

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.9
(1706 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
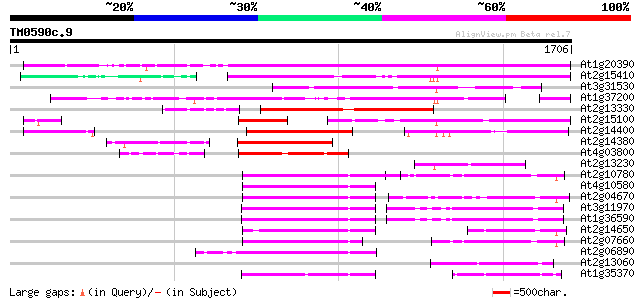
Score E
Sequences producing significant alignments: (bits) Value
At1g20390 hypothetical protein 1060 0.0
At2g15410 putative retroelement pol polyprotein 825 0.0
At3g31530 hypothetical protein 568 e-161
At1g37200 hypothetical protein 560 e-159
At2g13330 F14O4.9 463 e-130
At2g15100 putative retroelement pol polyprotein 387 e-107
At2g14400 putative retroelement pol polyprotein 386 e-107
At2g14380 putative retroelement pol polyprotein 357 3e-98
At4g03800 325 1e-88
At2g13230 pseudogene 231 3e-60
At2g10780 pseudogene 231 4e-60
At4g10580 putative reverse-transcriptase -like protein 224 3e-58
At2g04670 putative retroelement pol polyprotein 221 2e-57
At3g11970 hypothetical protein 213 8e-55
At1g36590 hypothetical protein 213 8e-55
At2g14650 putative retroelement pol polyprotein 213 1e-54
At2g07660 putative retroelement pol polyprotein 212 1e-54
At2g06890 putative retroelement integrase 188 2e-47
At2g13060 pseudogene 184 4e-46
At1g35370 hypothetical protein 180 6e-45
>At1g20390 hypothetical protein
Length = 1791
Score = 1060 bits (2740), Expect = 0.0
Identities = 653/1722 (37%), Positives = 934/1722 (53%), Gaps = 144/1722 (8%)
Query: 41 PFSEDVENVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKM----VIIAASDAVKCRMFPS 96
PF+ + NV+I K + L+SY+G +DPK+ L FN + + I DA +C++F
Sbjct: 156 PFTRRITNVSIRGAQK-IKLESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIE 214
Query: 97 TFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYM 156
A WF+ L SI +F +S FL ++ + DL++I Q ESL+ ++
Sbjct: 215 HLTGPAHNWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFV 274
Query: 157 ARYS--AASVKVEDEEPRACALAFKNGL-LPGGLNSKLTRKPARSMGEMRARASTYILDE 213
R+ ++ V DE A +A +N + +T ++ + RAS +I E
Sbjct: 275 DRFKLVVTNITVPDE---AAIVALRNAVWYDSRFRDDITLHAPSTLEDALHRASRFIELE 331
Query: 214 EDDAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTKEQLYPRRDDY 273
E+ ++ K V +D + +Q + KPT + D
Sbjct: 332 EEKLILARKHNSTKTPACKDAVVIKVGPDDSNEPRQHLDRNPSAGRKPTSFLVSTETPD- 390
Query: 274 EQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEA-PKYQPRDANPKKWCEF 332
+PW N + D D P A P Y CE+
Sbjct: 391 --AKPW---------------NKYIRD----------ADSPAAGPMY----------CEY 413
Query: 333 HRSAGHDTDDCRTLQREIDKLIRAGYQ-GNRQGQWRNGGDQNKAHKREEERA------DT 385
H+S H T++CR LQ L+ A Y+ G + NK +R E A T
Sbjct: 414 HKSRAHSTENCRFLQG----LLMAKYKSGGITIECDRPPINNKNQRRNETTARQYLNDQT 469
Query: 386 KGKKKQESAAIATKGADDTFA--QHSGPPVGT-------INTIAGGFGGGGDTHAARKRH 436
K E I + ADD A Q +G + ++ I GG D+ + K++
Sbjct: 470 KPPTPAEQGIITS--ADDPAAKRQRNGKAIAAEPVVVRQVHVIMGGLQNCSDSVRSIKQY 527
Query: 437 VRAVNSVHEVAFGFV-----HPD----ITISMADFEGIKPHKDDPIVVQLRMNSFNVRRV 487
+ V VA+ +P+ I+ + D EG+ +DP+VV+L ++ V RV
Sbjct: 528 RKKAEMV--VAWPSSTSTTRNPNQSAPISFTDVDLEGLDTPHNDPLVVELIISDSRVTRV 585
Query: 488 LLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECAR 547
L+D GSS D+I+ D + +TD + P + L GF G+ VM G I L G
Sbjct: 586 LIDTGSSVDLIFKDVLTAMNITDRQIKPVSKPLAGFDGDFVMTIGTIKLPIFVG----GL 641
Query: 548 VLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGKLKVDQKMAREC 607
+ V+++V+ A YNVI+G ++++ A+ ST H VK+P
Sbjct: 642 IAWVKFVVIGKPAVYNVILGTPWIHQMQAIPSTYHQCVKFPT------------------ 683
Query: 608 YNNCLNFYGKKSALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGDRTLKIGT 667
+N K A R YE + L ++ P R + +G
Sbjct: 684 HNGIFTLRAPKEAKTPSRSYE----ESELCRTEMVNIDESDPT----------RCVGVGA 729
Query: 668 RLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDK 727
++ L LL N FAWS +DM GIDP H L ++P+ KPV Q RR+LG ++
Sbjct: 730 EISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNVDPTFKPVKQKRRKLGPER 789
Query: 728 GKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLP 787
+AV +EV+KLL A I EVKYP WLAN V+VKK NGKWR+CVDYTDLNKACPKDSYPLP
Sbjct: 790 ARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLP 849
Query: 788 SIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGA 847
ID LV+ SGN LLS MDA+SGY+QI MH D++KT+F+T R YCY+ M FGLKNAGA
Sbjct: 850 HIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTYCYKVMSFGLKNAGA 909
Query: 848 TYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSF 907
TYQR ++++ A Q+GR +EVY+DDM+VKS++ DH + L + F + + M+LNP KC+F
Sbjct: 910 TYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCTF 969
Query: 908 GVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPQSGDRS 967
GV G+FLG+++T RGIE NP++ +AI ++ SP N +EVQRLTGRIAAL+RF+ +S D+
Sbjct: 970 GVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFISRSTDKC 1029
Query: 968 FPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVM 1027
PF+ L++ F+W + EEAF +LK+ LS+PPIL KP G L+LY AVSD A+SSV+
Sbjct: 1030 LPFYNLLKRRAQFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAVSSVL 1089
Query: 1028 LQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDLPLR 1087
++E GE R +++ S +L AE RY IEKAALAV+ +AR+LRPYFQS + V TD PLR
Sbjct: 1090 VREDRGEQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTDQPLR 1149
Query: 1088 QVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELTPDRFERV-----DTQ 1142
L P SGR+ W+VELSEY + + R + +Q LADF++EL ER +
Sbjct: 1150 VALHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQVLADFLIELPLQSAERAVSGNRGEE 1209
Query: 1143 WTLFVDGSSNSSGSGAGVTLEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKI 1202
W+L+VDGSS++ GSG G+ L P VLEQS + F ATNN AEYE LIAGL+LA ++I
Sbjct: 1210 WSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRFVATNNVAEYEVLIAGLRLAAGMQI 1269
Query: 1203 RSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADAL 1262
++ TDSQL+ Q+ G ++ K+ + YL+ V+ + F+ + +PR +N ADAL
Sbjct: 1270 TTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQLMTKDFENFKLSKIPRGDNAPADAL 1329
Query: 1263 AKLASTRKPGNNKSVIQETLAYPSIEG-ELMACVNRGRT-----------WMDPIISILA 1310
A LA T + + E++ PSI+ + + VN R+ W I L+
Sbjct: 1330 AALALTSDSDLRRIIPVESIDKPSIDSTDAVEIVNTIRSSNAPDPADPTDWRVEIRDYLS 1389
Query: 1311 GDPAEVEQCTKEQQR-EASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCAS 1369
++ T + R +A+ YTL+ HL + +L C+ + IM E HEG +
Sbjct: 1390 DGTLPSDKWTARRLRIKAAKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKETHEGAAGN 1449
Query: 1370 HIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAM 1429
H GGR+LA K+ + GFYWPT+ DC F +C++CQ A P + L AP+PF
Sbjct: 1450 HSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQPTELLRAGVAPYPFMR 1509
Query: 1430 WGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRA 1489
W +D+VGP P +R Q +FILV DYFTKW+EAE A I + + NF WK I+CR G+P
Sbjct: 1510 WAMDIVGPMPASR-QKRFILVMTDYFTKWVEAESYATIRANDVQNFVWKFIICRHGLPYE 1568
Query: 1490 IVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGA 1549
I++DNG+QF S FC I++ ++ +PQ NGQ E+ N+ IL GL++RL E KGA
Sbjct: 1569 IITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEATNKTILSGLKKRLDEKKGA 1628
Query: 1550 WLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFE--EENQANM 1607
W DEL VLWSY TT +S T +TPF YG++AM P E+ + R + E N M
Sbjct: 1629 WADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKNPELNDRMM 1688
Query: 1608 AVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLK--WRSGAPGN--KL 1663
LD L E R+ A R + A +N +V R VGDLVL+ + + A N KL
Sbjct: 1689 LDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGDLVLRKVFENTAEINAGKL 1748
Query: 1664 TPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1705
NWEG Y++ K++ G Y L + G +PR++N + L+ +Y
Sbjct: 1749 GANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSMHLKRYY 1790
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 825 bits (2132), Expect = 0.0
Identities = 464/1115 (41%), Positives = 637/1115 (56%), Gaps = 126/1115 (11%)
Query: 661 RTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLR 720
R + I L E + L L +N FAWS +DMPGID CH L ++P+ KP+ Q R
Sbjct: 728 RCVGIRIDLPSELQNELVNFLRQNAATFAWSVEDMPGIDSAVTCHELNVDPTYKPLKQKR 787
Query: 721 RRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKACP 780
R+LG D+ K V +EV KLL A I EV+YP WL N V+VKK NGKWR+C+D+TDLNKACP
Sbjct: 788 RKLGPDRTKDVNEEVKKLLDAGSIVEVRYPDWLRNPVVVKKKNGKWRVCIDFTDLNKACP 847
Query: 781 KDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPF 840
KDS+PLP ID LV+ +GNELLS MDA+SGY+QI MH D +KT F+T + YCY+ MPF
Sbjct: 848 KDSFPLPHIDRLVEATAGNELLSFMDAFSGYNQILMHQNDREKTVFITDQGTYCYKVMPF 907
Query: 841 GLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRL 900
GLKNAGATY RL++++F Q+ +MEVY+DDM+VKS+R +H L + F + +++M+L
Sbjct: 908 GLKNAGATYPRLVNQMFTDQLDHSMEVYIDDMLVKSLRAEEHITHLRQCFQVLNRYNMKL 967
Query: 901 NPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFL 960
NP KC+FGV G+FLG+++T RGIE NP++ AI + SP N +EVQRL GRIAAL+RF+
Sbjct: 968 NPSKCTFGVTSGEFLGYLVTRRGIEANPKQISAIIDLPSPRNTREVQRLIGRIAALNRFI 1027
Query: 961 PQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSD 1020
+S D+ PF++ LR N FEW +CEE G L+LY AVS
Sbjct: 1028 SRSTDKCLPFYQLLRANKRFEWDEKCEE--------------------GETLYLYIAVST 1067
