

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.12
(526 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
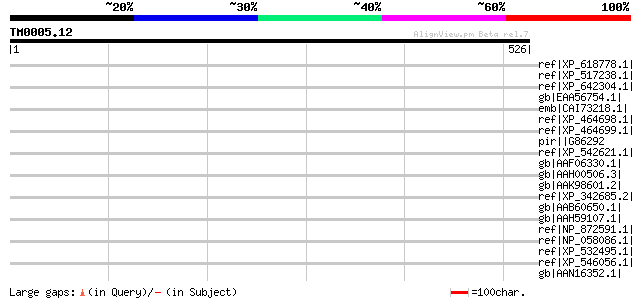
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_618778.1| PREDICTED: similar to heterogeneous nuclear rib... 45 0.006
ref|XP_517238.1| PREDICTED: similar to helix-destabilizing prote... 45 0.007
ref|XP_642304.1| hypothetical protein DDB0205391 [Dictyostelium ... 44 0.010
gb|EAA56754.1| hypothetical protein MG07109.4 [Magnaporthe grise... 44 0.013
emb|CAI73218.1| RNA-binding (SR) protein, putative [Theileria an... 44 0.016
ref|XP_464698.1| putative heterogeneous nuclearribonucleoprotein... 44 0.016
ref|XP_464699.1| putative heterogeneous nuclearribonucleoprotein... 44 0.016
pir||G86292 hypothetical protein F7H2.17 [imported] - Arabidopsi... 44 0.016
ref|XP_542621.1| PREDICTED: similar to Heterogeneous nuclear rib... 43 0.021
gb|AAF06330.1| vitamin D response element binding protein [Sagui... 43 0.021
gb|AAH00506.3| HNRPA2B1 protein [Homo sapiens] 42 0.037
gb|AAK98601.2| heterogeneous nuclear ribonucleoprotein A2/B1 [Mu... 42 0.037
ref|XP_342685.2| PREDICTED: similar to Heterogeneous nuclear rib... 42 0.037
gb|AAB60650.1| hnRNP protein A2 [Homo sapiens] gi|565644|dbj|BAA... 42 0.037
gb|AAH59107.1| Hnrpa2b1 protein [Mus musculus] 42 0.037
ref|NP_872591.1| heterogeneous nuclear ribonucleoprotein A2/B1 i... 42 0.037
ref|NP_058086.1| heterogeneous nuclear ribonucleoprotein A2/B1 i... 42 0.037
ref|XP_532495.1| PREDICTED: similar to Heterogeneous nuclear rib... 42 0.037
ref|XP_546056.1| PREDICTED: similar to 3,5 cyclic nucleotide pho... 42 0.037
gb|AAN16352.1| heterogeneous nuclear ribonucleoprotein A2/B1/B0 ... 42 0.037
>ref|XP_618778.1| PREDICTED: similar to heterogeneous nuclear ribonucleoprotein A2/B1
isoform 2 [Mus musculus]
Length = 281
Score = 45.1 bits (105), Expect = 0.006
Identities = 32/108 (29%), Positives = 48/108 (43%), Gaps = 8/108 (7%)
Query: 22 SRQAAGGTFGDLSSVDDEGDDEGEQ-DGGDARVARVVAPPAR-------TCFPLFVDGIA 73
SR+ TF ++ VD DG + R VAP T LFV G+
Sbjct: 61 SRRFGFVTFSSMAEVDAAMAARSHSIDGREVEPKRAVAPEESGKPGAHVTVKKLFVGGMK 120
Query: 74 DSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
+ H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 121 EDTEEHHLRDSFEEYGKIDTIEIITDRQPGKKRGFGFVTFDDHDPVDK 168
>ref|XP_517238.1| PREDICTED: similar to helix-destabilizing protein - rat [Pan
troglodytes]
Length = 494
Score = 44.7 bits (104), Expect = 0.007
Identities = 20/61 (32%), Positives = 34/61 (54%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRK 126
+FV GI + H+RD F Q+GK+ + + + G+K F FV + N + V K S++
Sbjct: 274 IFVGGIKEDTEEHHLRDYFEQYGKIEVIEIMTDQGSGKKRGFAFVTFDNHDSVRKALSKQ 333
Query: 127 Q 127
+
Sbjct: 334 E 334
>ref|XP_642304.1| hypothetical protein DDB0205391 [Dictyostelium discoideum]
gi|60470367|gb|EAL68347.1| hypothetical protein
DDB0205391 [Dictyostelium discoideum]
Length = 326
Score = 44.3 bits (103), Expect = 0.010
Identities = 33/114 (28%), Positives = 49/114 (42%), Gaps = 4/114 (3%)
Query: 14 SPKSGGITSRQAAGGTFGDLSSVDDEGDDEGEQDGGDARVARVVAPPARTCFPLFVDGIA 73
SP GG +R + G S D +GG+ +A A L V G+A
