enzymatic polyprotein [Contains: Aspartic protease ; Endonuclease;
Reverse transcriptase ]
Length = 679
Score = 293 bits (751), Expect = 3e-77
Identities = 204/600 (34%), Positives = 311/600 (51%), Gaps = 55/600 (9%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSA--AEGSRLKIK-------------- 1174
+E+ DTGA S+ +IP ++ + A+GS + I
Sbjct: 38 IELHCFVDTGASLCIASKFVIPEEHWVNAERPIMVKIADGSSITISKVCKDIDLIIAGEI 97
Query: 1175 YKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIK----------KLFPYNTDENGIT 1224
+KIP+ + ++ F++ N Q + PFI+ K +P + +
Sbjct: 98 FKIPTVYQQESGID----FIIGNNFCQ---LYEPFIQFTDRVIFTKNKSYPVHITKLTRA 150
Query: 1225 VQHLGQPIL------FKFSEP-PIDKTLNVISYKEKQINFLKEEISYRTIEEQLQQPSVK 1277
V+ + L K +P P++ + N I ++I L E R EE+L +
Sbjct: 151 VRVGIEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAILSE--GRRLSEEKLFITQQR 208
Query: 1278 -SRIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHF 1336
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 209 MQKIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREE 261
Query: 1337 CQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIP 1396
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y +P
Sbjct: 262 FDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIGDAYNLP 321
Query: 1397 NKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPS 1456
NK +LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS
Sbjct: 322 NKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPS 381
Query: 1457 EFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLF 1516
FQR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF
Sbjct: 382 IFQRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLF 441
Query: 1517 QTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTII 1576
+ KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I
Sbjct: 442 KKKINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIR 501
Query: 1577 KPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGIL 1635
KPL +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+L
Sbjct: 502 KPLQAKLKENVPWKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGML 561
Query: 1636 K----QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
K + + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 562 KAIKINEGTNTELICRYASGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 621
>emb|CAA23460.1| unnamed protein product [Cauliflower mosaic virus]
gi|130592|sp|P03554|POL_CAMVS Enzymatic polyprotein
[Contains: Aspartic protease ; Endonuclease; Reverse
transcriptase ] gi|9626943|ref|NP_056728.1| reverse
transcriptase [Cauliflower mosaic virus]
Length = 679
Score = 293 bits (749), Expect = 5e-77
Identities = 203/598 (33%), Positives = 307/598 (50%), Gaps = 51/598 (8%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSA--AEGSRLKIKYKIPSAIIKNGSLE 1188
+E+ DTGA S+ +IP ++ + A+GS + I S + K+ L
Sbjct: 38 IELHCFVDTGASLCIASKFVIPEEHWVNAERPIMVKIADGSSITI-----SKVCKDIDLI 92
Query: 1189 IETPFLLVRNLSQK-----IIIGTPFIKKLFPY-NTDENGITVQHLGQPILF-------- 1234
I + + Q+ IIG F + P+ + I ++ P+
Sbjct: 93 IAGEIFRIPTVYQQESGIDFIIGNNFCQLYEPFIQFTDRVIFTKNKSYPVHIAKLTRAVR 152
Query: 1235 --------------KFSEP-PIDKTLNVISYKEKQINFLKEEISYRTIEEQLQQPSVK-S 1278
K +P P++ + N I ++I L E R EE+L +
Sbjct: 153 VGTEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAILSE--GRRLSEEKLFITQQRMQ 210
Query: 1279 RIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQ 1338
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 211 KIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREEFD 263
Query: 1339 KEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNK 1398
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y +PNK
Sbjct: 264 KQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNLPNK 323
Query: 1399 KDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEF 1458
+LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS F
Sbjct: 324 DELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPSIF 383
Query: 1459 QRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQT 1518