Query: 1021 SALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKV 1080
SA+S V+++E GE +++VS TL GAE+RY +EK A AV+++AR+LRPYF+S+ V+V
Sbjct: 1068 SAVSGVLVREDRGEQHPIFYVSKTLDGAELRYPTLEKLAFAVVISARKLRPYFKSYTVEV 1127
Query: 1081 RTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELTPDRFERVD 1140
T+ PLR +L P SGRL W+VELSEY + Y R T + R
Sbjct: 1128 LTNQPLRTILHSPSQSGRLAKWAVELSEYDIAYKNR---------------TCAKSHRAP 1172
Query: 1141 TQWTLFVDGSSNSSGSGAGVTLEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREV 1200
T+ S V L F A+NN+AEYEALIAGL+LA +
Sbjct: 1173 TR-------PYRRSIPCRKVDLA-------------RFPASNNEAEYEALIAGLRLAHGI 1212
Query: 1201 KIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRAD 1260
+++ + DSQLV +Q G ++ K+ + YL+ VR L F+ + +PR++N AD
Sbjct: 1213 EVKKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVVRELSYNFEVFELTKIPRSDNAPAD 1272
Query: 1261 ALAKLASTRKPGNNKSV-----------IQETLAYPSI---------------------- 1287
ALA LAST P + + IQ T+ PS
Sbjct: 1273 ALAVLASTSDPDLRRVIPVECIDVPSIKIQGTITKPSEYTVIDLTAEQLSIVMVILPPAT 1332
Query: 1288 --------EGELMACVNRGRT----------------------WMDPIISILAGDPAEVE 1317
G L+ + G++ W++ I S +A +
Sbjct: 1333 DRSEDAGPSGNLVCSSDPGQSSGCDRSDDSNRSADSDSPASTNWIEEIRSYIADRIVPKD 1392
Query: 1318 QCTKEQQREAS-HYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSL 1376
+ + R S YTL HL R + LL C+ E+ + +M E+HEG +H GGR+L
Sbjct: 1393 KWAARRLRARSAQYTLPHEHLLRWSATGVLLSCLDDEEAQQVMREIHEGAGGNHSGGRAL 1452
Query: 1377 ACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVG 1436
A K+ + G YWPT+ DC FV +C++CQ A P + L T P+PF W +D++
Sbjct: 1453 ALKIRKHGQYWPTMNADCEKFVARCEKCQGHAPFIHIPSEVLQTAMPPYPFMRWAMDIIR 1512
Query: 1437 PFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGT 1496
PFP +R Q K+ILV DYFTKW EAE A+I S ++ NF WK I+C G+P IV+DNGT
Sbjct: 1513 PFPASR-QKKYILVMTDYFTKWFEAESYARIQSKEVQNFVWKNIICHHGLPYEIVTDNGT 1571
Query: 1497 QFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPA 1556
QF+S Q FC + I++ ++ +PQ NGQ E+ N+ IL GL++RL E KGAW DEL
Sbjct: 1572 QFTSLQFEGFCAKWKIRLSKSTPRYPQCNGQAEATNKTILDGLKKRLDEKKGAWADELDG 1631
Query: 1557 VLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEE--NQANMAVELDLL 1614
VLWSY TT + +T TPF +TYG++A+ P E T R + N + LD L
Sbjct: 1632 VLWSYRTTPRRSTDRTPFSLTYGMEALAPCEAGLPTVRRSMLINDPTLNDQMLLDNLDTL 1691
Query: 1615 SETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLKW----RSGAPGNKLTPNWEGP 1670
E RD+A +R +Q A +N +V+ R GDLVL+ KL +WEGP
Sbjct: 1692 EEQRDQALLRIQNYQQAAAKFYNKKVKNRLFAEGDLVLRKVFENTVEQDAGKLGAHWEGP 1751
Query: 1671 YRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1705
Y I KV+ G Y L +DG +PRS+N +L+ +Y
Sbjct: 1752 YLISKVVKPGVYELLTMDGTPIPRSWNSANLKRYY 1786
Score = 67.4 bits (163), Expect = 7e-11
Identities = 122/566 (21%), Positives = 200/566 (34%), Gaps = 120/566 (21%)
Query: 32 EADSVAEFR---PFSEDVENVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASD- 87
E D V E PF+ + +V I ++ + L +Y G DP+ L + + SD
Sbjct: 225 EVDKVLEETQRSPFTSCISDVRIR-HISKIKLANYEGLVDPRPFLTSVSIAIGRAHFSDE 283
Query: 88 ---AVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNI 144
A C++F A+ WF+ L SI + ++ FL + + + DL+ +
Sbjct: 284 DRDAGSCQLFVKHLSGAALTWFSRLEANSIDSVHALTTSFLKNYGVFMEKGASNVDLWTM 343
Query: 145 RQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTR-KPARSMGEMR 203
Q ESL+ ++ R+ V + A A A +N L G + T KP G +
Sbjct: 344 AQTAKESLRSFIGRFKEIVTSVATPDDAAIA-ALRNALWHGDSSRCTTLCKPFHRQGSAK 402
Query: 204 ARASTYILDEEDDAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQQRPGKGKSVFKPTK 263
A + + K ++G + R D K+++ KG S +
Sbjct: 403 APIA--------------KEKPKEGH------HESRQHYDADYAKEEKLKKGTSYYVGDT 442
Query: 264 EQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPRD 323
PR D +PW + + D
Sbjct: 443 A---PRSD-----KPWNKWNRSADAKGD-------------------------------- 462
Query: 324 ANPKKWCEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNG-GDQNKAHKREE-- 380
+K+ EFH+ GH TD+CR LQ + + G + R G D++ H+R E
Sbjct: 463 ---QKYYEFHKIPGHSTDECRQLQTLLLTKFKKGDLDIEPDRRRTGTNDRDNTHRRVENT 519
Query: 381 ERADTKGKKKQESA-------------AIATKGADDTFAQHSGPPVG--TINTIAGGFGG 425
+R + K++Q+ A K D Q P N I GG
Sbjct: 520 DRDNQDDKRRQDETDRRAERIHDDDRRAEPHKRNHDAPRQFEDEPASKRRRNMIMGGLTA 579
Query: 426 GGDTHAARKRHVRAVNSVH-----EVAFGFVHPDITISMADFEGIKPHKDDPIVVQLRMN 480
D+ + K + R + + IT S AD + +DP+VV+L++
Sbjct: 580 CRDSVRSIKSYRRQGEKQRAWIEKNTSAVECNEPITFSEADI-ALPGSHNDPLVVELKIG 638
Query: 481 SFNVRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAGEQVMVRGYIDLDTIF 540
+ ++ +D S D + L GF G+ +M G I L
Sbjct: 639 ENVLLQLEIDVRSFQDKVQ-------------------PLTGFDGDTIMTVGTITLPIYV 679
Query: 541 GEDECARVLKVRYLVLQVVASYNVII 566
G V + V+ YNVI+
Sbjct: 680 G----GTVSWFEFAVVDKPIIYNVIL 701
>At3g31530 hypothetical protein
Length = 831
Score = 568 bits (1463), Expect = e-161
Identities = 332/839 (39%), Positives = 476/839 (56%), Gaps = 105/839 (12%)
Query: 799 NELL--SLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRV 856
N+LL S +++ + + M+P D++KTAF T + +CYR MPFGLKNAGATY+R ++++
Sbjct: 77 NQLLETSYCHSWTLFVVMMMNPEDQEKTAFYTEQGIFCYRVMPFGLKNAGATYERFVNKI 136
Query: 857 FAGQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLG 916
FA Q+G+ MEVY+DDM+VKS+ DH L E F ++ ++++LNP KC FGV+ G
Sbjct: 137 FALQIGKTMEVYIDDMLVKSMTEKDHISHLRECFKQLNLYNVKLNPAKCRFGVRSG---- 192
Query: 917 FMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRK 976
IE NP++ +A+ M SP N +EVQ LTGR+AAL+RF+ +S ++ F+ LR+
Sbjct: 193 -------IEANPKQIEALLGMASPQNKREVQCLTGRVAALNRFISRSTEKCLAFYDVLRR 245
Query: 977 NVAFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHR 1036
N FEWT CEEAF LK+ L++PPIL+KP+ G P +LY AVSD+A+S +++E GE +
Sbjct: 246 NKKFEWTTRCEEAFQELKKYLATPPILAKPVIGEPQYLYVAVSDTAVSGELVREDRGEQK 305
Query: 1037 IVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLS 1096
++++VS T AE RY ++EK ALAV+++A++LRPYFQS + V +PLR +L P S
Sbjct: 306 LIFYVSQTFTSAESRYPQMEKLALAVVMSAQKLRPYFQSHSIIVMGSMPLRVILHSPSQS 365
Query: 1097 GRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELTPD--RFERVDTQWTLFVDGSSNSS 1154
GRL W++ELSEY ++Y + +Q LADF+VEL R +DT W L VDGSS+
Sbjct: 366 GRLAKWTIELSEYDIEYQNKTCAKSQVLADFIVELPTKEARENPLDTTWLLHVDGSSSKQ 425
Query: 1155 GSGAGVTLEGP-GELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQL 1213
GSG G+ L P GE ATNN AEYEAL+AGL LA +KI DSQL
Sbjct: 426 GSGVGIRLTSPTGE------------ATNNVAEYEALVAGLNLAWGLKIGKTRAFCDSQL 473
Query: 1214 VENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGN 1273
+ NQ G + +D + YL V+ L F E + +PR EN ADALA LAST
Sbjct: 474 IANQFNGEYTTQDKKMEAYLIHVQNLAKNFDEFELTRIPRGENTSADALAALASTSDTSL 533
Query: 1274 NKSVIQETLAYPSIEGELMACV-------------NRGRTWMDPIISILA-GDPAEVEQC 1319
+ + E + PSIE V + WM+PIIS ++ G +
Sbjct: 534 KRVIPVEFIEKPSIELGKEEHVLPIQISADQDDPDDCNSEWMEPIISYISEGKLPSDKWK 593
Query: 1320 TKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACK 1379
++ + +A+ + L+D LY+ S PL+ CV E IM E+H G SH
Sbjct: 594 ARKLKAQAARFVLVDTKLYKWRLSGPLMTCVEAEAICKIMKEIHSG---SHA-------- 642
Query: 1380 VLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFP 1439
PT+ + P + L ++++P+PF W +D++GP
Sbjct: 643 --------PTIHQ---------------------PAELLSSIASPYPFMRWSMDIIGPMH 673
Query: 1440 TARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFS 1499
++ Q K +LV DYF+KWIEAE A I A++ NF K I+CR GIP IV+DNG+QF
Sbjct: 674 PSK-QKKLVLVLTDYFSKWIEAESYASIKDAQVENFVLKHILCRHGIPYEIVTDNGSQFI 732
Query: 1500 SSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLW 1559
S++ + FC + GI++ ++ +PQ NGQ E+AN EL VLW
Sbjct: 733 STRFQGFCDKWGIRLSKSTPRYPQGNGQAEAAN--------------------ELEGVLW 772
Query: 1560 SYNTTEQSTTRETPFRMTYGVDAMLPVE--IDNFTWRTRPGFEEENQANMAVELDLLSE 1616
+ TT + TRETPF YG + ++P E + + P + +N ELDL+ E
Sbjct: 773 LHRTTPRRATRETPFASIYGTECVIPAEMIVPSLRRSLSPENDPDNTQTPLDELDLIDE 831
Score = 35.8 bits (81), Expect = 0.22
Identities = 21/51 (41%), Positives = 27/51 (52%)
Query: 562 YNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKVGKLKVDQKMARECYNNCL 612
YN IIG LN+ AV ST HL +K+P S KL ++ A + N L
Sbjct: 30 YNAIIGTPWLNQFRAVASTYHLCLKFPTSDSVWEKLGHERAEAVKAENQLL 80
>At1g37200 hypothetical protein
Length = 1564
Score = 560 bits (1444), Expect = e-159
Identities = 433/1463 (29%), Positives = 665/1463 (44%), Gaps = 272/1463 (18%)
Query: 124 FLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGL- 182
FL Q S Q + DL+++ + ES + Y+ ++ A K+ + A +NGL
Sbjct: 176 FLKQDSVLIEQRTSEADLWSLTLLQNESHRSYVEKFKAIKSKIANLNEEVAIAALRNGLW 235
Query: 183 LPGGLNSKLTRKPARSMGEMRARASTYILDEEDDAFKRKRAKLEKGDTSPKRVKKDRSGE 242
+LT + S+ + +A + EE+ A + K K +P +
Sbjct: 236 FSSRFREELTVRQPISLDDALHKALHFAKGEEELAVLALKFKESKTQNAPLAI------- 288
Query: 243 DKGDGKQQRPGKGKSVFKPTKEQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDML 302
+ P++ ++ Q + T + D
Sbjct: 289 ---------------------------------KTPFKKENQTQGQHTLFAIEEAAED-- 313
Query: 303 RGASDANLVDEPEAPKYQPRDANPKKWCEFHRSAGHDTDDCRTLQREIDKLIRAGYQGNR 362
E+P + + K+C++H GH T++CR ++ KLI AG + +
Sbjct: 314 ------------ESP-----ELDLGKYCKYHNKRGHSTEECRAVK----KLIAAGGKTKK 352
Query: 363 QGQWRNGGDQNKAHKREEERADTKGKKKQESAAIATKGADDTFAQHSGPPVGT--INTIA 420
