

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
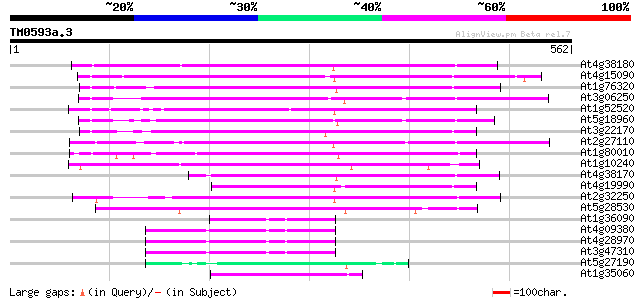
Query= TM0593a.3
(562 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g38180 unknown protein (At4g38180) 249 4e-66
At4g15090 unknown protein 226 3e-59
At1g76320 putative phytochrome A signaling protein 214 8e-56
At3g06250 unknown protein 198 8e-51
At1g52520 F6D8.26 187 1e-47
At5g18960 FAR1 - like protein 185 5e-47
At3g22170 far-red impaired response protein, putative 184 1e-46
At2g27110 Mutator-like transposase 182 4e-46
At1g80010 hypothetical protein 175 7e-44
At1g10240 unknown protein 166 2e-41
At4g38170 hypothetical protein 161 1e-39
At4g19990 putative protein 144 1e-34
At2g32250 Mutator-like transposase 144 1e-34
At5g28530 far-red impaired response protein (FAR1) - like 127 2e-29
At1g36090 hypothetical protein 61 2e-09
At4g09380 putative protein 59 5e-09
At4g28970 putative protein 57 2e-08
At3g47310 putative protein 57 3e-08
At5g27190 putative protein 54 2e-07
At1g35060 hypothetical protein 54 2e-07
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 249 bits (635), Expect = 4e-66
Identities = 133/431 (30%), Positives = 220/431 (50%), Gaps = 8/431 (1%)
Query: 63 IKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDK 122
IK + +TRV C A V+ S KW V F +HNH L P VH + S R ++ K
Sbjct: 139 IKRPRTITRVGCKASLSVKMQ-DSGKWLVSGFVKDHNHELVPPDQVHCLRSHRQISGPAK 197
Query: 123 AQVDSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSY 182
+D+L+ G+ I L+ + GG VGF + D N++ ++ S+ G+ L Y
Sbjct: 198 TLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRNYMRNNRQKSI-EGEIQLLLDY 256
Query: 183 LQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVV 242
L+ +P FF+ S ++++ N+FW D + +D+ FGD + F +TY+ N+Y P
Sbjct: 257 LRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDFTHFGDTVTFDTTYRSNRYRLPFA 316
Query: 243 NFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKV 302
F+G NHH + +F CA + +ET ++ W+ AM P ++ D D +RAA+
Sbjct: 317 PFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAAMSAHPPVSITTDHDAVIRAAIMH 376
Query: 303 VFPNATHRLCGWHIQQNCLEK-----IKIPDFLNEFKTLIYGNFTPERFETKWLQVIEKY 357
VFP A HR C WHI + C EK +K P F ++F + + E FE W +++KY
Sbjct: 377 VFPGARHRFCKWHILKKCQEKLSHVFLKHPSFESDFHKCVNLTESVEDFERCWFSLLDKY 436
Query: 358 GIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYN 417
+ + +W++ +Y R+ W ++R+ FFA + T + INS+ Y+ ++ F
Sbjct: 437 ELRDHEWLQAIYSDRRQWVPVYLRDTFFADMSLTHRSDSINSYFDGYINASTNLSQFFKL 496
Query: 418 FERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALNLN 477
+E+A+E E+ +D+ + PVL T S +E+ AS+L T +F + + L
Sbjct: 497 YEKALESRLEKEVKADYDTMNSPPVLKTP-SPMEKQASELYTRKLFMRFQEELVGTLTFM 555
Query: 478 LVERSEIGTTV 488
+ + G V
Sbjct: 556 ASKADDDGDLV 566
>At4g15090 unknown protein
Length = 768
Score = 226 bits (575), Expect = 3e-59
Identities = 136/478 (28%), Positives = 239/478 (49%), Gaps = 22/478 (4%)
Query: 69 VTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSL 128
V + C A V+ KW + F +HNH L PA HF R + +K +D L
Sbjct: 61 VKKTDCKASMHVK-RRPDGKWIIHEFVKDHNHELLPALAYHFRIQ-RNVKLAEKNNIDIL 118
Query: 129 KLYGVRSCNITGLLMGQKGGHESVG-FLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKG 187
R+ + + Q GG++++G L+ D+ + VDK + ++L GD+ L Y +
Sbjct: 119 HAVSERTKKMYVEMSRQSGGYKNIGSLLQTDVSSQVDKGRYLALEEGDSQVLLEYFKRIK 178
Query: 188 EKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGY 247
+++P FF+ + ++ L NLFW D SR DY +F DV+ F +TY K P+ F G
Sbjct: 179 KENPKFFYAIDLNEDQRLRNLFWADAKSRDDYLSFNDVVSFDTTYVKFNDKLPLALFIGV 238
Query: 248 NHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNA 307
