

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.8
(919 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
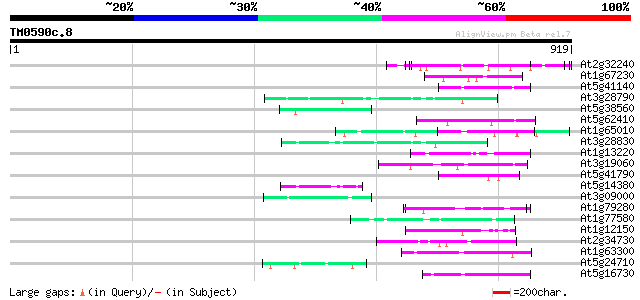
Sequences producing significant alignments: (bits) Value
At2g32240 putative myosin heavy chain 53 9e-07
At1g67230 unknown protein 51 3e-06
At5g41140 putative protein 50 4e-06
At3g28790 unknown protein 50 4e-06
At5g38560 putative protein 50 6e-06
At5g62410 chromosomal protein - like 50 8e-06
At1g65010 hypothetical protein 49 1e-05
At3g28830 hypothetical protein 48 2e-05
At1g13220 putative nuclear matrix constituent protein 48 2e-05
At3g19060 hypothetical protein 48 3e-05
At5g41790 myosin heavy chain-like protein 47 4e-05
At5g14380 agp6 47 5e-05
At3g09000 unknown protein 47 5e-05
At1g79280 hypothetical protein 47 6e-05
At1g77580 unknown protein 46 8e-05
At1g12150 hypothetical protein 46 8e-05
At2g34730 putative myosin heavy chain 45 1e-04
At1g63300 hypothetical protein 45 2e-04
At5g24710 unknown protein 45 2e-04
At5g16730 putative protein 45 2e-04
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 52.8 bits (125), Expect = 9e-07
Identities = 72/310 (23%), Positives = 127/310 (40%), Gaps = 27/310 (8%)
Query: 617 IEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWE 676
+E E L Q L++ E L E+F + + EA L + + ++E
Sbjct: 791 MELEALHQSLSIDSEHRLQ----------KAMEEFTSRDSEASSLTEKLRDLEGKIKSYE 840
Query: 677 KRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADEL-DASLQACKKEKEQA 735
++LA +GK +K+ T G ++ E +K+E D+ + SLQ+ + + A
Sbjct: 841 ---EQLAEASGKSSSLKEKLEQTLG-RLAAAESVNEKLKQEFDQAQEKSLQSSSESELLA 896
Query: 736 E-----KDLIARGEALIAKES-ELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVM 789
E K I E LI S E LE + ++E +S++ + K+ +
Sbjct: 897 ETNNQLKIKIQELEGLIGSGSVEKETALKRLEEAIERFNQKETESSDLVEKLKTHENQI- 955
Query: 790 QATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIALDN 849
EE KK + T+ E+ L+K+K+LE + AK +++ D+A N
Sbjct: 956 ----EEYKKLAHEASGVADTRKVELEDALSKLKNLESTIEELGAKCQGLEKESGDLAEVN 1011
Query: 850 RERGFYLAKDQAQHLFPNFDFSAMGVMKEITAAGLVGPDDPPLIDQNLWTATEEEEEEEE 909
+ LA ++ SA+ KE TA L + D +E E+ + +
Sbjct: 1012 LKLNLELANHGSEANELQTKLSALEAEKEQTANELEA-SKTTIEDLTKQLTSEGEKLQSQ 1070
Query: 910 EEEQEKENNE 919
+ENN+
Sbjct: 1071 ISSHTEENNQ 1080
Score = 43.5 bits (101), Expect = 5e-04
Identities = 41/200 (20%), Positives = 93/200 (46%), Gaps = 11/200 (5%)
Query: 655 NVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALM 714
N+E +E+ + Q +A E ++ A + K + + + +L+ Q++
Sbjct: 1015 NLELANHGSEANELQTKLSALEAEKEQTANELEASKTTIEDLTKQLTSEGEKLQSQISSH 1074
Query: 715 KEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKS 774
EE ++++A Q+ K+E + L E L + S+ L +E+E ++ AE+
Sbjct: 1075 TEENNQVNAMFQSTKEELQSVIAKL---EEQLTVESSKADTLVSEIEKLRAVAAEKSVLE 1131
Query: 775 AESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSL--EDELATEK 832
+ ++E + ++K+ E A A + K AE+ S+L + + + E ++ E+
Sbjct: 1132 SHF-----EELEKTLSEVKAQLKENVENAATA-SVKVAELTSKLQEHEHIAGERDVLNEQ 1185
Query: 833 AKAIEAREQAADIALDNRER 852
++ QAA ++D +++
Sbjct: 1186 VLQLQKELQAAQSSIDEQKQ 1205
Score = 43.1 bits (100), Expect = 7e-04
Identities = 59/280 (21%), Positives = 118/280 (42%), Gaps = 44/280 (15%)
Query: 659 ERLKAESAKHQEA----AAAWEKRFD-----KLATQAGKDKVYADKMIGTAGIKIGELED 709
++LK+ H EA AAA +K + + ++QA ++ A +I ELE
Sbjct: 452 QKLKSLEELHSEAGSAAAAATQKNLELEDVVRSSSQAAEE----------AKSQIKELET 501
Query: 710 QLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAE 769
+ +++ EL+ L + + AE++L E ++ + + E + + E
Sbjct: 502 KFTAAEQKNAELEQQLNLLQLKSSDAERELKELSEKSSELQTAIEVAEEEKKQATTQMQE 561
Query: 770 QEKKSAE-SLALAKS---------DMEAVMQATSEEIKKATETHAEALATKDAEIASQL- 818
++K++E L+L +S D+ +Q +E +A TH ++ + +SQ
Sbjct: 562 YKQKASELELSLTQSSARNSELEEDLRIALQKGAEHEDRANTTHQRSIELEGLCQSSQSK 621
Query: 819 -----AKIKSLEDELATEKAKAIEAREQAADIALDNRE-----RGFYLAKDQAQHLFPNF 868
++K LE L TEK + E EQ + + + E +G+ + Q F
Sbjct: 622 HEDAEGRLKDLELLLQTEKYRIQELEEQVSSLEKKHGETEADSKGYLGQVAELQSTLEAF 681
Query: 869 DFSAMGVMKEITAAGLVGPDDPPLIDQNLWTATEEEEEEE 908
+ + AA + ++ + +NL T E+++ E
Sbjct: 682 QVKS----SSLEAALNIATENEKELTENLNAVTSEKKKLE 717
Score = 43.1 bits (100), Expect = 7e-04
Identities = 70/276 (25%), Positives = 119/276 (42%), Gaps = 34/276 (12%)
Query: 649 EKFNAANVEAERLKAESAKHQEA---AAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIG 705
E F+A +E E + + + +E +A ++F++L Q+ +AD A ++
Sbjct: 184 EAFDALGIELESSRKKLIELEEGLKRSAEEAQKFEELHKQSAS---HADSESQKA-LEFS 239
Query: 706 ELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKK 765
EL +E +E ASLQ KE + + AL + ELA + EL L K
Sbjct: 240 ELLKSTKESAKEMEEKMASLQQEIKELNEKMSENEKVEAALKSSAGELAAVQEELALSKS 299
Query: 766 ALAEQEKKSAESLALAKSDMEAVMQATSE-EIKKATETHAEALATKDAEIASQLAKIKSL 824
L E E+K + + AL + + T E E KKA+E + +L+ ++ L
Sbjct: 300 RLLETEQKVSSTEAL-------IDELTQELEQKKASE----------SRFKEELSVLQDL 342
Query: 825 EDELATEKAKAIEAREQAADIALDNRERGFY--LAKDQAQHL-FPNFDFSAMGVMKEITA 881
+ + +AK E + +A + +E+ L+KDQ + L N + + KE
Sbjct: 343 DAQTKGLQAKLSEQEGINSKLAEELKEKELLESLSKDQEEKLRTANEKLAEVLKEKEALE 402
Query: 882 AGLVGPDDPPLIDQNLWTATEEEEEEEEEEEQEKEN 917
A + + N+ T TE E EE+ + EN
Sbjct: 403 ANVAE------VTSNVATVTEVCNELEEKLKTSDEN 432
Score = 42.0 bits (97), Expect = 0.002
Identities = 41/193 (21%), Positives = 89/193 (45%), Gaps = 16/193 (8%)
Query: 654 ANVEAERLKAESAKHQ-EAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLA 712
+ +EA ++K+ S + A EK + +K + + +KI E E+ L
Sbjct: 676 STLEAFQVKSSSLEAALNIATENEKELTENLNAVTSEKKKLEATVDEYSVKISESENLLE 735
Query: 713 LMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEK 772
++ E + L E E DL A G +ESE+ +L+ +++L EQ+
Sbjct: 736 SIRNELNVTQGKL-------ESIENDLKAAG----LQESEVM---EKLKSAEESL-EQKG 780
Query: 773 KSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEK 832
+ + + ++EA+ Q+ S + + + E ++D+E +S K++ LE ++ + +
Sbjct: 781 REIDEATTKRMELEALHQSLSIDSEHRLQKAMEEFTSRDSEASSLTEKLRDLEGKIKSYE 840
Query: 833 AKAIEAREQAADI 845
+ EA +++ +
Sbjct: 841 EQLAEASGKSSSL 853
Score = 41.2 bits (95), Expect = 0.003
Identities = 52/213 (24%), Positives = 87/213 (40%), Gaps = 14/213 (6%)
Query: 707 LEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKA 766
+E+Q +++ + L +++ ++ E +L L ESE L EL K+
Sbjct: 70 VEEQKEVIERSSSGSQRELHESQEKAKELELELERVAGELKRYESENTHLKDELLSAKEK 129
Query: 767 LAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLED 826
L E EKK D+E V + E+I + E H+ L + + + S AK K L +
Sbjct: 130 LEETEKK--------HGDLEVVQKKQQEKIVEGEERHSSQLKSLEDALQSHDAKDKELTE 181
Query: 827 ELATEKAKAIEAREQAADIALDNRERGFYLAKDQAQHLFPNFDFSAMGVMKEITAAGLVG 886
A IE +++ L E G + ++AQ SA E A
Sbjct: 182 VKEAFDALGIEL--ESSRKKLIELEEGLKRSAEEAQKFEELHKQSASHADSESQKA---- 235
Query: 887 PDDPPLIDQNLWTATEEEEEEEEEEEQEKENNE 919
+ L+ +A E EE+ +++ KE NE
Sbjct: 236 LEFSELLKSTKESAKEMEEKMASLQQEIKELNE 268
Score = 37.0 bits (84), Expect = 0.051
