

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.4
(283 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
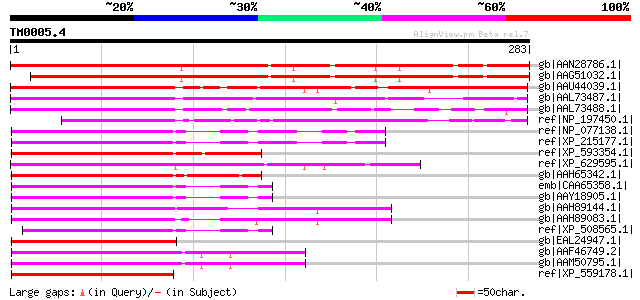
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN28786.1| At3g12480/MQC3.32 [Arabidopsis thaliana] gi|15795... 287 2e-76
gb|AAG51032.1| unknown protein; 69004-67516 [Arabidopsis thaliana] 266 4e-70
gb|AAU44039.1| unknown peotein [Oryza sativa (japonica cultivar-... 242 8e-63
gb|AAL73487.1| repressor protein [Oryza sativa] 202 9e-51
gb|AAL73488.1| repressor protein [Zea mays] 196 8e-49
ref|NP_197450.1| repressor protein-related [Arabidopsis thaliana] 156 7e-37
ref|NP_077138.1| Dr1 associated protein 1 (negative cofactor 2 a... 119 7e-26
ref|XP_215177.1| PREDICTED: similar to Dr1 associated protein 1 ... 116 6e-25
ref|XP_593354.1| PREDICTED: similar to Dr1 associated protein 1 ... 116 6e-25
ref|XP_629595.1| putative histone-like transcription factor [Dic... 116 8e-25
gb|AAH65342.1| Unknown (protein for MGC:77337) [Danio rerio] gi|... 115 1e-24
emb|CAA65358.1| NC2 [Homo sapiens] gi|14603112|gb|AAH10025.1| DR... 113 7e-24
gb|AAY18905.1| DR1-associated protein 1 [synthetic construct] 113 7e-24
gb|AAH89144.1| Unknown (protein for MGC:85186) [Xenopus laevis] 112 9e-24
gb|AAH89083.1| Unknown (protein for MGC:84860) [Xenopus laevis] 112 9e-24
ref|XP_508565.1| PREDICTED: similar to DR1-associated protein 1;... 107 4e-22
gb|EAL24947.1| GA10241-PA [Drosophila pseudoobscura] 106 7e-22
gb|AAF46749.2| CG10318-PA, isoform A [Drosophila melanogaster] g... 105 1e-21
gb|AAM50795.1| LD24434p [Drosophila melanogaster] gi|10242351|gb... 105 1e-21
ref|XP_559178.1| ENSANGP00000026107 [Anopheles gambiae str. PEST... 105 1e-21
>gb|AAN28786.1| At3g12480/MQC3.32 [Arabidopsis thaliana]
gi|15795167|dbj|BAB03155.1| unnamed protein product
[Arabidopsis thaliana] gi|18087597|gb|AAL58929.1|
At3g12480/MQC3.32 [Arabidopsis thaliana]
gi|30682195|ref|NP_187854.2| transcription factor,
putative [Arabidopsis thaliana]
Length = 293
Score = 287 bits (735), Expect = 2e-76
Identities = 168/300 (56%), Positives = 202/300 (67%), Gaps = 24/300 (8%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSK+LELFLQDLCDRTYEITL+RGAK
Sbjct: 1 MRKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKSLELFLQDLCDRTYEITLERGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSH--GHGHSEPSADDRTIPKRRKAAGDDCN 118
T++SLHLKHCV+ YNVFDFLR+VVSKVPDY H G GH + + DDR+I KRRK D+ N
Sbjct: 61 TVSSLHLKHCVERYNVFDFLREVVSKVPDYGHSQGQGHGDVTMDDRSISKRRKPISDEVN 120
Query: 119 DSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRG----RTAVRETPHQAVEPEPCASVQ 174
DSDEE K+ K E+ ++GRG GRGRGRGRGRG + A RE ++ +E E S Q
Sbjct: 121 DSDEEYKKSKTQEIGSAKTSGRG-GRGRGRGRGRGGRAAKAAEREGLNREMEVEAANSGQ 179
Query: 175 QGIQHDTNTDMTIHDSSETKELPK--ENAAVPAESAEFH--------NLDLNANTNE-NE 223