NHH++ + CALV DE++ET+ W++K AM G+ PK ++ D DK + +AV + PN
Sbjct: 239 NHHSQPMLLGCALVADESMETFVWLIKTWLRAMGGRAPKVILTDQDKFLMSAVSELLPNT 298
Query: 308 THRLCGWHIQQNCLEKI---------KIPDFLNEFKTLIYGNFTPERFETKWLQVIEKYG 358
H WH+ LEKI + +FL +F I+ ++T + F+ +W +++ ++G
Sbjct: 299 RHCFALWHV----LEKIPEYFSHVMKRHENFLLKFNKCIFRSWTDDEFDMRWWKMVSQFG 354
Query: 359 IGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNF 418
+ ++W+ ++E RQ W FM + F AG+ T+ E +NSF Y+ K ++ +F+ +
Sbjct: 355 LENDEWLLWLHEHRQKWVPTFMSDVFLAGMSTSQRSESVNSFFDKYIHKKITLKEFLRQY 414
Query: 419 ERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALNLNL 478
++ E +DF + + +P L + S E+ + T +F++ + + + +
Sbjct: 415 GVILQNRYEEESVADFDTCHKQPAL-KSPSPWEKQMATTYTHTIFKKFQVEVLGVVACHP 473
Query: 479 VERSEIGTTVMIKFSICRHPNTQLSSWSFMTRIRTC----LIVIVGFLNMLAYLVLTL 532
+ E + C + L +WS T+ C + GFL A ++L +
Sbjct: 474 RKEKEDENMATFRVQDCEKDDDFLVTWS-KTKSELCCFCRMFEYKGFLCRHALMILQM 530
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 214 bits (546), Expect = 8e-56
Identities = 124/428 (28%), Positives = 210/428 (48%), Gaps = 17/428 (3%)
Query: 71 RVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRG--LTDGDKAQVDSL 128
++ C A V+ KW V F EHNH L P H+ S R L + +++
Sbjct: 65 KIGCKASMHVK-RRPDGKWYVYSFVKEHNHDLLPE-QAHYFRSHRNTELVKSNDSRLRRK 122
Query: 129 KLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGE 188
K + C + + F+ + N DK +R+ L GDA L +L E
Sbjct: 123 KNTPLTDCK-------HLSAYHDLDFIDGYMRNQHDKGRRLVLDTGDAEILLEFLMRMQE 175
Query: 189 KDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYN 248
++P FFF S + L N+FW D DY++F DV+ F ++Y +KY P+V F G N
Sbjct: 176 ENPKFFFAVDFSEDHLLRNVFWVDAKGIEDYKSFSDVVSFETSYFVSKYKVPLVLFVGVN 235
Query: 249 HHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNAT 308
HH + + C L+ D+T+ TY W++++ AM G++PK ++ D + +++AA+ V P
Sbjct: 236 HHVQPVLLGCGLLADDTVYTYVWLMQSWLVAMGGQKPKVMLTDQNNAIKAAIAAVLPETR 295
Query: 309 HRLCGWHIQQNCLEKIKI-----PDFLNEFKTLIYGNFTPERFETKWLQVIEKYGIGEEK 363
H C WH+ + F+ + IY +++ E F+ +WL++I+K+ + +
Sbjct: 296 HCYCLWHVLDQLPRNLDYWSMWQDTFMKKLFKCIYRSWSEEEFDRRWLKLIDKFHLRDVP 355
Query: 364 WIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVE 423
W++ +YE R+ WA FMR FAG+ E +NS YV + S+ +F+ + +E
Sbjct: 356 WMRSLYEERKFWAPTFMRGITFAGLSMRCRSESVNSLFDRYVHPETSLKEFLEGYGLMLE 415
Query: 424 EYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALNLNLVERSE 483
+ E +DF + + P L + S E+ + + +FR + + A +L + SE
Sbjct: 416 DRYEEEAKADFDAWHEAPEL-KSPSPFEKQMLLVYSHEIFRRFQLEVLGAAACHLTKESE 474
Query: 484 IGTTVMIK 491
GTT +K
Sbjct: 475 EGTTYSVK 482
>At3g06250 unknown protein
Length = 764
Score = 198 bits (503), Expect = 8e-51
Identities = 127/476 (26%), Positives = 219/476 (45%), Gaps = 40/476 (8%)
Query: 70 TRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSLK 129
+R+ C A R++ S W V +HNH L P
Sbjct: 247 SRMGCGAYMRIK-RQDSGGWIVDRLNKDHNHDLEP------------------------- 280
Query: 130 LYGVRSCNITGLLMGQKGGHESVGFLK-KDLYNHVDKKKRISLGAGDAVSALSYLQGKGE 188
G ++ + + GG +SV ++ DL NH+ + ++G L Y Q K
Sbjct: 281 --GKKNAGMKKITDDVTGGLDSVDLIELNDLSNHISSTRENTIGKEWYPVLLDYFQSKQA 338
Query: 189 KDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYN 248
+D FF+ + ++FW D SR FGD +VF ++Y+K Y+ P F G+N
Sbjct: 339 EDMGFFYAIELDSNGSCMSIFWADSRSRFACSQFGDAVVFDTSYRKGDYSVPFATFIGFN 398
Query: 249 HHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNAT 308
HH + + ALV DE+ E + W+ + AM G++P+++V D D ++ AV VFP
Sbjct: 399 HHRQPVLLGGALVADESKEAFSWLFQTWLRAMSGRRPRSMVADQDLPIQQAVAQVFPGTH 458
Query: 309 HRLCGWHIQQNCLEKIKIPDFLNEFK----TLIYGNFTPERFETKWLQVIEKYGIGEEKW 364