QR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF+
Sbjct: 384 QRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKK 443
Query: 1519 KIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKP 1578
KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I KP
Sbjct: 444 KINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKP 503
Query: 1579 LHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILK- 1636
L +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+LK
Sbjct: 504 LQAKLKENVPWRWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKA 563
Query: 1637 ---QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
+ + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 564 IKINEGTNTELICRYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 621
>gb|AAW56089.1| polyprotein [Horseradish latent virus]
Length = 679
Score = 293 bits (749), Expect = 5e-77
Identities = 201/597 (33%), Positives = 296/597 (48%), Gaps = 53/597 (8%)
Query: 1130 VLEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSAAEGSRLKIKY----KIPSAIIKNG 1185
++E+ DTGA S+ +IP +E K++ + I + II
Sbjct: 45 IVELDCFVDTGASLCIASKHVIPEERWEAAPRKINVKIANESTITLDKICRNLDIIIAGE 104
Query: 1186 SLEIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDENGITVQHLGQPILFKFSEPPIDKTL 1245
I+ F + IIG F ++ P+ +Q G + + + T
Sbjct: 105 RFHIDVVFQQESGID--FIIGNNFCQEYSPF--------IQFTGHIVFTMKDQYQVPITK 154
Query: 1246 NVISYKEKQINFLKEEISYRTIEEQLQQPSVKS------------------------RIE 1281
++K FL E + R+ +Q + ++ + +IE
Sbjct: 155 LRRAFKRGIPGFL-ESMKKRSTAQQPEPLNISTNKTVSLSRGRRFGEKLRITQERMVKIE 213
Query: 1282 DILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQKEI 1341
++LE + CS+ P + K M SD K +P++ + +K+I
Sbjct: 214 ELLEKV----CSENPLDPEKSKGWMQA---SIKLSDPTKVIKVKPMKYSPMDREEFEKQI 266
Query: 1342 NDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDL 1401
+LL+ K+IR SKSP AF VN +AE RG R+V+NYK +N A Y +PNK +L
Sbjct: 267 QELLDLKVIRPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNDATVGDAYNLPNKDEL 326
Query: 1402 LARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRI 1461
L K+FS FD KSGFWQ++L E+ K TAFT P G YEWNV+PFG+K APS FQR
Sbjct: 327 LTLIRGKKIFSSFDCKSGFWQVRLDEESKSLTAFTCPQGHYEWNVVPFGMKQAPSIFQRH 386
Query: 1462 MNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIR 1521
M+E F + K +Y+DD+L+FS H H+ + ++G+ +SK K LF+ +I
Sbjct: 387 MDEAFKVFRKFCCIYVDDILVFSDNEQNHQLHVAMILQKCYQHGIILSKKKAQLFKERIN 446
Query: 1522 FLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHD 1581
FLG I QGT P + +E KFPD I K QLQRFLG L Y +D+ P+L+ I KPL
Sbjct: 447 FLGLEIDQGTHRPQSHILEHIQKFPDIIESKLQLQRFLGVLTYASDYIPKLAQIRKPLQA 506
Query: 1582 RLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVF 1640
+LK++ + T +K++K + P L+ P P+ IIETDASD +GGILK
Sbjct: 507 KLKENVQWRWTPEDTLYMKKVKKNLNGFPPLHHPLPEEKLIIETDASDNYWGGILKAIHI 566
Query: 1641 DK------EQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
D E + + S + PA+QNY + KE LA++ +I KF L +FLV D
Sbjct: 567 DLSTNESIELVCRYASGSFKPAEQNYHSNDKETLAVIRTIQKFSIYLTPVRFLVRTD 623
>ref|NP_619548.1| Enzymatic polyprotein [Contains: Aspartic protease; Endonuclease;
Reverse transcriptase] [Figwort mosaic virus]
gi|58813|emb|CAA29527.1| unnamed protein product [Figwort
mosaic virus] gi|130600|sp|P09523|POL_FMVD Enzymatic
polyprotein [Contains: Aspartic protease ; Endonuclease;
Reverse transcriptase ]
Length = 666
Score = 291 bits (746), Expect = 1e-76
Identities = 193/575 (33%), Positives = 292/575 (50%), Gaps = 32/575 (5%)
Query: 1138 DTGADSSCISEGLIPTRYFEKTTEKLSA--AEGSRLKIKYKIPSAIIKNGSLEIETPFLL 1195
DTGA S +IP +E + + + A +KI + +K E P +
Sbjct: 54 DTGASLCIASRYIIPEELWENSPKDIQVKIANQELIKITKVCKNLKVKFAGKSFEIPTVY 113
Query: 1196 VRNLSQKIIIGTPFIKKLFPYNTDENGITVQHLGQPIL-------FKFSEPPIDKTLNVI 1248
+ +IG F + P+ E+ I + +L F S P + +
Sbjct: 114 QQETGIDFLIGNNFCRLYNPFIQWEDRIAFHLKNEMVLIKKVTKAFSVSNPSFLENMKKD 173
Query: 1249 SYKEKQI-------NFLKEEISYRTIEEQLQQPSVKSRIEDILENIQSSICSDLPNAFWE 1301
S K +QI N + E Y I E+ Q+ +E + +CS+ P +
Sbjct: 174 S-KTEQIPGTNISKNIINPEERYFLITEKYQK----------IEQLLDKVCSENPIDPIK 222
Query: 1302 RKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAA 1361