Identities = 54/250 (21%), Positives = 99/250 (39%), Gaps = 29/250 (11%)
Query: 577 EAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLAC 636
EA + + A E ++ ++ E L + ++ E ++ TKEE +
Sbjct: 1036 EAEKEQTANELEASKTTIEDLTKQLTSEGEKLQSQISSHTEENNQVNAMFQSTKEELQS- 1094
Query: 637 LLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAA---AWEKRFDKLATQAGKDKVYA 693
+ A E+ + +A+ L +E K + AA E F++L + K
Sbjct: 1095 ------VIAKLEEQLTVESSKADTLVSEIEKLRAVAAEKSVLESHFEELEKTLSEVKAQL 1148
Query: 694 DKMI---GTAGIKIGELEDQLALMKE---EADELDASLQACKKEKEQAEKDLIARGEALI 747
+ + TA +K+ EL +L + E D L+ + +KE + A+ + + +A
Sbjct: 1149 KENVENAATASVKVAELTSKLQEHEHIAGERDVLNEQVLQLQKELQAAQSSIDEQKQAHS 1208
Query: 748 AKESELAILCAELELVKKALAEQ---EKKSAESLALAKSDMEAVMQATSEEIKKATETHA 804
K+SEL E K E+ +KK+ D+E +Q + K ET A
Sbjct: 1209 QKQSEL-------ESALKKSQEEIEAKKKAVTEFESMVKDLEQKVQLADAKTK---ETEA 1258
Query: 805 EALATKDAEI 814
+ K +I
Sbjct: 1259 MDVGVKSRDI 1268
Score = 36.6 bits (83), Expect = 0.067
Identities = 58/260 (22%), Positives = 104/260 (39%), Gaps = 15/260 (5%)
Query: 657 EAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKE 716
E++ E E A KR++ T + + A + + K G+LE +E
Sbjct: 90 ESQEKAKELELELERVAGELKRYESENTHLKDELLSAKEKLEETEKKHGDLEVVQKKQQE 149
Query: 717 EADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAE 776
+ E + + K E A + A+ + L + L ELE +K L E E+
Sbjct: 150 KIVEGEERHSSQLKSLEDALQSHDAKDKELTEVKEAFDALGIELESSRKKLIELEE---- 205
Query: 777 SLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAI 836
L +S EA EE+ K + +HA++ + K E + L K E+ EK ++
Sbjct: 206 --GLKRSAEEA---QKFEELHKQSASHADSESQKALEFSELLKSTKESAKEM-EEKMASL 259
Query: 837 EAREQAADIALDNRERGFYLAKDQAQHLFPNFDFSAMGVMKEITAAGLVGPDDPPLIDQN 896
+ + + + E+ K A L + A+ + + V + LID+
Sbjct: 260 QQEIKELNEKMSENEKVEAALKSSAGELAAVQEELALSKSRLLETEQKVSSTE-ALIDE- 317
Query: 897 LWTATEEEEEEEEEEEQEKE 916
T+E E+++ E + KE
Sbjct: 318 ---LTQELEQKKASESRFKE 334
>At1g67230 unknown protein
Length = 1132
Score = 50.8 bits (120), Expect = 3e-06
Identities = 51/188 (27%), Positives = 82/188 (43%), Gaps = 32/188 (17%)
Query: 680 DKLATQAGKDKVYADKMIGTAGIKIGELED-------QLALMKEEADELDASLQACKKEK 732
DK+ Q GK+ A K I A + + +LED LAL ++E D L S++ +E
Sbjct: 264 DKIIKQKGKELEEAQKKIDAANLAVKKLEDDVSSRIKDLALREQETDVLKKSIETKAREL 323
Query: 733 EQAEKDLIARGEALIAK-----ESELAILCAELEL------------VKKALAEQEKKSA 775
+ ++ L AR + + + +++L E EL +K +AE EK+ A
Sbjct: 324 QALQEKLEAREKMAVQQLVDEHQAKLDSTQREFELEMEQKRKSIDDSLKSKVAEVEKREA 383
Query: 776 ESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAE---IASQLAKIKSLEDELATEK 832
E ME + + + + E H E D I+ + +KS E L TEK
Sbjct: 384 E-----WKHMEEKVAKREQALDRKLEKHKEKENDFDLRLKGISGREKALKSEEKALETEK 438
Query: 833 AKAIEARE 840
K +E +E
Sbjct: 439 KKLLEDKE 446
Score = 36.2 bits (82), Expect = 0.087
Identities = 32/146 (21%), Positives = 69/146 (46%), Gaps = 10/146 (6%)
Query: 703 KIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEAL--IAKESE-----LAI 755
+I + Q L+++EA++L A ++ +KE E+ ++ G L I + E + +
Sbjct: 496 QIEKCRSQQELLQKEAEDLKAQRESFEKEWEELDERKAKIGNELKNITDQKEKLERHIHL 555
Query: 756 LCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIA 815
L+ K+A E ++ E+L +AK+ M+ + K E+ L +I
Sbjct: 556 EEERLKKEKQAANENMERELETLEVAKASFAETMEYERSMLSKKAESERSQLL---HDIE 612
Query: 816 SQLAKIKSLEDELATEKAKAIEAREQ 841
+ K++S + EK + ++A+++
Sbjct: 613 MRKRKLESDMQTILEEKERELQAKKK 638
>At5g41140 putative protein
Length = 983
Score = 50.4 bits (119), Expect = 4e-06
Identities = 41/156 (26%), Positives = 75/156 (47%), Gaps = 10/156 (6%)
Query: 703 KIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELEL 762
K+ EL + L +E + A L+ K++KE DL + ++ E+ IL +LE
Sbjct: 693 KLNELSGKTDLKTKEMKRMSADLEYQKRQKEDVNADLT---HEITRRKDEIEILRLDLEE 749
Query: 763 VKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKA---TETHAEALATKDAEIAS--- 816
+K+ E E +E L + EAV+ A +++ A + +L+ ++EI +
Sbjct: 750 TRKSSMETEASLSEELQRIIDEKEAVITALKSQLETAIAPCDNLKHSLSNNESEIENLRK 809
Query: 817 QLAKIKSLEDELATEKAKAIEAREQAADIALDNRER 852
Q+ +++S E E E+ +E RE +AD +R
Sbjct: 810 QVVQVRS-ELEKKEEEMANLENREASADNITKTEQR 844
Score = 35.4 bits (80), Expect = 0.15
Identities = 54/232 (23%), Positives = 96/232 (41%), Gaps = 17/232 (7%)
Query: 600 NAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAE 659
+++E L L R + EKE + L E +A ++ + +
Sbjct: 753 SSMETEASLSEELQRIIDEKEAVITALKSQLETAIAPCDNLKHSLSNNESEIENLRKQVV 812
Query: 660 RLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEAD 719
++++E K +E A E R +A D + + +I +LE Q+ L KE A
Sbjct: 813 QVRSELEKKEEEMANLENR------EASADNITKTEQRSNED-RIKQLEGQIKL-KENA- 863
Query: 720 ELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLA 779
L+A K + EKDL R E L K +E++ E + + + E L
Sbjct: 864 -----LEASSKIFIEKEKDLKNRIEELQTKLNEVSQNSQETDETLQGPEAIAMQYTEVLP 918
Query: 780 LAKSD--MEAVMQATS-EEIKKATETHAEALATKDAEIASQLAKIKSLEDEL 828
L+KSD + V + S E ET + + + +EI+ + A+++ +L
Sbjct: 919 LSKSDNLQDLVNEVASLREQNGLMETELKEMQERYSEISLRFAEVEGERQQL 970
Score = 33.5 bits (75), Expect = 0.56
Identities = 35/155 (22%), Positives = 73/155 (46%), Gaps = 9/155 (5%)
Query: 703 KIGELEDQLALMKEEADE----LDASLQACKKEKEQAEKDLIARGEALIAKESELAILCA 758
+I ELE Q+ M+EE ++ + ++A + K + E+ I EAL + A +
Sbjct: 572 RIKELETQIKGMEEELEKQAQIFEGDIEAVTRAKVEQEQRAIEAEEALRKTRWKNASVAG 631
Query: 759 ELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQL 818
+++ K ++EQ S+ A K M+A+ + + E++ E L + E+
Sbjct: 632 KIQDEFKRISEQ--MSSTLAANEKVTMKAMTE--TRELRMQKRQLEELLMNANDELRVNR 687
Query: 819 AKIKSLEDELATE-KAKAIEAREQAADIALDNRER 852
+ ++ +EL+ + K E + +AD+ R++
Sbjct: 688 VEYEAKLNELSGKTDLKTKEMKRMSADLEYQKRQK 722
>At3g28790 unknown protein
Length = 608
Score = 50.4 bits (119), Expect = 4e-06
Identities = 83/395 (21%), Positives = 145/395 (36%), Gaps = 40/395 (10%)
Query: 418 TEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVA 477
TE +K A+ + S + S +G S P PT + P ++
Sbjct: 239 TEAGSKSSSSAKTKEVSGGSSGNTYKDTTGSSSGAS-PSGSPTPTPSTPTPSTPTPSTPT 297
Query: 478 AATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP 537
+T + P S A +T A S G+ +AS + + S G K ET
Sbjct: 298 PSTPTPSTPTPS-TPAPSTPAAGKTSEKGSESASMKKESNSKSESESAASGSVSKTKETN 356
Query: 538 KSPPRQD-------APPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVG 590
K + SP + P PS G+ S G+A+ S A + + G
Sbjct: 357 KGSSGDTYKDTTGTSSGSPSGSPSGSPTPSTSTDGKASSKGSASASAGA----SASASAG 412
Query: 591 SSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEK 650
+S+ +E A ++ K S + +E F EK
Sbjct: 413 ASA----------SAEESAASQKKESNSKSSSSSSSTTSVKEVETQTSSEVNSFISNLEK 462
Query: 651 FNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQ 710
N E ++ E K +A+A KL+T K+ V + +A KI E
Sbjct: 463 KYTGNSEL-KVFFEKLKTSMSASA------KLSTSNAKELVTG---MRSAASKIAEAMMF 512
Query: 711 LALMKEEADELDASLQACKKEKEQAEKDL------IARGEALIA-KESELAILCAELELV 763
