

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0005.1
(346 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
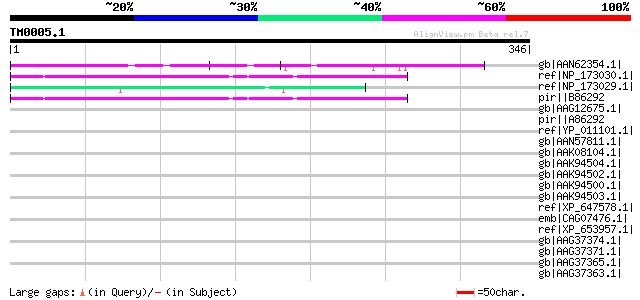
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN62354.1| CTV.22 [Poncirus trifoliata] 65 4e-09
ref|NP_173030.1| expressed protein [Arabidopsis thaliana] 52 2e-05
ref|NP_173029.1| expressed protein [Arabidopsis thaliana] 52 2e-05
pir||B86292 F7H2.12 protein - Arabidopsis thaliana gi|8927657|gb... 52 2e-05
gb|AAG12675.1| hypothetical protein; 58049-55604 [Arabidopsis th... 42 0.021
pir||A86292 protein F7H2.11 [imported] - Arabidopsis thaliana gi... 41 0.060
ref|YP_011101.1| methyl-accepting chemotaxis protein [Desulfovib... 39 0.17
gb|AAN57811.1| putative secreted antigen GbpB/SagA; putative pep... 39 0.17
gb|AAK08104.1| immunodominant glycoprotein IDG-60; general stres... 39 0.17
gb|AAK94504.1| glucan-binding protein B [Streptococcus mutans] 39 0.17
gb|AAK94502.1| glucan-binding protein B [Streptococcus mutans] 39 0.17
gb|AAK94500.1| glucan-binding protein B [Streptococcus mutans] 39 0.17
gb|AAK94503.1| glucan-binding protein B [Streptococcus mutans] 39 0.23
ref|XP_647578.1| hypothetical protein DDB0189801 [Dictyostelium ... 39 0.30
emb|CAG07476.1| unnamed protein product [Tetraodon nigroviridis] 39 0.30
ref|XP_653957.1| hypothetical protein 67.t00003 [Entamoeba histo... 37 0.66
gb|AAG37374.1| ACP36DE [Drosophila melanogaster] 37 0.66
gb|AAG37371.1| ACP36DE [Drosophila melanogaster] 37 0.66
gb|AAG37365.1| ACP36DE [Drosophila melanogaster] 37 0.66
gb|AAG37363.1| ACP36DE [Drosophila melanogaster] gi|11612216|gb|... 37 0.66
>gb|AAN62354.1| CTV.22 [Poncirus trifoliata]
Length = 1405
Score = 64.7 bits (156), Expect = 4e-09
Identities = 47/180 (26%), Positives = 86/180 (47%), Gaps = 12/180 (6%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+ + LN +YQ+I +KL+QH+S P +PK+ +EK + K + E I + ++ KS I
Sbjct: 626 MKEMYLPELNEMYQKIAAKLQQHDSLPQQPKSDQLEKLKIFKTMLERIISFLQVSKSNIL 685
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P E++ E I + I ++ K + +Q + Q+ P T + +Q SQ +
Sbjct: 686 PSFKEKLGSYEKQIVNFIS----TNRPRKPVSSMQQQGQLPP----THMHSMQQQQSQIS 737
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSAL 180
+ Q+ Q + + ++++ VQ ++ SGVST+ +S L
Sbjct: 738 QGQPHDNQMNSQIQSMNLAGSMVTMQPNNVTNVQHNSV----PSVSGVSTSQQNMLNSVL 793
Score = 58.9 bits (141), Expect = 2e-07
Identities = 62/205 (30%), Positives = 91/205 (44%), Gaps = 26/205 (12%)
Query: 134 GNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIK-------GSGN 186
G+SE+ ++ +S G +QT AQA A ++ G S+S L+ GN