HR W I+ E+ + F NEFK +Y + T F+T W ++ KYG+ + W
Sbjct: 459 HRFSAWQIRSK--ERENLRSFPNEFKYEYEKCLYQSQTTVEFDTMWSSLVNKYGLRDNMW 516
Query: 365 IKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVEE 424
++++YE R+ W A++R +FF GI + F + + S+ +FI +E+ +E+
Sbjct: 517 LREIYEKREKWVPAYLRASFFGGIHVDGT---FDPFYGTSLNSLTSLREFISRYEQGLEQ 573
Query: 425 YRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALNLNLVERSEI 484
R E DF S +P L T +E+ +L TL +FR + + ++ N ++ E
Sbjct: 574 RREEERKEDFNSYNLQPFLQTK-EPVEEQCRRLYTLTIFRIFQSELAQSYNYLGLKTYEE 632
Query: 485 GTTVMIKFSICRHPNTQLS-SWSFMTRIRTCLIVIVGFLNMLAYLVLTLYVL*ELR 539
G C + N + + ++S +C + + +L +L ++ L ++R
Sbjct: 633 GAISRFLVRKCGNENEKHAVTFSASNLNASCSCQMFEYEGLLCRHILKVFNLLDIR 688
>At1g52520 F6D8.26
Length = 703
Score = 187 bits (476), Expect = 1e-47
Identities = 115/412 (27%), Positives = 190/412 (45%), Gaps = 14/412 (3%)
Query: 60 ISRIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTD 119
I+ + + TR CPA R+R S +W+VV +HNH L + R
Sbjct: 141 INDVNRVRKETRTGCPAMIRMR-QVDSKRWRVVEVTLDHNH-LLGCKLYKSVKRKRKCVS 198
Query: 120 GDKAQVDSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSA 179
+ ++KLY R+C + G S L K N ++L GD+ +
Sbjct: 199 SPVSDAKTIKLY--RACVVDN---GSNVNPNST--LNKKFQNSTGSPDLLNLKRGDSAAI 251
Query: 180 LSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNK 239
+Y +P FF+ + E L N+FW D S++ FGDVI S+Y K+
Sbjct: 252 YNYFCRMQLTNPNFFYLMDVNDEGQLRNVFWADAFSKVSCSYFGDVIFIDSSYISGKFEI 311
Query: 240 PVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAA 299
P+V F+G NHH +TT+ +C + ET+E+Y W+LK M + P+ +V D K + AA
Sbjct: 312 PLVTFTGVNHHGKTTLLSCGFLAGETMESYHWLLKVWLSVM-KRSPQTIVTDRCKPLEAA 370
Query: 300 VKVVFPNATHRLCGWHIQQNCLEKI----KIPDFLNEFKTLIYGNFTPERFETKWLQVIE 355
+ VFP + R HI + EK+ F +Y FE W ++
Sbjct: 371 ISQVFPRSHQRFSLTHIMRKIPEKLGGLHNYDAVRKAFTKAVYETLKVVEFEAAWGFMVH 430
Query: 356 KYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFI 415
+G+ E +W++ +YE R WA ++++ FFAGI E + F + YV + + +F+
Sbjct: 431 NFGVIENEWLRSLYEERAKWAPVYLKDTFFAGIAAAHPGETLKPFFERYVHKQTPLKEFL 490
Query: 416 YNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMR 467
+E A+++ E SD +S+ + E S++ T +MF++ +
Sbjct: 491 DKYELALQKKHREETLSDIESQTLNTAELKTKCSFETQLSRIYTRDMFKKFQ 542
>At5g18960 FAR1 - like protein
Length = 788
Score = 185 bits (470), Expect = 5e-47
Identities = 120/422 (28%), Positives = 203/422 (47%), Gaps = 38/422 (9%)
Query: 70 TRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSLK 129
+R+ C A R++ S W V +HNH L P G K
Sbjct: 268 SRMGCGAYMRIK-RQDSGGWIVDRLNKDHNHDLEP---------------GKKNDA---- 307
Query: 130 LYGVRSCNITGLLMGQKGGHESVGFLKKDLY--NHVDKKKRISLGAGDAVSALSYLQGKG 187
G++ G GG +SV ++ + + NH+ K + +G L Y Q +
Sbjct: 308 --GMKKIPDDGT-----GGLDSVDLIELNDFGNNHIKKTRENRIGKEWYPLLLDYFQSRQ 360
Query: 188 EKDPMFFFKFTRS-GEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSG 246
+D FF+ + ++FW D +R FGD +VF ++Y+K Y+ P G
Sbjct: 361 TEDMGFFYAVELDVNNGSCMSIFWADSRARFACSQFGDSVVFDTSYRKGSYSVPFATIIG 420
Query: 247 YNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPN 306
+NHH + + CA+V DE+ E + W+ + AM G++P+++V D D ++ A+ VFP
Sbjct: 421 FNHHRQPVLLGCAMVADESKEAFLWLFQTWLRAMSGRRPRSIVADQDLPIQQALVQVFPG 480
Query: 307 ATHRLCGWHIQQNCLEKIKIP---DFLNEFKTLIYGNFTPERFETKWLQVIEKYGIGEEK 363
A HR W I++ E + IP +F E++ IY T F++ W +I KYG+ ++
Sbjct: 481 AHHRYSAWQIREKERENL-IPFPSEFKYEYEKCIYQTQTIVEFDSVWSALINKYGLRDDV 539
Query: 364 WIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVE 423
W++++YE R+ W A++R +FFAGI I F + + + +FI +E+A+E
Sbjct: 540 WLREIYEQRENWVPAYLRASFFAGIPINGT---IEPFFGASLDALTPLREFISRYEQALE 596