G K + + + KQ+ +G D S PP I+ +
Sbjct: 353 -------GSNPKVETPPPDEQEEEQTPKQKKRERTLEGGD------SPPPARRERIDLVF 399
Query: 421 GGFGGGGDTHAARKRHVRAVNSV-HEVAFGFVHPDITISMADFEGIKPHKDDPIVVQLRM 479
GG T R V S H+ F + E + P + D I+ ++
Sbjct: 400 TELDLGGKTG----RSVTTPPSAPHKKNMRFPLRLSKFCRSATEHLSPKRIDFIMGGSQL 455
Query: 480 NSFNVRRVLLDQGSSADIIYGDAF-----DKLGLTDNDLTPYAGTLVGFAGEQVMVRGYI 534
+ ++ + Q + G + K+ L +++ T
Sbjct: 456 CNDSINSIKTHQRKADSYTKGKSLMMGPDHKITLWESETT-------------------- 495
Query: 535 DLDTIFGEDECARVLKVRYLVLQVV---------ASYNVIIGRNTLNRLCAVISTAHLAV 585
DLD + R+ Y + +++ A +N I+GR L+ + AV ST H +
Sbjct: 496 DLDKPHDDAIVIRIDVGNYKLSRIMIDTGSSVDKAPFNAILGRPWLHAMKAVPSTYHQCI 555
Query: 586 KYPLSSGKVGKLKVDQKMARECYNNCLNFYGKKSALVGHRCYEIEASDENLDPRGEGRVN 645
K+P G + + Q+ +R CY KK+ LV + ++ L R +
Sbjct: 556 KFPSEKG-IAVIYGSQRSSRRCYMGSYELI-KKAELV------VLMIEDKLAEMKTVRSS 607
Query: 646 RPT---PIEETKALKFGDRT-----LKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPG 697
P+ P + + + D + + +G L L EN D FAWS ++ G
Sbjct: 608 DPSQCGPQKSSITQVYIDESDPKQCVGVGQDLDPAIREDFITFLKENKDSFAWSSANLQG 667
Query: 698 IDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVV 757
I H L ++P+ +P+ Q RR+LG ++ KAVQ EVD+LL IREVKYP WLAN V
Sbjct: 668 ISLEVTSHELNVDPTYRPIKQKRRKLGPERAKAVQDEVDRLLKIGSIREVKYPDWLANPV 727
Query: 758 MVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMH 817
+VK NGKWR+C+D+ DLNKACPKDS+PLP ID LV +G+ELLS MDA+SGY+QI M
Sbjct: 728 VVKNKNGKWRVCIDFMDLNKACPKDSFPLPHIDRLVKATAGHELLSFMDAFSGYNQILMR 787
Query: 818 PADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSV 877
P D++KTAF+T CY+ MPFGLKN GATYQRL++R+FA Q+G+ ME+Y+DDM+VKS
Sbjct: 788 PDDQEKTAFITD----CYKVMPFGLKNTGATYQRLVNRMFADQLGKTMELYIDDMLVKSA 843
Query: 878 RGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQM 937
DH L E F + K M+ +NPEKC
Sbjct: 844 HEKDHLPQLRECFKILNKFEMK--------------------------LNPEKCS----F 873
Query: 938 KSPSNVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELL 997
PS L + Q G + P K +A AF+ ++
Sbjct: 874 GVPSG-----------EFLGYLVTQRGIEANP------KQIA---------AFI---DMP 904
Query: 998 SSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEK 1057
S +P + +Y A D +G+ + +Y+VS TL AE Y+ +EK
Sbjct: 905 SPKMARDRPYRRPQQVIYRAPPDH----------EGQ-KPIYYVSKTLIDAETHYRAMEK 953
Query: 1058 AALAVLVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRG 1117
ALAV+++AR+LR Y QS P+ V T P+R +L S RL W++ELSEY ++Y R
Sbjct: 954 LALAVVMSARKLRQYLQSHPIVVMTSQPIRTILHSTTQSSRLAKWAIELSEYDIEYRTRT 1013
Query: 1118 TVGAQSLADFVVELTPDRFERVDT--QWTLFVDGSSNSSGSGAGVTLEGPGELVLEQSLK 1175
+ AQ LADFV+EL + ++ +W L VDGSSN GSG G+ L P V+EQS +
Sbjct: 1014 SFKAQVLADFVIELPLADLDGTNSNKKWLLHVDGSSNRQGSGEGIQLTSPTGEVIEQSFR 1073
Query: 1176 FEFKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLER 1235
F A+NN++EYEALI G+KLA+ ++IR + +DSQLV +Q G ++ KD + YLE
Sbjct: 1074 LGFNASNNESEYEALIDGIKLAQGMRIRDIHAHSDSQLVTSQFHGEYEAKDERMEAYLEL 1133
Query: 1236 VRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIE------- 1288
V+ L F+ + +PR EN DALA LAST P + + E + +PSI+
Sbjct: 1134 VKTLTQQFESFELTRIPRGENTSTDALAALASTLDPFVKRIIPVEGIEHPSIDLTVKHAG 1193
Query: 1289 --------------------GELMACVNRG----------------------RTWMDPII 1306
L A G W PII
Sbjct: 1194 METSATCNFTRVTRSTTAAARALAAAAEAGAAEEEAKLRTPEQLASYEPEPYNDWRIPII 1253
Query: 1307 S-ILAGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEG 1365
I G + T++ + +++ Y +++G L +R + P + C ++ + +M +HEG
Sbjct: 1254 DYIERGITPPDKWETRKLKAQSARYCIMEGRLMKRSVAGPYMVCTYGQQTKDLMKSMHEG 1313
Query: 1366 VCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPW 1425
C SH GR+L C +L DC+ +C +CQ A PP+E+ ++S+P+
Sbjct: 1314 QCGSHYSGRTLRCIML----------ADCIAHSLRCDKCQRHAPTLHQPPEEMSSISSPY 1363
Query: 1426 PFAMWGVDLVGPFPTA--RAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCR 1483
PF W +D+V P + + ++K +LV DY TKWIE + ++T ++ F + IV R
Sbjct: 1364 PFMKWSMDVVRPMEASGGKKRLKNLLVLTDYSTKWIEGKAFQQVTEKQVEVFLLENIVYR 1423
Query: 1484 FGIPRAIVSDNGTQFSSSQTREF 1506
GIP IV+DNG +++ F
Sbjct: 1424 HGIPYEIVTDNGLTSPREKSKHF 1446
Score = 56.2 bits (134), Expect = 2e-07
Identities = 33/99 (33%), Positives = 54/99 (54%), Gaps = 5/99 (5%)
Query: 1612 DLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLKW----RSGAPGNKLTPNW 1667
D + E RD A I+ +Q +NS +++R +VG+LVL+ KL NW
Sbjct: 1465 DFIDERRDHAMIQVQNYQQPAVRYYNSNIKIRRFEVGELVLRKVFSNTRELNAGKLGTNW 1524
Query: 1668 EGPYRIVKVLGNGAYHLEEL-DGRRLPRSFNGLSLRYFY 1705
EGPYRI +V+ +G Y L ++ +G R +N + L+ ++
Sbjct: 1525 EGPYRITEVVRDGIYKLVKVFNGVPELRPWNAMHLKKYH 1563
>At2g13330 F14O4.9
Length = 889
Score = 463 bits (1192), Expect = e-130
Identities = 244/528 (46%), Positives = 333/528 (62%), Gaps = 32/528 (6%)
Query: 763 NGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADED 822
NGKWR+C+D+ DLNKACPKDS+PLP ID LV+ +E LS MDA+ GY+QI M ++
Sbjct: 291 NGKWRVCIDFRDLNKACPKDSFPLPHIDRLVEATVEHEKLSFMDAFYGYNQILMRRDGQE 350
Query: 823 KTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDH 882
KTAF+T R Y Y+ MPFGLKNAG TYQRL++R+F Q+G+ +EVY+DDM+VKS DH
Sbjct: 351 KTAFITDRGTYYYKVMPFGLKNAGTTYQRLVNRMFVDQLGKTIEVYIDDMLVKSAHEKDH 410
Query: 883 HQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSN 942
L E F + K M+LNPEKCSF VQ +FL +++T RGIE NP++ A +M SP
Sbjct: 411 VPQLRECFKILIKFEMKLNPEKCSFEVQSREFLEYLVTERGIEANPKQIAAFIEMPSPKM 470
Query: 943 VKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPI 1002
+EVQRLTGRI AL+ F+ +S ++ PF++ LRK F+W +CE+AF +LK L+ PPI
Sbjct: 471 AREVQRLTGRIPALNGFISRSANKCVPFYQPLRKGKEFDWNKDCEQAFKQLKAYLTEPPI 530
Query: 1003 LSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAV 1062
L+KP +G PL+LY + R +EK AL V
Sbjct: 531 LAKPEKGEPLYLY------------------------------TNRDKRCPAMEKLALTV 560
Query: 1063 LVTARRLRPYFQSFPVKVRTDLPLRQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQ 1122
+++ R+LR YFQS P+ V T P+R +L SGRL W++EL EY ++Y R ++ AQ
Sbjct: 561 VMSVRKLRLYFQSHPIVVMTSQPIRTILHSLTQSGRLAKWAIELREYDIEYRTRTSLKAQ 620
Query: 1123 SLADFVVELTPDRFERVDT--QWTLFVDGSSNSSGSGAGVTLEGPGELVLEQSLKFEFKA 1180
LADFV++L + ++ +W L VDGSSN GSG G+ L P V+EQSL+ F A
Sbjct: 621 VLADFVIKLPLADLDGTNSNKKWLLHVDGSSNRQGSGVGIQLTSPTGEVIEQSLQLGFNA 680
Query: 1181 TNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLM 1240
+NN++EYEALIAG+KLA+E IR + +DSQLV +Q G ++ KD + YLE V+ L
Sbjct: 681 SNNESEYEALIAGIKLAQEKGIREIHAYSDSQLVTSQFHGEYEAKDERMEAYLELVKTLA 740
Query: 1241 TLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIE 1288
F+ + +PR EN AD LA LAST P + + E + + SI+
Sbjct: 741 QQFESFKLTRIPRGENTSADTLAALASTSDPFVKRIIPVEGIEHTSID 788
Score = 88.2 bits (217), Expect = 4e-17
Identities = 75/241 (31%), Positives = 117/241 (48%), Gaps = 21/241 (8%)
Query: 466 KPHKDDPIVVQLRMNSFNVRRVLLDQGSSADIIYGDAFDKLGLTDNDLTPYAGTLVGFAG 525
KPH DD +V+++ + ++ + +++D GSS D+++ DAF + G D+ L L GFAG
Sbjct: 59 KPH-DDALVIRIDVGNYELSCIMVDTGSSVDVLFYDAFKRTGHLDSKLQGRKTPLTGFAG 117
Query: 526 EQVMVRGYIDLDTIFGEDECARVLK--VRYLVLQVVASYNVIIGRNTLNRLCAVISTAHL 583
+ G I L TI AR ++ +LV+ A +N I+GR L+ + AV ST H
Sbjct: 118 DTTFSIGTIQLPTI------ARGVRQLTNFLVVDKKAPFNAILGRPWLHVMKAVPSTYHQ 171
Query: 584 AVKYPLSSGKVGKLKVDQKMARECYNNCLNFYGKKSALVGHRCYEIEASD----ENLDPR 639
+K+P G + + Q+ +R+CY K +V E E ++ +LDP
Sbjct: 172 CIKFPSYKG-IAVVYGSQRSSRKCYMGSYEDIKKADPVV--LMIEDELAEMKTVRSLDPS 228
Query: 640 GEGRVNRPTPIEETKALKFGD--RTLKIGTRLTEEQETRLTKLLGENLDLFAWSCKDMPG 697
G R + I + + D R + IG L L L EN D FAWS ++ G
Sbjct: 229 QRG--TRKSLITQV-CIDESDPKRCVGIGHDLDLTVREDLITFLKENKDSFAWSSANLQG 285
Query: 698 I 698
I
Sbjct: 286 I 286
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 387 bits (995), Expect = e-107
Identities = 255/763 (33%), Positives = 379/763 (49%), Gaps = 75/763 (9%)
Query: 968 FPFFKCLRK-NVAFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSV 1026
FPF K + F W CEEAF +LK LS P +L+K G L LY AVS+SA++ V
Sbjct: 616 FPFLHFTAKISKGFIWNESCEEAFKQLKRYLSEPSVLAKREFGEQLFLYIAVSESAVTGV 675
Query: 1027 MLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTDLPL 1086
++ + R +++V + RY +EK ALAV+ AR+LRPYFQS P+ V T LPL
Sbjct: 676 QVRVERSDKRPIFYV-------KTRYPMMEKLALAVVTAARKLRPYFQSHPIVVLTSLPL 728
Query: 1087 RQVLQKPDLSGRLVAWSVELSEYGLQYDKRGTVGAQSLADFVVELTPDRFERVDTQWTLF 1146
R +L P SGRL W++ELSE+ L++ R ++ + T D QW
Sbjct: 729 RTILHSPTQSGRLGKWAIELSEFDLEFRARTSLNS----------TMD----TSRQWGFI 774
Query: 1147 VDGSSNSSGSGAGVTLEGPGELVLEQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLL 1206
+ + S A GE +A+ + A L L REV R
Sbjct: 775 KTWVRSRNLSDAPTANRYSGEY---------------EAKDACMEAYLNLVREVSGRFEQ 819
Query: 1207 IR-TDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVV----VEYVP-RAENQRAD 1260
T EN L RV + T+ Q + + +V RA +R D
Sbjct: 820 FELTRIPRAENSAANALAALASTFEVTLPRVIPVETISQPSIRLDEISFVTTRAMRRRLD 879
Query: 1261 ALAKLASTRKPGNNKSVIQETLAYPSIEGELMACVNR-----------GRTWMDPIIS-I 1308
A + + G+++ + +E + + + G W +PI I
Sbjct: 880 AQSAENGLHQLGDDEEISDAVHPTEIVENQSLPDNHNAPLPDQPPHDWGADWREPIRDYI 939
Query: 1309 LAGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCA 1368
L G + ++ + + + + + LYRR FS P C+ E+ +M EVH+G C