+ N M +SS ++ K + A E + H + DLNA + + NE
Sbjct: 180 P--PPEDNVKMHASESSPQEDEKKGIDGTAASNEDTKQHLQSPKEGIDFDLNAESLDLNE 237
Query: 224 DKKASTTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
K A T + T+S EE GW + D+ KM D QLA+LG I+EDEEDYDEEG
Sbjct: 238 TKLAPATGTTTTTTAATDS--EEYSGWPMMDISKM--DPAQLASLGKRIDEDEEDYDEEG 293
>gb|AAG51032.1| unknown protein; 69004-67516 [Arabidopsis thaliana]
Length = 297
Score = 266 bits (681), Expect = 4e-70
Identities = 158/289 (54%), Positives = 191/289 (65%), Gaps = 24/289 (8%)
Query: 12 ARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCV 71
ARIKKIMQADEDVGKIALAVPVLVSK+LELFLQDLCDRTYEITL+RGAKT++SLHLKHCV
Sbjct: 16 ARIKKIMQADEDVGKIALAVPVLVSKSLELFLQDLCDRTYEITLERGAKTVSSLHLKHCV 75
Query: 72 QSYNVFDFLRDVVSKVPDYSH--GHGHSEPSADDRTIPKRRKAAGDDCNDSDEEVKRGKM 129
+ YNVFDFLR+VVSKVPDY H G GH + + DDR+I KRRK D+ NDSDEE K+ K
Sbjct: 76 ERYNVFDFLREVVSKVPDYGHSQGQGHGDVTMDDRSISKRRKPISDEVNDSDEEYKKSKT 135
Query: 130 PELSHTGSTGRGRGRGRGRGRGRG----RTAVRETPHQAVEPEPCASVQQGIQHDTNTDM 185
E+ ++GRG GRGRGRGRGRG + A RE ++ +E E S Q + N M
Sbjct: 136 QEIGSAKTSGRG-GRGRGRGRGRGGRAAKAAEREGLNREMEVEAANSGQP--PPEDNVKM 192
Query: 186 TIHDSSETKELPK--ENAAVPAESAEFH--------NLDLNANTNE-NEDKKASTTAKQE 234
+SS ++ K + A E + H + DLNA + + NE K A T
Sbjct: 193 HASESSPQEDEKKGIDGTAASNEDTKQHLQSPKEGIDFDLNAESLDLNETKLAPATGTTT 252
Query: 235 ISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEEDYDEEG 283
+ T+S EE GW + D+ KM D QLA+LG I+EDEEDYDEEG
Sbjct: 253 TTTAATDS--EEYSGWPMMDISKM--DPAQLASLGKRIDEDEEDYDEEG 297
>gb|AAU44039.1| unknown peotein [Oryza sativa (japonica cultivar-group)]
Length = 290
Score = 242 bits (618), Expect = 8e-63
Identities = 153/305 (50%), Positives = 192/305 (62%), Gaps = 39/305 (12%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKLDTRFPA RIKKIMQADEDVGKIALAVPVLVSKALELFLQDLC+RTY+IT+QRG K
Sbjct: 1 MRKKLDTRFPAPRIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCNRTYDITVQRGVK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDS 120
T++S HLK C+ SYNV+DFLRDVVSKVPD G S+ DD+ + KRRK A D DS
Sbjct: 61 TLSSSHLKQCIHSYNVYDFLRDVVSKVPDM----GTSDAGVDDK-LGKRRKTAED---DS 112
Query: 121 DEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRE--TPHQAVE-----PEPCASV 173
+EE KR + S T STGRGRGRGRGRGR GR + RE + ++ E P S
Sbjct: 113 EEESKRTRNEAASQT-STGRGRGRGRGRGRRGGRVSEREIISAYEKFEENHEFPPGQFSK 171
Query: 174 QQGIQHDTNTDMTIHDSSETKE-LPKENAAVPAESAEFHNLDLNANTNENEDKKA----- 227
++ D + D T D+ ETKE P NA A N+DLN + +D+ +
Sbjct: 172 PSQLKVDVSVDGT--DAIETKEATPLSNA-----RASLRNIDLNIELTDYDDEGSAPLEV 224
Query: 228 ----------STTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLANLGTHIEEDEE 277
+T++ +SE E++ ++ GW L ++ KMA+D VQ A H E++E
Sbjct: 225 QPPAPAAGVVTTSSGPLVSEVNEEAKTKDFLGWQLPELTKMAMDPVQFALSSNHRLEEDE 284
Query: 278 DYDEE 282
DYD E
Sbjct: 285 DYDNE 289