++ +++E S+ +C++E Q+ K+L I G+ + + +++EL + E V
Sbjct: 513 VSSRFSKSEETKTSMASCQQEVMQSLKELQDINSQIVSGKTVTSTQQTELKQTITKWEQV 572
Query: 764 KKALAEQEKKSAESLALAKSDMEAVMQATSEEIKK 798
E S+ S + + S + Q +++ K
Sbjct: 573 TTQFVETAASSSSSSSSSSSSSSSSSQGSAKMAMK 607
Score = 36.6 bits (83), Expect = 0.067
Identities = 37/190 (19%), Positives = 67/190 (34%), Gaps = 21/190 (11%)
Query: 404 ARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAA 463
A +R + + + D G+ + ++ + + + S +SK G++ T +
Sbjct: 171 AEKRGKSIDESSYGLDVDVNASIGSSSGSDGSSSSDNESSSNTKSQGTSSKSGSESTAGS 230
Query: 464 AAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASS 523
E + + TE + +SS + + + + T +S+G + + S
Sbjct: 231 I--------ETNTGSKTEAGSKSSSSAKTKEVSGGSSGNTYKDTTGSSSGASPS----GS 278
Query: 524 DTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQ 583
TP TP +P PS P+ P P S P A TSE
Sbjct: 279 PTP---------TPSTPTPSTPTPSTPTPSTPTPSTPTPSTPAPSTPAAGKTSEKGSESA 329
Query: 584 APAPEVGSSS 593
+ E S S
Sbjct: 330 SMKKESNSKS 339
Score = 34.3 bits (77), Expect = 0.33
Identities = 36/141 (25%), Positives = 61/141 (42%), Gaps = 18/141 (12%)
Query: 403 KARERRMARFSPKFNTEGDAKKRGGAENQAESTKAPKR-------RRLTKASSDAGTSKP 455
K E+ S K + ++ A TK + + T SS + + P
Sbjct: 320 KTSEKGSESASMKKESNSKSESESAASGSVSKTKETNKGSSGDTYKDTTGTSSGSPSGSP 379
Query: 456 GAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVN 515
PT + + GK ++ S +A + AS+ ASAGA+A +AE S A+ +
Sbjct: 380 SGSPTPSTST-DGKASSKGSASA-----SAGASASASAGASA--SAEES---AASQKKES 428
Query: 516 ATKAAASSDTPIGDKEKENET 536
+K+++SS + KE E +T
Sbjct: 429 NSKSSSSSSSTTSVKEVETQT 449
>At5g38560 putative protein
Length = 681
Score = 50.1 bits (118), Expect = 6e-06
Identities = 46/161 (28%), Positives = 65/161 (39%), Gaps = 11/161 (6%)
Query: 443 LTKASSDAGTSKPG---AQPTTAAAAPK-----GKNVAEASVAAATEPTAVPASSPASAG 494
L+ SS++ T+ P QPTT +A P + V +++ P V +S P S+
Sbjct: 10 LSPPSSNSSTTAPPPLQTQPTTPSAPPPVTPPPSPPQSPPPVVSSSPPPPVVSSPPPSSS 69
Query: 495 ATAAFAAESSIGATAASAGVNATKAAASS-DTPIGDKEKENETPKSPPRQDAPPSP--PS 551
+ +S T AS+ A+ TP +T PP DA PSP P+
Sbjct: 70 PPPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTPPAPPQTVSPPPPPDASPSPPAPT 129
Query: 552 THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSS 592
T + P PSP GE P T S P P +S
Sbjct: 130 TTNPPPKPSPSPPGETPSPPGETPSPPKPSPSTPTPTTTTS 170
Score = 35.0 bits (79), Expect = 0.19
Identities = 26/142 (18%), Positives = 49/142 (34%), Gaps = 5/142 (3%)
Query: 446 ASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSI 505
+S +S P + P + P + V A+ P + PA++P + T +
Sbjct: 62 SSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTPPAPPQTVSPPPPPDA 121
Query: 506 GATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQG 565
+ + S P ETP P + P+P +T PP
Sbjct: 122 SPSPPAPTTTNPPPKPSPSPPGETPSPPGETPSPPKPSPSTPTPTTT-----TSPPPPPA 176
Query: 566 EKSCPGAATTSEAAQIEQAPAP 587
+ P ++ ++ + + P P
Sbjct: 177 TSASPPSSNPTDPSTLAPPPTP 198
Score = 30.8 bits (68), Expect = 3.7
Identities = 33/138 (23%), Positives = 49/138 (34%), Gaps = 16/138 (11%)
Query: 485 VPASSPASAGATAAFAAESSIGATAASAG--VNATKAAASSDTPIGDKEKENETPKSPPR 542
+P SP S+ ++ T SA V + S P+ SPP
Sbjct: 7 LPILSPPSSNSSTTAPPPLQTQPTTPSAPPPVTPPPSPPQSPPPVVSSSPPPPVVSSPPP 66
Query: 543 QDAPP-------SPPSTHDAGPMP-----SPPHQGEKSCPGA--ATTSEAAQIEQAPAPE 588
+PP SPP T + P P SPP + P A T S + +P+P
Sbjct: 67 SSSPPPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTPPAPPQTVSPPPPPDASPSPP 126
Query: 589 VGSSSYYNMLPNAIEPSE 606
+++ P+ P E
Sbjct: 127 APTTTNPPPKPSPSPPGE 144
>At5g62410 chromosomal protein - like
Length = 1175
Score = 49.7 bits (117), Expect = 8e-06
Identities = 52/214 (24%), Positives = 106/214 (49%), Gaps = 20/214 (9%)
Query: 667 KHQEAAAAWEKRFDKLATQAGK-DKVYADKMIG-TAGIKIGELEDQLALMK-------EE 717
K +E AA ++RF +L+T + +K + + G ++G + LEDQL K E
Sbjct: 354 KSEEGAADLKQRFQELSTTLEECEKEHQGVLAGKSSGDEEKCLEDQLRDAKIAVGTAGTE 413
Query: 718 ADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAES 777
+L ++ C+KE ++ + L+++ E I E+EL ++E VKKAL +
Sbjct: 414 LKQLKTKIEHCEKELKERKSQLMSKLEEAIEVENELGARKNDVEHVKKALESIPYNEGQM 473
Query: 778 LALAK---SDMEAV------MQATSEEIKKATETHAEALATKD-AEIASQLAKIKSLEDE 827
AL K +++E V ++ S ++ T+++ + D +++ +AK+ ++D
Sbjct: 474 EALEKDRGAELEVVQRLEDKVRGLSAQLANFQFTYSDPVRNFDRSKVKGVVAKLIKVKDR 533
Query: 828 LATEKAKAIEAREQAADIALDNRERGFYLAKDQA 861
++ A + A + D+ +D+ + G L ++ A
Sbjct: 534 -SSMTALEVTAGGKLYDVVVDSEDTGKQLLQNGA 566
Score = 33.1 bits (74), Expect = 0.74
Identities = 50/243 (20%), Positives = 96/243 (38%), Gaps = 61/243 (25%)
Query: 659 ERLKAESAKHQEAAAAWEKRFDKLA--------TQAGKDKVYADKMIGTAGIKIGELEDQ 710
E+L+ E +++ + A D+L QA K + A +G K+G+++ +
Sbjct: 208 EKLRKEKSQYMQWANG-NAELDRLRRFCIAFEYVQAEKIRDNAVLGVGEMKAKLGKIDAE 266
Query: 711 LALMKEEADELDASLQACKKEKE----------------------QAEKDLIARGEALIA 748
+EE E + ++A + KE + L + + L+
Sbjct: 267 TEKTQEEIQEFEKQIKALTQAKEASMGGEVKTLSEKVDSLAQEMTRESSKLNNKEDTLLG 326
Query: 749 KESELAILCAELELVKKALAEQE---KKSAESLALAKSDMEAVMQATSEEIKKATETHAE 805
++ + + +E +KK++ E+ KKS E A D++ Q S +++ + H
Sbjct: 327 EKENVEKIVHSIEDLKKSVKERAAAVKKSEEGAA----DLKQRFQELSTTLEECEKEHQG 382
Query: 806 ALATK--------------DAEIASQLA---------KIKSLEDELATEKAKAIEAREQA 842
LA K DA+IA A KI+ E EL K++ + E+A
Sbjct: 383 VLAGKSSGDEEKCLEDQLRDAKIAVGTAGTELKQLKTKIEHCEKELKERKSQLMSKLEEA 442
Query: 843 ADI 845
++
Sbjct: 443 IEV 445
Score = 31.2 bits (69), Expect = 2.8
Identities = 36/175 (20%), Positives = 74/175 (41%), Gaps = 10/175 (5%)
Query: 659 ERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEA 718
E L+ ++ +E A++ FD ++ K + G ++ +LE + +K +
Sbjct: 748 EELEEAKSQIKEKELAYKNCFDAVSKLENSIKDHDKNREG----RLKDLEKNIKTIKAQM 803
Query: 719 DELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELEL-VKKALAEQEKKSAES 777
L++ + EKE+ L+ EA+ ++S L LE + +E +++ A+
Sbjct: 804 QAASKDLKSHENEKEK----LVMEEEAMKQEQSSLESHLTSLETQISTLTSEVDEQRAKV 859
Query: 778 LALAKSDMEAVMQATSEEIK-KATETHAEALATKDAEIASQLAKIKSLEDELATE 831
AL K E++ + K K +T T + +L+ +K +L E
Sbjct: 860 DALQKIHDESLAELKLIHAKMKECDTQISGFVTDQEKCLQKLSDMKLERKKLENE 914
>At1g65010 hypothetical protein
Length = 1318
Score = 49.3 bits (116), Expect = 1e-05
Identities = 49/168 (29%), Positives = 80/168 (47%), Gaps = 21/168 (12%)
Query: 702 IKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELE 761
+K EL+ QL ++E+ + D ++ KK+K +A DL KESE + A E
Sbjct: 51 VKGTELQTQLNQIQEDLKKADEQIELLKKDKAKAIDDL---------KESEKLVEEAN-E 100
Query: 762 LVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKI 821
+K+ALA Q K++ ES + K + QA E ++K T L + ++ A ++ +
Sbjct: 101 KLKEALAAQ-KRAEESFEVEKFRAVELEQAGLEAVQKKDVTSKNELESIRSQHALDISAL 159
Query: 822 KSLEDEL----------ATEKAKAIEAREQAADIALDNRERGFYLAKD 859
S +EL A K KA+ E+A IA + E+ LA +
Sbjct: 160 LSTTEELQRVKHELSMTADAKNKALSHAEEATKIAEIHAEKAEILASE 207
Score = 45.8 bits (107), Expect = 1e-04
Identities = 106/442 (23%), Positives = 178/442 (39%), Gaps = 75/442 (16%)