Sbjct: 940 LNGTLPAEKWAARKLKATCARFCIANDILYRRIFSAPDAVCIFGEQTRTVMKEVHDGTCG 999
Query: 1369 SHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFA 1428
+H GGRSLA KV + G+YWPTL DC + ++C++CQ A L P + L T+SAP+PF
Sbjct: 1000 NHTGGRSLAFKVRKYGYYWPTLVADCEAYARKCEQCQKHAPLILQPAELLTTVSAPYPFM 1059
Query: 1429 MWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPR 1488
W +D+VGP + T+ +EA + IT ++ NF WK I+CR G+P
Sbjct: 1060 KWLMDIVGPLHVS--------------TRGVEAAAYSNITHVQVWNFIWKDIICRHGLPY 1105
Query: 1489 AIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKG 1548
IV+DNG+QF S Q FC+E I++ ++ +PQ NGQ E+ N+ I+ L+++L KG
Sbjct: 1106 EIVTDNGSQFISEQFEVFCEEWQIRLSHSTPRYPQGNGQAEAMNKTIISNLKKKLNAYKG 1165
Query: 1549 AWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWR--TRPGFEEENQAN 1606
AW EL VLW+ TT + T ETPF + YG++A++P EI + R P E EN
Sbjct: 1166 AWFGELQNVLWAVRTTPRRATDETPFSLIYGMEAVIPAEIKVPSARRIRNPQNETENNEM 1225
Query: 1607 MAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLKW----RSGAPGNK 1662
+ +D + E R+ A R A +NS VR R +VG LVL+ ++ K
Sbjct: 1226 IIDVIDTIDERRNRALARMQNYHNAGARYYNSNVRNRSFEVGTLVLRRVQQNKAEKGAGK 1285
Query: 1663 LTPNWEGPYRIVKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 1705
L +WEGPY+I V+ NG Y L ++G+ + R++N + L+ FY
Sbjct: 1286 LGISWEGPYKITHVVRNGVYRLINMEGKTVRRAWNSMHLKRFY 1328
Score = 188 bits (477), Expect = 3e-47
Identities = 90/151 (59%), Positives = 113/151 (74%)
Query: 695 MPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLA 754
M GI I H L ++P+ KPV Q RR+LG D+ +AV EV +LL IREVKYP WLA
Sbjct: 412 MTGISTEVISHELNVDPTFKPVKQKRRKLGPDRAQAVNIEVVRLLEVGRIREVKYPEWLA 471
Query: 755 NVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQI 814
N V+VKK NGKWR+CVD+TDLNKAC KD +PLP ID LV+ +G+E+LS MDA+SGY+QI
Sbjct: 472 NPVVVKKKNGKWRVCVDFTDLNKACSKDFFPLPHIDRLVESTTGHEMLSFMDAFSGYNQI 531
Query: 815 RMHPADEDKTAFMTARVNYCYRTMPFGLKNA 845
M+P D++KT+F+T Y Y+ MPFGLKNA
Sbjct: 532 LMNPEDQEKTSFITECGTYYYKVMPFGLKNA 562
Score = 62.0 bits (149), Expect = 3e-09
Identities = 39/126 (30%), Positives = 58/126 (45%), Gaps = 13/126 (10%)
Query: 41 PFSEDVENVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAA--------SDAVKCR 92
PF+ + + I + + L Y+G D K+HL F +IA DA C+
Sbjct: 157 PFTPRISKLRIRE-FRDFKLPVYNGKGDLKEHLTSFQ----VIAGRVPLEPHEEDAGLCK 211
Query: 93 MFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESL 152
+F A+ WFT L GSI NF+ S+ F+ Q+ N +T L+N Q E L
Sbjct: 212 LFSENLFGLALTWFTQLEEGSIDNFKQLSTAFIKQYEYFINSDITEAHLWNFSQSADEPL 271
Query: 153 KEYMAR 158
+ Y+ R
Sbjct: 272 RTYIYR 277
Score = 34.3 bits (77), Expect = 0.64
Identities = 19/47 (40%), Positives = 25/47 (52%)
Query: 548 VLKVRYLVLQVVASYNVIIGRNTLNRLCAVISTAHLAVKYPLSSGKV 594
V V + V A+YNVI+G L + V ST H VK+P GK+
Sbjct: 366 VKMVEFTVFDRPAAYNVILGTPWLYEMKVVPSTYHQCVKFPTPVGKM 412
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 386 bits (992), Expect = e-107
Identities = 182/322 (56%), Positives = 245/322 (75%)
Query: 720 RRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKWRMCVDYTDLNKAC 779
RR+LG ++ KAV EVDKLL IREV+YP WLAN V+VKK NGK R+C+D+TDLNKAC
Sbjct: 469 RRKLGVERAKAVNDEVDKLLKIGSIREVQYPDWLANTVVVKKKNGKDRVCIDFTDLNKAC 528
Query: 780 PKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMP 839
PKDS+PLP ID LV+ +GNELLS MDA+SGY+QI M+P D++KT F+T R YCY+ MP
Sbjct: 529 PKDSFPLPHIDRLVESTAGNELLSFMDAFSGYNQIMMNPEDQEKTLFITDRGIYCYKVMP 588
Query: 840 FGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMR 899
FGL+NAGATY RL++++F+ VG+ MEVY+DDM++KS++ DH + LEE F + ++ M+
Sbjct: 589 FGLRNAGATYPRLVNKMFSEHVGKTMEVYIDDMLIKSLKKEDHVKHLEECFAILNQYQMK 648
Query: 900 LNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRF 959
LNP KC+FGV G+FLG+++T RGIE NP + A M SP N KEVQRLTGRIAAL+RF
Sbjct: 649 LNPAKCTFGVPSGEFLGYIVTKRGIEANPNQINAFLNMPSPKNFKEVQRLTGRIAALNRF 708
Query: 960 LPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVS 1019
+ +S D+S PF++ L+ N F W +CEEAF +LK L+SPP+L KP L+LY +VS
Sbjct: 709 ISRSTDKSLPFYQILKGNKEFLWDEKCEEAFGQLKAYLTSPPVLFKPELDEKLYLYVSVS 768
Query: 1020 DSALSSVMLQEIDGEHRIVYFV 1041
+ A+S V+++E GE + +Y++
Sbjct: 769 NHAVSGVLIREDRGEQKPIYYI 790
Score = 278 bits (711), Expect = 2e-74
Identities = 178/548 (32%), Positives = 273/548 (49%), Gaps = 94/548 (17%)
Query: 1202 IRSLLIRTDS-------QLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRA 1254
+ +LIR D ++ NQ G ++ KD + YL V+ L F++ + +PR
Sbjct: 772 VSGVLIREDRGEQKPIYYIIVNQFNGDYEAKDSRMEAYLHVVKDLAKNFRKFELIRIPRG 831
Query: 1255 ENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMACV---------NRGRTWMD-P 1304
+N ADALA LAST P N + E ++ SI+ E A V +RG T ++ P
Sbjct: 832 QNTTADALAALASTSDPEVNSIIPVECISERSIKEEKEAFVVTRSRAAARDRGDTAVELP 891
Query: 1305 IISILAGDPAEV--------------EQCTKEQQREASHYTLID---------------G 1335
+ A V E+ + + EA +D
Sbjct: 892 PVKRRKSKEATVPKPAVQENIGIELIEEVISDAKIEAETEPWVDEDILAPNEHPEPEALN 951
Query: 1336 HLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCM 1395
LYRRG S P L + E +MSEVHEG+C SH GR++A K+ + +YWPT+ DC+
Sbjct: 952 SLYRRGVSDPYLLSIFGPVVEIVMSEVHEGLCGSHSSGRAMAFKIKKMCYYWPTMITDCV 1011
Query: 1396 DFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYF 1455
+ ++CK CQ+ A L P + ++SAP+PF W +D++GP + ++++LV DYF
Sbjct: 1012 KYAQRCKRCQLHAPLIHQPSELFSSISAPYPFMRWSMDIIGPLHRSTRGVQYLLVLTDYF 1071
Query: 1456 TKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMR 1515
+KWIEAE + ++N+ GI++
Sbjct: 1072 SKWIEAEAI-------LLNW-----------------------------------GIKVS 1089
Query: 1516 FASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFR 1575
++++ +PQ NGQ E+AN+ IL L++RL KG W DEL VLW+Y TT + +T ETPF
Sbjct: 1090 YSTLRYPQGNGQAEAANKTILSNLKKRLNHLKGGWYDELQPVLWAYRTTPRRSTGETPFS 1149
Query: 1576 MTYGVDAMLPVEID--NFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVA 1633
+ YG+ A++P E++ P E+EN A + LD ++E RD+A I+ + A
Sbjct: 1150 LVYGMKAVVPAELNVPGLRRTEAPLNEKENSAMLDDSLDTINERRDQALIQIQNYQHAAA 1209
Query: 1634 AKFNSRVRVRDMQVGDLVLKW----RSGAPGNKLTPNWEGPYRIVKVLGNGAYHLEELDG 1689
+NS+V+ R VGD VLK + KL NWEGPY + +V+ NG Y L++L+
Sbjct: 1210 RYYNSKVKSRPFFVGDYVLKRVFDNKKEEGAGKLGINWEGPYIVTEVVRNGVYRLKDLED 1269
Query: 1690 RRLPRSFN 1697
R + R +N
Sbjct: 1270 RPVQRPWN 1277
Score = 79.0 bits (193), Expect = 2e-14
Identities = 62/234 (26%), Positives = 111/234 (46%), Gaps = 20/234 (8%)
Query: 41 PFSEDVENVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVII----AASDAVKCRMFPS 96
PF+ + ++ I D+ K L L+SY+G DPK +L F + A D C++F
Sbjct: 144 PFTPQITSLRIRDSRK-LNLESYNGLEDPKGYLAAFLIAAGRVDLNEADEDVRYCKLFSE 202
Query: 97 TFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYM 156
A+ WFT L GSISNF + S FL Q+S ++ ++ DL+N+ Q E+L+ ++
Sbjct: 203 NLCGQALMWFTQLEPGSISNFNELSVVFLKQYSILMDKSISDTDLWNLSQGPNETLRAFI 262
Query: 157 ARYSAASVKVEDEEPRACALAFKNGL-LPGGLNSKLTRKPARSMGEMRARASTYI-LDEE 214
++ K+ ++ A + GL L ++ + RA+ ++ +++E
Sbjct: 263 TKFKYVLSKLSRISQQSALSALRKGLWYDSRFKEDLILHKLDTIQDALFRANNWMEVEDE 322
Query: 215 DDAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGK-----------QQRPGKGKS 257
++F ++ + + T P KK E++G K +Q GKG+S
Sbjct: 323 KESFAKRDKQAKPAVTFPP--KKFEPRENQGPRKFGLLPFNNNVGKQFQGKGRS 374
Score = 43.5 bits (101), Expect = 0.001
Identities = 36/148 (24%), Positives = 66/148 (44%), Gaps = 13/148 (8%)
Query: 379 EEERADTKGKKKQESAAIATKGADDTFAQHSGP-PVGTI---NTIAGGFGGGG------- 427
E+E+ + KQ A+ ++ GP G + N + F G G
Sbjct: 320 EDEKESFAKRDKQAKPAVTFPPKKFEPRENQGPRKFGLLPFNNNVGKQFQGKGRSNTWIR 379
Query: 428 DTHAARKRHVRAVNSVHEVA--FGFVHPDITISMADFEGIKPHKDDPIVVQLRMNSFNVR 485
D + K+H R V+ ++ F I + + ++ DD +VV L + +F V
Sbjct: 380 DESSVIKKHSRNVHLKLSLSEEVDFQSTSILFDEKETQHLERSHDDALVVTLDVANFEVS 439
Query: 486 RVLLDQGSSADIIYGDAFDKLGLTDNDL 513
R+L+D GSS D+I+ +++G++ D+
Sbjct: 440 RILIDTGSSVDLIFLSTLERMGISRADV 467
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 357 bits (917), Expect = 3e-98
Identities = 170/289 (58%), Positives = 219/289 (74%), Gaps = 1/289 (0%)
Query: 694 DMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWL 753
DM GIDP CH L ++P+ K V Q RR+LG ++ KAV EVDKLL A I EVKYP WL
Sbjct: 476 DMVGIDPEVACHELNVDPTFKLVKQKRRKLGPERSKAVNDEVDKLLDAGSIVEVKYPEWL 535
Query: 754 ANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQ 813
AN V+VKK N KWR+C+D+TDLNKACPKDS+PLP ID +V+ +GNELLS MDA+SGY+Q
Sbjct: 536 ANPVVVKKKNDKWRVCIDFTDLNKACPKDSFPLPHIDRMVEATTGNELLSFMDAFSGYNQ 595
Query: 814 IRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMI 873
I MH D++KT+F+ R YCY+ MPFGLKN GA YQRL++++FA Q+G+ MEVY+DDM+
Sbjct: 596 IPMHKDDQEKTSFIIDRGTYCYKVMPFGLKNVGARYQRLVNQMFAPQLGKTMEVYIDDML 655
Query: 874 VKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKA 933
VKS R DH L+ F + K++M+LNP KC FGV G+FLG+++T RGIE NP++ +A
Sbjct: 656 VKSTRSADHIDHLKACFETLNKYNMKLNPAKCLFGVTSGEFLGYIVTKRGIEANPKQIRA 715
Query: 934 IQQMKSPSNVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRK-NVAFE 981
I ++SP N KEVQRLTGRIA L+RF+ +S D+S PF++ LR N FE
Sbjct: 716 ILDLQSPRNKKEVQRLTGRIAGLNRFIARSTDKSLPFYQLLRSANKTFE 764
Score = 98.6 bits (244), Expect = 3e-20
Identities = 83/335 (24%), Positives = 144/335 (42%), Gaps = 34/335 (10%)