>gb|AAL73487.1| repressor protein [Oryza sativa]
Length = 258
Score = 202 bits (514), Expect = 9e-51
Identities = 127/285 (44%), Positives = 161/285 (55%), Gaps = 31/285 (10%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKL TRFPAARIKKIMQADEDVGKIALAVPVLVS+ALELFLQDL DRTYEITLQ GAK
Sbjct: 1 MRKKLGTRFPAARIKKIMQADEDVGKIALAVPVLVSRALELFLQDLIDRTYEITLQSGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDS 120
T+NS HLK CV+ Y+ FDFL +VV+KVPD G ++ DDR +P+RRKA + +
Sbjct: 61 TLNSFHLKQCVRRYSSFDFLTEVVNKVPDL----GGADSCGDDRALPRRRKALPNGSDPE 116
Query: 121 DEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQG--IQ 178
+EE + KM S + RGRGRGRGRGRGR T +E + E E QG +
Sbjct: 117 NEESRSSKMAVRS-ANISPRGRGRGRGRGRGRPPTKRKEVGYVQFEDESSMFADQGEALP 175
Query: 179 HDTNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEP 238
+ TIH + A P+ +AE + +N+D +
Sbjct: 176 GEETVPETIHGTESVPPSTHPPAEAPS-AAEIPAPNPKVEEAKNDDHQ------------ 222
Query: 239 PTESQHEEIPGWSLSD-VDKMAIDSVQLANLGTHIEEDEEDYDEE 282
P W + D + + + +L ++ED EDYD E
Sbjct: 223 ---------PDWPMPDAIGNIGVGPSGFGHLTVQVDED-EDYDNE 257
>gb|AAL73488.1| repressor protein [Zea mays]
Length = 254
Score = 196 bits (497), Expect = 8e-49
Identities = 125/285 (43%), Positives = 169/285 (58%), Gaps = 35/285 (12%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
M+KKL TRFPAARIKKIMQADEDVGKIALAVPVLVS++LELFLQDL DRTYEITLQ GAK
Sbjct: 1 MRKKLGTRFPAARIKKIMQADEDVGKIALAVPVLVSRSLELFLQDLIDRTYEITLQSGAK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDS 120
T+NS HLK CV+ Y+ FDFL +VVSKVPD G ++ D+R +P+RRK+ G D
Sbjct: 61 TLNSFHLKQCVKRYSSFDFLTEVVSKVPDL----GGADSCGDERGLPRRRKSNGSD--PE 114
Query: 121 DEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHD 180
++E + KM + + + RGRGRGRGRGRGR T +E + E E +QG
Sbjct: 115 NDESRSSKM-AIRNANISPRGRGRGRGRGRGRPPTKRKEVGYVQFEDESSMFAEQG--EP 171
Query: 181 TNTDMTIHDSSETKELPKENAAVPAESAEFHNLDLNANTNENEDKKASTTAKQEISEPPT 240
+ T+ + + + +P ++ P E+ +A+T++K E
Sbjct: 172 LPGEETVQEINGNETMP-QSTQPPVEAP------------PTALAQATTSSKAE------ 212
Query: 241 ESQHEEIPGWSLSDVDKMAIDSVQLANLG---THIEEDEEDYDEE 282
E+ + W + D AI S+ + G ++ ++EDYD E
Sbjct: 213 EANSDHQSDWPMPD----AIGSIGVVPSGFGHLTVQVEDEDYDNE 253
>ref|NP_197450.1| repressor protein-related [Arabidopsis thaliana]
Length = 236
Score = 156 bits (394), Expect = 7e-37
Identities = 107/255 (41%), Positives = 137/255 (52%), Gaps = 25/255 (9%)
Query: 29 LAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHLKHCVQSYNVFDFLRDVVSKVP 88
+AVP+LVSKALELFLQDLC+ TY++TL RGAKT+N+ HLK CVQ+ NVFDFLRD V+KVP
Sbjct: 1 MAVPLLVSKALELFLQDLCNHTYDVTLSRGAKTVNAFHLKQCVQATNVFDFLRDTVAKVP 60
Query: 89 DYSHGHGHSEPSADDRTIPKRRKAA-GDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRG 147
D G S+ +D++ KRRK G CND D +K +M E+ HT S GRGR RGRG
Sbjct: 61 DL----GGSD--TEDQSATKRRKVVDGSSCNDED-MIKTTQMHEVKHT-SCGRGR-RGRG 111