Query: 535 ETPKSPPRQDAP------PSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPE 588
ETP+S P P S ++ A P+P+ ++S P T +++ + +
Sbjct: 2 ETPRSKPSPPPPRLSKLSASKSDSNSASPVPNTRLSLDRSPP----TKVHSRLVKGTELQ 57
Query: 589 VGSSSYYNMLPNAIEPSEFLLAGLNR---DVIEKEVLSQGLNVTKEETLACLLRAGCIFA 645
+ L A E E L + D+ E E L + N +E LA RA F
Sbjct: 58 TQLNQIQEDLKKADEQIELLKKDKAKAIDDLKESEKLVEEANEKLKEALAAQKRAEESFE 117
Query: 646 HTFEKFNAANVEAERLKA------------ESAKHQEAA--AAWEKRFDKLATQAGKDKV 691
EKF A +E L+A ES + Q A +A ++L + +
Sbjct: 118 --VEKFRAVELEQAGLEAVQKKDVTSKNELESIRSQHALDISALLSTTEELQRVKHELSM 175
Query: 692 YAD---KMIGTA--GIKIGELEDQLA-LMKEEADELDASLQACKKEKEQAEKDLIARGEA 745
AD K + A KI E+ + A ++ E L A L + K+EKE E G
Sbjct: 176 TADAKNKALSHAEEATKIAEIHAEKAEILASELGRLKALLGS-KEEKEAIE------GNE 228
Query: 746 LIAK-ESELAILCAELELVKKALAEQEKKSAESLA-LAKSDMEAVMQATS---------- 793
+++K +SE+ +L ELE K ++ E K E L K D+EA A S
Sbjct: 229 IVSKLKSEIELLRGELE--KVSILESSLKEQEGLVEQLKVDLEAAKMAESCTNSSVEEWK 286
Query: 794 ---EEIKKATE--THAEALATKDAE-IASQLAKIKSLEDELAT------EKAKAIEAREQ 841
E++K E +++ A++ E + QLA++ + E + EK + +E +
Sbjct: 287 NKVHELEKEVEESNRSKSSASESMESVMKQLAELNHVLHETKSDNAAQKEKIELLEKTIE 346
Query: 842 AADIALDNRERGFYLAKDQA-------QHLFPNFDFSAMGVMKEITAAGLVGPDDPPLID 894
A L+ R +AK++A + + + S + + + L+D
Sbjct: 347 AQRTDLEEYGRQVCIAKEEASKLENLVESIKSELEISQEEKTRALDNEKAATSNIQNLLD 406
Query: 895 QNLWTATEEEEEEEEEEEQEKE 916
Q + E E + EEE+ +K+
Sbjct: 407 QRTELSIELERCKVEEEKSKKD 428
Score = 33.9 bits (76), Expect = 0.43
Identities = 30/118 (25%), Positives = 54/118 (45%), Gaps = 15/118 (12%)
Query: 716 EEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSA 775
++ EL L+ CK E+E+++KD+ + AL +E + A L + ++ L E +
Sbjct: 406 DQRTELSIELERCKVEEEKSKKDMESLTLALQEASTESSEAKATLLVCQEELKNCESQ-V 464
Query: 776 ESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKA 833
+SL LA K+ E + + L EI S + + S+++E KA
Sbjct: 465 DSLKLAS--------------KETNEKYEKMLEDARNEIDSLKSTVDSIQNEFENSKA 508
>At3g28830 hypothetical protein
Length = 536
Score = 48.1 bits (113), Expect = 2e-05
Identities = 71/343 (20%), Positives = 136/343 (38%), Gaps = 28/343 (8%)
Query: 446 ASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSI 505
ASS++ ++K G+ ++ + + K K E+S + ++ +V S S+G +AA ++ S
Sbjct: 197 ASSESSSTKSGSVSSSGSVSTKSK---ESSSSGSSASGSVATKSKESSGGSAATKSKESS 253
Query: 506 GATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQG 565
G +AA+ ++ +A++ K + +P P+ A PS + + S QG
Sbjct: 254 GGSAATKSKESSGGSATTG-------KTSGSPSGSPK--ASPSGSVSGKSSSKGSASAQG 304
Query: 566 EKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVIEKEVLSQG 625
S G+A+ +A + + + + S M + E + + E S
Sbjct: 305 SASAQGSASAQGSASAQGSASAQRRESGAMAM----SKSRETKTSSQRQSKSSSESSSSS 360
Query: 626 LNVTK-EETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLAT 684
T ++ + + F EK AA E ++ ES K A+A + +
Sbjct: 361 TTTTTVKQVESETSKEVMSFIMQLEKKYAAKAEL-KVFFESLKSSMQASA------SVGS 413
Query: 685 QAGKDKVYADK----MIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLI 740
+ KD V A K + A + + A MK D + C K+ + ++
Sbjct: 414 KTAKDYVSASKAATGKLSEAMASVSSKNVKSAKMKSNLDTSKDEMLKCVKQIQDINGKMV 473
Query: 741 ARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKS 783
+ ++SEL + E V E S+ S + + S
Sbjct: 474 SGKTVSSTQQSELKQTITKWEKVTTQFVETAASSSSSSSSSSS 516
Score = 33.9 bits (76), Expect = 0.43
Identities = 53/300 (17%), Positives = 105/300 (34%), Gaps = 45/300 (15%)
Query: 413 SPKFNTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVA 472
S ++ G + + + S+ + +K SS + + + +AA K K +
Sbjct: 206 SGSVSSSGSVSTKSKESSSSGSSASGSVATKSKESSGGSAATKSKESSGGSAATKSKESS 265
Query: 473 EASVAAAT---EPTAVPASSP--------ASAGATAAFAAESSIGATAASAGVNATKAAA 521
S P+ P +SP +S G+ +A + S+ G+ +A +A +A+
Sbjct: 266 GGSATTGKTSGSPSGSPKASPSGSVSGKSSSKGSASAQGSASAQGSASAQGSASAQGSAS 325
Query: 522 SSDTPIG--DKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAA 579
+ G K ET S RQ S S+ + E S S
Sbjct: 326 AQRRESGAMAMSKSRETKTSSQRQSKSSSESSSSSTTTTTVKQVESETS---KEVMSFIM 382
Query: 580 QIEQAPAPEVGSSSYYNMLPNAIEPSEFL----------------------LAGLNRDVI 617
Q+E+ A + ++ L ++++ S + +A ++ +
Sbjct: 383 QLEKKYAAKAELKVFFESLKSSMQASASVGSKTAKDYVSASKAATGKLSEAMASVSSKNV 442
Query: 618 EKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEK 677
+ + L+ +K+E L C+ + I ++ + LK K WEK
Sbjct: 443 KSAKMKSNLDTSKDEMLKCVKQIQDINGKMVSGKTVSSTQQSELKQTITK-------WEK 495
>At1g13220 putative nuclear matrix constituent protein
Length = 1128
Score = 48.1 bits (113), Expect = 2e-05
Identities = 44/198 (22%), Positives = 86/198 (43%), Gaps = 22/198 (11%)
Query: 657 EAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKM--IGTAGIKIGELEDQLALM 714
E E + K +E WEK+ GK++ ++ + K+ E+E +L L
Sbjct: 250 ERESYEGTFQKQREYLNEWEKKLQ------GKEESITEQKRNLNQREEKVNEIEKKLKLK 303
Query: 715 KEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKS 774
++E +E + + + ++ E+D+ R E L KE E L ++ L+ K E E ++
Sbjct: 304 EKELEEWNRKVDLSMSKSKETEEDITKRLEELTTKEKEAHTL--QITLLAK---ENELRA 358
Query: 775 AESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAK 834
E +A+ EI+K + E L +K E + +I+ D+ K +
Sbjct: 359 FEEKLIARE---------GTEIQKLIDDQKEVLGSKMLEFELECEEIRKSLDKELQRKIE 409
Query: 835 AIEAREQAADIALDNRER 852
+E ++ D + + E+
Sbjct: 410 ELERQKVEIDHSEEKLEK 427
Score = 29.6 bits (65), Expect = 8.1
Identities = 45/205 (21%), Positives = 83/205 (39%), Gaps = 21/205 (10%)
Query: 649 EKFNAANVEAERLKAESAK---HQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIG 705
E+ E +RL E + +E+ ++ +K+ + K + ++ + IK
Sbjct: 455 EREKIIQAEEKRLSLEKQQLLSDKESLEDLQQEIEKIRAEMTKKEEMIEEECKSLEIKKE 514
Query: 706 ELEDQLALMKEEADELDAS----------LQACKKEKEQAEKDLIARGEALIAKESELAI 755
E E+ L L E +++ S ++ K+EKE+ EK+ E E
Sbjct: 515 EREEYLRLQSELKSQIEKSRVHEEFLSKEVENLKQEKERFEKEWEILDEKQAVYNKERIR 574
Query: 756 LCAELELVKK-ALAEQEKKSAESLALAKSDMEAV----MQATSEEIKKATETHA--EALA 808
+ E E ++ L E E+ E AL M+ + +Q S E E A E +
Sbjct: 575 ISEEKEKFERFQLLEGERLKKEESALRVQIMQELDDIRLQRESFEANMEHERSALQEKVK 634
Query: 809 TKDAEIASQLAKI-KSLEDELATEK 832
+ +++ L + ++LE EL K
Sbjct: 635 LEQSKVIDDLEMMRRNLEIELQERK 659
Score = 29.6 bits (65), Expect = 8.1
Identities = 40/185 (21%), Positives = 84/185 (44%), Gaps = 30/185 (16%)
Query: 666 AKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEAD-ELDAS 724
AK E A EK + T+ K D G K+ E E + +++ D EL
Sbjct: 351 AKENELRAFEEKLIAREGTEIQK---LIDDQKEVLGSKMLEFELECEEIRKSLDKELQRK 407
Query: 725 LQACKKEK---EQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALA 781
++ +++K + +E+ L R +A+ K + +LE K + E+EK
Sbjct: 408 IEELERQKVEIDHSEEKLEKRNQAMNKKFDRVNEKEMDLEAKLKTIKEREK--------- 458
Query: 782 KSDMEAVMQATSEEIKKATETHAEALATKDA--EIASQLAKIK---SLEDELATEKAKAI 836
++QA E K+ + + L+ K++ ++ ++ KI+ + ++E+ E+ K++
Sbjct: 459 ------IIQA---EEKRLSLEKQQLLSDKESLEDLQQEIEKIRAEMTKKEEMIEEECKSL 509