Query: 424 EYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALNLNLVERSE 483
+ R E DF S +P L T +E+ +L TL +FR ++ + ++ N ++ E
Sbjct: 597 QRREEERKEDFNSYNLQPFLQTK-EPVEEQCRRLYTLTVFRIFQNELVQSYNYLCLKTYE 655
Query: 484 IG 485
G
Sbjct: 656 EG 657
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 184 bits (466), Expect = 1e-46
Identities = 108/402 (26%), Positives = 196/402 (47%), Gaps = 30/402 (7%)
Query: 71 RVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSLKL 130
+ C A V+ KW + F EHNH L PA V + K+
Sbjct: 154 KTDCKASMHVK-RRPDGKWVIHSFVREHNHELLPAQAV---------------SEQTRKI 197
Query: 131 YGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKD 190
Y + Q +++V LK D + +K + +S+ GD L +L +
Sbjct: 198 YAA--------MAKQFAEYKTVISLKSDSKSSFEKGRTLSVETGDFKILLDFLSRMQSLN 249
Query: 191 PMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHH 250
FF+ ++ ++N+FW D SR +Y +F DV+ +TY +NKY P+ F G N H
Sbjct: 250 SNFFYAVDLGDDQRVKNVFWVDAKSRHNYGSFCDVVSLDTTYVRNKYKMPLAIFVGVNQH 309
Query: 251 TETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHR 310
+ + CAL+ DE+ TY W+++ A+ G+ PK ++ + D M + V +FPN H
Sbjct: 310 YQYMVLGCALISDESAATYSWLMETWLRAIGGQAPKVLITELDVVMNSIVPEIFPNTRHC 369
Query: 311 LCGWH----IQQNCLEKIKIPD-FLNEFKTLIYGNFTPERFETKWLQVIEKYGIGEEKWI 365
L WH + +N + +K D F+ +F+ IY + E F KW + + ++G+ +++W+
Sbjct: 370 LFLWHVLMKVSENLGQVVKQHDNFMPKFEKCIYKSGKDEDFARKWYKNLARFGLKDDQWM 429
Query: 366 KQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVEEY 425
+YE R+ WA +M + AG+ T+ + IN+F Y+ K S+ +F+ ++ +++
Sbjct: 430 ISLYEDRKKWAPTYMTDVLLAGMSTSQRADSINAFFDKYMHKKTSVQEFVKVYDTVLQDR 489
Query: 426 RHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMR 467
E +D + +P + + S E+ S++ T +F++ +
Sbjct: 490 CEEEAKADSEMWNKQPAM-KSPSPFEKSVSEVYTPAVFKKFQ 530
>At2g27110 Mutator-like transposase
Length = 851
Score = 182 bits (462), Expect = 4e-46
Identities = 126/486 (25%), Positives = 214/486 (43%), Gaps = 22/486 (4%)
Query: 61 SRIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDG 120
S K +K SC A R+ KW V F EH HGL + +H + R +
Sbjct: 95 SSSKRSKRRLSESCDAMVRIELQGHE-KWVVTKFVKEHTHGLASSNMLHCLRPRRHFANS 153
Query: 121 DKAQVDSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSAL 180
+K+ N+ +M S G + + + ++G DA + L
Sbjct: 154 EKSSYQE-------GVNVPSGMMYVSMDANSRGARNASMATNTKR----TIGR-DAHNLL 201
Query: 181 SYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKP 240
Y + ++P FF+ + + N+FW D SR+ Y FGD + + Y+ N++ P
Sbjct: 202 EYFKRMQAENPGFFYAVQLDEDNQMSNVFWADSRSRVAYTHFGDTVTLDTRYRCNQFRVP 261
Query: 241 VVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAV 300
F+G NHH + +F CAL+ DE+ ++ W+ K AM + P ++V D D++++ A
Sbjct: 262 FAPFTGVNHHGQAILFGCALILDESDTSFIWLFKTFLTAMRDQPPVSLVTDQDRAIQIAA 321
Query: 301 KVVFPNATHRLCGWHIQQNCLEK-----IKIPDFLNEFKTLIYGNFTPERFETKWLQVIE 355
VFP A H + W + + EK + P F E I T E FE+ W VI+
Sbjct: 322 GQVFPGARHCINKWDVLREGQEKLAHVCLAYPSFQVELYNCINFTETIEEFESSWSSVID 381
Query: 356 KYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFI 415
KY +G +W+ +Y R W + R++FFA + + G SF YV + ++ F
Sbjct: 382 KYDLGRHEWLNSLYNARAQWVPVYFRDSFFAAVFPSQGYSG--SFFDGYVNQQTTLPMFF 439
Query: 416 YNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALN 475
+ERA+E + E+ +D + PVL T S +E A+ L T +F + + +
Sbjct: 440 RLYERAMESWFEMEIEADLDTVNTPPVLKTP-SPMENQAANLFTRKIFGKFQEELVETFA 498
Query: 476 LNLVERSEIGTTVMIKFSICRHPN-TQLSSWSFMTRIRTCLIVIVGFLNMLAYLVLTLYV 534
+ GTT + + + N + ++ + C + +L VLT++
Sbjct: 499 HTANRIEDDGTTSTFRVANFENDNKAYIVTFCYPEMRANCSCQMFEHSGILCRHVLTVFT 558
Query: 535 L*ELRT 540
+ + T