Query: 148 RGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDTNTDMTIHDSSETKELPKENAAVPAES 207
RGR GRT + + E + N ++ D+S K N
Sbjct: 112 RGRSSGRTGSGLSLKFEEDLEDGSPESSRTPSPENGSLSHDDTSWKKVASHNNHHSSNSE 171
Query: 208 AEFHNLDLNANTNENEDKKASTTAKQEISEPPTESQHEEIPGWSLSDVDKMAIDSVQLAN 267
+ N DLN +EN D E+Q E P + L ++++M ID
Sbjct: 172 VKVRNFDLNVELDENGDNATW-----------LETQLERSPDYPL-EINEMKIDPDDQQQ 219
Query: 268 LGTHIEEDEEDYDEE 282
DEEDYDEE
Sbjct: 220 ASA---SDEEDYDEE 231
>ref|NP_077138.1| Dr1 associated protein 1 (negative cofactor 2 alpha) [Mus musculus]
gi|12805255|gb|AAH02090.1| Dr1 associated protein 1
(negative cofactor 2 alpha) [Mus musculus]
gi|56404664|sp|Q9D6N5|DRAP1_MOUSE Dr1-associated
corepressor (Dr1-associated protein 1) (Negative
co-factor 2 alpha) (NC2 alpha)
gi|26352504|dbj|BAC39882.1| unnamed protein product [Mus
musculus] gi|12845397|dbj|BAB26737.1| unnamed protein
product [Mus musculus]
Length = 205
Score = 119 bits (299), Expect = 7e-26
Identities = 76/204 (37%), Positives = 107/204 (52%), Gaps = 41/204 (20%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHVDGD 105
Query: 122 EEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDT 181
+ +RG+ P GS+GR G +G+ + + S Q+ DT
Sbjct: 106 KGPRRGRKP-----GSSGRKNGGTGSKGKDKKLSGT-------------DSEQEDESEDT 147
Query: 182 NTDMTIHDSSETKELPKENAAVPA 205
+TD ET +LP + + PA
Sbjct: 148 DTD----GEEETPQLPPQASHPPA 167
>ref|XP_215177.1| PREDICTED: similar to Dr1 associated protein 1 (negative cofactor 2
alpha) [Rattus norvegicus]
Length = 205
Score = 116 bits (291), Expect = 6e-25
Identities = 74/204 (36%), Positives = 105/204 (51%), Gaps = 41/204 (20%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHTDGD 105
Query: 122 EEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVEPEPCASVQQGIQHDT 181
+ +RG+ P GS+GR G + + + + S Q+ DT
Sbjct: 106 KGPRRGRKP-----GSSGRKNGGTGSKSKDKKLSGT-------------DSEQEDESEDT 147
Query: 182 NTDMTIHDSSETKELPKENAAVPA 205
+TD ET + P + + PA
Sbjct: 148 DTD----GEEETPQAPPQASHPPA 167
>ref|XP_593354.1| PREDICTED: similar to Dr1 associated protein 1 (negative cofactor 2
alpha) [Bos taurus]
Length = 205
Score = 116 bits (291), Expect = 6e-25
Identities = 62/136 (45%), Positives = 86/136 (62%), Gaps = 2/136 (1%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ P+R + +G +
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGEDNHMDGDKG-PRRGRKSGSSGRKNG 122
Query: 122 EEVKRGKMPELSHTGS 137
+GK +LS T S
Sbjct: 123 GMGSKGKDKKLSGTDS 138
>ref|XP_629595.1| putative histone-like transcription factor [Dictyostelium
discoideum] gi|60463038|gb|EAL61234.1| putative
histone-like transcription factor [Dictyostelium
discoideum]
Length = 550
Score = 116 bits (290), Expect = 8e-25
Identities = 79/256 (30%), Positives = 120/256 (46%), Gaps = 35/256 (13%)
Query: 1 MKKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAK 60
