

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.3.5
(80 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
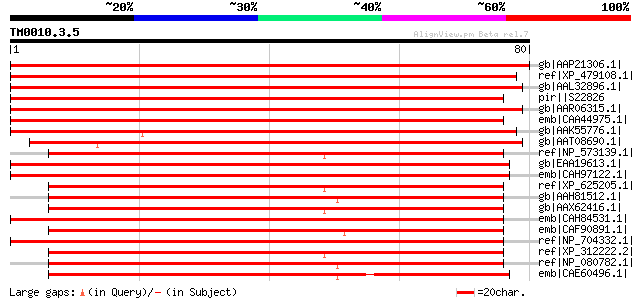
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP21306.1| At2g23930 [Arabidopsis thaliana] gi|21593829|gb|A... 145 3e-34
ref|XP_479108.1| putative small nuclear ribonucleoprotein E [Ory... 143 1e-33
gb|AAL32896.1| putative small nuclear ribonucleoprotein polypept... 143 1e-33
pir||S22826 small nuclear ribonucleoprotein E homolog - alfalfa 141 4e-33
gb|AAR06315.1| putative small nuclear ribonucleoprotein polypept... 140 6e-33
emb|CAA44975.1| snRNP-related protein [Medicago sativa] gi|13412... 138 3e-32
gb|AAK55776.1| Putative small nuclear ribonucleoprotein G [Oryza... 133 1e-30
gb|AAT08690.1| small nuclear ribonucleoprotein polypeptide G [Hy... 120 9e-27
ref|NP_573139.1| CG9742-PA [Drosophila melanogaster] gi|7293253|... 92 4e-18
gb|EAA19613.1| putative small nuclear ribonucleoprotein polypept... 91 6e-18
emb|CAH97122.1| small nuclear ribonucleoprotein, putative [Plasm... 91 8e-18
ref|XP_625205.1| PREDICTED: similar to ENSANGP00000010868 [Apis ... 91 1e-17
gb|AAH81512.1| Zgc:103688 [Danio rerio] gi|52219152|ref|NP_00100... 90 1e-17
gb|AAX62416.1| small nuclear ribonucleoprotein G [Lysiphlebus te... 90 2e-17
emb|CAH84531.1| small nuclear ribonucleoprotein, putative [Plasm... 89 2e-17
emb|CAF90891.1| unnamed protein product [Tetraodon nigroviridis] 89 3e-17
ref|NP_704332.1| small nuclear ribonucleoprotein, putative [Plas... 89 4e-17
ref|XP_312222.2| ENSANGP00000010868 [Anopheles gambiae str. PEST... 88 5e-17
ref|NP_080782.1| small nuclear ribonucleoprotein polypeptide G [... 87 8e-17
emb|CAE60496.1| Hypothetical protein CBG04114 [Caenorhabditis br... 86 2e-16
>gb|AAP21306.1| At2g23930 [Arabidopsis thaliana] gi|21593829|gb|AAM65796.1|
putative small nuclear ribonucleoprotein E [Arabidopsis
thaliana] gi|3738322|gb|AAC63663.1| putative small
nuclear ribonucleoprotein E [Arabidopsis thaliana]
gi|25083131|gb|AAN72046.1| putative small nuclear
ribonucleoprotein E [Arabidopsis thaliana]
gi|15224075|ref|NP_179971.1| small nuclear
ribonucleoprotein G, putative / snRNP-G, putative / Sm
protein G, putative [Arabidopsis thaliana]
gi|25329697|pir||E84630 probable small nuclear
ribonucleoprotein E [imported] - Arabidopsis thaliana
gi|6094211|sp|O82221|RUXG_ARATH Probable small nuclear
ribonucleoprotein G (snRNP-G) (Sm protein G) (Sm-G)
(SmG)
Length = 80
Score = 145 bits (366), Expect = 3e-34
Identities = 70/80 (87%), Positives = 76/80 (94%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRM+ GTLRG+DQFMN+VVDNTVEVNGN+K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRMVTGTLRGFDQFMNLVVDNTVEVNGNDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRTS 80
VIRGNS+VTVEALEPV R+S
Sbjct: 61 VIRGNSIVTVEALEPVGRSS 80
>ref|XP_479108.1| putative small nuclear ribonucleoprotein E [Oryza sativa