Query: 837 EAREQ 841
E +++
Sbjct: 510 EIKKE 514
>At3g19060 hypothetical protein
Length = 1647
Score = 47.8 bits (112), Expect = 3e-05
Identities = 66/263 (25%), Positives = 116/263 (44%), Gaps = 37/263 (14%)
Query: 605 SEFLLA---GLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAA-----NV 656
+E LLA L ++I+ + +S+ + + L + A T K N A N+
Sbjct: 1326 AEDLLAEKCSLEEEMIQTKKVSESMEMELFNLRNALGQLNDTVAFTQRKLNDAIDERDNL 1385
Query: 657 EAE--RLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALM 714
+ E LK E K + A E R+ + A K YAD E E+++ L+
Sbjct: 1386 QDEVLNLKEEFGKMKSEAKEMEARYIEAQQIAESRKTYAD-----------EREEEVKLL 1434
Query: 715 KEEADELDASLQACKKE----KEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQ 770
+ +EL+ ++ + + K++AE+ + R E E EL + ++E + A E
Sbjct: 1435 EGSVEELEYTINVLENKVNVVKDEAERQRLQREEL----EMELHTIRQQMESARNADEEM 1490
Query: 771 EKKSAE---SLALAKSDMEAVMQATSE---EIKKATETHAEALATKDAEIASQLAKIKSL 824
++ E LA AK +EA+ + T++ EI + +E +E +A+ + + K K L
Sbjct: 1491 KRILDEKHMDLAQAKKHIEALERNTADQKTEITQLSEHISELNLHAEAQASEYMHKFKEL 1550
Query: 825 EDELATEKAKAIEAREQAADIAL 847
E E+ K QA D +L
Sbjct: 1551 --EAMAEQVKPEIHVSQAIDSSL 1571
Score = 34.3 bits (77), Expect = 0.33
Identities = 49/213 (23%), Positives = 88/213 (41%), Gaps = 17/213 (7%)
Query: 656 VEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMK 715
VE LK E + + + + LA Q M K+ LE ++A MK
Sbjct: 604 VEMANLKGEKEELKASEKRSLSNLNDLAAQICNLNTVMSNMEEQYEHKMETLEHEIAKMK 663
Query: 716 EEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAI--LCAELELVKKALAEQEKK 773
EAD+ K+ E+A+ + +E+++ + L E ++ L +Q+K+
Sbjct: 664 IEADQEYVENLCILKKFEEAQGTI---------READITVNELVIANEKMRFDLEKQKKR 714
Query: 774 SAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKA 833
+ K+ +E + + S +K+ E LA + S L I +L +ELAT
Sbjct: 715 GISLVGEKKALVEKLQELESINVKE-----NEKLAYLEKLFESSLMGIGNLVEELATVVR 769
Query: 834 KAIEAREQA-ADIALDNRERGFYLAKDQAQHLF 865
K + A +A D E ++++ + LF
Sbjct: 770 KLQDESSVALTGMAKDLSELKSWVSETNSARLF 802
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 47.4 bits (111), Expect = 4e-05
Identities = 32/148 (21%), Positives = 78/148 (52%), Gaps = 16/148 (10%)
Query: 703 KIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELEL 762
++ ELE QL L+++ +L ASL A ++EK+ ++ + L +S++ L EL
Sbjct: 486 RLSELETQLKLLEQRVVDLSASLNAAEEEKKSLSSMILEITDELKQAQSKVQELVTELAE 545
Query: 763 VKKALAEQEKKSAESLALAKS----------DMEAVMQATSEEIKK------ATETHAEA 806
K L ++E + + + + ++ ++EA +++ E++K+ ++E +
Sbjct: 546 SKDTLTQKENELSSFVEVHEAHKRDSSSQVKELEARVESAEEQVKELNQNLNSSEEEKKI 605
Query: 807 LATKDAEIASQLAKIKSLEDELATEKAK 834
L+ + +E++ ++ + +S EL++E +
Sbjct: 606 LSQQISEMSIKIKRAESTIQELSSESER 633
Score = 42.7 bits (99), Expect = 0.001
Identities = 57/284 (20%), Positives = 120/284 (42%), Gaps = 45/284 (15%)
Query: 575 TSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLA----GLNRDVIEKEVLSQGLNVTK 630
+ ++ +E E SSS L IE SE L+A LN EK++LSQ +
Sbjct: 48 SEHSSLVELHKTHERESSSQVKELEAHIESSEKLVADFTQSLNNAEEEKKLLSQKI---- 103
Query: 631 EETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDK 690
A + A + L +ES + +E+ + E+ L +
Sbjct: 104 --------------AELSNEIQEAQNTMQELMSESGQLKESHSVKERELFSL-------R 142
Query: 691 VYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKE 750
+ + + ELE QL K++ +L ASL+A ++E + + L +
Sbjct: 143 DIHEIHQRDSSTRASELEAQLESSKQQVSDLSASLKAAEEENKAISSKNVETMNKLEQTQ 202
Query: 751 SELAILCAELELVKKALAEQEKKSAESLALAKS----------DMEAVMQATSEEIKKAT 800
+ + L AEL +K + E+E + + + + ++ ++E ++++ + + +
Sbjct: 203 NTIQELMAELGKLKDSHREKESELSSLVEVHETHQRDSSIHVKELEEQVESSKKLVAELN 262
Query: 801 ET------HAEALATKDAEIASQLAKIKSLEDELATEKAKAIEA 838
+T + L+ K AE+++++ + ++ EL +E + E+
Sbjct: 263 QTLNNAEEEKKVLSQKIAELSNEIKEAQNTIQELVSESGQLKES 306
Score = 41.2 bits (95), Expect = 0.003
Identities = 44/207 (21%), Positives = 92/207 (44%), Gaps = 26/207 (12%)
Query: 659 ERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVY-----ADKMIGTAGIKIGELEDQLAL 713
+ LK +K QE + D L + + + A K ++ +K ELE ++
Sbjct: 527 DELKQAQSKVQELVTELAESKDTLTQKENELSSFVEVHEAHKRDSSSQVK--ELEARVES 584
Query: 714 MKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKK 773
+E+ EL+ +L + ++EK+ + + + ES + L +E E +K + AE++ +
Sbjct: 585 AEEQVKELNQNLNSSEEEKKILSQQISEMSIKIKRAESTIQELSSESERLKGSHAEKDNE 644
Query: 774 ----------SAESLALAKSDMEAVMQATS------EEIKKATETHAEALATKDAEIASQ 817
L+ +EA ++++ E KA E + ++TK +E + +
Sbjct: 645 LFSLRDIHETHQRELSTQLRGLEAQLESSEHRVLELSESLKAAEEESRTMSTKISETSDE 704
Query: 818 LAKIKSLEDELATEKAKAIEAREQAAD 844
L + + + EL + +K +EQ A+
Sbjct: 705 LERTQIMVQELTADSSK---LKEQLAE 728
Score = 40.4 bits (93), Expect = 0.005
Identities = 57/257 (22%), Positives = 104/257 (40%), Gaps = 24/257 (9%)
Query: 620 EVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAAN--VEAERLKAESAKHQEAAAAWEK 677
E LSQ N+ +E ++ + E+ N + ++ L+ E+ Q + E
Sbjct: 904 EYLSQITNLKEE-----IINKVKVHESILEEINGLSEKIKGRELELETLGKQRSELDEEL 958
Query: 678 RFDKLATQAGKDKV-YADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAE 736
R K DK+ A I I L+++L ++ + E +A L+ K+EK +
Sbjct: 959 RTKKEENVQMHDKINVASSEIMALTELINNLKNELDSLQVQKSETEAELEREKQEKSELS 1018
Query: 737 KDLIARGEALIAKESELAILCAELELVKKALAEQEK---------KSAESLALAKSDMEA 787
+ +AL+ +E+ L E + + + E E K A+ L +
Sbjct: 1019 NQITDVQKALVEQEAAYNTLEEEHKQINELFKETEATLNKVTVDYKEAQRLLEERGKEVT 1078
Query: 788 VMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIAL 847
+T ++ E+ L K EI + + KI ++E +L K + EQ L
Sbjct: 1079 SRDSTIGVHEETMESLRNELEMKGDEIETLMEKISNIEVKLRLSNQK-LRVTEQ----VL 1133
Query: 848 DNRERGFYLAKDQAQHL 864
+E F K++A+HL
Sbjct: 1134 TEKEEAF--RKEEAKHL 1148
Score = 37.4 bits (85), Expect = 0.039
Identities = 42/185 (22%), Positives = 83/185 (44%), Gaps = 33/185 (17%)
Query: 700 AGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAE 759
+ I + ELE+Q+ K+ EL+ +L ++EK+ + + + ++ + L +E
Sbjct: 240 SSIHVKELEEQVESSKKLVAELNQTLNNAEEEKKVLSQKIAELSNEIKEAQNTIQELVSE 299
Query: 760 LELVKK----------ALAEQEKKSAESLALAKSDMEAVMQATSEEIK------KATETH 803
+K+ +L + + + S++EA ++++ + I K E
Sbjct: 300 SGQLKESHSVKDRDLFSLRDIHETHQRESSTRVSELEAQLESSEQRISDLTVDLKDAEEE 359
Query: 804 AEALATKDAEIASQLAK----IKSLEDELATEKAKAIE-----------AREQAADI--A 846
+A+++K+ EI +L + IK L DEL K + E A +Q AD+ +
Sbjct: 360 NKAISSKNLEIMDKLEQAQNTIKELMDELGELKDRHKEKESELSSLVKSADQQVADMKQS 419
Query: 847 LDNRE 851
LDN E
Sbjct: 420 LDNAE 424
Score = 36.6 bits (83), Expect = 0.067
Identities = 87/371 (23%), Positives = 148/371 (39%), Gaps = 58/371 (15%)
Query: 591 SSSYYNMLPNAIEPSEFLLAGLNRDVI----EKEVLSQGL----NVTKE--ETLACLLRA 640
SS + L +E S+ L+A LN+ + EK+VLSQ + N KE T+ L+
Sbjct: 240 SSIHVKELEEQVESSKKLVAELNQTLNNAEEEKKVLSQKIAELSNEIKEAQNTIQELVSE 299
Query: 641 GCIFAHTF-----EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLA--TQAGKDKVYA 693
+ + F+ ++ + S + E A E +++ T KD
