

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0627.15
(557 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
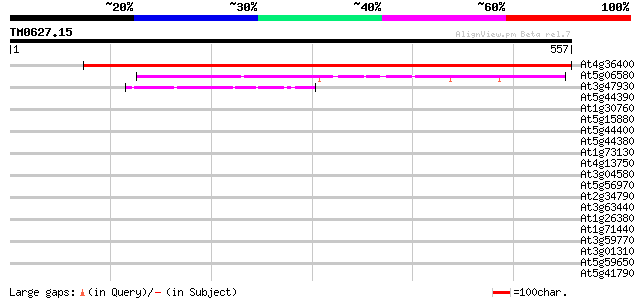
Score E
Sequences producing significant alignments: (bits) Value
At4g36400 unknown protein 758 0.0
At5g06580 unknown protein 147 2e-35
At3g47930 L-galactono-1,4-lactone dehydrogenase - like protein 46 6e-05
At5g44390 berberine bridge enzyme-like protein 34 0.19
At1g30760 putative reticuline oxidase-like protein, 3' partial 34 0.24
At5g15880 unknown protein 33 0.42
At5g44400 berberine bridge enzyme 33 0.54
At5g44380 berberine bridge enzyme-like protein 33 0.54
At1g73130 hypothetical protein 33 0.54
At4g13750 hypothetical protein 32 0.93
At3g04580 putative ethylene receptor 32 0.93
At5g56970 cytokinin oxidase 31 1.6
At2g34790 putative berberine bridge enzyme 31 2.1
At3g63440 cytokinin oxidase -like protein 30 2.7
At1g26380 unknown protein 30 2.7
At1g71440 unknown protein 30 4.6
At3g59770 hypothetical protein 29 6.0
At3g01310 unknown protein 29 6.0
At5g59650 serine/threonine-specific protein kinase - like 29 7.9
At5g41790 myosin heavy chain-like protein 29 7.9
>At4g36400 unknown protein
Length = 559
Score = 758 bits (1956), Expect = 0.0
Identities = 368/485 (75%), Positives = 430/485 (87%), Gaps = 1/485 (0%)
Query: 74 QHKCY-ASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGS 132
Q+KC+ +S L+QRNP FS L+ DV YF+EILG+KNVV+D+E+L T NTDWMHKY+GS
Sbjct: 74 QYKCFGSSAASLIQRNPLFSSLDSKDVSYFKEILGEKNVVEDKERLETANTDWMHKYKGS 133
Query: 133 SKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISF 192
SKL+L P NT +VSQIL+YC+SR LAVVPQGGNTGLVGGSVPVFDEVIV++ MNK++SF
Sbjct: 134 SKLMLLPKNTQEVSQILEYCDSRRLAVVPQGGNTGLVGGSVPVFDEVIVNVGLMNKILSF 193
Query: 193 DKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSL 252
D+VSG+LVCEAGCILEN+ +FLD +GFIMPLDLGAKGSC IGGNVSTNAGGLRL+RYGSL
Sbjct: 194 DEVSGVLVCEAGCILENLATFLDTKGFIMPLDLGAKGSCHIGGNVSTNAGGLRLIRYGSL 253
Query: 253 HGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSV 312
HG+VLG+EAV A+G VLDML TLRKDNTGYDLKHLFIGSEGSLGIVTKV+ILT PKLSSV
Sbjct: 254 HGTVLGLEAVTANGNVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIVTKVSILTQPKLSSV 313
Query: 313 NVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFY 372
N+A +ACKDY CQKLL EAKR LGEILSAFEFLD+ SM+LV+NHL+G RNP+ + +FY
Sbjct: 314 NLAFIACKDYLSCQKLLVEAKRNLGEILSAFEFLDNNSMDLVLNHLDGVRNPVSSSENFY 373
Query: 373 VLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVG 432
+LIETTGSDE++D++KLEAFLL S+E L+SDGV+AQDINQASSFWR+REGITEAL + G
Sbjct: 374 ILIETTGSDETNDREKLEAFLLKSLEKGLVSDGVIAQDINQASSFWRIREGITEALQKAG 433
Query: 433 AVYKYDLSIPLENLYNLVEEMRSRLGNAANVVGYGHLGDGNLHLNISVPQYDDKILSQIE 492
AVYKYDLS+P+E +YN+V ++R RLG+ ANV+GYGHLGDGNLHLNIS +Y+DK+L IE
Sbjct: 434 AVYKYDLSLPVEEIYNIVNDLRGRLGDLANVMGYGHLGDGNLHLNISAAEYNDKLLGLIE 493