Sbjct: 1045 GDSEKPISGISSLSNAGNIGHQQTTSAQAA-APSLAIGTPGISASPLLAEFTGPDGAHGN 1103
Query: 187 LNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAK----- 241
A ++E+P ++RLI SMS +AL AS+ I VV + D I +
Sbjct: 1104 ALTAISIKASVTEQP---LERLIKAVKSMSPKALSASVSDIGSVVSMIDRIAGSAPGNGS 1160
Query: 242 --FIDEPLEMAGQQNQPSR--IPQG-----RKMARNINAMAFDTSRSCARTHESFNQLTD 292
+ E L + +R I Q RKM R +AM S ++SF QLT
Sbjct: 1161 RAAVGEDLVAMTKCRLQARNFITQDGSSGPRKMRRYTSAMPLSVVSSAGSMNDSFKQLTG 1220
Query: 293 VEEPDLDLVA-SKVKRSRIEVIYLL 316
E DL+ A S +KR R+E + L
Sbjct: 1221 SETSDLESTATSSIKRPRMEANHAL 1245
>ref|NP_173030.1| expressed protein [Arabidopsis thaliana]
Length = 1335
Score = 52.4 bits (124), Expect = 2e-05
Identities = 59/268 (22%), Positives = 119/268 (44%), Gaps = 9/268 (3%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+T+ LN IYQ++ +KL+Q +S P + ++ +EK R K + E + + KS I
Sbjct: 604 MKETYLPDLNEIYQRVAAKLQQ-DSMPQQQRSDQLEKLRQFKTMLERMIQFLSVSKSNIM 662
Query: 61 PEISERVDRVE-TLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTS-Q 118
P + ++V E +I + H+ Q Q + PQ Q+ QT+ Q
Sbjct: 663 PALKDKVAYYEKQIIGFLNMHRPRKPVQQGQLPQSQMQPMQQPQSQTVQDQSHDNQTNPQ 722
Query: 119 RNNMCLQ-EQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSS 177
+M +Q A+Q + N LS + PG + +Q + + AS + + +
Sbjct: 723 MQSMSMQGAGPRAQQSSMTNMQSNVLSSR--PGVSAPQQNI-PSSIPASSLESGQGNTLN 779
Query: 178 SALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
+ G++ + + L++ ++A L + ++++ L +S+ + + + D
Sbjct: 780 NGQQVAMGSMQQ--NTSQLVNNSSASAQSGLSTLQSNVNQPQLSSSLLQHQHLKQQQDQQ 837
Query: 238 PAAKFIDEPLEMAGQQNQPSRIPQGRKM 265
K + +M QQ Q + Q +++
Sbjct: 838 MQLKQQFQQRQMQQQQLQARQQQQQQQL 865
Score = 35.0 bits (79), Expect = 3.3
Identities = 57/257 (22%), Positives = 98/257 (37%), Gaps = 38/257 (14%)
Query: 96 SKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSI---------- 145
S Q+L + HL+ Q Q+N + + Q NS V S
Sbjct: 919 SSPQLLQGASPQMSQHLSPQVDQKNTV--NKMGTPLQPANSPFVVPSPSSTPLAPSPMQV 976
Query: 146 -KEGPGKA------VQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLN-EVSHKPALI 197
E PG + + RQ ++ G S+S L++ + + + + +
Sbjct: 977 DSEKPGSSSLSMGNIARQQATGMQGVVQSLAIGTPGISASPLLQEFTSPDGNILNSSTIT 1036
Query: 198 SEKPSAA---MQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQ-- 252
S KPSA ++RLI S+S +AL +++ I VV + D I + + G+
Sbjct: 1037 SGKPSATELPIERLIRAVKSISPQALSSAVSDIGSVVSMVDRIAGSAPGNGSRASVGEDL 1096
Query: 253 --------QNQPSRIPQG----RKMARNINAMAFDTSRSCARTHESFNQLTDVEEPDLDL 300
Q + +G +KM R+ AM + +++ Q E DL+
Sbjct: 1097 VAMTKCRLQARNFMTQEGMMATKKMKRHTTAMPLSVASLGGSVGDNYKQFAGSETSDLES 1156