Sbjct: 559 VTNILT 564
>At1g80010 hypothetical protein
Length = 696
Score = 175 bits (443), Expect = 7e-44
Identities = 125/423 (29%), Positives = 197/423 (46%), Gaps = 27/423 (6%)
Query: 61 SRIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPA------AHVHFIPSF 114
SR KE TR C A R+R +WKV + +HNH P +H S
Sbjct: 127 SRRKE----TRTGCQAMIRLRL-IHFDRWKVDQVKLDHNHSFDPQRAHNSKSHKKSSSSA 181
Query: 115 RGLTDGDK-----AQVDSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRI 169
T + QV ++KLY + + L G S G +H +R+
Sbjct: 182 SPATKTNPEPPPHVQVRTIKLYRTLALDTPPAL----GTSLSSGETSDLSLDHFQSSRRL 237
Query: 170 SLGAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFY 229
L G + Q + P F + + + +L N+FW D +R Y FGDV++F
Sbjct: 238 ELRGGFRALQDFFFQIQ-LSSPNFLYLMDLADDGSLRNVFWIDARARAAYSHFGDVLLFD 296
Query: 230 STYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVV 289
+T N Y P+V F G NHH +T + C L+ D++ ETY W+ +A M G+ P+ +
Sbjct: 297 TTCLSNAYELPLVAFVGINHHGDTILLGCGLLADQSFETYVWLFRAWLTCMLGRPPQIFI 356
Query: 290 IDGDKSMRAAVKVVFPNATHRLCGWHIQQN-CLEKIKIPD---FLNEFKTLIYGNFTPER 345
+ K+MR AV VFP A HRL H+ N C +++ D F ++YG E
Sbjct: 357 TEQCKAMRTAVSEVFPRAHHRLSLTHVLHNICQSVVQLQDSDLFPMALNRVVYGCLKVEE 416
Query: 346 FETKWLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKS-Y 404
FET W ++I ++G+ + I+ M++ R++WA ++++ F AG T L FI S Y
Sbjct: 417 FETAWEEMIIRFGMTNNETIRDMFQDRELWAPVYLKDTFLAGALTFPLGNVAAPFIFSGY 476
Query: 405 VLCKNSILDFIYNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFR 464
V S+ +F+ +E +++ E D +S P L T + E +K+ T+ +FR
Sbjct: 477 VHENTSLREFLEGYESFLDKKYTREALCDSESLKLIPKLKTTHPY-ESQMAKVFTMEIFR 535
Query: 465 EMR 467
+
Sbjct: 536 RFQ 538
>At1g10240 unknown protein
Length = 680
Score = 166 bits (421), Expect = 2e-41
Identities = 107/431 (24%), Positives = 200/431 (45%), Gaps = 29/431 (6%)
Query: 60 ISRIKEAKPV-----TRVSCPAKFRVRFHAK--SSKWKVVLFEPEHNHGLTPAAHVHFIP 112
I + E KP +R C A R+ + S++W+V F HNH L V F+P
Sbjct: 105 IKTLSEGKPQRNRRSSRCGCQAYLRISKLTELGSTEWRVTGFANHHNHELLEPNQVRFLP 164
Query: 113 SFRGLTDGDKAQVDSLKLYGVRSCNITGLLMGQKGGHES-VGFLKKDLYNHVDKKKRISL 171
++R ++D DK+++ G+ + LL +K + F +KD+ N + K++
Sbjct: 165 AYRSISDADKSRILMFSKTGISVQQMMRLLELEKCVEPGFLPFTEKDVRNLLQSFKKLD- 223
Query: 172 GAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYST 231
+ + L Q EKDP F F+FT + LEN+ W +S Y+ FGD +VF +T
Sbjct: 224 PEDENIDFLRMCQSIKEKDPNFKFEFTLDANDKLENIAWSYASSIQSYELFGDAVVFDTT 283
Query: 232 YKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVID 291
++ + P+ + G N++ F C L+ DE + ++ W L+A M GK P+ ++ D
Sbjct: 284 HRLSAVEMPLGIWVGVNNYGVPCFFGCVLLRDENLRSWSWALQAFTGFMNGKAPQTILTD 343
Query: 292 GDKSMRAAVKVVFPNATHRLCGWHIQQNCLE--KIKIPDFLNEFKTLIYGNF---TPERF 346
+ ++ A+ P H LC W + + + N++K Y + + E F
Sbjct: 344 HNMCLKEAIAGEMPATKHALCIWMVVGKFPSWFNAGLGERYNDWKAEFYRLYHLESVEEF 403
Query: 347 ETKWLQVIEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVL 406
E W ++ +G+ + I +Y +R +W+ ++R +F AG+ T + IN+FI+ ++
Sbjct: 404 ELGWRDMVNSFGLHTNRHINNLYASRSLWSLPYLRSHFLAGMTLTGRSKAINAFIQRFLS 463
Query: 407 CKNSILDFIYNF-------ERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLT 459
+ + F+ ++A E+ + + + G P +E A+ +LT
Sbjct: 464 AQTRLAHFVEQVAVVVDFKDQATEQQTMQQNLQNISLKTGAP--------MESHAASVLT 515
Query: 460 LNMFREMRHMI 470
F +++ +
Sbjct: 516 PFAFSKLQEQL 526
>At4g38170 hypothetical protein
Length = 531
Score = 161 bits (407), Expect = 1e-39
Identities = 90/319 (28%), Positives = 163/319 (50%), Gaps = 13/319 (4%)
Query: 180 LSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNK-YN 238
L+YL+ + ++P F + E++ N+FW D T RL+Y FGD +VF +TY++ K Y