Query: 493 PFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
P+VYEWTSKH GSISAEHGLG+MKANEI YSKS ETV LMASIK LLDP ILNPYKVLP
Sbjct: 494 PYVYEWTSKHRGSISAEHGLGVMKANEIFYSKSPETVALMASIKKLLDPKGILNPYKVLP 553
Query: 553 HSLIA 557
HSL +
Sbjct: 554 HSLFS 558
>At5g06580 unknown protein
Length = 567
Score = 147 bits (370), Expect = 2e-35
Identities = 121/437 (27%), Positives = 198/437 (44%), Gaps = 25/437 (5%)
Query: 127 HKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSM 186
HK +++ P + ++VS+ILK CN + +VP GG T + G ++ V + +S M
Sbjct: 140 HKAVNIPDVVVFPRSEEEVSKILKSCNEYKVPIVPYGGATSIEGHTLAPKGGVCIDMSLM 199
Query: 187 NKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRL 246
+V + ++ E G + +L+ G PLD G S IGG +T G
Sbjct: 200 KRVKALHVEDMDVIVEPGIGWLELNEYLEEYGLFFPLDPGPGAS--IGGMCATRCSGSLA 257
Query: 247 VRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTP 306
VRYG++ +V+ ++ VL +G V+ RK GYDL L IGSEG+LG++T++ +
Sbjct: 258 VRYGTMRDNVISLKVVLPNGDVVKTASRARKSAAGYDLTRLIIGSEGTLGVITEITLRLQ 317
Query: 307 --PKLSSVNVA-LLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGAR- 362
P+ S V V KD + A G +S E LD + IN G
Sbjct: 318 KIPQHSVVAVCNFPTVKD----AADVAIATMMSGIQVSRVELLDEVQIR-AINMANGKNL 372
Query: 363 NPLPTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLRE 422
PTL + E G++ + +Q + S N SD + A++ W++R+
Sbjct: 373 TEAPTL-----MFEFIGTEAYTREQTQIVQQIASKHNG--SDFMFAEEPEAKKELWKIRK 425
Query: 423 GITEALMRVGAVYK---YDLSIPLENLYNLVEEMRSRL-GNAANVVGYGHLGDGNLHLNI 478
A + ++ D+ +PL +L L+ + L ++ H GDGN H I
Sbjct: 426 EALWACYAMAPGHEAMITDVCVPLSHLAELISRSKKELDASSLLCTVIAHAGDGNFHTCI 485
Query: 479 SVPQYDD---KILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASI 535
+ + ++ F+ +G+ + EHG+G K + +E +Q M I
Sbjct: 486 MFDPSSEEQRREAERLNHFMVHSALSMDGTCTGEHGVGTGKMKYLEKELGIEALQTMKRI 545
Query: 536 KNLLDPNHILNPYKVLP 552
K LDPN I+NP K++P
Sbjct: 546 KKTLDPNDIMNPGKLIP 562
>At3g47930 L-galactono-1,4-lactone dehydrogenase - like protein
Length = 610
Score = 45.8 bits (107), Expect = 6e-05
Identities = 44/188 (23%), Positives = 87/188 (45%), Gaps = 10/188 (5%)
Query: 116 EKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPV 175
E L TV+ +W +E ++ QP N + ++K + + L + P G +GL + +
Sbjct: 107 EDLHTVS-NWSGTHEVQTRNFNQPENLADLEALVKESHEKKLRIRPVG--SGLSPNGIGL 163
Query: 176 FDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGG 235
+V+L+ M+KV+ DK + +AG ++ ++ + + G + + + QIGG
Sbjct: 164 SRSGMVNLALMDKVLEVDKEKKRVTVQAGIRVQQLVDAIKDYGLTLQ-NFASIREQQIGG 222
Query: 236 NVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSL 295
+ A G R + V+ ++ V +++ + KD +L HL G L
Sbjct: 223 IIQVGAHGTG-ARLPPIDEQVISMKLVTPAKGTIELSR--EKDP---ELFHLARCGLGGL 276
Query: 296 GIVTKVAI 303
G+V +V +
Sbjct: 277 GVVAEVTL 284
>At5g44390 berberine bridge enzyme-like protein
Length = 542
Score = 34.3 bits (77), Expect = 0.19