Query: 301 VA-SKVKRSRIEVIYLL 316
A S K++R E + L
Sbjct: 1157 TATSDGKKARTETEHAL 1173
>ref|NP_173029.1| expressed protein [Arabidopsis thaliana]
Length = 332
Score = 52.4 bits (124), Expect = 2e-05
Identities = 57/247 (23%), Positives = 98/247 (39%), Gaps = 12/247 (4%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
M++ + + +YQ++ KL ES P + ++ + +KF+ K E + + LRK I
Sbjct: 20 MRERYLPHVTDVYQRLADKLNHEESLPQQQRSKHFDKFQSMKTNIEQLIQVLSLRKRNIM 79
Query: 61 PEISE-RVDRVET-----LINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAK 114
P + + R D E LI ++ HQ + Q L Q L L
Sbjct: 80 PILKDCRWDYYEKRIIYFLITLRTGWEDNQIHQRDLNEIYQRVAAKLQQSLRKTVQKLQL 139
Query: 115 QTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMG 174
S+ M Q + + +Q Q+ G + Q + ++ + T G
Sbjct: 140 TKSEIQPMQQPLSQTVQDQSHDDQTTLQMQSMSMQGAGSRVQQIRQGVLQSLEIGT--PG 197
Query: 175 FSSSALI----KGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREV 230
S+S L+ GN+ S ++RLI S+S +AL +++ IR V
Sbjct: 198 ISASPLLPELTSPDGNIINPLTSTCGKSSATELPIERLIRAMKSISPQALSSAVCDIRSV 257
Query: 231 VHLNDAI 237
V + D I
Sbjct: 258 VSMVDRI 264
>pir||B86292 F7H2.12 protein - Arabidopsis thaliana gi|8927657|gb|AAF82148.1|
EST gb|N38213 comes from this gene. [Arabidopsis
thaliana]
Length = 1366
Score = 52.4 bits (124), Expect = 2e-05
Identities = 59/268 (22%), Positives = 119/268 (44%), Gaps = 9/268 (3%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
MK+T+ LN IYQ++ +KL+Q +S P + ++ +EK R K + E + + KS I
Sbjct: 635 MKETYLPDLNEIYQRVAAKLQQ-DSMPQQQRSDQLEKLRQFKTMLERMIQFLSVSKSNIM 693
Query: 61 PEISERVDRVE-TLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTS-Q 118
P + ++V E +I + H+ Q Q + PQ Q+ QT+ Q
Sbjct: 694 PALKDKVAYYEKQIIGFLNMHRPRKPVQQGQLPQSQMQPMQQPQSQTVQDQSHDNQTNPQ 753
Query: 119 RNNMCLQ-EQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSS 177
+M +Q A+Q + N LS + PG + +Q + + AS + + +
Sbjct: 754 MQSMSMQGAGPRAQQSSMTNMQSNVLSSR--PGVSAPQQNI-PSSIPASSLESGQGNTLN 810
Query: 178 SALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLNDAI 237
+ G++ + + L++ ++A L + ++++ L +S+ + + + D
Sbjct: 811 NGQQVAMGSMQQ--NTSQLVNNSSASAQSGLSTLQSNVNQPQLSSSLLQHQHLKQQQDQQ 868
Query: 238 PAAKFIDEPLEMAGQQNQPSRIPQGRKM 265
K + +M QQ Q + Q +++
Sbjct: 869 MQLKQQFQQRQMQQQQLQARQQQQQQQL 896
Score = 35.0 bits (79), Expect = 3.3
Identities = 57/257 (22%), Positives = 98/257 (37%), Gaps = 38/257 (14%)
Query: 96 SKQQVLPQVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSI---------- 145
S Q+L + HL+ Q Q+N + + Q NS V S
Sbjct: 950 SSPQLLQGASPQMSQHLSPQVDQKNTV--NKMGTPLQPANSPFVVPSPSSTPLAPSPMQV 1007