MKKK T+FP ARIKKIMQ DE+VGKIA A P+L+S+ LELF+ DL +T +IT + K
Sbjct: 1 MKKKYKTKFPMARIKKIMQKDEEVGKIASATPILISQCLELFMADLVMKTCKITQAKKGK 60
Query: 61 TMNSLHLKHCVQSYNVFDFLRDVVSKVPD------------YSHGHGHSEPSADDRTIPK 108
++ HLK C++ + FDFL ++V ++PD G E ++
Sbjct: 61 VISVNHLKECIKQESTFDFLTEIVDRIPDDKNEKRGRPKKTEGEDGGEEEEEEEEEMDMG 120
Query: 109 RRKAAGDDCNDSDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRE--------- 159
+ +D +D D + + + P+ GS G G+G GRG RG + R+
Sbjct: 121 EEEEEEEDEDDDDSDEEEEEAPKKGGKGSRG-GKGSRGGRGGARGGASTRKVKSETSPVL 179
Query: 160 ------TPHQAVEPEPC-----ASVQQGIQHDTNTDMTIHDSSETKELPKENAAVPAESA 208
T P P A+ + IQ + + S+ T PK + + + +
Sbjct: 180 KNTTITTTTATTTPTPTPTPNFANSPKDIQSTSLKKPSARKSNTTS--PKSSPFLSSSAG 237
Query: 209 EFHNLDLNANTNENED 224
+ LN N N N +
Sbjct: 238 SIQSPPLNNNNNNNNN 253
>gb|AAH65342.1| Unknown (protein for MGC:77337) [Danio rerio]
gi|28279647|gb|AAH45853.1| Drap1 protein [Danio rerio]
gi|45387529|ref|NP_991104.1| DR1-associated protein 1
(negative cofactor 2 alpha) [Danio rerio]
Length = 211
Score = 115 bits (288), Expect = 1e-24
Identities = 64/136 (47%), Positives = 86/136 (63%), Gaps = 3/136 (2%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLTKACDVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G E + IP+R + G +
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVATVPD-MQGDG-EENHTEGEKIPRRGRKPGSGRKNGG 122
Query: 122 EEVKRGKMPELSHTGS 137
K GK +LS T S
Sbjct: 123 AGAK-GKDKKLSGTES 137
>emb|CAA65358.1| NC2 [Homo sapiens] gi|14603112|gb|AAH10025.1| DR1-associated
protein 1 [Homo sapiens]
gi|56404465|sp|Q14919|DRAP1_HUMAN Dr1-associated
corepressor (Dr1-associated protein 1) (Negative
co-factor 2 alpha) (NC2 alpha)
gi|18426973|ref|NP_006433.2| DR1-associated protein 1
[Homo sapiens]
Length = 205
Score = 113 bits (282), Expect = 7e-24
Identities = 63/142 (44%), Positives = 84/142 (58%), Gaps = 24/142 (16%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHMDGD 105
Query: 122 EEVKRGKMPELSHTGSTGRGRG 143
+ +RG+ P GS GR G
Sbjct: 106 KGARRGRKP-----GSGGRKNG 122
>gb|AAY18905.1| DR1-associated protein 1 [synthetic construct]
Length = 229
Score = 113 bits (282), Expect = 7e-24
Identities = 63/142 (44%), Positives = 84/142 (58%), Gaps = 24/142 (16%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKT
Sbjct: 29 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKT 88
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G G D+ D D
Sbjct: 89 MTTSHLKQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHMDGD 129
Query: 122 EEVKRGKMPELSHTGSTGRGRG 143
+ +RG+ P GS GR G
Sbjct: 130 KGARRGRKP-----GSGGRKNG 146
>gb|AAH89144.1| Unknown (protein for MGC:85186) [Xenopus laevis]
Length = 213
Score = 112 bits (281), Expect = 9e-24