Sbjct: 300 SGQLKESHSVKDRDLFSLRDIHETHQRESSTRVSELEAQLESSEQRISDLTVDLKDAEEE 359
Query: 694 DKMIGTAGIKI-GELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESE 752
+K I + ++I +LE +KE DEL L+ KEKE L+ + +A
Sbjct: 360 NKAISSKNLEIMDKLEQAQNTIKELMDEL-GELKDRHKEKESELSSLVKSADQQVAD--- 415
Query: 753 LAILCAELELVKKAL--AEQEKKSAESLALAKSDMEAVMQAT-------SEEIKKA---- 799
+K++L AE+EKK L S+ Q T SE++K++
Sbjct: 416 ----------MKQSLDNAEEEKKMLSQRILDISNEIQEAQKTIQEHMSESEQLKESHGVK 465
Query: 800 ----------TETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIALDN 849
ETH +T+ +E+ +QL ++ +L+ A E ++ + + L+
Sbjct: 466 ERELTGLRDIHETHQRESSTRLSELETQLKLLEQRVVDLSASLNAAEEEKKSLSSMILEI 525
Query: 850 RERGFYLAKDQAQHLFPNFDFSAMGV-MKEITAAGLVGPDDPPLIDQNLWTATEEEEEEE 908
+ A+ + Q L S + KE + V + D + E E E
Sbjct: 526 TDE-LKQAQSKVQELVTELAESKDTLTQKENELSSFVEVHEAHKRDSSS-QVKELEARVE 583
Query: 909 EEEEQEKENNE 919
EEQ KE N+
Sbjct: 584 SAEEQVKELNQ 594
Score = 31.2 bits (69), Expect = 2.8
Identities = 39/172 (22%), Positives = 78/172 (44%), Gaps = 29/172 (16%)
Query: 700 AGIKIGELEDQLALMKEEADELDA-----SLQACKKEKEQAE---------KDLIAR--- 742
A +KI L+D++ ++++ LD+ +Q KK +E +E +++I +
Sbjct: 863 ASVKIKRLDDEVNGLRQQVASLDSQRAELEIQLEKKSEEISEYLSQITNLKEEIINKVKV 922
Query: 743 GEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSD---MEAVMQATSEEIKKA 799
E+++ + + L+ EL + L +Q + E L K + M + S EI
Sbjct: 923 HESILEEINGLSEKIKGRELELETLGKQRSELDEELRTKKEENVQMHDKINVASSEIMAL 982
Query: 800 TE------THAEALATKDAEIASQLAKIKSLEDELA---TEKAKAIEAREQA 842
TE ++L + +E ++L + K + EL+ T+ KA+ +E A
Sbjct: 983 TELINNLKNELDSLQVQKSETEAELEREKQEKSELSNQITDVQKALVEQEAA 1034
>At5g14380 agp6
Length = 150
Score = 47.0 bits (110), Expect = 5e-05
Identities = 39/135 (28%), Positives = 62/135 (45%), Gaps = 19/135 (14%)
Query: 444 TKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAES 503
T ++DA ++ P P+ AA K A+ A + PT PA++P S+ A++ A+
Sbjct: 17 TAFAADAPSASPKKSPSPTAAPTKAPT---ATTKAPSAPTKAPAAAPKSSSASSPKASSP 73
Query: 504 SIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDA-GPMPSPP 562
+ +A+ + S++ P T SPP P+P ST A GP P
Sbjct: 74 AAEGPVPEDDYSASSPSDSAEAP---------TVSSPP----APTPDSTSAADGPSDGPT 120
Query: 563 HQGEKSCPGAATTSE 577
+ KS GA TT++
Sbjct: 121 AESPKS--GAVTTAK 133
Score = 31.2 bits (69), Expect = 2.8
Identities = 24/99 (24%), Positives = 41/99 (41%), Gaps = 7/99 (7%)
Query: 498 AFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSP---PRQDAPPSPPSTHD 554
AFAA+ A +AS + + AA + P + + K+P P+ + SP ++
Sbjct: 18 AFAAD----APSASPKKSPSPTAAPTKAPTATTKAPSAPTKAPAAAPKSSSASSPKASSP 73
Query: 555 AGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
A P P S P + + AP P+ S++
Sbjct: 74 AAEGPVPEDDYSASSPSDSAEAPTVSSPPAPTPDSTSAA 112
>At3g09000 unknown protein
Length = 541
Score = 47.0 bits (110), Expect = 5e-05
Identities = 39/177 (22%), Positives = 66/177 (37%), Gaps = 6/177 (3%)
Query: 417 NTEGDAKKRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASV 476
N D + Q S+ RR + + S TS+P A PT + P +
Sbjct: 121 NCREDIVSGNNNKPQTSSSSVAGLRRPSSSGSSRSTSRP-ATPTRRSTTPTTSTSRPVTT 179
Query: 477 AAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTP-IGDKEKENE 535
A+ ++ P S A A + + T +S + S+ P +K
Sbjct: 180 RASNSRSSTPTSRATLTAARATTSTAAPRTTTTSSGSARSATPTRSNPRPSSASSKKPVS 239
Query: 536 TPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSS 592
P +P R+ + P+ PS + P +G P + S+A +P+P + SS
Sbjct: 240 RPATPTRRPSTPTGPSIVSS----KAPSRGTSPSPTVNSLSKAPSRGTSPSPTLNSS 292
>At1g79280 hypothetical protein
Length = 2111
Score = 46.6 bits (109), Expect = 6e-05
Identities = 48/212 (22%), Positives = 91/212 (42%), Gaps = 34/212 (16%)
Query: 646 HTFEKFNAANVEAERLKAESAKHQEAAAAWE-------KRFDKLATQAGKDKVYADKMIG 698
H FEK A++ + ES + + K +KL + K D++
Sbjct: 1308 HNFEKCQEMREVAQKARMESENFENLLKTKQTELDLCMKEMEKLRMETDLHKKRVDELRE 1367
Query: 699 T-AGIKIGE---LEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELA 754
T I I + L+D++ ++E+ DA + CKK L+ K+++++
Sbjct: 1368 TYRNIDIADYNRLKDEVRQLEEKLKAKDAHAEDCKK--------------VLLEKQNKIS 1413
Query: 755 ILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEI 814
+L EL KK L+E+EK+ + A++ M++ +E++K + H TK
Sbjct: 1414 LLEKELTNCKKDLSEREKR-LDDAQQAQATMQSEFNKQKQELEKNKKIHYTLNMTK---- 1468
Query: 815 ASQLAKIKSLEDELATEKAKAIEAREQAADIA 846
K + +DEL+ + + E+A + A
Sbjct: 1469 ----RKYEKEKDELSKQNQSLAKQLEEAKEEA 1496
Score = 45.8 bits (107), Expect = 1e-04
Identities = 50/207 (24%), Positives = 86/207 (41%), Gaps = 13/207 (6%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELE 708
EK A + AE K + Q + EK T KD +K + A ++
Sbjct: 1389 EKLKAKDAHAEDCKKVLLEKQNKISLLEKEL----TNCKKDLSEREKRLDDAQQAQATMQ 1444
Query: 709 DQLALMKEEADE---LDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKK 765
+ K+E ++ + +L K++ E+ EKD +++ +AK+ E A A
Sbjct: 1445 SEFNKQKQELEKNKKIHYTLNMTKRKYEK-EKDELSKQNQSLAKQLEEAKEEAGKRTTTD 1503
Query: 766 ALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLE 825
A+ EQ K E ++ + +E++K TE L KD E+ + ++ KS+E
Sbjct: 1504 AVVEQSVKEREEKEKRIQILDKYVHQLKDEVRKKTED----LKKKDEELTKERSERKSVE 1559
Query: 826 DELATEKAKAIEAREQAADIALDNRER 852
E+ K I+ + D L ER
Sbjct: 1560 KEVGDSLTK-IKKEKTKVDEELAKLER 1585
Score = 38.1 bits (87), Expect = 0.023
Identities = 46/175 (26%), Positives = 80/175 (45%), Gaps = 17/175 (9%)
Query: 700 AGIKIGE-LEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCA 758
AG +IG + +L KEE ++L +++ K Q + IA+ K+ E A
Sbjct: 955 AGFRIGSAMSIELRTAKEEIEKLRGEVESSKSHMLQYKS--IAQVNETALKQMESAHENF 1012
Query: 759 ELELVKKALAEQEKKSAESLALAK--SDMEAVMQATSEEIKKATETHAEALATKDAEIAS 816
LE K+ Q AE ++L + S++E SE++ A +AL + AEIAS
Sbjct: 1013 RLEAEKR----QRSLEAELVSLRERVSELENDCIQKSEQLATAAAGKEDALLSASAEIAS 1068
Query: 817 -------QLAKIKSLEDELATEKAKAIEAREQAADIALDNRERGFYLAKDQAQHL 864
+ ++I+++ +++T K +E + +A N ER L + Q L
Sbjct: 1069 LREENLVKKSQIEAMNIQMSTLK-NDLETEHEKWRVAQRNYERQVILLSETIQEL 1122
Score = 35.4 bits (80), Expect = 0.15
Identities = 37/157 (23%), Positives = 69/157 (43%), Gaps = 10/157 (6%)
Query: 700 AGIKIGELEDQLALMKEEADELDASLQACKKEKEQA----EKDLIARGEALIAKESELAI 755
A + ++ + + E D + A A EQ E+ ++ + + ES+ A
Sbjct: 18 ASVVAERADEYIRKIYAELDSVRAKADAASITAEQTCSLLEQKYLSLSQDFSSLESQNAK 77
Query: 756 LCAELE--LVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAE 813
L ++ + L + A ++ +K ++ K M E+ K+ E L KDAE
Sbjct: 78 LQSDFDDRLAELAQSQAQKHQLHLQSIEKDGEVERMSTEMSELHKSKRQLMELLEQKDAE 137
Query: 814 IASQLAKIKSLEDELA--TEKAKAIEAR--EQAADIA 846
I+ + + IKS D++ T+ + EAR E A++A
Sbjct: 138 ISEKNSTIKSYLDKIVKLTDTSSEKEARLAEATAELA 174
Score = 30.8 bits (68), Expect = 3.7
Identities = 44/195 (22%), Positives = 91/195 (46%), Gaps = 25/195 (12%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLA-TQAGKDKVYADKMIGTAGIKIGEL 707
+K+ + + + L++++AK Q + ++ R +LA +QA K +++ + E
Sbjct: 59 QKYLSLSQDFSSLESQNAKLQ---SDFDDRLAELAQSQAQKHQLHLQSI---------EK 106