Query: 146 -KEGPGKA------VQRQTLCAQATEASGVSTNDMGFSSSALIKGSGNLN-EVSHKPALI 197
E PG + + RQ ++ G S+S L++ + + + + +
Sbjct: 1008 DSEKPGSSSLSMGNIARQQATGMQGVVQSLAIGTPGISASPLLQEFTSPDGNILNSSTIT 1067
Query: 198 SEKPSAA---MQRLIGVFTSMSSEALGASIGKIREVVHLNDAIPAAKFIDEPLEMAGQ-- 252
S KPSA ++RLI S+S +AL +++ I VV + D I + + G+
Sbjct: 1068 SGKPSATELPIERLIRAVKSISPQALSSAVSDIGSVVSMVDRIAGSAPGNGSRASVGEDL 1127
Query: 253 --------QNQPSRIPQG----RKMARNINAMAFDTSRSCARTHESFNQLTDVEEPDLDL 300
Q + +G +KM R+ AM + +++ Q E DL+
Sbjct: 1128 VAMTKCRLQARNFMTQEGMMATKKMKRHTTAMPLSVASLGGSVGDNYKQFAGSETSDLES 1187
Query: 301 VA-SKVKRSRIEVIYLL 316
A S K++R E + L
Sbjct: 1188 TATSDGKKARTETEHAL 1204
>gb|AAG12675.1| hypothetical protein; 58049-55604 [Arabidopsis thaliana]
Length = 634
Score = 42.4 bits (98), Expect = 0.021
Identities = 32/138 (23%), Positives = 62/138 (44%), Gaps = 6/138 (4%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEP-KTFNVEKFRGHKRLYEDIFAMFRLRKSQI 59
+K+ HF L+ ++++I KLRQ ES P +P + ++EK + K E + +++S +
Sbjct: 33 LKEMHFPVLSLMHKRIAEKLRQTESLPPQPMQAQSIEKLKAGKLSMEHLMFFLSVQRSNV 92
Query: 60 TPEISERVDRVETLINSIIQHQNL-----SSHQGKHSADVQSKQQVLPQVLGTQNDHLAK 114
+ + ++ E I + Q + QG+ + Q PQV +Q+ +
Sbjct: 93 SEKHRDKFSLYEHHILKFTKSQTMVQRSTQQQQGQFPPSQTAMQSQSPQVHVSQSLDKEQ 152
Query: 115 QTSQRNNMCLQEQQVAKQ 132
S+ C E + Q
Sbjct: 153 MRSRLMPSCQNEASSSLQ 170
Score = 35.0 bits (79), Expect = 3.3
Identities = 41/170 (24%), Positives = 73/170 (42%), Gaps = 17/170 (10%)
Query: 79 QHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNNMCLQ--------EQQV- 129
QHQ S+Q DV+ +Q++L +++ + + KQ S+ ++ +Q +QQ+
Sbjct: 234 QHQMQRSNQTNEMNDVRIRQRLLEKLVSSSQLQVPKQVSKVSSPQIQNHSSPQLVDQQIL 293
Query: 130 ---AKQYGNSEQGVNQLSIKEGPGKAV-QRQTLCAQATEASGVSTNDMGFSSSALIKGSG 185
+ G + + P + + + SGV N SSS L
Sbjct: 294 PATVNKTGTPLKSSGSPFVAPAPSPVPGDSEMPISVESPVSGVEINSTLDSSSKLGTQET 353
Query: 186 NLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVHLND 235
L V P I+E+P + RLI F + S ++L S+ +I V+ + D
Sbjct: 354 PLLSVP-PPEPITERP---IDRLIKAFQAASPKSLAESVSEISSVISMVD 399
>pir||A86292 protein F7H2.11 [imported] - Arabidopsis thaliana
gi|8927656|gb|AAF82147.1| F7H2.11 [Arabidopsis thaliana]
Length = 392
Score = 40.8 bits (94), Expect = 0.060
Identities = 57/307 (18%), Positives = 111/307 (35%), Gaps = 72/307 (23%)
Query: 1 MKDTHFARLNAIYQQICSKLRQHESSPHEPKTFNVEKFRGHKRLYEDIFAMFRLRKSQIT 60
M++ + + +YQ++ KL ES P + ++ + +KF+ K E + + LRK I
Sbjct: 20 MRERYLPHVTDVYQRLADKLNHEESLPQQQRSKHFDKFQSMKTNIEQLIQVLSLRKRNIM 79
Query: 61 P----------------------------EISER-VDRVETLINSIIQHQNLSSHQGKHS 91
P +I +R ++ + + + +Q ++ SHQ + S