Identities = 71/210 (33%), Positives = 103/210 (48%), Gaps = 20/210 (9%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A VPV++S+ALELFL+ L +T +T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAPVPVIISRALELFLESLLKKTCHVTQSRSAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G + +R + RK N
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVATVPDI-QGDAEDNHTEGERVSRRSRKPGSSRKNG-- 121
Query: 122 EEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVE---PEPCASVQQGIQ 178
GST +G+ + + + T + E P+P Q +
Sbjct: 122 --------------GSTMKGKDKKQSETESEQEESSEGTESEGEEDMCPDPPVGNQSQVT 167
Query: 179 HDTNTDMTIHDSSETKELPKENAAVPAESA 208
+ + + LP+ AA+PA A
Sbjct: 168 FQSPQAPSFINFHSGTALPQVPAALPAMPA 197
>gb|AAH89083.1| Unknown (protein for MGC:84860) [Xenopus laevis]
Length = 212
Score = 112 bits (281), Expect = 9e-24
Identities = 72/212 (33%), Positives = 107/212 (49%), Gaps = 24/212 (11%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + +T R AKT
Sbjct: 5 KKKYNARFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACHVTQSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSD 121
M + HLK C++ FDFL+D+V+ VPD G +E D+ + +
Sbjct: 65 MTTSHLKQCIELEQQFDFLKDLVAAVPDM---QGETE----------------DNHTEGE 105
Query: 122 EEVKRGKMPELS--HTGSTGRGRGRGRGRGRGRGRTAVRETPHQAVE---PEPCASVQQG 176
+RG+ P S + GST +G+ + + T + E P+P Q
Sbjct: 106 RVSRRGRKPGSSRKNGGSTMKGKDKKQSETESEQEEDSEGTESEGEEDMCPDPPVGNQSQ 165
Query: 177 IQHDTNTDMTIHDSSETKELPKENAAVPAESA 208
+ + + + LP+ AA+P A
Sbjct: 166 VTFQSPQAPSFMNFHSGTALPQVPAALPVMPA 197
>ref|XP_508565.1| PREDICTED: similar to DR1-associated protein 1; negative cofactor 2
alpha; DR1-associated corepressor [Pan troglodytes]
Length = 336
Score = 107 bits (267), Expect = 4e-22
Identities = 60/136 (44%), Positives = 80/136 (58%), Gaps = 24/136 (17%)
Query: 8 RFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKTMNSLHL 67
RFP ARIKKIMQ DE++GK+A AVPV++S+ALELFL+ L + ++T R AKTM + HL
Sbjct: 142 RFPPARIKKIMQTDEEIGKVAAAVPVIISRALELFLESLLKKACQVTQSRNAKTMTTSHL 201
Query: 68 KHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADDRTIPKRRKAAGDDCNDSDEEVKRG 127
K C++ FDFL+D+V+ VPD G G D+ D D+ +RG
Sbjct: 202 KQCIELEQQFDFLKDLVASVPD-MQGDGE------------------DNHMDGDKGARRG 242
Query: 128 KMPELSHTGSTGRGRG 143
+ P GS GR G
Sbjct: 243 RKP-----GSGGRKNG 253
>gb|EAL24947.1| GA10241-PA [Drosophila pseudoobscura]
Length = 316
Score = 106 bits (265), Expect = 7e-22
Identities = 48/90 (53%), Positives = 68/90 (75%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ+DE++GK+A AVPV++S+ LELF++ L +T IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQSDEEIGKVAQAVPVIISRTLELFVESLLTKTMRITNSRNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYS 91
+++ H+K C+ S FDFL+++V +PD S
Sbjct: 65 LSTSHMKQCIMSEQRFDFLKELVRNIPDIS 94
>gb|AAF46749.2| CG10318-PA, isoform A [Drosophila melanogaster]
gi|24657218|ref|NP_611601.2| CG10318-PA, isoform A
[Drosophila melanogaster]
Length = 341