Query: 295 NTDVSDMLRGASDANLVDEPEAPKYQPRDANPK----KWCEFHRSAGHDTDDCRTLQR-- 348
NTD L S A D + P +A + ++C+FH+ +GH T CR LQ
Sbjct: 153 NTDPQQFLSSFSVATQRDH-----FHPSEAEVENRLDQYCDFHKRSGHSTAACRHLQSIL 207
Query: 349 -------EIDKLIRAGYQGNRQGQWRNGGDQNKA------HKREEERADTKGKKKQESAA 395
+I+ R N R G D N H ++ A ++++
Sbjct: 208 LNKYKKGDIEVQHRQYKSHNNTYAARGGRDGNNVFHRLGPHTGRQQEAPPANEEERHPDM 267
Query: 396 IATKGADDTFAQHSGPPVGT--INTIAGGFGGGGDTHAARKRHVR--AVNSVHEVAFGFV 451
K + QH+ PV +N I GG D+ + K +++ A A +
Sbjct: 268 EPPKKNRENDQQHNDAPVPRRRVNMIMGGLTACRDSFRSIKEYIKSGAATLWSSPATKEM 327
Query: 452 HPDITISMADFEGIKPHKDDPIVVQLRMNSFNVRRVLLDQGSSADIIYGDAFDKLGLTDN 511
P +T + D G+ +DP+V++L + V R+L+D GSS ++++ D K+ + D
Sbjct: 328 TP-LTFTSEDLFGVDLPHNDPLVIELHIGESEVTRILIDTGSSVNVVFKDVLQKMKVHDR 386
Query: 512 DLTPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNTL 571
+ P L GF G +M G I L G +++V+ YN+I+G +
Sbjct: 387 HIKPSVRPLTGFDGNTMMTNGTIKLPIYLG----GAATWHKFVVVDKPTIYNIILGTPWI 442
Query: 572 NRLCAVISTAHLAVKYPLSSGKVGKLKVDQKMARE 606
+ + A+ S+ H +K P S G + ++ +Q +A +
Sbjct: 443 HDMQAIPSSYHQCIKIPTSIG-IETIRGNQNLAHD 476
>At4g03800
Length = 637
Score = 325 bits (833), Expect = 1e-88
Identities = 165/336 (49%), Positives = 230/336 (68%), Gaps = 21/336 (6%)
Query: 695 MPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLA 754
MP IDP+ ICH L ++P KP+ Q RR+LG ++ KAV ++DKLL IREV+YP W+A
Sbjct: 321 MPDIDPSIICHELNVDPRFKPLKQKRRKLGVERAKAVNGKIDKLLKIGSIREVQYPDWVA 380
Query: 755 NVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQI 814
V+VKK NGK R+C+D+TDLNKACPKDS+PLP ID LV+ +GNELL+ MDA+ GY+QI
Sbjct: 381 ITVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHIDRLVESTAGNELLTFMDAFLGYNQI 440
Query: 815 RMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIV 874
M+P D++KT+F+T R GATYQ L++++F + + MEV +DD +V
Sbjct: 441 MMNPEDQEKTSFITDR---------------GATYQWLVNKMFNEHLRKTMEVSIDDTLV 485
Query: 875 KSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAI 934
KS++ DH + L E F + ++ M+LN KC+FGV G+FLG+++T RGIE NP + A
Sbjct: 486 KSLKKEDHVKHLGECFEILNQYQMKLNLAKCTFGVPSGEFLGYIVTKRGIEANPNQINAF 545
Query: 935 QQMKSPSNVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLK 994
+ S N KEVQRLTGRIA+ +S D+S PF++ L+ N F W +CEEAF +LK
Sbjct: 546 LKTPSLRNFKEVQRLTGRIAS------RSTDKSLPFYQILKGNNGFLWDEKCEEAFRQLK 599
Query: 995 ELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQE 1030
L++PP+LSKP L+LY VS+ A+S V+++E
Sbjct: 600 AYLTTPPVLSKPEADEKLYLYVFVSNHAVSGVLVRE 635
Score = 63.9 bits (154), Expect = 8e-10
Identities = 67/262 (25%), Positives = 115/262 (43%), Gaps = 37/262 (14%)
Query: 334 RSAGHDTDDCRTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKKKQES 393
R H T DC L+R + +L +G N D K + + ++T+ K+
Sbjct: 77 RVTEHLTKDCIVLKRYLAELWASGDLSKF-----NIDDFVKQYHEARDNSETQNSKRPRL 131
Query: 394 AAIATKGADDTFAQHSGPPVGTINTIAGGFGGGGDTHAARKRH---VRAVNSVHEVAFGF 450
T + G IN I GG ++ +A K+H V +S+ E +
Sbjct: 132 ----------TNEEEPRSSKGKINVILGGSKLCHNSVSAIKKHRPNVLLKSSLSEEV-NY 180
Query: 451 VHPDITISMADFEGIKPHKDDPIVVQLRMNSFNVRRVLLDQGSSADIIYGDAFDKLGLTD 510
++ + I+ DD +++ L + +F + R+L+D GSS D+I+
Sbjct: 181 QCSSVSFDEEETRHIERPHDDALIITLDVANFKISRILVDTGSSVDLIF----------- 229
Query: 511 NDLTPYAGTLVGFAGEQVMVRGYIDLDTIFGEDECARVLKVRYLVLQVVASYNVIIGRNT 570
L P LV F E M G I L + G + +++ V ++V A+YN+I+G
Sbjct: 230 --LGP-PSPLVAFTSESAMSLGTIKLPVLAG--KMSKI--VDFVVFDKPATYNIILGTPW 282
Query: 571 LNRLCAVISTAHLAVKYPLSSG 592
+ ++ AV ST H +K+P S+G
Sbjct: 283 IFQMKAVPSTYHQWLKFPTSNG 304
>At2g13230 pseudogene
Length = 353
Score = 231 bits (589), Expect = 3e-60
Identities = 126/358 (35%), Positives = 200/358 (55%), Gaps = 28/358 (7%)
Query: 1232 YLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIE--- 1288
YL V+ F E + +PR EN ADALA LAST + + E + PSIE
Sbjct: 4 YLALVQDHAKQFHEFKLTRIPRGENTSADALAALASTSDLNMRRVIPVEFIEKPSIELDK 63
Query: 1289 ----------------GELMACVNRGRTWMDPIISILAGDPAEVEQC-TKEQQREASHYT 1331
+ + W++PI + ++ +++ ++ + EA+ +
Sbjct: 64 VEHAFPIQVVAGYEDASDNSPSLGCDSEWIEPIRNFISKRKVRLDKWEARKLKAEAARFV 123
Query: 1332 LIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLR 1391
L++ L++ S PL+ CV E +M EVH G C ++ GGR+LA K+ R G++WPT+
Sbjct: 124 LVEEKLFKWRLSGPLMTCVEKEAARKVMKEVHGGSCGNNSGGRALAIKIKRLGYFWPTMI 183
Query: 1392 KDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVA 1451
KDC ++CQ A P + L ++++P+P W +D++GP ++ Q K +LV
Sbjct: 184 KDC-------EKCQRNAPTIHLPAELLSSIASPYPLMRWSMDIIGPMHASK-QKKLVLVL 235
Query: 1452 VDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMG 1511
DYF+K EAE A I A++ +F WK I+CR G+P IV+DNG+QF S++ + FC + G
Sbjct: 236 TDYFSKLREAESYASIKEAQVESFMWKHILCRHGVPYEIVTDNGSQFISTRFQGFCDKWG 295
Query: 1512 IQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTT 1569
I++ ++ + Q NGQ ++AN+ IL GL++RL KG+W+DEL VLWS+ TT + T
Sbjct: 296 IRLSKSTPRYLQGNGQADAANKTILDGLKKRLTAKKGSWVDELEGVLWSHRTTPRRAT 353
>At2g10780 pseudogene
Length = 1611
Score = 231 bits (588), Expect = 4e-60
Identities = 148/493 (30%), Positives = 244/493 (49%), Gaps = 16/493 (3%)
Query: 707 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKW 766
+ L P P+S+ R+ + +++++++LL FIR P W A V+ VKK +G +
Sbjct: 651 IELEPGTTPISKAPYRMAPAEMAKLKKQLEELLDKGFIRPSSSP-WGAPVLFVKKKDGSF 709
Query: 767 RMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAF 826
R+C+DY LNK K+ YPLP ID L+D G + S +D SGYHQI + P D KTAF
Sbjct: 710 RLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAF 769
Query: 827 MTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDL 886
T ++ + MPFGL NA A + ++M+ VF + + ++++D++V S H + L
Sbjct: 770 RTRYDHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHL 829
Query: 887 EEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEV 946
+R+H + KCSF + FLG +I+ +G+ ++PEK ++I++ P N E+
Sbjct: 830 RAVLERLREHELFAKLSKCSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEI 889
Query: 947 QRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKP 1006
+ G RF+ + P + K+ AF W+ ECE++F+ LK +L++ P+L P
Sbjct: 890 RSFLGLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLP 949
Query: 1007 IQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTA 1066
+G P +Y S L V++Q + ++ + S L+ E Y + AV+
Sbjct: 950 EEGEPYTVYTDASIVGLGCVLMQ----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFL 1005
Query: 1067 RRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSEYGLQY-----DKRGTVG 1120
+ R Y V++ TD L+ + +P+L+ R W +++Y L
Sbjct: 1006 KIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLRQRRWMELVADYNLDIAYHPGKANQVAD 1065
Query: 1121 AQSLADFVVELTPDRFERVDTQWTLFVDGSSNS---SGSGAGVTLEGPGELVLEQSLKFE 1177
A S VE + + V+ TL V+ S G GA + + L Q E
Sbjct: 1066 ALSRRRSEVEAERSQVDLVNMMGTLHVNALSKEVEPLGLGAADQADLLSRIRLAQERDEE 1125
Query: 1178 FK--ATNNQAEYE 1188
K A NN+ EY+
Sbjct: 1126 IKGWAQNNKTEYQ 1138
Score = 106 bits (264), Expect = 1e-22
Identities = 138/564 (24%), Positives = 235/564 (41%), Gaps = 49/564 (8%)
Query: 1143 WTLFVDGSSNSSGSGAGVTLEGPGELVL---EQSLKFEFKATNNQAEYEALIAGLKLARE 1199
+T++ D S G G L G ++ Q K E + E A++ LK+ R
Sbjct: 955 YTVYTDASI----VGLGCVLMQKGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRS 1010
Query: 1200 VKIRSLL-IRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQE--VVVEYVPRAEN 1256
+ + I TD + +K F + NL R R M L + + + Y P N
Sbjct: 1011 YLYGAKVQIYTDHK----SLKYIFTQPELNL-----RQRRWMELVADYNLDIAYHPGKAN 1061
Query: 1257 QRADALAKLASTRKPGNNKSVIQE---TLAYPSIEGELMACVNRGRTWMDPIISILAGDP 1313
Q ADAL++ S + ++ + TL ++ E+ D + I
Sbjct: 1062 QVADALSRRRSEVEAERSQVDLVNMMGTLHVNALSKEVEPLGLGAADQADLLSRIRLAQE 1121
Query: 1314 AEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKY--EAIMSEVHEGVCASHI 1371
+ E Q + + T +G + G CV ++ E I+ E H+ + H
Sbjct: 1122 RDEEIKGWAQNNKTEYQTSNNGTIVVNG-----RVCVPNDRALKEEILREAHQSKFSIHP 1176
Query: 1372 GGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAP-WPFAMW 1430
G + + L+ ++W ++KD +V +C CQ+ + P L + P W +
Sbjct: 1177 GSNKMY-RDLKRYYHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPIPEWKWDHI 1235
Query: 1431 GVDLVGPFPTA-RAQMKFILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPR 1488
+D V PT +++ + V VD TK ++ A+I+ Y IV GIP
Sbjct: 1236 TMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIVRLHGIPV 1295
Query: 1489 AIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKG 1548
+IVSD T+F+S + F K +G ++ ++ HPQT+ Q E + + LR + + G
Sbjct: 1296 SIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQTDEQSERTIQTLEDMLRACVLDWGG 1355
Query: 1549 AWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMA 1608
W L V ++YN + Q++ +P+ YG P+ W T G E +
Sbjct: 1356 NWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACRTPL-----CW-TPVG---ERRLFGP 1406
Query: 1609 VELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVLKWRSGAPG-------N 1661
+D +E I+ + R + N R + + QVGDLV G
Sbjct: 1407 TIVDETTERMKFLKIKLKEAQDRQKSYANKRRKELEFQVGDLVYLKAMTYKGAGRFTSRK 1466
Query: 1662 KLTPNWEGPYRIVKVLGNGAYHLE 1685
KL+P + GPY++++ +G AY L+
Sbjct: 1467 KLSPRYVGPYKVIERVGAVAYKLD 1490