Sbjct: 80 PILKDCRWDYYEKRIIYFLITLRTGWEDNQIHQRDLNEIYQRVAAKLQQEDSLSHQKQRS 139
Query: 92 ADVQ------------------SKQQVLP-----------QVLGTQNDHLAKQTSQRNNM 122
+ SK + P ++ N ++T Q+ +
Sbjct: 140 DQFEKLKRGKTVLEGMLRFLSLSKSNIKPDLKDSMDYRKNNIMNFLNMQSLRKTVQKLQL 199
Query: 123 CLQE--------QQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMG 174
E Q + + +Q Q+ G + Q + ++ + T G
Sbjct: 200 TKSEIQPMQQPLSQTVQDQSHDDQTTLQMQSMSMQGAGSRVQQIRQGVLQSLEIGT--PG 257
Query: 175 FSSSALI----KGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREV 230
S+S L+ GN+ S ++RLI S+S +AL +++ IR V
Sbjct: 258 ISASPLLPELTSPDGNIINPLTSTCGKSSATELPIERLIRAMKSISPQALSSAVCDIRSV 317
Query: 231 VHLNDAI 237
V + D I
Sbjct: 318 VSMVDRI 324
>ref|YP_011101.1| methyl-accepting chemotaxis protein [Desulfovibrio vulgaris subsp.
vulgaris str. Hildenborough] gi|46449710|gb|AAS96360.1|
methyl-accepting chemotaxis protein [Desulfovibrio
vulgaris subsp. vulgaris str. Hildenborough]
Length = 800
Score = 39.3 bits (90), Expect = 0.17
Identities = 46/187 (24%), Positives = 84/187 (44%), Gaps = 42/187 (22%)
Query: 125 QEQQVAKQYGNSEQGVNQ-LSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSALIKG 183
+ +++A+Q + G Q +S E G AV++ A+ EA+ ++ +
Sbjct: 535 ETEELARQVEHGADGARQQVSCLENTGAAVRQ---LAELLEAAAQHADEA-------VNS 584
Query: 184 SGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSS---------EALGASIGKIREV---- 230
+G E + + I E+ +AAMQR+ G+ S+SS E++G+ +G I ++
Sbjct: 585 AGEAGEKARQGTEIMERSAAAMQRMHGLSMSLSSSMHQLGEQTESIGSILGVINDIADQT 644
Query: 231 --VHLNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCARTHESFN 288
+ LN AI AA+ AG+ GR A + + R+ A T E N
Sbjct: 645 NLLALNAAIEAAR--------AGE--------SGRGFAVVADEVRKLAERTVAATGEVRN 688
Query: 289 QLTDVEE 295
+ +++E
Sbjct: 689 SIRNIQE 695
>gb|AAN57811.1| putative secreted antigen GbpB/SagA; putative peptidoglycan
hydrolase [Streptococcus mutans UA159]
gi|24378550|ref|NP_720505.1| putative secreted antigen
GbpB/SagA; putative peptidoglycan hydrolase
[Streptococcus mutans UA159]
Length = 431
Score = 39.3 bits (90), Expect = 0.17
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>gb|AAK08104.1| immunodominant glycoprotein IDG-60; general stress protein GSP-781
[Streptococcus mutans] gi|15341182|gb|AAK94501.1|
glucan-binding protein B [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.17
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>gb|AAK94504.1| glucan-binding protein B [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.17
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>gb|AAK94502.1| glucan-binding protein B [Streptococcus mutans]