Sbjct: 8 LNYLKRRQLENPGFLYAI----EDDCGNVFWADPTCRLNYTYFGDTLVFDTTYRRGKRYQ 63
Query: 239 KPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRA 298
P F+G+NHH + +F CAL+ +E+ ++ W+ + +AM P ++ ++ D+ ++
Sbjct: 64 VPFAAFTGFNHHGQPVLFGCALILNESESSFAWLFQTWLQAMSAPPPPSITVEPDRLIQV 123
Query: 299 AVKVVFPNATHRLCGWHIQQNCLEKIKI-----PDFLNEFKTLIYGNFTPERFETKWLQV 353
AV VF R I + EK+ P F +EF + T FE W +
Sbjct: 124 AVSRVFSQTRLRFSQPLIFEETEEKLAHVFQAHPTFESEFINCVTETETAAEFEASWDSI 183
Query: 354 IEKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILD 413
+ +Y + + W++ +Y RQ W F+R+ F+ + T +NSF + +V ++
Sbjct: 184 VRRYYMEDNDWLQSIYNARQQWVRVFIRDTFYGELSTNEGSSILNSFFQGFVDASTTMQM 243
Query: 414 FIYNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRA 473
I +E+A++ +R EL +D+++ PV+ T S +E+ A+ L T F + +
Sbjct: 244 LIKQYEKAIDSWREKELKADYEATNSTPVMKTP-SPMEKQAASLYTRAAFIKFQEEFVET 302
Query: 474 LNL--NLVERSEIGTTVMI 490
L + N++ S TT +
Sbjct: 303 LAIPANIISDSGTHTTYRV 321
>At4g19990 putative protein
Length = 672
Score = 144 bits (364), Expect = 1e-34
Identities = 76/274 (27%), Positives = 145/274 (52%), Gaps = 13/274 (4%)
Query: 203 ENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVC 262
++L N+FW D R DY F DV+ +T+ KN+Y P+V F+G NHH + + L+
Sbjct: 184 QSLRNIFWVDAKGRFDYTCFSDVVSIDTTFIKNEYKLPLVAFTGVNHHGQFLLLGFGLLL 243
Query: 263 -DETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCL 321
DE+ + W+ +A +AM G +P+ ++ D+ ++ AV VFP++ H W
Sbjct: 244 TDESKSGFVWLFRAWLKAMHGCRPRVILTKHDQMLKEAVLEVFPSSRHCFYMWDTLGQMP 303
Query: 322 EKI--------KIPDFLNEFKTLIYGNFTPERFETKWLQVIEKYGIGEEKWIKQMYETRQ 373
EK+ K+ D +N+ IYG+ E FE W +V++++ + + W++ +YE R+
Sbjct: 304 EKLGHVIRLEKKLVDEIND---AIYGSCQSEDFEKNWWEVVDRFHMRDNVWLQSLYEDRE 360
Query: 374 MWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFERAVEEYRHNELASD 433
W +M++ AG+ T + +NS + Y+ K + F+ +++ ++E E S+
Sbjct: 361 YWVPVYMKDVSLAGMCTAQRSDSVNSGLDKYIQRKTTFKAFLEQYKKMIQERYEEEEKSE 420
Query: 434 FKSRYGEPVLITALSHIEQGASKLLTLNMFREMR 467
++ Y +P L + S + +++ T MF++ +
Sbjct: 421 IETLYKQPGL-KSPSPFGKQMAEVYTREMFKKFQ 453
>At2g32250 Mutator-like transposase
Length = 684
Score = 144 bits (363), Expect = 1e-34
Identities = 97/437 (22%), Positives = 183/437 (41%), Gaps = 50/437 (11%)
Query: 64 KEAKPVTRVSCPAK-FRVRFHAK---SSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTD 119
++A + SCP + H K KW + F EHNH + P
Sbjct: 94 EKATAINPRSCPKTGCKAGLHMKRKEDEKWVIYNFVKEHNHEICP--------------- 138
Query: 120 GDKAQVDSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSA 179
+ +G ++ G L + K +++L D
Sbjct: 139 -------------------DDFYVSVRGKNKPAGALA------IKKGLQLALEEEDLKLL 173
Query: 180 LSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNK 239
L + +K P FF+ ++ + N+FW D ++ DY +F DV++F + Y +N Y
Sbjct: 174 LEHFMEMQDKQPGFFYAVDFDSDKRVRNVFWLDAKAKHDYCSFSDVVLFDTFYVRNGYRI 233
Query: 240 PVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAA 299
P F G +HH + + CAL+ + + TY W+ + +A+ G+ P ++ D DK +
Sbjct: 234 PFAPFIGVSHHRQYVLLGCALIGEVSESTYSWLFRTWLKAVGGQAPGVMITDQDKLLSDI 293
Query: 300 VKVVFPNATHRLCGWHIQQNCLEKI-----KIPDFLNEFKTLIYGNFTPERFETKWLQVI 354
V VFP+ H C W + E + + F+ F + ++T E FE +W +I
Sbjct: 294 VVEVFPDVRHIFCLWSVLSKISEMLNPFVSQDDGFMESFGNCVASSWTDEHFERRWSNMI 353
Query: 355 EKYGIGEEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDF 414
K+ + E +W++ ++ R+ W + AG+ I S Y+ + + DF
Sbjct: 354 GKFELNENEWVQLLFRDRKKWVPHYFHGICLAGLSGPERSGSIASHFDKYMNSEATFKDF 413
Query: 415 IYNFERAVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRAL 474
+ + ++ E D + + +P L ++L+ E+ S + T F++ + + +