K M D + +P+ + + K+I +LL+ LI SKS A
Sbjct: 223 SKQWMKA---SIKLIDPLKVIRVKPMSYSPQDREGFAKQIKELLDLGLIIPSKSQHMSPA 279
Query: 1362 FYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFW 1421
F V +AE RG R+V+NYK +NQA + +PN ++LL +FS FD KSGFW
Sbjct: 280 FLVENEAERRRGKKRMVVNYKAINQATIGDSHNLPNMQELLTLLRGKSIFSSFDCKSGFW 339
Query: 1422 QIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKLTIVYIDDVL 1481
Q+ L E+ + TAFT P G ++W V+PFGLK APS FQR M N K +VY+DD++
Sbjct: 340 QVVLDEESQKLTAFTCPQGHFQWKVVPFGLKQAPSIFQRHMQTALNGADKFCMVYVDDII 399
Query: 1482 IFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEF 1541
+FS + H+ H+ + ++++ G+ +SK K +LF+ KI FLG I +GT P N +E
Sbjct: 400 VFSNSELDHYNHVYAVLKIVEKYGIILSKKKANLFKEKINFLGLEIDKGTHCPQNHILEN 459
Query: 1542 TDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQ 1600
KFPD++ DK LQRFLG L Y + P+L+ I KPL +LKKD + ++ VK+
Sbjct: 460 IHKFPDRLEDKKHLQRFLGVLTYAETYIPKLAEIRKPLQVKLKKDVTWNWTQSDSDYVKK 519
Query: 1601 IKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIA-FTSKHWNPAQQN 1659
IK + + P LYLP P+ IIETDASD +GG+LK + D ++I ++S + A++N
Sbjct: 520 IKKNLGSFPKLYLPKPEDHLIIETDASDSFWGGVLKARALDGVELICRYSSGSFKQAEKN 579
Query: 1660 YSTVKKEVLAIVLSISKFQSDLINQKFLVCVDCKS 1694
Y + KE+LA+ I+KF + L +F V D K+
Sbjct: 580 YHSNDKELLAVKQVITKFSAYLTPVRFTVRTDNKN 614
>emb|CAA28360.1| unnamed protein product [Carnation etched ring virus]
gi|19919894|ref|NP_612577.1| Enzymatic polyprotein
[Contains: Aspartic protease; Endonuclease; Reverse
transcriptase] [Carnation etched ring virus]
gi|130593|sp|P05400|POL_CERV Enzymatic polyprotein
[Contains: Aspartic protease ; Endonuclease; Reverse
transcriptase ] gi|225356|prf||1301227E ORF 5
Length = 659
Score = 290 bits (743), Expect = 2e-76
Identities = 199/591 (33%), Positives = 305/591 (50%), Gaps = 40/591 (6%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEK---LSAAEGSRLKIKYKIPSAIIKNGSL 1187
L++ DTG+ S+ +IP Y++ T EK + A G +++ I+ G
Sbjct: 27 LDLHCYVDTGSSLCMASKYVIPEEYWQ-TAEKPLNIKIANGKIIQLTKVCSKLPIRLGGE 85
Query: 1188 EIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDENGITVQHLGQPILF------------- 1234
P L + +++G F + P+ + I Q ++
Sbjct: 86 RFLIPTLFQQESGIDLLLGNNFCQLYSPFIQYTDRIYFHLNKQSVIIGKITKAYQYGVKG 145
Query: 1235 ------KFSEPPIDKTLNVISYKEKQINFLKEEISYRTIEEQLQQPSVK--SRIEDILEN 1286
K S+ + +N+ S Q FL+E ++ ++E L + + S IE++LE
Sbjct: 146 FLESMKKKSKVNRPEPINITS---NQHLFLEEGGNH--VDEMLYEIQISKFSAIEEMLER 200
Query: 1287 IQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQKEINDLLE 1346
+ S D P + + +EL D + K +P+ + ++I +LLE
Sbjct: 201 VSSENPID-PEKSKQWMTATIEL------IDPKTVVKVKPMSYSPSDREEFDRQIKELLE 253
Query: 1347 KKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQH 1406
K+I+ SKS AF V +AE RG R+V+NYK +N+A + +PNK +LL
Sbjct: 254 LKVIKPSKSTHMSPAFLVENEAERRRGKKRMVVNYKAMNKATKGDAHNLPNKDELLTLVR 313
Query: 1407 DAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQR-IMNEI 1465
K++S FD KSG WQ+ L ++ + TAFT P G Y+WNV+PFGLK APS F + N
Sbjct: 314 GKKIYSSFDCKSGLWQVLLDKESQLLTAFTCPQGHYQWNVVPFGLKQAPSIFPKTYANSH 373
Query: 1466 FNPYSKLTIVYIDDVLIFSQT-LDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLG 1524
N YSK VY+DD+L+FS T +H+ H+ + ++ G+ +SK K LF+ KI FLG
Sbjct: 374 SNQYSKYCCVYVDDILVFSNTGRKEHYIHVLNILRRCEKLGIILSKKKAQLFKEKINFLG 433
Query: 1525 HNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLK 1584
I QGT P N +E KFPD+I DK QLQRFLG L Y +D+ P+L++I KPL +LK
Sbjct: 434 LEIDQGTHCPQNHILEHIHKFPDRIEDKKQLQRFLGILTYASDYIPKLASIRKPLQSKLK 493
Query: 1585 KDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKE 1643
+D +D + + +IK +K+ P LY P P +IETDAS+ +GGILK E
Sbjct: 494 EDSTWTWNDTDSQYMAKIKKNLKSFPKLYHPEPNDKLVIETDASEEFWGGILKAIHNSHE 553
Query: 1644 QIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVDCKS 1694
I + S + A++NY + +KE+LA++ I KF L +FL+ D K+
Sbjct: 554 YICRYASGSFKAAERNYHSNEKELLAVIRVIKKFSIYLTPSRFLIRTDNKN 604
>gb|AAD37341.1| reverse transcriptase [Cauliflower mosaic virus]
Length = 674
Score = 289 bits (739), Expect = 7e-76
Identities = 167/418 (39%), Positives = 243/418 (57%), Gaps = 12/418 (2%)