Identities = 36/137 (26%), Positives = 63/137 (45%), Gaps = 10/137 (7%)
Query: 179 VIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVS 238
V++ LS + ++ K + V EAG + + + + G S IGG+++
Sbjct: 135 VLIDLSKLRQINVDIKDTSAWV-EAGATVGELYYRIAEKSKFHGFPAGVYPSLGIGGHIT 193
Query: 239 TNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFI-GSEGSLGI 297
A G + +YG +VL + V A+G +LD + + G DL GS GS GI
Sbjct: 194 GGAYGSLMRKYGLAADNVLDAKIVDANGKLLD------RASMGEDLFWAIRGGSGGSFGI 247
Query: 298 VT--KVAILTPPKLSSV 312
+ K+ ++ P+ +V
Sbjct: 248 ILSWKIKLVPVPETLTV 264
>At1g30760 putative reticuline oxidase-like protein, 3' partial
Length = 431
Score = 33.9 bits (76), Expect = 0.24
Identities = 34/136 (25%), Positives = 62/136 (45%), Gaps = 8/136 (5%)
Query: 179 VIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVS 238
VIV LS + + IS D S AG + + + + I G S IGG++
Sbjct: 133 VIVDLSKLRQ-ISVDIESNSAWVHAGASIGEVYYRIQEKSKIHGFPAGLCTSLGIGGHII 191
Query: 239 TNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIV 298
A G + ++G +VL V ADG +L+ + ++ + ++ G GS G++
Sbjct: 192 GGAYGSMMRKFGLGADNVLDARIVDADGKILN--RAAMGEDVFWAIRG---GGGGSFGVI 246
Query: 299 T--KVAILTPPKLSSV 312
K+ ++ P++ +V
Sbjct: 247 LAWKIKLVPVPEIVTV 262
>At5g15880 unknown protein
Length = 348
Score = 33.1 bits (74), Expect = 0.42
Identities = 24/153 (15%), Positives = 74/153 (47%), Gaps = 7/153 (4%)
Query: 325 CQKLLQEAKRKLGEILSAFEFLDSQSMNLVI--NHLEGARNPLPTLHDFYVLIETTGSD- 381
C+ ++E + + +L + L++ NL ++ + + + + +I+ G+
Sbjct: 117 CETKIEEKRNEADSLLRKLKELEAVEENLKTEQDNAQASLDARQSKSSSETVIQPDGNGK 176
Query: 382 ESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSI 441
+ +D + +++F+L +E++ ++ + + W + I E ++ + + + ++
Sbjct: 177 DGADTEAMKSFMLEKLESKKNDMSLMEEKVQDLERSWAV---IQERALKQPSPAQREKTL 233
Query: 442 PLENLYNLVEEMRSRLGNAANVVGYGHLGDGNL 474
+ L++L+E++ ++ A +VG HL + L
Sbjct: 234 D-KQLHSLIEQLAAKQAQAEGIVGEIHLNEMEL 265
>At5g44400 berberine bridge enzyme
Length = 537
Score = 32.7 bits (73), Expect = 0.54
Identities = 26/89 (29%), Positives = 47/89 (52%), Gaps = 7/89 (7%)
Query: 226 GAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLK 285
G S IGG+++ A G + +YG +VL + V A+G +LD + ++T + ++
Sbjct: 179 GLCSSLGIGGHITGGAYGSMMRKYGLGADNVLDAKIVDANGKLLD--RAAMGEDTFWAIR 236
Query: 286 HLFIGSEGSLGIVT--KVAILTPPKLSSV 312
G+ GS GI+ K+ ++ PK +V
Sbjct: 237 G---GAGGSFGIILAWKIKLVPVPKTVTV 262
>At5g44380 berberine bridge enzyme-like protein
Length = 541
Score = 32.7 bits (73), Expect = 0.54
Identities = 38/137 (27%), Positives = 60/137 (43%), Gaps = 10/137 (7%)
Query: 179 VIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVS 238
+++ LS + K I+ D S + G L + + + I G S IGG ++
Sbjct: 136 ILLDLSKL-KQINVDIESNSAWVQPGATLGELYYRIAEKSKIHGFPAGLCTSVGIGGYMT 194
Query: 239 TNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG-SLGI 297