(japonica cultivar-group)] gi|34394185|dbj|BAC84637.1|
putative small nuclear ribonucleoprotein E [Oryza
sativa (japonica cultivar-group)]
Length = 80
Score = 143 bits (361), Expect = 1e-33
Identities = 68/78 (87%), Positives = 75/78 (95%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANR+++GTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRVVIGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTR 78
VIRGNSVV +EALEPV +
Sbjct: 61 VIRGNSVVMIEALEPVPK 78
>gb|AAL32896.1| putative small nuclear ribonucleoprotein polypeptide G
[Arabidopsis thaliana] gi|12322914|gb|AAG51452.1|
putative small nuclear ribonucleoprotein polypeptide G;
65009-64161 [Arabidopsis thaliana]
gi|15229773|ref|NP_187757.1| small nuclear
ribonucleoprotein G, putative / snRNP-G, putative / Sm
protein G, putative [Arabidopsis thaliana]
gi|24899751|gb|AAN65090.1| putative small nuclear
ribonucleoprotein polypeptide G [Arabidopsis thaliana]
Length = 79
Score = 143 bits (361), Expect = 1e-33
Identities = 69/79 (87%), Positives = 76/79 (95%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANRM+VGTLRG+DQFMN+VVDNTVEVNG++K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRMVVGTLRGFDQFMNLVVDNTVEVNGDDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRT 79
VIRGNS+VTVEALEPV R+
Sbjct: 61 VIRGNSIVTVEALEPVGRS 79
>pir||S22826 small nuclear ribonucleoprotein E homolog - alfalfa
Length = 81
Score = 141 bits (356), Expect = 4e-33
Identities = 70/76 (92%), Positives = 72/76 (94%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MS SGQPPDLKKYMDK LQI L ANRMIVGTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 1 MSTSGQPPDLKKYMDKQLQINLKANRMIVGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 60
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNSVVTVEALEPV
Sbjct: 61 VIRGNSVVTVEALEPV 76
>gb|AAR06315.1| putative small nuclear ribonucleoprotein polypeptide G [Oryza
sativa (japonica cultivar-group)]
gi|50916481|ref|XP_468637.1| putative small nuclear
ribonucleoprotein polypeptide G [Oryza sativa (japonica
cultivar-group)]
Length = 80
Score = 140 bits (354), Expect = 6e-33
Identities = 67/79 (84%), Positives = 75/79 (94%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MSRSGQPPDLKKYMDK LQIKLNANR+IVGTLRG+DQFMN+VVDNTVEVNGN+K DIGMV
Sbjct: 1 MSRSGQPPDLKKYMDKKLQIKLNANRVIVGTLRGFDQFMNLVVDNTVEVNGNDKTDIGMV 60
Query: 61 VIRGNSVVTVEALEPVTRT 79
V+RGNSVV +EALEPV ++
Sbjct: 61 VVRGNSVVMIEALEPVPKS 79
>emb|CAA44975.1| snRNP-related protein [Medicago sativa]
gi|134126|sp|P24715|RUXG_MEDSA Probable small nuclear
ribonucleoprotein G (snRNP-G) (Sm protein G) (Sm-G)
(SmG)
Length = 81
Score = 138 bits (348), Expect = 3e-32
Identities = 69/76 (90%), Positives = 71/76 (92%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
MS SGQPP LKKYMDK LQI L ANRMIVGTLRG+DQFMN+VVDNTVEVNGNEKNDIGMV
Sbjct: 1 MSTSGQPPALKKYMDKQLQINLKANRMIVGTLRGFDQFMNLVVDNTVEVNGNEKNDIGMV 60
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNSVVTVEALEPV
Sbjct: 61 VIRGNSVVTVEALEPV 76
>gb|AAK55776.1| Putative small nuclear ribonucleoprotein G [Oryza sativa]
Length = 94
Score = 133 bits (335), Expect = 1e-30
Identities = 67/92 (72%), Positives = 75/92 (80%), Gaps = 14/92 (15%)
Query: 1 MSRSGQPPDLKKYMDKNLQ--------------IKLNANRMIVGTLRGYDQFMNMVVDNT 46
MSRSGQPPDLKKYMDK LQ +KLNANR+++GTLRG+DQFMN+VVDNT
Sbjct: 1 MSRSGQPPDLKKYMDKKLQTLMVYSFTLFSILPVKLNANRVVIGTLRGFDQFMNLVVDNT 60
Query: 47 VEVNGNEKNDIGMVVIRGNSVVTVEALEPVTR 78
VEVNGNEKNDIGMVVIRGNSVV +EALEPV +
Sbjct: 61 VEVNGNEKNDIGMVVIRGNSVVMIEALEPVPK 92
>gb|AAT08690.1| small nuclear ribonucleoprotein polypeptide G [Hyacinthus
orientalis]
Length = 96
Score = 120 bits (301), Expect = 9e-27
Identities = 59/77 (76%), Positives = 69/77 (88%), Gaps = 1/77 (1%)
Query: 4 SGQPPDLKK-YMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMVVI 62
SGQPP LK+ + LQIKLNANR++VGTLRG+DQFMN+V+DNT+EVNGN+KNDIGMVVI
Sbjct: 19 SGQPPALKQVHGTSKLQIKLNANRVVVGTLRGFDQFMNLVIDNTMEVNGNDKNDIGMVVI 78
Query: 63 RGNSVVTVEALEPVTRT 79
RGNSVV +EALEPV RT
Sbjct: 79 RGNSVVMIEALEPVART 95
>ref|NP_573139.1| CG9742-PA [Drosophila melanogaster] gi|7293253|gb|AAF48634.1|
CG9742-PA [Drosophila melanogaster]
gi|17944563|gb|AAL48169.1| RH35475p [Drosophila
melanogaster] gi|29428065|sp|Q9VXE0|RUXG_DROME Probable
small nuclear ribonucleoprotein G (snRNP-G) (Sm protein
G) (Sm-G) (SmG)
Length = 76
Score = 91.7 bits (226), Expect = 4e-18
Identities = 44/71 (61%), Positives = 56/71 (77%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP++KKYMDK + +KLN R + G LRG+D FMN+V+D+TV E N KN+IGMVVIRGN
Sbjct: 6 PPEVKKYMDKRMMLKLNGGRAVTGILRGFDPFMNVVLDDTVEECKDNTKNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S+V VEAL+ V
Sbjct: 66 SIVMVEALDRV 76
>gb|EAA19613.1| putative small nuclear ribonucleoprotein polypeptide G
[Plasmodium yoelii yoelii]
Length = 83
Score = 91.3 bits (225), Expect = 6e-18
Identities = 43/77 (55%), Positives = 58/77 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR+IVG LRGYD FMN+V+DNTVE+ +E +IG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRLIVGVLRGYDTFMNLVLDNTVEIKKDENIEIGIV 64
Query: 61 VIRGNSVVTVEALEPVT 77
VIRGNS+ E L+ VT
Sbjct: 65 VIRGNSISYWECLDKVT 81
>emb|CAH97122.1| small nuclear ribonucleoprotein, putative [Plasmodium berghei]
Length = 83
Score = 90.9 bits (224), Expect = 8e-18
Identities = 42/77 (54%), Positives = 58/77 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR+IVG LRGYD FMN+V+DNT+E+ +E +IG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRLIVGVLRGYDTFMNLVLDNTIEIKKDENIEIGIV 64
Query: 61 VIRGNSVVTVEALEPVT 77
VIRGNS+ E L+ VT
Sbjct: 65 VIRGNSISYWECLDKVT 81
>ref|XP_625205.1| PREDICTED: similar to ENSANGP00000010868 [Apis mellifera]
Length = 76
Score = 90.5 bits (223), Expect = 1e-17
Identities = 42/71 (59%), Positives = 55/71 (77%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP+LKKYMDK L +KLN R ++G LRG+D FMNMV+D ++ E KN+IGMVVIRGN