Sbjct: 66 SPNRGGSPNRGGSPNRGGSPS----RDDKRRYGNGGNGETRNRLANTASPSNVLGVFGLA 121
Query: 74 DSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRKQ 127
++D F + GK+ +V L R GR FGFV + N+ED ++ + Q
Sbjct: 122 PQTEERDLKDEFSRFGKIDHVDLIMDRKTGRSKCFGFVYFENKEDAVRAKEECQ 175
>gb|EAA56754.1| hypothetical protein MG07109.4 [Magnaporthe grisea 70-15]
gi|39971587|ref|XP_367184.1| hypothetical protein
MG07109.4 [Magnaporthe grisea 70-15]
Length = 303
Score = 43.9 bits (102), Expect = 0.013
Identities = 37/117 (31%), Positives = 55/117 (46%), Gaps = 8/117 (6%)
Query: 18 GGITSRQ-AAGGTFGDLSSVDD---EGD-DEGEQDGGDARVARVVAPPARTCFPLFVDGI 72
GGI S AAGG S GD EGE+ GG A T L V +
Sbjct: 174 GGILSAPGAAGGAGAKKGSYVPPALRGDRKEGEKMGGGAGGKYGERDDLAT---LRVTNV 230
Query: 73 ADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRKQSY 129
++ +RD+F + G++T VFL + R+ G F F+ +A+RED +K ++ +
Sbjct: 231 SEMAEEQELRDMFERFGRVTRVFLAKDRDTGLAKGFAFISFADREDAVKACNKMDGW 287
>emb|CAI73218.1| RNA-binding (SR) protein, putative [Theileria annulata]
Length = 303
Score = 43.5 bits (101), Expect = 0.016
Identities = 32/103 (31%), Positives = 50/103 (48%), Gaps = 12/103 (11%)
Query: 29 TFGDLSSVDDEGDDEGEQDGGDARV--------ARVVAPPARTCFPLFVDGIADSVNYFH 80
TF D SV+ + DG +A V ++++AP +FV G+++ + +
Sbjct: 75 TFADRDSVNTVLRKVHKIDGVEADVKLAVRKEKSKILAPQYDQTKRIFVGGVSEKITESY 134
Query: 81 VRDLFLQHGKLT--NVFLQRKRNRGRKFRFGFVRYANREDVMK 121
RD F ++G +T N + R NR R + GFV Y N ED K
Sbjct: 135 FRDYFARYGPITSYNYLVDRATNRPRGY--GFVIYENMEDAEK 175
Score = 38.9 bits (89), Expect = 0.41
Identities = 17/53 (32%), Positives = 32/53 (60%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDV 119
+F G+ S +++ F ++G++T+ + R +N GR FGFV +A+R+ V
Sbjct: 30 IFAGGLTRSTTPEDLKEYFSKYGEVTHTEIVRDKNTGRSRGFGFVTFADRDSV 82
>ref|XP_464698.1| putative heterogeneous nuclearribonucleoprotein A2 [Oryza sativa
(japonica cultivar-group)] gi|46806518|dbj|BAD17631.1|
putative heterogeneous nuclearribonucleoprotein A2
[Oryza sativa (japonica cultivar-group)]
gi|46806499|dbj|BAD17623.1| putative heterogeneous
nuclearribonucleoprotein A2 [Oryza sativa (japonica
cultivar-group)]
Length = 374
Score = 43.5 bits (101), Expect = 0.016
Identities = 43/151 (28%), Positives = 61/151 (39%), Gaps = 23/151 (15%)
Query: 29 TFGDLSSVDDEGDDEGEQDGGDARVARVV---------APPARTCFPLFVDGIADSVNYF 79
TF D S +D +DE DG V R V P R +FV G+ S+
Sbjct: 113 TFSDPSVIDKVLEDEHVIDGRTVEVKRTVPREEMSSKDGPKTRK---IFVGGLPSSLTED 169
Query: 80 HVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDV----MKGRSR----KQSYVS 131
+R+ F +GK+ + + GR FGFV + + + V +GR R KQ +
Sbjct: 170 ELREHFSPYGKIVEHQIMLDHSTGRSRGFGFVTFESEDSVERVISEGRMRDLGGKQVEIK 229
Query: 132 MAPPTT---VKDSRAKERLHLSGGFASSSRS 159
A P S + GG+ SS RS
Sbjct: 230 KAEPKKHGGDHSSNGRSSHGSGGGYRSSYRS 260
>ref|XP_464699.1| putative heterogeneous nuclearribonucleoprotein A2 [Oryza sativa
(japonica cultivar-group)] gi|46806519|dbj|BAD17632.1|
putative heterogeneous nuclearribonucleoprotein A2
[Oryza sativa (japonica cultivar-group)]
gi|46806500|dbj|BAD17624.1| putative heterogeneous
nuclearribonucleoprotein A2 [Oryza sativa (japonica
cultivar-group)]
Length = 397