G + +YG +VL V+ V A+G +LD + G DL G G S GI
Sbjct: 195 GGGYGTLMRKYGLAGDNVLDVKMVDANGKLLD------RAAMGEDLFWAIRGGGGASFGI 248
Query: 298 VT--KVAILTPPKLSSV 312
V K+ ++ PK +V
Sbjct: 249 VLAWKIKLVPVPKTVTV 265
>At1g73130 hypothetical protein
Length = 646
Score = 32.7 bits (73), Expect = 0.54
Identities = 46/212 (21%), Positives = 85/212 (39%), Gaps = 36/212 (16%)
Query: 42 IAHGEHRVSSLFNPLRFSVKNQQMGVGISTGIQHKCYASMPG--LVQRNPRFSELNDDDV 99
+ E ++++ P +FS +N + G + + + +S+ ++ ++P E D ++
Sbjct: 290 VTDDESQITNTLTPQKFSAENSSVFPGEESVQEVRVESSLSDEEILSKSPLLEEHCDANL 349
Query: 100 RYFQEILGQKNVVQDEEKLSTVNTDWMHKYE-----GSSKLLLQ---------------- 138
+ G+ +V D+E L T NT K+ S LL +
Sbjct: 350 TSTSTLGGEGPIVTDDESLIT-NTLTPQKFSEEEILSKSPLLEEHCDANLTSDTTLGDEK 408
Query: 139 PWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEV-------IVSLSSMNKVIS 191
P TD S I S+ + GN+ + G V +EV +VS S + S
Sbjct: 409 PIITDDESWISNSLTSQKSSA----GNSWVFSGEDSV-EEVKVKSCRDVVSTESQSTQSS 463
Query: 192 FDKVSGILVCEAGCILENIISFLDNEGFIMPL 223
+ ++ C G +L + F DN+ + PL
Sbjct: 464 MESFGTVVKCNDGPVLAALGCFRDNDSSLNPL 495
>At4g13750 hypothetical protein
Length = 2137
Score = 32.0 bits (71), Expect = 0.93
Identities = 32/114 (28%), Positives = 55/114 (48%), Gaps = 16/114 (14%)
Query: 298 VTKVAILTPPKLSSVNV----ALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNL 353
V ++ + KLS NV + + K++ C L +A L LDS S+ +
Sbjct: 1764 VKRLQVFVVDKLSYRNVIPQYGISSKKEFKCSSLLQDKA-------LYTTPSLDSHSLFM 1816
Query: 354 VINHLEGARNPLPTLH--DFYVLIETTGSDESSDKQKLEAFLLHSMENELISDG 405
++ L N +P LH +F LI+T S++Q +E+F+L+S + + DG
Sbjct: 1817 ELSRL--FFNGVPDLHLANFLHLIKTMAESGLSEEQ-MESFILNSQKVHQVPDG 1867
>At3g04580 putative ethylene receptor
Length = 766
Score = 32.0 bits (71), Expect = 0.93
Identities = 29/126 (23%), Positives = 53/126 (42%), Gaps = 4/126 (3%)
Query: 278 DNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQEA-KRKL 336
D + H + E L + K+ I L + A++A + N CQK++ +R +
Sbjct: 325 DQVAVAISHASVLEESQL-MREKLGIQNRALLRAKQNAMMASQARNTCQKVMSHGMRRPM 383
Query: 337 GEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYV--LIETTGSDESSDKQKLEAFLL 394
IL S+SM+L + A T+ + +I+ + D +++ F L
Sbjct: 384 HTILGLLSMFQSESMSLDQKIIVDALMKTSTVLSALINDVIDISPKDNGKSALEVKRFQL 443
Query: 395 HSMENE 400
HS+ E
Sbjct: 444 HSLIRE 449
>At5g56970 cytokinin oxidase
Length = 523
Score = 31.2 bits (69), Expect = 1.6
Identities = 40/201 (19%), Positives = 77/201 (37%), Gaps = 18/201 (8%)
Query: 111 VVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAV-VPQGGNTGLV 169
+ + + TD+ H + +L P + + ++ ++K L+ + G+
Sbjct: 48 LTSSSSSVESAATDFGHVTKIFPSAVLIPSSVEDITDLIKLSFDSQLSFPLAARGHGHSH 107
Query: 170 GGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKG 229
G D V+V++ SM ++ GI V ++ ++L E L+LG
Sbjct: 108 RGQASAKDGVVVNMRSM-----VNRDRGIKVSRTCLYVDVDAAWLWIEVLNKTLELGLTP 162