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 224 bits (572), Expect = 3e-58
Identities = 133/407 (32%), Positives = 212/407 (51%), Gaps = 6/407 (1%)
Query: 707 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKW 766
+ L P P+S+ R+ + +++++ LL FIR P W A V+ VKK +G +
Sbjct: 483 IELEPGTAPLSKAPYRMAPAEMAELKKQLKDLLGKGFIRPSTSP-WGAPVLFVKKKDGSF 541
Query: 767 RMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAF 826
R+C+DY +LN+ K+ YPLP ID L+D G S +D SGYHQI + AD KTAF
Sbjct: 542 RLCIDYRELNRVTVKNRYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAF 601
Query: 827 MTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDL 886
T ++ + MPFGL NA A + RLM+ VF + + +++DD++V S + L
Sbjct: 602 RTRYGHFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEQEVHL 661
Query: 887 EEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEV 946
++R+ + KCSF + FLG ++++ G+ ++PEK +AI+ P+N E+
Sbjct: 662 RRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEI 721
Query: 947 QRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKP 1006
+ G RF+ + P K K+V F W+ ECEE FV LKE+L+S P+L+ P
Sbjct: 722 RSFLGWAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALP 781
Query: 1007 IQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTA 1066
G P +Y S L V++Q +++ + S L E Y + AV+
Sbjct: 782 EHGQPYMVYTDASRVGLGCVLMQ----HGKVIAYASRQLMKHEGNYPTHDLEMAAVIFAL 837
Query: 1067 RRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSEYGLQ 1112
+ R Y V+V TD L+ + +P+L+ R W +++Y L+
Sbjct: 838 KIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLE 884
Score = 30.4 bits (67), Expect = 9.2
Identities = 33/148 (22%), Positives = 62/148 (41%), Gaps = 7/148 (4%)
Query: 1381 LRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAP-WPFAMWGVDLVGPFP 1439
L+ + W ++ D ++V +C CQ+ + L ++ P W + +DLV
Sbjct: 965 LKRYYQWVGMKMDVANWVAECDVCQLVKAEHQVLGGMLQSLPIPEWKWDFITMDLVVGLR 1024
Query: 1440 TARAQMKFILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAI--VSDNGT 1496
+R + I V VD TK + K A ++ + IV G+P + D
Sbjct: 1025 VSRTK-DAIWVIVDRLTKSAHFLAIRKTDGAAVLAKKFVSEIVKLHGVPLNMKEAQDRQR 1083
Query: 1497 QFSSSQTREFCKEMG--IQMRFASVEHP 1522
++ + RE E+G + ++ A + P
Sbjct: 1084 SYADKRRRELEFEVGDRVYLKMAMLRGP 1111
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 221 bits (564), Expect = 2e-57
Identities = 131/407 (32%), Positives = 211/407 (51%), Gaps = 6/407 (1%)
Query: 707 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKW 766
+ L P P+S+ R+ + ++++++ LL FIR P W A V+ VKK +G +
Sbjct: 509 IELEPGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGFIRPSTSP-WGAPVLFVKKKDGSF 567
Query: 767 RMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAF 826
R+C+DY LN K+ YPLP ID L+D G S +D SGYHQI + AD KTAF
Sbjct: 568 RLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAF 627
Query: 827 MTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDL 886
T ++ + MPF L NA A + RLM+ VF + + +++DD++V S +H L
Sbjct: 628 RTRYGHFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVHL 687
Query: 887 EEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEV 946
++R+ + KCSF + FLG ++++ G+ ++PEK +AI+ P+N E+
Sbjct: 688 RRVMEKLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEI 747
Query: 947 QRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKP 1006
+ RF+ + P K K+V F W+ ECEE FV LKE+L+S P+L+ P
Sbjct: 748 RSFLRLTGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALP 807
Query: 1007 IQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTA 1066
G P +Y S L V++Q +++ + S L+ E Y + V+
Sbjct: 808 EHGQPYMVYTDASRVGLGCVLMQ----RGKVIAYASRQLRKHEGNYPTHDLEMAVVIFAL 863
Query: 1067 RRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSEYGLQ 1112
+ R Y V+V TD L+ + +P+L+ R + W +++Y L+
Sbjct: 864 KIWRSYLYGGKVQVFTDHKSLKYIFNQPELNLRQMRWMELVADYDLE 910
Score = 103 bits (257), Expect = 9e-22
Identities = 140/577 (24%), Positives = 234/577 (40%), Gaps = 87/577 (15%)
Query: 1151 SNSSGSGAGVTLEGPGELVL---EQSLKFEFKATNNQAEYEALIAGLKLAREVKIRSLLI 1207
+++S G G L G+++ Q K E + E +I LK+ R S L
Sbjct: 817 TDASRVGLGCVLMQRGKVIAYASRQLRKHEGNYPTHDLEMAVVIFALKIWR-----SYLY 871
Query: 1208 RTDSQLVENQVKGTFQVKDPNLIKYLERVRYL-MTLFQEVVVEYVPRAENQRADALAKLA 1266
Q+ + + P L L ++R++ + ++ + Y P N ADAL++
Sbjct: 872 GGKVQVFTDHKSLKYIFNQPEL--NLRQMRWMELVADYDLEIAYHPGKANVVADALSRKR 929
Query: 1267 STRKPGNNKSVIQETLAYPSIEGELMACVNRGRTWMDPIISILAGDPAEV---EQCTKEQ 1323
PG Q A S G L CV +P+ + A D A++ + +E+
Sbjct: 930 VGAAPG------QSVEALVSEIGALRLCV----VAREPL-GLEAVDRADLLTRARLAQEK 978
Query: 1324 QREASHYTLIDGHLYRRGFSTPLLK----CV--SPEKYEAIMSEVHEGVCASHIGGRSLA 1377
+ +G Y+ + + CV E I+SE H + + H G +
Sbjct: 979 DEGLIAASKAEGSEYQFAANGTIFVYGRVCVPKDEELRREILSEAHASMFSIHPGATKMY 1038
Query: 1378 CKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGP 1437
+ L+ + W +++D ++V +C CQ LV P A+W
Sbjct: 1039 -RDLKRYYQWVGMKRDVANWVAECDVCQ------------LVKAEHQVPDAIW------- 1078
Query: 1438 FPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGT 1496
V +D TK + K A ++ Y IV G+P +IVSD +
Sbjct: 1079 ------------VIMDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDS 1126
Query: 1497 QFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPA 1556
+F+ + R F +MG +++ ++ HPQT+GQ E + + LR + + G W D L
Sbjct: 1127 KFTFAFWRAFQAKMGTKVQMSTAYHPQTDGQSERTIQTLEDMLRMCVLDWGGHWADHLSL 1186
Query: 1557 VLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSE 1616
V ++YN + Q++ PF YG P+ R+ G D + E
Sbjct: 1187 VEFAYNNSYQASIGMAPFEALYGRPCWTPLRWTQVEERSIYG------------ADYVQE 1234
Query: 1617 TRDEAHIRETAMKQ---RVAAKFNSRVRVRDMQVGDLVLKWRSGAPG-------NKLTPN 1666
T + + + MK+ R + + R R + +VGD V + G KL+P
Sbjct: 1235 TTERIRVLKLNMKEAQARQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSILETKLSPR 1294
Query: 1667 WEGPYRIVKVLGNGAYHLEELD-GRRLPRSFNGLSLR 1702
+ GP+RIV+ +G AY LE D R + F+ L LR
Sbjct: 1295 YMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVLMLR 1331
>At3g11970 hypothetical protein
Length = 1499
Score = 213 bits (542), Expect = 8e-55
Identities = 136/412 (33%), Positives = 218/412 (52%), Gaps = 9/412 (2%)
Query: 705 HRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG 764
H++ L PV+Q R + + + V+ LL ++ P + + VV+VKK +G
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSP-YASPVVLVKKKDG 649
Query: 765 KWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKT 824
WR+CVDY +LN KDS+P+P I+ L+D G + S +D +GYHQ+RM P D KT
Sbjct: 650 TWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKT 709
Query: 825 AFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQ 884
AF T ++ Y MPFGL NA AT+Q LM+ +F + + + V+ DD++V S +H Q
Sbjct: 710 AFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQ 769
Query: 885 DLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVK 944
L++ F +R + + KC+F V ++LG I+++GIE +P K KA+++ P+ +K
Sbjct: 770 HLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLK 829
Query: 945 EVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILS 1004
+++ G RF+ G + P L K AFEWTA ++AF LK L P+LS
Sbjct: 830 QLRGFLGLAGYYRRFVRSFGVIAGP-LHALTKTDAFEWTAVAQQAFEDLKAALCQAPVLS 888
Query: 1005 KPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLV 1064
P+ + + +V++QE H + Y +S L+G ++ EK LAV+
Sbjct: 889 LPLFDKQFVVETDACGQGIGAVLMQE---GHPLAY-ISRQLKGKQLHLSIYEKELLAVIF 944
Query: 1065 TARRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSE--YGLQY 1113
R+ R Y ++TD L+ +L++ + W +L E Y +QY
Sbjct: 945 AVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQY 996
Score = 101 bits (251), Expect = 4e-21
Identities = 133/565 (23%), Positives = 234/565 (40%), Gaps = 80/565 (14%)
Query: 1147 VDGSSNSSGSGAGVTLEGPGELVLEQSLKF-EFKATNNQAEYEALIAGLKLAREVKIRS- 1204
V+ + G GA + EG + + LK + + + E A+I ++ R ++S
Sbjct: 898 VETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSH 957
Query: 1205 LLIRTDSQ----LVENQVKGTFQVKD-PNLIKYLERVRYLMTLFQEVVVEYVPRAENQRA 1259
+I+TD + L+E ++ Q + P L+++ + ++Y EN A
Sbjct: 958 FIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEF------------DYEIQYRQGKENVVA 1005
Query: 1260 DALAKLASTRKPGNNKSVIQETLAYPSIEGELMACVNRGR---TWMDPIISILAGDPAEV 1316
DAL+++ + V+ +A +E +L+ + G + + II+ L DP
Sbjct: 1006 DALSRVEGSE-------VLH--MAMTVVECDLLKDIQAGYANDSQLQDIITALQRDP--- 1053
Query: 1317 EQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSL 1376
++ Y ++ RR + ++ + I+ +H H GR +
Sbjct: 1054 ---------DSKKYFSWSQNILRR--KSKIVVPANDNIKNTILLWLHGSGVGGH-SGRDV 1101
Query: 1377 ACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWG---VD 1433
+ ++ FYW + KD +++ C CQ A P L + P P +W +D
Sbjct: 1102 THQRVKGLFYWKGMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPL--PIPDTIWSEVSMD 1159
Query: 1434 LVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSA-KIVNFYWKRIVCRFGIPRAIVS 1492
+ P + + I+V VD +K L+ SA + + Y + G P +IVS
Sbjct: 1160 FIEGLPVSGGKT-VIMVVVDRLSKAAHFIALSHPYSALTVAHAYLDNVFKLHGCPTSIVS 1218
Query: 1493 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLD 1552
D F+S REF G+ ++ S HPQ++GQ E NR + LR + W
Sbjct: 1219 DRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSK 1278
Query: 1553 ELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELD 1612