Length = 432
Score = 39.3 bits (90), Expect = 0.17
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>gb|AAK94500.1| glucan-binding protein B [Streptococcus mutans]
Length = 431
Score = 39.3 bits (90), Expect = 0.17
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLQQQEQDKAAVEQKQQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>gb|AAK94503.1| glucan-binding protein B [Streptococcus mutans]
Length = 432
Score = 38.9 bits (89), Expect = 0.23
Identities = 46/197 (23%), Positives = 82/197 (41%), Gaps = 13/197 (6%)
Query: 112 LAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTN 171
+A Q S+ NN+ Q+Q Q + V+ L ++ +A ++ AT + T
Sbjct: 34 IASQDSKINNLTAQQQAAQAQVNTIQGQVSALQTQQAELQAENQRLEAQSATLGQQIQT- 92
Query: 172 DMGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVV 231
SS + + NE + A ++K +AA + + S S + IREVV
Sbjct: 93 ---LSSKIVAR-----NESLKQQARSAQKSNAATSYINAIINSKSVSDAINRVSAIREVV 144
Query: 232 HLNDAIPAAKFIDE-PLEMAGQQNQP---SRIPQGRKMARNINAMAFDTSRSCARTHESF 287
N+ + + D+ +E Q+NQ + +A+N NA+ ++ A
Sbjct: 145 SANEKMLHQQEQDKAAVEQKHQENQAAINTVAANQETIAQNTNALNTQQAQLEAAQLNLQ 204
Query: 288 NQLTDVEEPDLDLVASK 304
+LT ++ LVA K
Sbjct: 205 AELTTAQDQKATLVAQK 221
>ref|XP_647578.1| hypothetical protein DDB0189801 [Dictyostelium discoideum]
gi|60475585|gb|EAL73520.1| hypothetical protein
DDB0189801 [Dictyostelium discoideum]
Length = 1537
Score = 38.5 bits (88), Expect = 0.30
Identities = 31/120 (25%), Positives = 54/120 (44%), Gaps = 12/120 (10%)
Query: 52 FRLRKSQITPEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSK------QQVLPQVL 105
F ++ +TP+ E + ++++ + H N +++ K S+ +Q K QQ Q
Sbjct: 49 FNIQYKDLTPQQKELLKVHNSIVSDLNNHTN-NNNLNKSSSSIQKKPQQQKQQQAQTQQA 107
Query: 106 GTQND-HLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATE 164
TQ HL +Q NN Q+QQ A+Q + Q ++ A Q+Q L Q +
Sbjct: 108 QTQQQQHLHQQQQHLNNQLFQQQQQAQQQAQQQAQQQQAQQQQ----AQQQQALQQQGQQ 163
Score = 34.7 bits (78), Expect = 4.3
Identities = 27/119 (22%), Positives = 49/119 (40%), Gaps = 3/119 (2%)
Query: 57 SQITPEISERVDRVETLINSIIQHQ---NLSSHQGKHSADVQSKQQVLPQVLGTQNDHLA 113
S+I P+ R+ ++ + +Q+Q N Q + Q +QQ Q Q A
Sbjct: 883 SEIAPQSQNRIVAPQSPPSPPVQNQQQQNQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQA 942
Query: 114 KQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTND 172
+Q +Q+ Q+QQ +Q +Q N ++ K Q Q Q + + + N+
Sbjct: 943 QQQAQQQQQAQQQQQQQQQLAAQQQQQNNQQYQQYQQKFYQYQLNLQQQQQQANNNNNN 1001
>emb|CAG07476.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1034
Score = 38.5 bits (88), Expect = 0.30