Query: 708 EDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKAL 767
+ ++ M E EL K K Q + L + + K S + ++ +
Sbjct: 107 DGEVERMSTEMSELH-------KSKRQLMELLEQKDAEISEKNSTIKSYLDKIVKLTDTS 159
Query: 768 AEQEKKSAESLA-LAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLED 826
+E+E + AE+ A LA+S +A+ S+E K+ TE HA+ L + A+++
Sbjct: 160 SEKEARLAEATAELARS--QAMCSRLSQE-KELTERHAKWLDEELTAKVDSYAELRRRHS 216
Query: 827 ELATE-KAKAIEARE 840
+L +E AK ++ +
Sbjct: 217 DLESEMSAKLVDVEK 231
>At1g77580 unknown protein
Length = 779
Score = 46.2 bits (108), Expect = 8e-05
Identities = 65/275 (23%), Positives = 111/275 (39%), Gaps = 33/275 (12%)
Query: 558 MPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSEFLLAGLNRDVI 617
+P P KS P + T E + A E+ +L + I+ E L L
Sbjct: 314 LPETPDGNGKSGPESVTEEVVVPSENSLASEI------EVLTSRIKELEEKLEKLEA--- 364
Query: 618 EKEVLSQGLNVTKEETLACLLRAGCIFAHTFE---KFNAANVEAERLKAESAKHQEAAAA 674
EK L + +EE + + + + + T E K E E LK+E ++E A
Sbjct: 365 EKHELENEVKCNREEAVVHIENSEVLTSRTKELEEKLEKLEAEKEELKSEVKCNREKAVV 424
Query: 675 WEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEK-- 732
+ + + A+ + T+ K ELE+QL ++ E EL++ ++ C +E+
Sbjct: 425 HVE-----------NSLAAEIEVLTSRTK--ELEEQLEKLEAEKVELESEVK-CNREEAV 470
Query: 733 EQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQAT 792
Q E L E L + +L +LE+ K L + K + E + + ++EA+
Sbjct: 471 AQVENSLATEIEVLTCRIKQLEEKLEKLEVEKDELKSEVKCNREVESTLRFELEAIACE- 529
Query: 793 SEEIKKATETHAEALATKDAEIASQLAKIKSLEDE 827
K E E L + AE+ IK +E
Sbjct: 530 ----KMELENKLEKLEVEKAELQISFDIIKDKYEE 560
Score = 38.1 bits (87), Expect = 0.023
Identities = 50/223 (22%), Positives = 96/223 (42%), Gaps = 43/223 (19%)
Query: 629 TKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGK 688
T+ E L C ++ EK VE + LK+E ++E + + +A
Sbjct: 479 TEIEVLTCRIK------QLEEKLEKLEVEKDELKSEVKCNREVESTLRFELEAIA----- 527
Query: 689 DKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIA 748
+KM ELE++L ++ E EL S K + E+++ L
Sbjct: 528 ----CEKM---------ELENKLEKLEVEKAELQISFDIIKDKYEESQV-------CLQE 567
Query: 749 KESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALA 808
E++L + E++LV + AE E ++ A AK+ A +++ E+++K E A
Sbjct: 568 IETKLGEIQTEMKLVNELKAEVESQTIAMEADAKTK-SAKIESLEEDMRK------ERFA 620
Query: 809 TKDAEIASQLAKIKSLEDELATEKAKAIEAREQAADIALDNRE 851
+ K ++LE+E++ K +I++ + I ++ E
Sbjct: 621 FDELR-----RKCEALEEEISLHKENSIKSENKEPKIKQEDIE 658
Score = 33.1 bits (74), Expect = 0.74
Identities = 47/235 (20%), Positives = 92/235 (39%), Gaps = 4/235 (1%)
Query: 605 SEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAANVEAERLKAE 664
+E L A L + +++ Q + V EE +A +A EK AA+ + R+ +
Sbjct: 66 AEKLSAALANVSAKDDLVKQHVKVA-EEAVAGWEKAENEVVELKEKLEAAD-DKNRVLED 123
Query: 665 SAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGE--LEDQLALMKEEADELD 722
H + A R + A + ++ + T ++ LE+Q+ +++EL
Sbjct: 124 RVSHLDGALKECVRQLRQARDEQEQRIQDAVIERTQELQSSRTSLENQIFETATKSEELS 183
Query: 723 ASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAK 782
++ KE +L+AR E L + E + E K + KK A+ A +
Sbjct: 184 QMAESVAKENVMLRHELLARCEELEIRTIERDLSTQAAETASKQQLDSIKKVAKLEAECR 243
Query: 783 SDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATEKAKAIE 837
+ S ++T++H++ D + A +E + +IE
Sbjct: 244 KFRMLAKSSASFNDHRSTDSHSDGGERMDVSCSDSWASSTLIEKRSLQGTSSSIE 298
>At1g12150 hypothetical protein
Length = 548
Score = 46.2 bits (108), Expect = 8e-05
Identities = 50/194 (25%), Positives = 88/194 (44%), Gaps = 33/194 (17%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELE 708
E +++ E++K E+ + + AA ++ + L + + A++ + I E+E
Sbjct: 362 EALKQESLKLEQMKTEAIEARNEAANMNRKIESLKKETEAAMIAAEEAEKRLELVIREVE 421
Query: 709 DQLALMKEEADELD--ASLQACKKEKEQAEKDLI------------ARGEALIAKESELA 754
+ + ++ +E+ + Q KK+ E++ I GE A E +LA
Sbjct: 422 EAKSAEEKVREEMKMISQKQESKKQDEESSGSKIKITIQEFESLKRGAGETEAAIEKKLA 481
Query: 755 ILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEI 814
+ AELE + K AE + K +EA ++A EE+K+ATE LA K AE
Sbjct: 482 TIAAELEEINKRRAEADNK-----------LEANLKAI-EEMKQATE-----LAQKSAES 524
Query: 815 ASQLAKIKSLEDEL 828
A A + +E EL
Sbjct: 525 AE--AAKRMVESEL 536
Score = 32.7 bits (73), Expect = 0.96
Identities = 41/207 (19%), Positives = 89/207 (42%), Gaps = 18/207 (8%)
Query: 649 EKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELE 708
E++ + VE + K + K +++ ++ D AT A A + + K+ EL
Sbjct: 152 EQYISTTVELDAAKQQLNKIRQS---FDSAMDFKAT-ALNQAAEAQRALQVNSAKVNELS 207
Query: 709 DQLALMKEEADELDASLQACKKEKEQA----EKDLIARGEALIAKESELAILCAELELVK 764
+++ MK+ +L L A + +E A EKD + +E+E +L +++
Sbjct: 208 KEISDMKDAIHQL--KLAAAQNLQEHANIVKEKDDLRECYRTAVEEAEKKLL-----VLR 260
Query: 765 KALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSL 824
K E E + + +L + + ++ EE+KKA E+ + E+ +++
Sbjct: 261 K---EYEPELSRTLEAKLLETTSEIEVLREEMKKAHESEMNTVKIITNELNEATMRLQEA 317
Query: 825 EDELATEKAKAIEAREQAADIALDNRE 851
D+ + ++ R + D+ + E
Sbjct: 318 ADDECSLRSLVNSLRMELEDLRREREE 344
>At2g34730 putative myosin heavy chain
Length = 829
Score = 45.4 bits (106), Expect = 1e-04
Identities = 55/247 (22%), Positives = 103/247 (41%), Gaps = 27/247 (10%)
Query: 601 AIEPSEFLLAGLNRDVIEKEVLSQGLNVTKEETLACLLRAGCIFAHTFEKFNAA--NVEA 658
A++ + + LN V EKE + V KE + R GC+ A N+
Sbjct: 544 AVKEAHKKIVELNLHVTEKEGTLRSEMVDKERLKEEIHRLGCLVKEKENLVQTAENNLAT 603
Query: 659 ERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGI--------KIGELEDQ 710
ER K E Q + + ++ T+ +DK+ A ++ + KI L ++
Sbjct: 604 ERKKIEVVSQQ--INDLQSQVERQETEI-QDKIEALSVVSARELEKVKGYETKISSLREE 660
Query: 711 LAL-------MKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELV 763
L L MK+E + + L K EKE +K L++ + + L +++
Sbjct: 661 LELARESLKEMKDEKRKTEEKLSETKAEKETLKKQLVSLDLVVPPQ------LIKGFDIL 714
Query: 764 KKALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKS 823
+ +AE+ +K+ L +S + + + E+K T+ + L K ++ A++
Sbjct: 715 EGLIAEKTQKTNSRLKNMQSQLSDLSHQIN-EVKGKASTYKQRLEKKCCDLKKAEAEVDL 773
Query: 824 LEDELAT 830
L DE+ T
Sbjct: 774 LGDEVET 780
Score = 36.6 bits (83), Expect = 0.067
Identities = 53/243 (21%), Positives = 95/243 (38%), Gaps = 34/243 (13%)
Query: 656 VEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMK 715
V+ ERLK E H+ EK + L A + K I +I +L+ Q+ +
Sbjct: 571 VDKERLKEEI--HRLGCLVKEK--ENLVQTAENNLATERKKIEVVSQQINDLQSQVERQE 626
Query: 716 EEA-DELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAE---QE 771
E D+++A +E E+ + E++++ L ELEL +++L E ++
Sbjct: 627 TEIQDKIEALSVVSARELEKVK-----------GYETKISSLREELELARESLKEMKDEK 675
Query: 772 KKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSLEDELATE 831
+K+ E L+ E K ET + L + D + QL K + + L E
Sbjct: 676 RKTEEKLS---------------ETKAEKETLKKQLVSLDLVVPPQLIKGFDILEGLIAE 720
Query: 832 KAKAIEAREQAADIALDNRERGFYLAKDQAQHLFPNFDFSAMGVMKEITAAGLVGPDDPP 891
K + +R + L + K +A + + K L+G +
Sbjct: 721 KTQKTNSRLKNMQSQLSDLSHQINEVKGKASTYKQRLEKKCCDLKKAEAEVDLLGDEVET 780