Query: 230 SC-------QIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGY 282
+GG +S + RYG +VL ++ + G + K +
Sbjct: 163 VSWTDYLYLTVGGTLSNGGISGQTFRYGPQITNVLEMDVITGKGEIATCSKDMNS----- 217
Query: 283 DLKHLFIGSEGSLGIVTKVAI 303
DL +G G GI+T+ I
Sbjct: 218 DLFFAVLGGLGQFGIITRARI 238
>At2g34790 putative berberine bridge enzyme
Length = 532
Score = 30.8 bits (68), Expect = 2.1
Identities = 33/136 (24%), Positives = 57/136 (41%), Gaps = 6/136 (4%)
Query: 179 VIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVS 238
VIV LS + +V D S AG + + + + G S IGG++
Sbjct: 131 VIVDLSKLRQV-DVDLDSNSAWAHAGATIGEVYYRIQEKSQTHGFPAGLCSSLGIGGHLV 189
Query: 239 TNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIV 298
A G + ++G +VL V A+G +LD + ++ + ++ G GS G++
Sbjct: 190 GGAYGSMMRKFGLGADNVLDARIVDANGQILD--RAAMGEDVFWAIRG---GGGGSFGVI 244
Query: 299 TKVAILTPPKLSSVNV 314
I P ++V V
Sbjct: 245 LAWKIKLVPVPATVTV 260
>At3g63440 cytokinin oxidase -like protein
Length = 504
Score = 30.4 bits (67), Expect = 2.7
Identities = 39/199 (19%), Positives = 80/199 (39%), Gaps = 13/199 (6%)
Query: 116 EKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYC---NSRSLAVVPQGGNTGLVGGS 172
E + + D+ ++Y+ +L P + ++ +++ + S V G + G
Sbjct: 42 EHVHHASKDFGNRYQLIPLAVLHPKSVSDIASTIRHIWMMGTHSQLTVAARGRGHSLQGQ 101
Query: 173 VPVFDEVIVSLSSMN----KVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAK 228
+++ + S++ +V S D + + G + NI+ G + P
Sbjct: 102 AQTRHGIVIHMESLHPQKLQVYSVDSPAPYVDVSGGELWINILHETLKYG-LAPKSWTDY 160
Query: 229 GSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLF 288
+GG +S + R+G +V +E V G +L+ K + N+ DL +
Sbjct: 161 LHLTVGGTLSNAGISGQAFRHGPQISNVHQLEIVTGKGEILNCTK---RQNS--DLFNGV 215
Query: 289 IGSEGSLGIVTKVAILTPP 307
+G G GI+T+ I P
Sbjct: 216 LGGLGQFGIITRARIALEP 234
>At1g26380 unknown protein
Length = 535
Score = 30.4 bits (67), Expect = 2.7
Identities = 27/114 (23%), Positives = 48/114 (41%), Gaps = 9/114 (7%)
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+AG L + +D + G + GG++S G + ++G+ V+ E
Sbjct: 146 QAGATLGELYVKIDEASQTLAFPAGICATVGAGGHISGGGYGNLMRKFGTTVDHVIDAEL 205
Query: 262 VLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG-SLGIVT--KVAILTPPKLSSV 312
V +G K L + G DL G G S G++ K+ ++ PK+ +V
Sbjct: 206 VDVNG------KLLNRSTMGEDLFWAIRGGGGASFGVILSWKINLVEVPKIFTV 253
>At1g71440 unknown protein
Length = 531
Score = 29.6 bits (65), Expect = 4.6
Identities = 59/230 (25%), Positives = 92/230 (39%), Gaps = 29/230 (12%)
Query: 182 SLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKG-SCQIGGNVSTN 240
S SS + V S + GI + +A + IS D E + L G + S Q+ G
Sbjct: 68 SQSSASFVRSQNLSRGITLLQALELRYRTISTKDEEDEMYVLSAGNRRVSIQLLGGDKIQ 127
Query: 241 AGGLRLVRYGSLHGSVLGVEA--VLAD-GTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGI 297
R S S LGV + V +D G++L LK L DL I +G