Identities = 50/251 (19%), Positives = 104/251 (40%), Gaps = 16/251 (6%)
Query: 62 EISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRNN 121
EI ER+ ++ ++ + N + KH ++++ K++ L Q + T+ D L + +
Sbjct: 611 EIDERLKALQKETAALDRRDNELLAEKKHLSELKGKRRQLEQKISTKQDSLRQMEQNITD 670
Query: 122 M-CLQEQQVAKQYGNSEQGVNQL-----SIKEGPGKAVQRQTLCAQATEASGVST---ND 172
+ ++E+ K + Q V + SIK +++ L + E + T ND
Sbjct: 671 LKKIEEETKGKVSEVNSQKVAMVKVFIDSIKLKAKLTMEKVYLALKMVELTAEKTKLEND 730
Query: 173 MGFSSSALIKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALGASIGKIREVVH 232
+S L +++ + A ++E+ M+R + + S ++L + +R
Sbjct: 731 FREGASLLRSMDQKCSQLEQRRAQLTEQCKGHMKRAMSICRMQSKDSLPEDLRNVRLCFF 790
Query: 233 LNDAIPAAKFIDEPLEMAGQQNQPSRIPQGRKMARNINAMAFDTSRSCARTHESFNQL-T 291
AK D P E+ SR+ + R A +++ + R + QL
Sbjct: 791 GGGKQAFAKLPDTPDEI------ESRLSEERSRAECFTSLSENVRMRWFRRDQEIKQLEK 844
Query: 292 DVEEPDLDLVA 302
++EE + L A
Sbjct: 845 ELEEKENALEA 855
>ref|XP_653957.1| hypothetical protein 67.t00003 [Entamoeba histolytica HM-1:IMSS]
gi|56470965|gb|EAL48571.1| hypothetical protein
67.t00003 [Entamoeba histolytica HM-1:IMSS]
Length = 378
Score = 37.4 bits (85), Expect = 0.66
Identities = 31/120 (25%), Positives = 56/120 (45%), Gaps = 15/120 (12%)
Query: 47 DIFAMFRLRKSQITPEISERVDRVETLINSIIQHQNLSSHQGKHSADVQ----SKQQVLP 102
DI L SQ ++ + V ++T +NS+IQ + + K ++ +K VLP
Sbjct: 3 DITPQQYLELSQKIDQLQKMVQSIQTSVNSLIQITEIQNTLPKTIENIVKATFNKYLVLP 62
Query: 103 --------QVLGTQNDHLAKQTSQRNNMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQ 154
+++ +N +++ ++S +QQ KQ S +NQ S+KE P K +Q
Sbjct: 63 EDYDASIQEMIDKKNKNISTRSSSSYKDTEHKQQEVKQ---SSSSINQNSVKEEPKKVLQ 119
>gb|AAG37374.1| ACP36DE [Drosophila melanogaster]
Length = 756
Score = 37.4 bits (85), Expect = 0.66
Identities = 35/161 (21%), Positives = 67/161 (40%), Gaps = 10/161 (6%)
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P +SE + E+ S Q Q+ S Q + +Q++ Q+L Q+ N+ A Q++ +
Sbjct: 84 PPVSESQSQSESQSQSESQKQSQSQSQSQRQQQIQTQLQILRQLQQKSNEQSAAQSASQ- 142
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSAL 180
+Q Q+ + N + S E GK +Q Q Q G+ ++ S++
Sbjct: 143 ---IQSQRQSDSQSNLQLQEQSQSQSE-QGKPIQSQIQILQ-----GLQQKELDDKSASQ 193
Query: 181 IKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALG 221
+ + K + A ++L + +S+S LG
Sbjct: 194 SQSESKTRQEQQKQLNLQLLREAQQKQLEELSSSLSQSRLG 234
>gb|AAG37371.1| ACP36DE [Drosophila melanogaster]
Length = 756
Score = 37.4 bits (85), Expect = 0.66
Identities = 35/161 (21%), Positives = 67/161 (40%), Gaps = 10/161 (6%)
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P +SE + E+ S Q Q+ S Q + +Q++ Q+L Q+ N+ A Q++ +