Query: 1279 RIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQ 1338
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 206 KIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREEFD 258
Query: 1339 KEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNK 1398
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y +PNK
Sbjct: 259 KQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNLPNK 318
Query: 1399 KDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEF 1458
+LL K+FS FD KSGFWQ+ ++ + TAFT P G YEWNV+PFGLK APS F
Sbjct: 319 DELLTLIRGKKIFSSFDCKSGFWQVLPDQESRPLTAFTCPQGHYEWNVVPFGLKQAPSIF 378
Query: 1459 QRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQT 1518
QR M+E F + K VY+DD+L+FS T + H H+ + ++G+ +SK K LF+
Sbjct: 379 QRHMDEAFRVFRKFCCVYVDDILVFSNTEEDHLLHVAMILQKCNQHGIILSKKKAQLFKK 438
Query: 1519 KIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKP 1578
KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I KP
Sbjct: 439 KINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKP 498
Query: 1579 LHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILK- 1636
L +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+LK
Sbjct: 499 LQAKLKENVPWKWTKEDTLYMQKVKKNLQGFPPLHRPLPEEKLIIETDASDDYWGGMLKA 558
Query: 1637 ---QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
+ + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 559 IQINEGTNTELICRYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 616
>sp|Q00962|POL_CAMVN Enzymatic polyprotein [Contains: Aspartic protease ; Endonuclease;
Reverse transcriptase ] gi|331571|gb|AAA46358.1| reverse
transcriptase gi|445598|prf||1909346E reverse
transcriptase
Length = 680
Score = 288 bits (736), Expect = 1e-75
Identities = 203/602 (33%), Positives = 311/602 (50%), Gaps = 59/602 (9%)
Query: 1131 LEIPALFDTGADSSCISEGLIPTRYFEKTTEKLSA--AEGSRLKIK-------------- 1174
+E+ DTGA S+ +IP ++ + A+GS + I
Sbjct: 39 IELHCFVDTGASLCIASKFVIPEEHWVNAERPIMVKIADGSSITISKVCKDIDLIIVGVI 98
Query: 1175 YKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIK----------KLFPYNTDENGIT 1224
+KIP+ + ++ F++ N Q + PFI+ K +P + +
Sbjct: 99 FKIPTVYQQESGID----FIIGNNFCQ---LYEPFIQFTDRVIFTKNKSYPVHIAKLTRA 151
Query: 1225 VQHLGQPIL------FKFSEP-PIDKTLNVISYKEKQINFLKEEISYRTIEEQL---QQP 1274
V+ + L K +P P++ + N I ++I L E R EE+L QQ
Sbjct: 152 VRVGTEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAILSE--GRRLSEEKLFITQQR 209
Query: 1275 SVKSRIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELL 1334
K+ E++LE + S D PN + ++L SD + K +P++ +
Sbjct: 210 MQKT--EELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDR 260
Query: 1335 HFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYP 1394
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y
Sbjct: 261 EEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAENGRGNKRMVVNYKAMNKATVGDAYN 320
Query: 1395 IPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNA 1454
+PNK +LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK A
Sbjct: 321 LPNKDELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQA 380
Query: 1455 PSEFQRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVS 1514
PS FQR M+E F + K VY+DD+++FS + H H+ + ++G+ +SK K
Sbjct: 381 PSIFQRHMDEAFRVFRKFCCVYVDDIVVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQ 440
Query: 1515 LFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLST 1574
LF+ KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P L+
Sbjct: 441 LFKKKINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPNLAQ 500
Query: 1575 IIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGG 1633
+ +PL +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG
Sbjct: 501 MRQPLQAKLKENVPWKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGG 560
Query: 1634 ILK----QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVC 1689
+LK + + E I + S + A++NY + KE LA++ +I KF L FL+
Sbjct: 561 MLKAIKINEGTNTELICRYRSGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIR 620
Query: 1690 VD 1691
D
Sbjct: 621 TD 622
>sp|P03556|POL_CAMVD Enzymatic polyprotein [Contains: Aspartic protease ; Endonuclease;