Sbjct: 414 FELYMKFLQYRCDVEAKDDLEYQSKQPTLRSSLA-FEKQLSLIYTDAAFKKFQAEVPGVV 472
Query: 475 NLNLVERSEIGTTVMIK 491
+ L + E GTT + +
Sbjct: 473 SCQLQKEREDGTTAIFR 489
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 127 bits (319), Expect = 2e-29
Identities = 99/400 (24%), Positives = 172/400 (42%), Gaps = 24/400 (6%)
Query: 87 SKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSLKLYGVRSCNITGLLMGQK 146
S W V F HNH L V +P++R + D+ ++ L G I LL +K
Sbjct: 159 SHWYVSQFSNVHNHELLEDDQVRLLPAYRKIQQSDQERILLLSKAGFPVNRIVKLLELEK 218
Query: 147 GGHES-VGFLKKDLYNHVDKKKR---------ISLGAGDAVSALSYLQGKGEKDPMFFFK 196
G + F++KD+ N V K+ D + L +G E+D F +
Sbjct: 219 GVVSGQLPFIEKDVRNFVRACKKSVQENDAFMTEKRESDTLELLECCKGLAERDMDFVYD 278
Query: 197 FTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIF 256
T + +EN+ W G S Y FGDV+VF ++Y+ Y + F G +++ + +
Sbjct: 279 CTSDENQKVENIAWAYGDSVRGYSLFGDVVVFDTSYRSVPYGLLLGVFFGIDNNGKAMLL 338
Query: 257 ACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHI 316
C L+ DE+ ++ W L+ M G+ P+ ++ D D ++ A+ PN H + HI
Sbjct: 339 GCVLLQDESCRSFTWALQTFVRFMRGRHPQTILTDIDTGLKDAIGREMPNTNHVVFMSHI 398
Query: 317 QQNCLEKIK--IPDFLNEFKT---LIYGNFTPERFETKWLQVIEKYGIGEEKWIKQMYET 371
+ EF+ ++ + FE +W ++ ++G+ ++ +Y
Sbjct: 399 VSKLASWFSQTLGSHYEEFRAGFDMLCRAGNVDEFEQQWDLLVTRFGLVPDRHAALLYSC 458
Query: 372 RQMWATAFMRENFFAGIRTTSLCEGINSFIKSYV---LCKNSILDFIYNFERAVEEYRHN 428
R W +RE+F A T+ I+SF+K V C +L+ E A++
Sbjct: 459 RASWLPCCIREHFVAQTMTSEFNLSIDSFLKRVVDGATCMQLLLE-----ESALQVSAAA 513
Query: 429 ELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRH 468
LA R+ P L T + +E A +LT F +++
Sbjct: 514 SLAKQILPRFTYPSLKTCMP-MEDHARGILTPYAFSVLQN 552
>At1g36090 hypothetical protein
Length = 645
Score = 60.8 bits (146), Expect = 2e-09
Identities = 41/129 (31%), Positives = 67/129 (51%), Gaps = 6/129 (4%)
Query: 201 GEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGY--NHHTETTIFAC 258
GE+ + LF G ++A VIV +T+ K Y +V + + NHH F
Sbjct: 346 GEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLVIATAHDPNHHHYPLAFG- 404
Query: 259 ALVCDETIETYKWVLKALDEAMFGKQPKAVVI-DGDKSMRAAVKVVFPNATHRLCGWHIQ 317
++ E ++ W L+ L + ++ P+ V I D +S++ AVK V+PNA H C WH+
Sbjct: 405 -IIDSEKDVSWIWFLEKL-KTVYSDVPRLVFISDRHQSIKKAVKTVYPNALHAACIWHLC 462
Query: 318 QNCLEKIKI 326
QN +++KI
Sbjct: 463 QNMRDRVKI 471
>At4g09380 putative protein
Length = 960
Score = 59.3 bits (142), Expect = 5e-09
Identities = 51/193 (26%), Positives = 87/193 (44%), Gaps = 8/193 (4%)
Query: 137 NITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFK 196
NI ++ G+ G H S + HV + + + + Y+ K + + +
Sbjct: 324 NIMSIVRGRLGVHCSYSTALRGKMLHVSDVRGTPERSYTMLFSYLYMLEKVNPGTVTYVE 383
Query: 197 FTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGY--NHHTETT 254
GE+ + LF G ++A VIV +T+ K Y +V + NHH
Sbjct: 384 L--EGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPL 441
Query: 255 IFACALVCDETIETYKWVLKALDEAMFGKQPKAVVI-DGDKSMRAAVKVVFPNATHRLCG 313
F ++ E ++ W L+ L + ++ P V I D +S++ AVK V+PNA H C
Sbjct: 442 AFG--IIDSEKDVSWIWFLENL-KTVYSDVPGLVFISDRHQSIKKAVKTVYPNALHAACI 498
Query: 314 WHIQQNCLEKIKI 326
WH+ QN +++KI
Sbjct: 499 WHLCQNMRDRVKI 511
>At4g28970 putative protein
Length = 914
Score = 57.4 bits (137), Expect = 2e-08
Identities = 50/193 (25%), Positives = 86/193 (43%), Gaps = 8/193 (4%)
Query: 137 NITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFK 196
NI ++ G+ G H S + HV + + + + Y+ K + + +
Sbjct: 324 NIMSIVRGRLGVHCSYSTALRGKMLHVSDVRGTPERSYTMLFSYLYMLEKVNPGTVTYVE 383
Query: 197 FTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGY--NHHTETT 254