Score = 43.5 bits (101), Expect = 0.016
Identities = 43/151 (28%), Positives = 61/151 (39%), Gaps = 23/151 (15%)
Query: 29 TFGDLSSVDDEGDDEGEQDGGDARVARVV---------APPARTCFPLFVDGIADSVNYF 79
TF D S +D +DE DG V R V P R +FV G+ S+
Sbjct: 113 TFSDPSVIDKVLEDEHVIDGRTVEVKRTVPREEMSSKDGPKTRK---IFVGGLPSSLTED 169
Query: 80 HVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDV----MKGRSR----KQSYVS 131
+R+ F +GK+ + + GR FGFV + + + V +GR R KQ +
Sbjct: 170 ELREHFSPYGKIVEHQIMLDHSTGRSRGFGFVTFESEDSVERVISEGRMRDLGGKQVEIK 229
Query: 132 MAPPTT---VKDSRAKERLHLSGGFASSSRS 159
A P S + GG+ SS RS
Sbjct: 230 KAEPKKHGGDHSSNGRSSHGSGGGYRSSYRS 260
>pir||G86292 hypothetical protein F7H2.17 [imported] - Arabidopsis thaliana
gi|8927662|gb|AAF82153.1| Contains similarity to
extensin (atExt1) from Arabidopsis thaliana gb|U43627
and is rich in proline and glycine. EST gb|Z37262 comes
from this gene
Length = 1006
Score = 43.5 bits (101), Expect = 0.016
Identities = 26/61 (42%), Positives = 30/61 (48%), Gaps = 8/61 (13%)
Query: 431 DDPVLDFSPPRP--RPRGRPKRKKEVKRGLEGSRRPQGHPRKSGTMDLDAAPSPSHPSPP 488
++P L F PPRP RPR RP+ + L S P HPR S P P PSPP
Sbjct: 90 ENPFLPFQPPRPPPRPRPRPRPSPRLPPPLVPSPPPPLHPRPS------PCPPPLMPSPP 143
Query: 489 P 489
P
Sbjct: 144 P 144
>ref|XP_542621.1| PREDICTED: similar to Heterogeneous nuclear ribonucleoprotein A1
(Helix-destabilizing protein) (Single-strand binding
protein) (hnRNP core protein A1) (HDP) [Canis
familiaris]
Length = 337
Score = 43.1 bits (100), Expect = 0.021
Identities = 23/65 (35%), Positives = 34/65 (51%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRK 126
+FV GI + H+RD F QHGK+ + + R G+K F FV + + + V K RK
Sbjct: 85 IFVGGIKEDTEEHHLRDYFEQHGKIEVLEIMTDRGSGKKRGFAFVTFDDHDSVDKIVIRK 144
Query: 127 QSYVS 131
V+
Sbjct: 145 SYTVN 149
>gb|AAF06330.1| vitamin D response element binding protein [Saguinus oedipus]
Length = 341
Score = 43.1 bits (100), Expect = 0.021
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 102 LFVGGIKEDTEEHHLRDYFAEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 156
>gb|AAH00506.3| HNRPA2B1 protein [Homo sapiens]
Length = 249
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 102 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 156
>gb|AAK98601.2| heterogeneous nuclear ribonucleoprotein A2/B1 [Mus musculus]
Length = 341
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 102 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 156
>ref|XP_342685.2| PREDICTED: similar to Heterogeneous nuclear ribonucleoproteins
A2/B1 (hnRNP A2 / hnRNP B1) [Rattus norvegicus]
gi|133257|sp|P22626|ROA2_HUMAN Heterogeneous nuclear
ribonucleoproteins A2/B1 (hnRNP A2 / hnRNP B1)
gi|565643|dbj|BAA06031.1| hnRNP B1 protein [Homo
sapiens] gi|337453|gb|AAA60271.1| hnRNP B1 protein
gi|14043072|ref|NP_112533.1| heterogeneous nuclear
ribonucleoprotein A2/B1 isoform B1 [Homo sapiens]
Length = 353
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 114 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 168
>gb|AAB60650.1| hnRNP protein A2 [Homo sapiens] gi|565644|dbj|BAA06032.1| hnRNP A2
protein [Homo sapiens] gi|337449|gb|AAA36574.1| hnRNP A2