Reverse transcriptase ]
Length = 674
Score = 287 bits (735), Expect = 2e-75
Identities = 167/418 (39%), Positives = 242/418 (56%), Gaps = 12/418 (2%)
Query: 1279 RIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQ 1338
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 206 KIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREEFD 258
Query: 1339 KEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNK 1398
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y PNK
Sbjct: 259 KQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNPPNK 318
Query: 1399 KDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEF 1458
+LL K+FS FD KSGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS F
Sbjct: 319 DELLTLIRGKKIFSSFDCKSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPSIF 378
Query: 1459 QRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQT 1518
QR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF+
Sbjct: 379 QRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKK 438
Query: 1519 KIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKP 1578
KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I KP
Sbjct: 439 KINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKP 498
Query: 1579 LHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILK- 1636
L +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+LK
Sbjct: 499 LQAKLKENVPWKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKA 558
Query: 1637 ---QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
+ + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 559 IKINEGTNTELICRYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 616
>gb|AAA46350.1| ORF5; putative
Length = 675
Score = 282 bits (721), Expect = 8e-74
Identities = 165/418 (39%), Positives = 240/418 (56%), Gaps = 12/418 (2%)
Query: 1279 RIEDILENIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQ 1338
+IE++LE + S D PN + ++L SD + K +P++ +
Sbjct: 207 KIEELLEKVCSENPLD-PNKTKQWMKASIKL------SDPSKAIKVKPMKYSPMDREEFD 259
Query: 1339 KEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNK 1398
K+I +LL+ K+I+ SKSP AF VN +AE RG R+V+NYK +N+A Y PNK
Sbjct: 260 KQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNPPNK 319
Query: 1399 KDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEF 1458
+LL K+FS F SGFWQ+ L ++ + TAFT P G YEWNV+PFGLK APS F
Sbjct: 320 DELLTLIRGKKIFSSFHCNSGFWQVLLDQESRPLTAFTCPQGHYEWNVVPFGLKQAPSIF 379
Query: 1459 QRIMNEIFNPYSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQT 1518
QR M+E F + K VY+DD+L+FS + H H+ + ++G+ +SK K LF+
Sbjct: 380 QRHMDEAFRVFRKFCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKK 439
Query: 1519 KIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKP 1578
KI FLG I +GT P +E +KFPD + DK QLQRFLG L Y +D+ P+L+ I KP
Sbjct: 440 KINFLGLEIDEGTHKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKP 499
Query: 1579 LHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILK- 1636
L +LK++ P + T ++++K ++ P L+ P P+ IIETDASD +GG+LK
Sbjct: 500 LQAKLKENVPWKWTKEDTLYMQKVKKNLQAFPPLHHPLPEEKLIIETDASDDYWGGMLKA 559
Query: 1637 ---QKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVD 1691
+ + E I + S + A++NY + KE LA++ +I KF L FL+ D
Sbjct: 560 IKINEGTNTELICRYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTD 617
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,979,226,365
Number of Sequences: 2540612
Number of extensions: 135987769
Number of successful extensions: 749309
Number of sequences better than 10.0: 36915
Number of HSP's better than 10.0 without gapping: 3248
Number of HSP's successfully gapped in prelim test: 33721
Number of HSP's that attempted gapping in prelim test: 682837
Number of HSP's gapped (non-prelim): 64427
length of query: 1703
length of database: 863,360,394
effective HSP length: 142
effective length of query: 1561
effective length of database: 502,593,490
effective search space: 784548437890
effective search space used: 784548437890
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0019a.7