GE+ + LF G ++A VIV +T+ K Y +V + NHH
Sbjct: 384 L--EGEKKFKYLFIALGACIEGFRAMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPL 441
Query: 255 IFACALVCDETIETYKWVLKALDEAMFGKQPKAVVI-DGDKSMRAAVKVVFPNATHRLCG 313
F ++ E ++ W L+ L + ++ P V I D +S++ AVK V+PNA H C
Sbjct: 442 AFG--IIDSEKDVSWIWFLEKL-KTVYSDVPGLVFISDRHQSIKKAVKTVYPNALHAACI 498
Query: 314 WHIQQNCLEKIKI 326
WH+ QN +++ I
Sbjct: 499 WHLCQNMRDRVTI 511
>At3g47310 putative protein
Length = 735
Score = 57.0 bits (136), Expect = 3e-08
Identities = 49/193 (25%), Positives = 85/193 (43%), Gaps = 8/193 (4%)
Query: 137 NITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFK 196
NI ++ G+ G H S + HV + + + + Y+ K + + +
Sbjct: 273 NIMSIVRGRLGVHCSYSTALRGKMLHVSDVRGTPERSYTMLFSYLYMLEKVNPGTVTYVE 332
Query: 197 FTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGY--NHHTETT 254
GE+ + LF G ++ VIV +T+ K Y +V + NHH
Sbjct: 333 L--EGEKKFKYLFIALGACIEGFRTMRKVIVVDATHLKTVYGGMLVIATAQDPNHHHYPL 390
Query: 255 IFACALVCDETIETYKWVLKALDEAMFGKQPKAVVI-DGDKSMRAAVKVVFPNATHRLCG 313
F ++ E ++ W L+ L + ++ P V I D +S++ VK V+PNA H C
Sbjct: 391 AFG--IIDSENDVSWIWFLEKL-KTVYSDVPGLVFISDRHQSIKKVVKTVYPNALHAACI 447
Query: 314 WHIQQNCLEKIKI 326
WH+ QN +++KI
Sbjct: 448 WHLCQNMRDRVKI 460
>At5g27190 putative protein
Length = 779
Score = 54.3 bits (129), Expect = 2e-07
Identities = 58/267 (21%), Positives = 105/267 (38%), Gaps = 30/267 (11%)
Query: 137 NITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFK 196
+I + G+ G H S DLY+++ K ++ G +SY+ E D FK
Sbjct: 325 SIMSIARGRMGLHCSPEKSYTDLYSYLHVLKEVNPGT------ISYV----EVDAQQKFK 374
Query: 197 FTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIF 256
+ LF G ++ V++ +T+ K Y +V + + + I
Sbjct: 375 Y----------LFVALGACIEGFKVMRKVVIVDATFLKTVYGGMLVFATAQDPNHHNYII 424
Query: 257 ACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHI 316
A A++ E ++ W L + V D +S+ ++ VFPNA H C WH+
Sbjct: 425 ASAVIDRENDASWSWFFNKLKTVIPDVPGLVFVSDRHQSIIKSIMQVFPNARHGHCVWHL 484
Query: 317 QQNCLEKIKIPDFLNEFKTL----IYGNFTPERFETKWLQVIEKYGIGEEKWIKQMYETR 372
QN ++K + +Y F R ++ + G ++W
Sbjct: 485 SQNVKVRVKTEKEEAAANFIACAHVYTQFEFTREYARFRRRFPNVGPYLDRWAPV----- 539
Query: 373 QMWATAFMRENFFAGIRTTSLCEGINS 399
+ WA + + + I T++ CE +NS
Sbjct: 540 ENWAMCYFEGDRY-NIDTSNACESLNS 565
>At1g35060 hypothetical protein
Length = 873
Score = 54.3 bits (129), Expect = 2e-07
Identities = 38/152 (25%), Positives = 66/152 (43%), Gaps = 1/152 (0%)
Query: 202 EENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALV 261
+E +F G S ++ V+V T+ K KY ++ SG + + + A A+V
Sbjct: 502 KERFLYMFLAFGASIQGFKHLRRVLVVDGTHLKGKYKGVLLTASGQDANFQVYPLAFAVV 561
Query: 262 CDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRAAVKVVFPNATHRLCGWHIQQNCL 321
E + + W L+ + ++ D +S++ VK VFP A H C H+ +N
Sbjct: 562 DSENDDAWTWFFTKLERIIADNNTLTILSDRHESIKVGVKKVFPQAHHGACIIHLCRNIQ 621
Query: 322 EKIKIPDFLNEFKTLIYGNFTPERFETKWLQV 353
+ K K Y FT +F+T + Q+
Sbjct: 622 ARFKNRGLTQLVKNAGY-EFTSGKFKTLYNQI 652
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.331 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,057,168
Number of Sequences: 26719
Number of extensions: 508054
Number of successful extensions: 1589
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 1474
Number of HSP's gapped (non-prelim): 93
length of query: 562
length of database: 11,318,596
effective HSP length: 104
effective length of query: 458
effective length of database: 8,539,820
effective search space: 3911237560
effective search space used: 3911237560
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0593a.3