Sbjct: 128 DKLSRFEELTSASLSYLGVSSLGVSSDLGSILPNLKLL-------DLTGNLISDWEEIGA 180
Query: 298 VTKVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINH 357
+ + P L+++N L+C + K L + K L + L
Sbjct: 181 LCEQL----PALTTLN---LSCNSLSSDIKSLPQLKN--------IRVLVLNNSGLSWTQ 225
Query: 358 LEGARNPLPTLHDFYVL---IETTGSDESSDKQKLEAFLLHSMENELISD 404
+E R LP + + +++ I T S SSD Q + L ++++ ISD
Sbjct: 226 VEILRRSLPGIEELHLMGNMISTITSTSSSDDQAFNSLRLLNLDDNCISD 275
>At3g59770 hypothetical protein
Length = 1616
Score = 29.3 bits (64), Expect = 6.0
Identities = 33/117 (28%), Positives = 49/117 (41%), Gaps = 19/117 (16%)
Query: 319 CKDYNCCQKLLQEAK-----------RKLGEILSAF-EFLDSQSMNLVINHLEGARNPLP 366
CK CC ++ EA R+ G + +A E L+ + + NHL P P
Sbjct: 1146 CK--KCCSSIVLEALIVDYVRVMVSLRRSGRVDNAGREALNEVFGSNITNHLAVRGQPSP 1203
Query: 367 TLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGV--LAQDINQASS--FWR 419
DF L + G +ES + +F LH +E S L +N ASS +W+
Sbjct: 1204 NREDFNFLRQILGKEESLSEFPFASF-LHKVETATDSAPFFSLLTPLNLASSNAYWK 1259
>At3g01310 unknown protein
Length = 498
Score = 29.3 bits (64), Expect = 6.0
Identities = 16/52 (30%), Positives = 28/52 (53%)
Query: 244 LRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSL 295
+ ++RY +L S+LG E+++ + + KT D Y + LF +E SL
Sbjct: 337 MNVLRYCNLDESLLGEESLICQNALERLCKTKELDYMSYIVLRLFENTEVSL 388
>At5g59650 serine/threonine-specific protein kinase - like
Length = 892
Score = 28.9 bits (63), Expect = 7.9
Identities = 14/42 (33%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Query: 306 PPKLSSV---NVALLACKDYNCCQKLLQEAKRKLGEILSAFE 344
PPKL N++ + CK NC +L++ ++ L +L+A E
Sbjct: 309 PPKLGVTTIHNLSPVTCKGENCIYQLIKTSRSTLPSLLNALE 350
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 28.9 bits (63), Expect = 7.9
Identities = 30/133 (22%), Positives = 56/133 (41%), Gaps = 10/133 (7%)
Query: 326 QKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESSD 385
Q +QE +LGE+ ++ +S+ +LV H R + + IE++
Sbjct: 26 QTTIQELISELGEMKEKYKEKESEHSSLVELHKTHERESSSQVKELEAHIESS------- 78
Query: 386 KQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPLEN 445
+KL A S+ N +L+Q I + S+ + + + LM K S+
Sbjct: 79 -EKLVADFTQSLNNAEEEKKLLSQKIAELSNEIQEAQNTMQELMSESGQLKESHSVKERE 137
Query: 446 LYNL--VEEMRSR 456
L++L + E+ R
Sbjct: 138 LFSLRDIHEIHQR 150
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,672,207
Number of Sequences: 26719
Number of extensions: 556755
Number of successful extensions: 1373
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 1362
Number of HSP's gapped (non-prelim): 25
length of query: 557
length of database: 11,318,596
effective HSP length: 104
effective length of query: 453
effective length of database: 8,539,820
effective search space: 3868538460
effective search space used: 3868538460
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0627.15