L + YNT S++R TPF + YG + PV + PG ++ +AV
Sbjct: 1279 WLALAEYWYNTNYHSSSRMTPFEIVYG--QVPPVHLPYL-----PG-----ESKVAVVAR 1326
Query: 1613 LLSETRD-----EAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL----KWRSGA----P 1659
L E D + H+ + + A + R+ ++GD V +R +
Sbjct: 1327 SLQEREDMLLFLKFHLMRAQHRMKQFA--DQHRTEREFEIGDYVYVKLQPYRQQSVVMRA 1384
Query: 1660 GNKLTPNWEGPYRIVKVLGNGAYHL 1684
KL+P + GPY+I+ G AY L
Sbjct: 1385 NQKLSPKYFGPYKIIDRCGEVAYKL 1409
>At1g36590 hypothetical protein
Length = 1499
Score = 213 bits (542), Expect = 8e-55
Identities = 136/412 (33%), Positives = 218/412 (52%), Gaps = 9/412 (2%)
Query: 705 HRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG 764
H++ L PV+Q R + + + V+ LL ++ P + + VV+VKK +G
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSP-YASPVVLVKKKDG 649
Query: 765 KWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKT 824
WR+CVDY +LN KDS+P+P I+ L+D G + S +D +GYHQ+RM P D KT
Sbjct: 650 TWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKT 709
Query: 825 AFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQ 884
AF T ++ Y MPFGL NA AT+Q LM+ +F + + + V+ DD++V S +H Q
Sbjct: 710 AFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQ 769
Query: 885 DLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVK 944
L++ F +R + + KC+F V ++LG I+++GIE +P K KA+++ P+ +K
Sbjct: 770 HLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLK 829
Query: 945 EVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILS 1004
+++ G RF+ G + P L K AFEWTA ++AF LK L P+LS
Sbjct: 830 QLRGFLGLAGYYRRFVRSFGVIAGP-LHALTKTDAFEWTAVAQQAFEDLKAALCQAPVLS 888
Query: 1005 KPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLV 1064
P+ + + +V++QE H + Y +S L+G ++ EK LAV+
Sbjct: 889 LPLFDKQFVVETDACGQGIGAVLMQE---GHPLAY-ISRQLKGKQLHLSIYEKELLAVIF 944
Query: 1065 TARRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSE--YGLQY 1113
R+ R Y ++TD L+ +L++ + W +L E Y +QY
Sbjct: 945 AVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQY 996
Score = 95.5 bits (236), Expect = 2e-19
Identities = 132/565 (23%), Positives = 232/565 (40%), Gaps = 80/565 (14%)
Query: 1147 VDGSSNSSGSGAGVTLEGPGELVLEQSLKF-EFKATNNQAEYEALIAGLKLAREVKIRS- 1204
V+ + G GA + EG + + LK + + + E A+I ++ R ++S
Sbjct: 898 VETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSH 957
Query: 1205 LLIRTDSQ----LVENQVKGTFQVKD-PNLIKYLERVRYLMTLFQEVVVEYVPRAENQRA 1259
+I+TD + L+E ++ Q + P L+++ + ++Y EN A
Sbjct: 958 FIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEF------------DYEIQYRQGKENVVA 1005
Query: 1260 DALAKLASTRKPGNNKSVIQETLAYPSIEGELMACVNRGR---TWMDPIISILAGDPAEV 1316
DAL+++ + V+ +A +E +L+ + G + + II+ L DP
Sbjct: 1006 DALSRVEGSE-------VLH--MAMTVVECDLLKDIQAGYANDSQLQDIITALQRDP--- 1053
Query: 1317 EQCTKEQQREASHYTLIDGHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSL 1376
++ Y ++ RR + ++ + I+ +H H GR +
Sbjct: 1054 ---------DSKKYFSWSQNILRR--KSKIVVPANDNIKNTILLWLHGSGVGGH-SGRDV 1101
Query: 1377 ACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWG---VD 1433
+ ++ FY + KD +++ C CQ A P L + P P +W +D
Sbjct: 1102 THQRVKGLFYSKGMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPL--PIPDTIWSEVSMD 1159
Query: 1434 LVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSA-KIVNFYWKRIVCRFGIPRAIVS 1492
+ P + + I+V VD +K L+ SA + Y + G P +IVS
Sbjct: 1160 FIEGLPVSGGKT-VIMVVVDRLSKAAHFIALSHPYSALTVAQAYLDNVFKLHGCPTSIVS 1218
Query: 1493 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLD 1552
D F+S REF G+ ++ S HPQ++GQ E NR + LR + W
Sbjct: 1219 DRDVVFTSEFWREFFTLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSK 1278
Query: 1553 ELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELD 1612
L + YNT S++R TPF + YG + PV + PG ++ +AV
Sbjct: 1279 WLALAEYWYNTNYHSSSRMTPFEIVYG--QVPPVHLPYL-----PG-----ESKVAVVAR 1326
Query: 1613 LLSETRD-----EAHIRETAMKQRVAAKFNSRVRVRDMQVGDLVL----KWRSGA----P 1659
L E D + H+ + + A + R+ ++GD V +R +
Sbjct: 1327 SLQEREDMLLFLKFHLMRAQHRMKQFA--DQHRTEREFEIGDYVYVKLQPYRQQSVVMRA 1384
Query: 1660 GNKLTPNWEGPYRIVKVLGNGAYHL 1684
KL+P + GPY+I+ G AY L
Sbjct: 1385 NQKLSPKYFGPYKIIDRCGEVAYKL 1409
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 213 bits (541), Expect = 1e-54
Identities = 128/407 (31%), Positives = 208/407 (50%), Gaps = 18/407 (4%)
Query: 707 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKW 766
+ L P P+S+ R+ + ++++++ LL A V+ VKK +G +
Sbjct: 470 IELEPGTAPLSKAPYRMAPAEMAELKKQLEDLLGAP-------------VLFVKKKDGSF 516
Query: 767 RMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAF 826
R+C+DY LN K+ YPLP ID L+D G S +D SGYH I + AD KTAF
Sbjct: 517 RLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADVRKTAF 576
Query: 827 MTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDL 886
T ++ + MPFGL NA A + RLM+ VF + + +++DD++V S +H L
Sbjct: 577 RTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHEVHL 636
Query: 887 EEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEV 946
++R+ + KCSF + FLG ++++ G+ ++PEK +AI+ +P+N E+
Sbjct: 637 RRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTPTNATEI 696
Query: 947 QRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKP 1006
+ G RF+ + P K K+V F W+ ECEE FV LKE+L+S P+L+ P
Sbjct: 697 RSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALP 756
Query: 1007 IQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTA 1066
G P +Y S L V++Q +++ + S L+ E Y + AV+
Sbjct: 757 EHGEPYMVYTDASGVGLGCVLMQ----RGKVIAYASRQLRKHEGNYPTHDLEMAAVIFAL 812
Query: 1067 RRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSEYGLQ 1112
+ R Y V+V TD L+ + +P+L+ R W +++Y L+
Sbjct: 813 KIWRSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLE 859
Score = 100 bits (250), Expect = 6e-21
Identities = 87/312 (27%), Positives = 140/312 (43%), Gaps = 25/312 (8%)
Query: 1392 KDCMDFVKQCKECQVFADLSKAPPKELVTMSAP-WPFAMWGVDLVGPFPTARAQMKFILV 1450
+D ++V +C CQ+ + P L ++ P W + +D V P +R + I V
Sbjct: 941 EDVANWVAECDVCQLVKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLPVSRTK-DAIWV 999
Query: 1451 AVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKE 1509
VD TK + K A ++ Y IV G+P +IVSD ++F+S+ R F E
Sbjct: 1000 IVDRLTKSAHFLAIRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAE 1059
Query: 1510 MGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTT 1569
MG +++ ++ HPQT GQ E + + LR + + G W D L V ++YN + ++
Sbjct: 1060 MGTKVQMSTAYHPQTYGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPASI 1119
Query: 1570 RETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMK 1629
PF Y P+ + R+ G D + ET + + + MK
Sbjct: 1120 GMAPFEALYERPCRTPLCLTQVGERSIYG------------ADYVQETTERIRVLKLNMK 1167
Query: 1630 Q---RVAAKFNSRVRVRDMQVGDLVLKWRSGAPG-------NKLTPNWEGPYRIVKVLGN 1679
+ R + + R R + +VGD V + G KL+P + GP+RIV+ +G
Sbjct: 1168 EAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSISETKLSPRYMGPFRIVERVGP 1227
Query: 1680 GAYHLEELDGRR 1691
AY LE D R
Sbjct: 1228 VAYRLELPDVMR 1239
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 212 bits (540), Expect = 1e-54
Identities = 118/366 (32%), Positives = 194/366 (52%), Gaps = 5/366 (1%)
Query: 707 LALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANGKW 766
+ L P P+S+ R+ + +++++++LLA FIR P W A V+ VKK +G +
Sbjct: 146 IELEPGTTPISKAPYRMAPAEMAELKKQLEELLAKGFIRPSSSP-WGAPVLFVKKKDGSF 204
Query: 767 RMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKTAF 826
R+C+DY LNK K+ YPLP ID L+D G + S +D SGYHQI + P D KTAF
Sbjct: 205 RLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAF 264
Query: 827 MTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQDL 886
T ++ + MPFGL NA A + ++M+ VF + + +++DD++V S H + L
Sbjct: 265 RTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFIDDILVHSKSWEAHQEHL 324
Query: 887 EEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVKEV 946
+R+H + K SF + FLG +I+ +G+ ++PEK ++I++ P N E+
Sbjct: 325 RAVLERLREHELFAKLSKFSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEI 384
Query: 947 QRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILSKP 1006
+ G RF+ + P + K+ AF W+ ECE++F+ LK +L + P+L P
Sbjct: 385 RSFLGLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLINAPVLVLP 444
Query: 1007 IQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLVTA 1066
+G P +Y S L V++Q + ++ + S L+ E Y + AV+
Sbjct: 445 EEGEPYTVYTDASIVGLGCVLMQ----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFAL 500
Query: 1067 RRLRPY 1072
+ R Y
Sbjct: 501 KIWRSY 506
Score = 109 bits (272), Expect = 2e-23
Identities = 105/414 (25%), Positives = 183/414 (43%), Gaps = 29/414 (7%)
Query: 1284 YPSIEGELMACVNRGRTWMDPIISILAGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFS 1343
YP+ + E+ A V + W +I + E++ T + E + T +G + G
Sbjct: 486 YPTHDLEMAAVVFALKIWRSYLIRLAQERDEEIKGWTLNNKTE--YQTSNNGTIVVNG-- 541
Query: 1344 TPLLKCVSPEKY--EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQC 1401
CV ++ E I+ E H+ + H G + + L+ ++W +RKD +V +C
Sbjct: 542 ---RVCVPNDRALKEEILREAHQSKFSIHPGSNKMY-RDLKRYYHWVGMRKDVARWVAKC 597