Query: 892 LID 894
L+D
Sbjct: 781 LLD 783
>At1g63300 hypothetical protein
Length = 1029
Score = 45.1 bits (105), Expect = 2e-04
Identities = 47/222 (21%), Positives = 102/222 (45%), Gaps = 21/222 (9%)
Query: 643 IFAHTFEKFNAANVEAERLKAESAKHQEAAAAWEKRFDKLATQAGKDK---VYADKMIGT 699
+ A+ ++ E E LK K+Q++ ++ + L K K + A+ +
Sbjct: 722 VTANLNQEIKILKEEIENLK----KNQDSLMLQAEQAENLRVDLEKTKKSVMEAEASLQR 777
Query: 700 AGIKIGELEDQLALMKEEADELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAE 759
+K ELE +++LM++E++ L A LQ K K++ E A+ ++EL + ++
Sbjct: 778 ENMKKIELESKISLMRKESESLAAELQVIKLAKDEKE-------TAISLLQTELETVRSQ 830
Query: 760 LELVKKALAEQE---KKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIAS 816
+ +K +L+E + +K + +A KS+++ + + KK E+ T +
Sbjct: 831 CDDLKHSLSENDLEMEKHKKQVAHVKSELKKKEETMANLEKKLKESRTAITKTAQRNNIN 890
Query: 817 QLAKI----KSLEDELATEKAKAIEAREQAADIALDNRERGF 854
+ + + S E + +K K +E + + + AL++ F
Sbjct: 891 KGSPVGAHGGSKEVAVMKDKIKLLEGQIKLKETALESSSNMF 932
Score = 35.8 bits (81), Expect = 0.11
Identities = 45/228 (19%), Positives = 95/228 (40%), Gaps = 41/228 (17%)
Query: 659 ERLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEA 718
E +K + + + A +E + +L+ + ++M+ K E+++Q ++
Sbjct: 664 EMIKDANDELRANQAEYEAKLHELSEKLSFKTSQMERMLENLDEKSNEIDNQKRHEEDVT 723
Query: 719 DELDASLQACKKEKEQAEKDLIARGEALIAKESELAILCAELELVKKALAEQE------- 771
L+ ++ K+E E +K+ ++L+ + + L +LE KK++ E E
Sbjct: 724 ANLNQEIKILKEEIENLKKNQ----DSLMLQAEQAENLRVDLEKTKKSVMEAEASLQREN 779
Query: 772 -------------KKSAESLA-------LAKSDME---AVMQATSEEIKKATETHAEALA 808
+K +ESLA LAK + E +++Q E ++ + +L+
Sbjct: 780 MKKIELESKISLMRKESESLAAELQVIKLAKDEKETAISLLQTELETVRSQCDDLKHSLS 839
Query: 809 TKDAE-------IASQLAKIKSLEDELATEKAKAIEAREQAADIALDN 849
D E +A +++K E+ +A + K E+R A N
Sbjct: 840 ENDLEMEKHKKQVAHVKSELKKKEETMANLEKKLKESRTAITKTAQRN 887
>At5g24710 unknown protein
Length = 1003
Score = 44.7 bits (104), Expect = 2e-04
Identities = 54/199 (27%), Positives = 76/199 (38%), Gaps = 32/199 (16%)
Query: 414 PKFNTEGDAKKRG-------GAENQAESTKAPKRRRLTKASSDAGTSKPGA--QPTTAAA 464
PK + DA ++ G + S APK+ LT+ ++ +KP A +PT A
Sbjct: 785 PKLTSLSDASRKPPIEILPPGMSSIFASITAPKKPLLTQKTAQPEVAKPLALEEPTKPLA 844
Query: 465 --------APKGKNVAEASVAAATEP---TAVPASSPASAGATAAFAAESSIGATAASAG 513
AP+ ++ E + AAA P TA A SPA A A A S A
Sbjct: 845 IEAPPSSEAPQTESAPETA-AAAESPAPETAAVAESPAPGTAAVAEAPASETAAAPVDGP 903
Query: 514 VNATKAAASSDTPIGDKE---KENETPKSPPRQDAPPSPPSTHDAGPMP-----SPPHQG 565
V T S P+ +E +E P S P + S +T P +PP
Sbjct: 904 VTET---VSEPPPVEKEETSLEEKSDPSSTPNTETATSTENTSQTTTTPESVTTAPPEPI 960
Query: 566 EKSCPGAATTSEAAQIEQA 584
+ P TT+ E A
Sbjct: 961 TTAPPETVTTTAVKPTENA 979
Score = 35.0 bits (79), Expect = 0.19
Identities = 38/152 (25%), Positives = 49/152 (32%), Gaps = 10/152 (6%)
Query: 465 APKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSD 524
AP K A ++ A+ E + + S AS A+ +
Sbjct: 767 APSSKTDAASAFLASLEDPKLTSLSDASRKPPIEILPPGMSSIFASITAPKKPLLTQKTA 826
Query: 525 TPIGDKEKENETPKSPPRQDAPPSP--PSTHDA--------GPMPSPPHQGEKSCPGAAT 574
P K E P P +APPS P T A P P E PG A
Sbjct: 827 QPEVAKPLALEEPTKPLAIEAPPSSEAPQTESAPETAAAAESPAPETAAVAESPAPGTAA 886
Query: 575 TSEAAQIEQAPAPEVGSSSYYNMLPNAIEPSE 606
+EA E A AP G + P +E E
Sbjct: 887 VAEAPASETAAAPVDGPVTETVSEPPPVEKEE 918
>At5g16730 putative protein
Length = 853
Score = 44.7 bits (104), Expect = 2e-04
Identities = 43/180 (23%), Positives = 83/180 (45%), Gaps = 13/180 (7%)
Query: 676 EKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACKKEKEQA 735
E+R KL T K + ++ G + + +L+ +KE+ + + + + +K+K +A
Sbjct: 68 ERRSPKLPTPPEKSQA---RVAAVKGTESPQTTTRLSQIKEDLKKANERISSLEKDKAKA 124
Query: 736 EKDLIARGEALIAKESELAILCAELELVKKALAEQEKKSAESLALAKSDMEAVMQATSEE 795
+L KE+E L + + +K +E E ++ +EAV Q EE
Sbjct: 125 LDEL-----KQAKKEAEQVTLKLD-DALKAQKHVEENSEIEKFQAVEAGIEAV-QNNEEE 177
Query: 796 IKKATETHAEALATKDAEIASQLAKIKSLEDELAT---EKAKAIEAREQAADIALDNRER 852
+KK ET A+ A + + +++ + +ELA K+KA+ E A+ A + E+
Sbjct: 178 LKKELETVKNQHASDSAALVAVRQELEKINEELAAAFDAKSKALSQAEDASKTAEIHAEK 237
Score = 38.1 bits (87), Expect = 0.023
Identities = 50/248 (20%), Positives = 103/248 (41%), Gaps = 36/248 (14%)
Query: 618 EKEVLSQGLNVTKEETLACLLRAGCIFAHTFEK--------FNAANVEAERLKAESAKHQ 669
EK +S V K E +L+ A FE NV+ E K +
Sbjct: 257 EKTAISDNEMVAKLEDEIVVLKRDLESARGFEAEVKEKEMIVEKLNVDLEAAKMAESNAH 316
Query: 670 EAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMKEEADELDASLQACK 729
+ W+ + +L Q + A+K+ +A + + + QL ++ + + + K
Sbjct: 317 SLSNEWQSKAKELEEQLEE----ANKLERSASVSLESVMKQLEGSNDKLHDTETEITDLK 372
Query: 730 KEKEQAEKDLIARGEALIAKESELAILCAELELVKKALA---EQEKKSAESLALAKSDME 786
+ I E +AK+ E +LE+ ++ L E+ K+ + + KS++E
Sbjct: 373 ER--------IVTLETTVAKQKE------DLEVSEQRLGSVEEEVSKNEKEVEKLKSELE 418
Query: 787 AVMQATSEEIKKATE--THAEALATKDAEIASQLAKIKSLEDELATEKAKAIEAREQAA- 843
V + + +KK + + + L+ + +++ S L K E+ + KA+E+ A
Sbjct: 419 TVKEEKNRALKKEQDATSRVQRLSEEKSKLLSDLESSKEEEE----KSKKAMESLASALH 474
Query: 844 DIALDNRE 851
+++ + RE
Sbjct: 475 EVSSEGRE 482
Score = 35.4 bits (80), Expect = 0.15
Identities = 67/284 (23%), Positives = 115/284 (39%), Gaps = 68/284 (23%)
Query: 660 RLKAESAKHQEAAAAWEKRFDKLATQAGKDKVYADKMIGTAGIKIGELEDQLALMK---- 715
++K + K E ++ EK K + + K A+++ +L+D L K
Sbjct: 102 QIKEDLKKANERISSLEKDKAKALDELKQAKKEAEQVTL-------KLDDALKAQKHVEE 154
Query: 716 ----EEADELDASLQACKKEKEQAEKDL-------IARGEALIAKESELAILCAELELVK 764
E+ ++A ++A + +E+ +K+L + AL+A EL
Sbjct: 155 NSEIEKFQAVEAGIEAVQNNEEELKKELETVKNQHASDSAALVAVRQEL----------- 203
Query: 765 KALAEQEKKSAESLALAKSDMEAVMQATSEEIKKATETHAEALATKDAEIASQLAKIKSL 824
EK + E A + +A+ QA E+ K E HAE K ++S+L ++K+L
Sbjct: 204 ------EKINEELAAAFDAKSKALSQA--EDASKTAEIHAE----KVDILSSELTRLKAL 251
Query: 825 EDELATEKAKAIEAREQAA----DIALDNRE----RGFYLAKDQAQHLFP--NFDFSAMG 874
D +T + AI E A +I + R+ RGF + + + N D A
Sbjct: 252 LD--STREKTAISDNEMVAKLEDEIVVLKRDLESARGFEAEVKEKEMIVEKLNVDLEA-A 308
Query: 875 VMKEITAAGLVGPDDPPLIDQNLWTATEEEEEEEEEEEQEKENN 918
M E A L N W + +E EE+ EE + E +
Sbjct: 309 KMAESNAHSL----------SNEWQSKAKELEEQLEEANKLERS 342
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.128 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,993,427
Number of Sequences: 26719
Number of extensions: 976447
Number of successful extensions: 11408
Number of sequences better than 10.0: 474
Number of HSP's better than 10.0 without gapping: 146
Number of HSP's successfully gapped in prelim test: 340
Number of HSP's that attempted gapping in prelim test: 8215
Number of HSP's gapped (non-prelim): 2012
length of query: 919
length of database: 11,318,596
effective HSP length: 108
effective length of query: 811
effective length of database: 8,432,944
effective search space: 6839117584
effective search space used: 6839117584
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0590c.8