protein gi|4504447|ref|NP_002128.1| heterogeneous
nuclear ribonucleoprotein A2/B1 isoform A2 [Homo
sapiens]
Length = 341
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 102 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 156
>gb|AAH59107.1| Hnrpa2b1 protein [Mus musculus]
Length = 219
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 20 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 74
>ref|NP_872591.1| heterogeneous nuclear ribonucleoprotein A2/B1 isoform 2 [Mus
musculus] gi|26354140|dbj|BAC40700.1| unnamed protein
product [Mus musculus]
Length = 301
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 102 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 156
>ref|NP_058086.1| heterogeneous nuclear ribonucleoprotein A2/B1 isoform 1 [Mus
musculus] gi|3329498|gb|AAC26867.1| heterogenous nuclear
ribonucleoprotein A2/B1 [Mus musculus]
gi|6647752|sp|O88569|ROA2_MOUSE Heterogeneous nuclear
ribonucleoproteins A2/B1 (hnRNP A2 / hnRNP B1)
Length = 341
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 102 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 156
>ref|XP_532495.1| PREDICTED: similar to Heterogeneous nuclear ribonucleoproteins
A2/B1 (hnRNP A2 / hnRNP B1) [Canis familiaris]
Length = 391
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 167 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 221
>ref|XP_546056.1| PREDICTED: similar to 3,5 cyclic nucleotide phosphodiesterase 8B
[Canis familiaris]
Length = 1541
Score = 42.4 bits (98), Expect = 0.037
Identities = 36/129 (27%), Positives = 56/129 (42%), Gaps = 15/129 (11%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMKGRSRK 126
+FV GI + H+RD F Q+GK+ + + R G+K F FV + + + V K +K
Sbjct: 113 IFVGGIREDTEEHHLRDYFEQYGKIEVIEIMTDRGSGKKRGFAFVTFDDHDSVDKIVIQK 172
Query: 127 ------------QSYVSMAPPTTVKDSRAKERLHLSGGFASSSRSW--CDVLKGCSAKWI 172
+ V +AP +D + LH GG + S S D L CS+ +
Sbjct: 173 YHTVNGHSCVVSSAEVRIAPMRLTQDP-IQVDLHQRGGVSESGPSMGSKDCLYFCSSAYT 231
Query: 173 EKQPLLDSE 181
L S+
Sbjct: 232 TVPRLFPSK 240
>gb|AAN16352.1| heterogeneous nuclear ribonucleoprotein A2/B1/B0 [Mus musculus]
Length = 353
Score = 42.4 bits (98), Expect = 0.037
Identities = 20/55 (36%), Positives = 31/55 (56%)
Query: 67 LFVDGIADSVNYFHVRDLFLQHGKLTNVFLQRKRNRGRKFRFGFVRYANREDVMK 121
LFV GI + H+RD F ++GK+ + + R G+K FGFV + + + V K
Sbjct: 114 LFVGGIKEDTEEHHLRDYFEEYGKIDTIEIITDRQSGKKRGFGFVTFDDHDPVDK 168
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 966,469,590
Number of Sequences: 2540612
Number of extensions: 44479887
Number of successful extensions: 163719
Number of sequences better than 10.0: 524
Number of HSP's better than 10.0 without gapping: 284
Number of HSP's successfully gapped in prelim test: 254
Number of HSP's that attempted gapping in prelim test: 162332
Number of HSP's gapped (non-prelim): 1387
length of query: 526
length of database: 863,360,394
effective HSP length: 133
effective length of query: 393
effective length of database: 525,458,998
effective search space: 206505386214
effective search space used: 206505386214
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0005.12