Query: 1402 KECQVFADLSKAPPKELVTMS-APWPFAMWGVDLVGPFPTA-RAQMKFILVAVDYFTKWI 1459
CQ+ + P L + + W + +D V PT +++ + V VD TK
Sbjct: 598 PTCQLVKAEHQVPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSA 657
Query: 1460 EAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFAS 1518
++ A+I+ Y I+ GIP +IVSD T+F+S F K +G ++ ++
Sbjct: 658 HFMAISDKDGAEIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLST 717
Query: 1519 VEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTY 1578
HPQT+GQ E + + LR + + G W L + ++YN + Q++ +P+ Y
Sbjct: 718 AYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALY 777
Query: 1579 GVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNS 1638
G P+ W T G E + +D +E I+ + R + N
Sbjct: 778 GRACRTPL-----CW-TPVG---ERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANK 828
Query: 1639 RVRVRDMQVGDLVLKWRSGAPG-------NKLTPNWEGPYRIVKVLGNGAYHLE 1685
R + + QVGDLV G KL+P + GPY++++ +G AY L+
Sbjct: 829 RRKELEFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLD 882
>At2g06890 putative retroelement integrase
Length = 1215
Score = 188 bits (478), Expect = 2e-47
Identities = 146/560 (26%), Positives = 252/560 (44%), Gaps = 37/560 (6%)
Query: 564 VIIGRNTLNRLCAVI--STAHLAVKYPLSSGKVGKLKVDQKMARECYNN--CLNFYGKKS 619
++IG+ LC V+ H+ + P S D+K+ + + N F G K+
Sbjct: 287 IVIGKYEDEILCDVLPMEAGHILLGRPWQS--------DRKVMHDGFTNRHSFEFKGGKT 338
Query: 620 ALVGHRCYEIEASDENLDPRGEGRVNRPTPIEETKALKFGDRTLKIGTRLTEEQETRLTK 679
LV +E+ +L + E V +P F ++ ++ + + +Q L
Sbjct: 339 ILVSMTPHEVYQDQIHLKQKKEQVVKQPN---------FFAKSGEVKSAYSSKQPMLLF- 388
Query: 680 LLGENLDLFAWSCKDMPGIDPNFICHRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLL 739
+F + + P +L K V G + ++ ++D +
Sbjct: 389 -------VFKEALTSLTNFAPVLPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVP 441
Query: 740 AAEFIREVKYPTW-LANVVMVKKANGKWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASG 798
A Y T + + ++ +G WRMC D +N K +P+P +D ++D G
Sbjct: 442 GASLPNRPAYRTNPVETKELQRQKDGSWRMCFDCRAINNVTVKYCHPIPRLDDMLDELHG 501
Query: 799 NELLSLMDAYSGYHQIRMHPADEDKTAFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFA 858
+ + S +D SGYHQIRM+ DE KTAF T Y + MPFGL +A +T+ RLM+ V
Sbjct: 502 SSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKHGLYEWLVMPFGLTHAPSTFMRLMNHVLR 561
Query: 859 GQVGRNMEVYVDDMIVKSVRGLDHHQDLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFM 918
+G + VY DD++V S +H + L+ +RK + N +KC+F FLGF+
Sbjct: 562 AFIGIFVIVYFDDILVYSESLREHIEHLDSVLNVLRKEELYANLKKCTFCTDNLVFLGFV 621
Query: 919 ITSRGIEINPEKCKAIQQMKSPSNVKEVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNV 978
+++ G++++ EK KAI+ SP V EV+ G RF P + ++K+V
Sbjct: 622 VSADGVKVDEEKVKAIRDWPSPKTVGEVRSFHGLAGFYRRFFKDFSTIVAPLTEVMKKDV 681
Query: 979 AFEWTAECEEAFVRLKELLSSPPILSKPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIV 1038
F+W EEAF LK+ L++ P+L + S + +V++Q + +++
Sbjct: 682 GFKWEKAQEEAFQSLKDKLTNAPVLILSEFLKTFEIECDASGIGIGAVLMQ----DQKLI 737
Query: 1039 YFVSHTLQGAEVRYQKIEKAALAVLVTARRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSG 1097
F S L GA + Y +K A++ +R + Y + TD L+ + + L+
Sbjct: 738 AFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLWPKVFVIHTDHESLKHLKGQQKLNK 797
Query: 1098 RLVAW--SVELSEYGLQYDK 1115
R W +E Y ++Y K
Sbjct: 798 RHARWVEFIETFAYVIKYKK 817
Score = 38.1 bits (87), Expect = 0.044
Identities = 30/122 (24%), Positives = 57/122 (46%), Gaps = 11/122 (9%)
Query: 1572 TPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQR 1631
+PF++ YG + + P ++ R + + +A +L+ + + A +
Sbjct: 988 SPFQIVYGFNPISPFDLIPLPLSERVSIDGKKKA------ELVQQIHENARRNIEEKTKL 1041
Query: 1632 VAAKFNSRVRVRDMQVGDLVL----KWRSGAPG-NKLTPNWEGPYRIVKVLGNGAYHLEE 1686
A + N R + +VGD+V K R A +KL P +GP++I+K + + AY L+
Sbjct: 1042 YAKQANKGRREQIFEVGDMVWIHLRKERFPAQRKSKLMPRIDGPFKIIKRINDNAYQLDL 1101
Query: 1687 LD 1688
D
Sbjct: 1102 QD 1103
>At2g13060 pseudogene
Length = 930
Score = 184 bits (467), Expect = 4e-46
Identities = 117/384 (30%), Positives = 185/384 (47%), Gaps = 17/384 (4%)
Query: 1279 QETLAYPSIEGELMACVNRGRTWMDPIISILAGDPAEVEQCT----KEQQREASHYTLID 1334
+ T + S E + V + W I++ LA D E + T K RE Y +
Sbjct: 116 ESTRKWSSTENCAVTAVEKDYPWYADIVNYLAAD-VEPDNFTDYNKKRFLREIRRYQWDE 174
Query: 1335 GHLYRRGFSTPLLKCVSPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDC 1394
+LY+ + +C++ + +I+S H H KVL+A F+WPT+ +D
Sbjct: 175 PYLYKHSYDGIYRRCIAATEVPSILSHCHSSSYGGHFATFKTVSKVLQADFWWPTMFRDT 234
Query: 1395 MDFVKQCKECQVFADLSKA---PPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVA 1451
F+ QC CQ +SK PP + + F WG+D +GPFP + + +ILV
Sbjct: 235 QKFISQCDPCQRRGKISKRNEMPPNFRLEVEV---FDRWGIDFMGPFPPSNKNL-YILVD 290
Query: 1452 VDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMG 1511
VDY +KW+EA K SA ++ + I RFG+PR ++SD F + + + G
Sbjct: 291 VDYVSKWVEAIASLKNDSAVVMKLFKSIIFPRFGVPRIVISDGDKHFINKILEKLLLQYG 350
Query: 1512 IQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRE 1571
+Q R A+ HPQT+GQVE +NR I L + + +AK W +L LW+Y T ++
Sbjct: 351 VQHRVATPYHPQTSGQVEVSNRQIKEILEKTVGKAKKEWSYKLYDALWAYKTAFKTPLGT 410
Query: 1572 TPFRMTYGVDAMLPVEIDN-FTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQ 1630
TPF + YG LPVE+++ W + + A + ELD E R A+ K+
Sbjct: 411 TPFHLLYGKACHLPVELEHKAAWAVKMMNFDIKSAGES-ELD---EIRIHAYDNSKLYKE 466
Query: 1631 RVAAKFNSRVRVRDMQVGDLVLKW 1654
R A + ++ R + D +L++
Sbjct: 467 RTKAYHDKKILTRTFEPNDQILRF 490
>At1g35370 hypothetical protein
Length = 1447
Score = 180 bits (457), Expect = 6e-45
Identities = 125/412 (30%), Positives = 208/412 (50%), Gaps = 17/412 (4%)
Query: 705 HRLALNPSVKPVSQLRRRLGGDKGKAVQQEVDKLLAAEFIREVKYPTWLANVVMVKKANG 764
H++ L PV+Q R + + + V ++ + I+ P + + VV+VKK +G
Sbjct: 570 HKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQVSSSP-FASPVVLVKKKDG 628
Query: 765 KWRMCVDYTDLNKACPKDSYPLPSIDSLVDGASGNELLSLMDAYSGYHQIRMHPADEDKT 824
WR+CVDYT+LN KD + +P I+ L+D G+ + S +D +GYHQ+RM P D KT
Sbjct: 629 TWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDPDDIQKT 688
Query: 825 AFMTARVNYCYRTMPFGLKNAGATYQRLMDRVFAGQVGRNMEVYVDDMIVKSVRGLDHHQ 884
AF T ++ Y M FGL NA AT+Q LM+ VF + + + V+ DD+++ S +H +
Sbjct: 689 AFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSSIEEHKE 748
Query: 885 DLEEAFGEIRKHSMRLNPEKCSFGVQGGKFLGFMITSRGIEINPEKCKAIQQMKSPSNVK 944
L F +R H + F + LG I++R IE +P K +A+++ +P+ VK
Sbjct: 749 HLRLVFEVMRLHKL--------FAKGSKEHLGHFISAREIETDPAKIQAVKEWPTPTTVK 800
Query: 945 EVQRLTGRIAALSRFLPQSGDRSFPFFKCLRKNVAFEWTAECEEAFVRLKELLSSPPILS 1004
+V+ G RF+ G + P L K F W+ E + AF LK +L + P+L+
Sbjct: 801 QVRGFLGFAGYYRRFVRNFGVIAGP-LHALTKTDGFCWSLEAQSAFDTLKAVLCNAPVLA 859
Query: 1005 KPIQGHPLHLYFAVSDSALSSVMLQEIDGEHRIVYFVSHTLQGAEVRYQKIEKAALAVLV 1064
P+ + + +V++Q+ H + Y +S L+G ++ EK LA +
Sbjct: 860 LPVFDKQFMVETDACGQGIRAVLMQK---GHPLAY-ISRQLKGKQLHLSIYEKELLAFIF 915
Query: 1065 TARRLRPYFQSFPVKVRTD-LPLRQVLQKPDLSGRLVAWSVELSE--YGLQY 1113
R+ R Y ++TD L+ +L++ + W +L E Y +QY
Sbjct: 916 AVRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQY 967
Score = 101 bits (252), Expect = 3e-21
Identities = 90/347 (25%), Positives = 158/347 (44%), Gaps = 30/347 (8%)
Query: 1348 KCVSPEKYEAIMSEVHEGVCASHIGGRS---LACKVLRAGFYWPTLRKDCMDFVKQCKEC 1404
K V P E I +++ + + S +GGRS + + +++ FYW + KD F++ C C
Sbjct: 1042 KIVVPNDVE-ITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFIRSCGTC 1100
Query: 1405 QVFADLSKAPPKELVTMSAPWPFAMW---GVDLVGPFPTARAQMKFILVAVDYFTKWIEA 1461
Q + A P L + P P +W +D + P + + I+V VD +K
Sbjct: 1101 QQCKSDNAAYPGLLQPL--PIPDKIWCDVSMDFIEGLPNSGGK-SVIMVVVDRLSKAAHF 1157
Query: 1462 EPLAKITSA-KIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVE 1520
LA SA + + + G P +IVSD F+S +EF K G+++R +S
Sbjct: 1158 VALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKLQGVELRMSSAY 1217
Query: 1521 HPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGV 1580
HPQ++GQ E NR + LR W LP + YNT S+++ TPF + YG
Sbjct: 1218 HPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSSQMTPFELVYG- 1276
Query: 1581 DAMLPVEIDNFTWRTR-----PGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAK 1635
P+ + +++ +E + ++ L+ R + +++ A + R
Sbjct: 1277 -QAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLM---RAQHRMKQFADQHRTERT 1332
Query: 1636 FN----SRVRVRDMQVGDLVLKWRSGAPGNKLTPNWEGPYRIVKVLG 1678
F+ V+++ + +VL+ KL+P + GPY+I++ G
Sbjct: 1333 FDIGDFVYVKLQPYRQQSVVLR-----VNQKLSPKYFGPYKIIEKCG 1374
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,525,336
Number of Sequences: 26719
Number of extensions: 1706825
Number of successful extensions: 5118
Number of sequences better than 10.0: 205
Number of HSP's better than 10.0 without gapping: 137
Number of HSP's successfully gapped in prelim test: 69
Number of HSP's that attempted gapping in prelim test: 4604
Number of HSP's gapped (non-prelim): 415
length of query: 1706
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1593
effective length of database: 8,299,349
effective search space: 13220862957
effective search space used: 13220862957
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0590c.9