Sbjct: 84 PPVSESQSQSESQSQSESQKQSQSQSQSQRQQQIQTQLQILRQLQQKSNEQSAAQSASQ- 142
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSAL 180
+Q Q+ + N + S E GK +Q Q Q G+ ++ S++
Sbjct: 143 ---IQSQRQSDSQSNLQLQEQSQSQSE-QGKPIQSQIQILQ-----GLQQKELDDKSASQ 193
Query: 181 IKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALG 221
+ + K + A ++L + +S+S LG
Sbjct: 194 SQSESKTRQEQQKQLNLQLLREAQQKQLEELSSSLSQSRLG 234
>gb|AAG37365.1| ACP36DE [Drosophila melanogaster]
Length = 751
Score = 37.4 bits (85), Expect = 0.66
Identities = 35/161 (21%), Positives = 67/161 (40%), Gaps = 10/161 (6%)
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P +SE + E+ S Q Q+ S Q + +Q++ Q+L Q+ N+ A Q++ +
Sbjct: 84 PPVSESQSQSESQSQSESQKQSQSQSQSQRQQQIQTQLQILRQLQQKSNEQSAAQSASQ- 142
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSAL 180
+Q Q+ + N + S E GK +Q Q Q G+ ++ S++
Sbjct: 143 ---IQSQRQSDSQSNLQLQEQSQSQSE-QGKPIQSQIQILQ-----GLQQKELDDKSASQ 193
Query: 181 IKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALG 221
+ + K + A ++L + +S+S LG
Sbjct: 194 SQSESKTRQEQQKQLNLQLLREAQQKQLEELSSSLSQSRLG 234
>gb|AAG37363.1| ACP36DE [Drosophila melanogaster] gi|11612216|gb|AAG37359.1|
ACP36DE [Drosophila melanogaster]
Length = 756
Score = 37.4 bits (85), Expect = 0.66
Identities = 35/161 (21%), Positives = 67/161 (40%), Gaps = 10/161 (6%)
Query: 61 PEISERVDRVETLINSIIQHQNLSSHQGKHSADVQSKQQVLPQVLGTQNDHLAKQTSQRN 120
P +SE + E+ S Q Q+ S Q + +Q++ Q+L Q+ N+ A Q++ +
Sbjct: 84 PPVSESQSQSESQSQSESQKQSQSQSQSQRQQQIQTQLQILRQLQQKSNEQSAAQSASQ- 142
Query: 121 NMCLQEQQVAKQYGNSEQGVNQLSIKEGPGKAVQRQTLCAQATEASGVSTNDMGFSSSAL 180
+Q Q+ + N + S E GK +Q Q Q G+ ++ S++
Sbjct: 143 ---IQSQRQSDSQSNLQLQEQSQSQSE-QGKPIQSQIQILQ-----GLQQKELDDKSASQ 193
Query: 181 IKGSGNLNEVSHKPALISEKPSAAMQRLIGVFTSMSSEALG 221
+ + K + A ++L + +S+S LG
Sbjct: 194 SQSESKTRQEQQKQLNLQLLREAQQKQLEELSSSLSQSRLG 234
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.130 0.362
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 520,308,011
Number of Sequences: 2540612
Number of extensions: 19693205
Number of successful extensions: 65199
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 103
Number of HSP's that attempted gapping in prelim test: 64726
Number of HSP's gapped (non-prelim): 328
length of query: 346
length of database: 863,360,394
effective HSP length: 129
effective length of query: 217
effective length of database: 535,621,446
effective search space: 116229853782
effective search space used: 116229853782
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0005.1