Sbjct: 6 PPELKKYMDKKLSLKLNGGRHVIGILRGFDPFMNMVIDESIEECKDGTKNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
SV+ +EAL+ +
Sbjct: 66 SVIMLEALDRI 76
>gb|AAH81512.1| Zgc:103688 [Danio rerio] gi|52219152|ref|NP_001004660.1|
zgc:103688 [Danio rerio]
Length = 76
Score = 90.1 bits (222), Expect = 1e-17
Identities = 43/71 (60%), Positives = 56/71 (78%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+VVD+T+E+ G + N IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVVDDTIEMAPGGQMNTIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>gb|AAX62416.1| small nuclear ribonucleoprotein G [Lysiphlebus testaceipes]
Length = 76
Score = 89.7 bits (221), Expect = 2e-17
Identities = 42/71 (59%), Positives = 55/71 (77%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP+LKKYMDK L +KLN R ++G LRG+D FMNMV+D +V E +N+IGMVVIRGN
Sbjct: 6 PPELKKYMDKKLSLKLNGGRHVIGILRGFDPFMNMVIDESVEECKDGSQNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
SV+ +EAL+ +
Sbjct: 66 SVIMLEALDRI 76
>emb|CAH84531.1| small nuclear ribonucleoprotein, putative [Plasmodium chabaudi]
Length = 83
Score = 89.4 bits (220), Expect = 2e-17
Identities = 42/76 (55%), Positives = 57/76 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR+IVG LRGYD FMN+V+DNTVE+ +E +IG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRLIVGVLRGYDTFMNLVLDNTVEIKKDENIEIGIV 64
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNS+ E L+ V
Sbjct: 65 VIRGNSISYWECLDKV 80
>emb|CAF90891.1| unnamed protein product [Tetraodon nigroviridis]
Length = 76
Score = 89.0 bits (219), Expect = 3e-17
Identities = 42/71 (59%), Positives = 56/71 (78%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVN-GNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+VVD T+E+ G +++ IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVVDETLEMGPGGQQSSIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>ref|NP_704332.1| small nuclear ribonucleoprotein, putative [Plasmodium falciparum
3D7] gi|23499071|emb|CAD51151.1| small nuclear
ribonucleoprotein, putative [Plasmodium falciparum 3D7]
Length = 83
Score = 88.6 bits (218), Expect = 4e-17
Identities = 41/76 (53%), Positives = 57/76 (74%)
Query: 1 MSRSGQPPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEVNGNEKNDIGMV 60
+ ++G D +K+M+K LQI LN NR +VG LRGYD FMN+V+DNT+E+ +E+ DIG+V
Sbjct: 5 VGKAGPASDFRKFMEKRLQIYLNGNRQVVGILRGYDTFMNLVLDNTMEIKKDEQIDIGVV 64
Query: 61 VIRGNSVVTVEALEPV 76
VIRGNS+ E L+ V
Sbjct: 65 VIRGNSISYWECLDKV 80
>ref|XP_312222.2| ENSANGP00000010868 [Anopheles gambiae str. PEST]
gi|55242066|gb|EAA07689.2| ENSANGP00000010868
[Anopheles gambiae str. PEST]
Length = 76
Score = 88.2 bits (217), Expect = 5e-17
Identities = 41/71 (57%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTV-EVNGNEKNDIGMVVIRGN 65
PP+LKKYMDK L +KLN R++ G LRG+D FMN+VVD ++ E +N+IGMVVIRGN
Sbjct: 6 PPELKKYMDKRLSLKLNGGRVVSGILRGFDPFMNVVVDESIEECKDGTRNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ VEAL+ +
Sbjct: 66 SIIMVEALDRI 76