Score = 105 bits (262), Expect = 1e-21
Identities = 62/171 (36%), Positives = 94/171 (54%), Gaps = 12/171 (7%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ+DE++GK+A AVPV++S+ LELF++ L +T IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQSDEEIGKVAQAVPVIISRTLELFVESLLTKTLRITNARNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADD--RTIPKRRKAAGDDCND 119
++ H++ C+ S FDFL+++V +PD S + + DD R+ P+ + D D
Sbjct: 65 LSPSHMRQCIVSEKRFDFLKELVRNIPDISVAE-EAAYNEDDVLRSSPEEQYPDSDTPYD 123
Query: 120 ---------SDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETP 161
S M +S G G G + + + + + +ETP
Sbjct: 124 LSLPSTSMRSQANGTAAYMRSMSLNNGAGSGGGAAATKRQFQSQHSTQETP 174
>gb|AAM50795.1| LD24434p [Drosophila melanogaster] gi|10242351|gb|AAG15389.1|
NC2alpha [Drosophila melanogaster]
Length = 341
Score = 105 bits (262), Expect = 1e-21
Identities = 62/171 (36%), Positives = 94/171 (54%), Gaps = 12/171 (7%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ+DE++GK+A AVPV++S+ LELF++ L +T IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQSDEEIGKVAQAVPVIISRTLELFVESLLTKTLRITNARNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPDYSHGHGHSEPSADD--RTIPKRRKAAGDDCND 119
++ H++ C+ S FDFL+++V +PD S + + DD R+ P+ + D D
Sbjct: 65 LSPSHMRQCIVSEKRFDFLKELVRNIPDISVAE-EAAYNEDDVLRSSPEEQYPDSDTPYD 123
Query: 120 ---------SDEEVKRGKMPELSHTGSTGRGRGRGRGRGRGRGRTAVRETP 161
S M +S G G G + + + + + +ETP
Sbjct: 124 LSLPSTSMRSQANGTAAYMRSMSLNNGAGSGGGAAATKRQFQSQHSTQETP 174
>ref|XP_559178.1| ENSANGP00000026107 [Anopheles gambiae str. PEST]
gi|55242883|gb|EAL41071.1| ENSANGP00000026107
[Anopheles gambiae str. PEST]
Length = 93
Score = 105 bits (262), Expect = 1e-21
Identities = 50/88 (56%), Positives = 66/88 (74%)
Query: 2 KKKLDTRFPAARIKKIMQADEDVGKIALAVPVLVSKALELFLQDLCDRTYEITLQRGAKT 61
KKK + RFPA RIKKIMQ DE+VGK+A AVPV+V + LELF++ L +T +IT R AKT
Sbjct: 5 KKKYNARFPAGRIKKIMQTDEEVGKVAQAVPVIVPRTLELFVESLLTKTLKITNARNAKT 64
Query: 62 MNSLHLKHCVQSYNVFDFLRDVVSKVPD 89
++ H+K C+ S + FDFLRD+V +PD
Sbjct: 65 LSPSHMKQCIISESRFDFLRDLVKNIPD 92
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.310 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 499,439,811
Number of Sequences: 2540612
Number of extensions: 22791156
Number of successful extensions: 116624
Number of sequences better than 10.0: 1089
Number of HSP's better than 10.0 without gapping: 514
Number of HSP's successfully gapped in prelim test: 620
Number of HSP's that attempted gapping in prelim test: 111053
Number of HSP's gapped (non-prelim): 2744
length of query: 283
length of database: 863,360,394
effective HSP length: 126
effective length of query: 157
effective length of database: 543,243,282
effective search space: 85289195274
effective search space used: 85289195274
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0005.4