>ref|NP_080782.1| small nuclear ribonucleoprotein polypeptide G [Mus musculus]
gi|62420278|gb|AAX81996.1| unknown [Homo sapiens]
gi|47682759|gb|AAH70166.1| Small nuclear
ribonucleoprotein polypeptide G [Homo sapiens]
gi|42542824|gb|AAH66302.1| Small nuclear
ribonucleoprotein polypeptide G [Homo sapiens]
gi|20380602|gb|AAH27499.1| Small nuclear
ribonucleoprotein polypeptide G [Mus musculus]
gi|30185984|gb|AAH51470.1| Small nuclear
ribonucleoprotein polypeptide G [Mus musculus]
gi|12652645|gb|AAH00070.1| Small nuclear
ribonucleoprotein polypeptide G [Homo sapiens]
gi|18490257|gb|AAH22432.1| Small nuclear
ribonucleoprotein polypeptide G [Homo sapiens]
gi|59800217|sp|P62309|RUXG_MOUSE Small nuclear
ribonucleoprotein G (snRNP-G) (Sm protein G) (Sm-G)
(SmG) gi|59800216|sp|P62308|RUXG_HUMAN Small nuclear
ribonucleoprotein G (snRNP-G) (Sm protein G) (Sm-G)
(SmG) gi|62948132|gb|AAH94411.1| Small nuclear
ribonucleoprotein polypeptide G [Mus musculus]
gi|806566|emb|CAA59689.1| Sm protein G [Homo sapiens]
gi|48145953|emb|CAG33199.1| SNRPG [Homo sapiens]
gi|12850471|dbj|BAB28732.1| unnamed protein product
[Mus musculus] gi|12849786|dbj|BAB28480.1| unnamed
protein product [Mus musculus]
gi|4507133|ref|NP_003087.1| small nuclear
ribonucleoprotein polypeptide G [Homo sapiens]
Length = 76
Score = 87.4 bits (215), Expect = 8e-17
Identities = 41/71 (57%), Positives = 55/71 (76%), Gaps = 1/71 (1%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV-NGNEKNDIGMVVIRGN 65
PP+LKK+MDK L +KLN R + G LRG+D FMN+V+D VE+ ++N+IGMVVIRGN
Sbjct: 6 PPELKKFMDKKLSLKLNGGRHVQGILRGFDPFMNLVIDECVEMATSGQQNNIGMVVIRGN 65
Query: 66 SVVTVEALEPV 76
S++ +EALE V
Sbjct: 66 SIIMLEALERV 76
>emb|CAE60496.1| Hypothetical protein CBG04114 [Caenorhabditis briggsae]
Length = 77
Score = 85.9 bits (211), Expect = 2e-16
Identities = 44/73 (60%), Positives = 54/73 (73%), Gaps = 3/73 (4%)
Query: 7 PPDLKKYMDKNLQIKLNANRMIVGTLRGYDQFMNMVVDNTVEV--NGNEKNDIGMVVIRG 64
PP+LKKYMDK +++KLN NR + G LRG+D FMNMV+D VE NGN N +GM VIRG
Sbjct: 6 PPELKKYMDKEIELKLNGNRHVSGILRGFDPFMNMVIDEAVEYPKNGNSIN-LGMTVIRG 64
Query: 65 NSVVTVEALEPVT 77
NSVV +E E V+
Sbjct: 65 NSVVIMEPKERVS 77
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.133 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 124,303,862
Number of Sequences: 2540612
Number of extensions: 3968554
Number of successful extensions: 8298
Number of sequences better than 10.0: 451
Number of HSP's better than 10.0 without gapping: 226
Number of HSP's successfully gapped in prelim test: 225
Number of HSP's that attempted gapping in prelim test: 7929
Number of HSP's gapped (non-prelim): 452
length of query: 80
length of database: 863,360,394
effective HSP length: 56
effective length of query: 24
effective length of database: 721,086,122
effective search space: 17306066928
effective search space used: 17306066928
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0010.3.5


