

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.10
(993 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
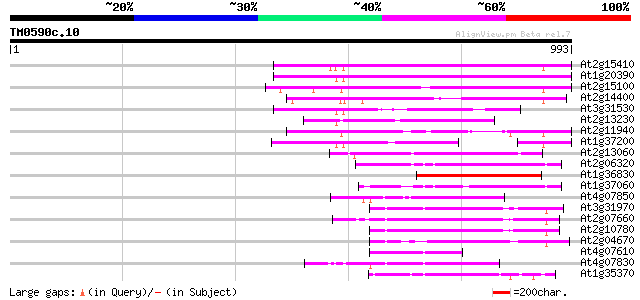
Score E
Sequences producing significant alignments: (bits) Value
At2g15410 putative retroelement pol polyprotein 368 e-102
At1g20390 hypothetical protein 367 e-101
At2g15100 putative retroelement pol polyprotein 289 6e-78
At2g14400 putative retroelement pol polyprotein 278 1e-74
At3g31530 hypothetical protein 234 1e-61
At2g13230 pseudogene 231 2e-60
At2g11940 putative retroelement gag/pol polyprotein 191 1e-48
At1g37200 hypothetical protein 185 1e-46
At2g13060 pseudogene 181 1e-45
At2g06320 putative retroelement pol polyprotein 172 6e-43
At1g36830 hypothetical protein 169 5e-42
At1g37060 Athila retroelment ORF 1, putative 159 6e-39
At4g07850 putative polyprotein 119 6e-27
At3g31970 hypothetical protein 117 4e-26
At2g07660 putative retroelement pol polyprotein 115 1e-25
At2g10780 pseudogene 109 7e-24
At2g04670 putative retroelement pol polyprotein 103 4e-22
At4g07610 103 6e-22
At4g07830 putative reverse transcriptase 102 1e-21
At1g35370 hypothetical protein 102 1e-21
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 368 bits (945), Expect = e-102
Identities = 220/596 (36%), Positives = 312/596 (51%), Gaps = 71/596 (11%)
Query: 468 ATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYL 527
A+NN+AEYEALIAGL+LA ++++ + DSQLV +Q G ++ K+ + YL+ VR L
Sbjct: 1192 ASNNEAEYEALIAGLRLAHGIEVKKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVVREL 1251
Query: 528 MTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSV-----------IQETLAYPSI- 575
F+ + +PR++N ADALA LAST P + + IQ T+ PS
Sbjct: 1252 SYNFEVFELTKIPRSDNAPADALAVLASTSDPDLRRVIPVECIDVPSIKIQGTITKPSEY 1311
Query: 576 -----------------------------EGELMACVSRGRT------------------ 588
G L+ G++
Sbjct: 1312 TVIDLTAEQLSIVMVILPPATDRSEDAGPSGNLVCSSDPGQSSGCDRSDDSNRSADSDSP 1371
Query: 589 ----WMDPIISILEGDPAEVEQCTKEQQREAS-HYTLIDGHLYRRGFSTPLLKCVPPEKY 643
W++ I S + ++ + R S YTL HL R + LL C+ E+
Sbjct: 1372 ASTNWIEEIRSYIADRIVPKDKWAARRLRARSAQYTLPHEHLLRWSATGVLLSCLDDEEA 1431
Query: 644 EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPP 703
+ +M E+HEG +H GGR+LA K+ + G YWPT+ DC FV +C++CQ A P
Sbjct: 1432 QQVMREIHEGAGGNHSGGRALALKIRKHGQYWPTMNADCEKFVARCEKCQGHAPFIHIPS 1491
Query: 704 KELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNF 763
+ L T P+PF W +D++ PFP +R Q K+ILV DYFTKW EAE A+I S ++ NF
Sbjct: 1492 EVLQTAMPPYPFMRWAMDIIRPFPASR-QKKYILVMTDYFTKWFEAESYARIQSKEVQNF 1550
Query: 764 YWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVI 823
WK I+C G+P IV+DNGTQF+S Q FC + I++ ++ +PQ NGQ E+ N+ I
Sbjct: 1551 VWKNIICHHGLPYEIVTDNGTQFTSLQFEGFCAKWKIRLSKSTPRYPQCNGQAEATNKTI 1610
Query: 824 LRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRT 883
L GL++RL E KGAW DEL VLWSY TT + +T TPF +TYG++A+ P E T R
Sbjct: 1611 LDGLKKRLDEKKGAWADELDGVLWSYRTTPRRSTDRTPFSLTYGMEALAPCEAGLPTVRR 1670
Query: 884 RPGFEEE--NQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVREMQVGDLVLK 941
+ N + LD L E RD+A +R +Q A +N +V+ R GDLVL+
Sbjct: 1671 SMLINDPTLNDQMLLDNLDTLEEQRDQALLRIQNYQQAAAKFYNKKVKNRLFAEGDLVLR 1730
Query: 942 W----RSGAPGNKLTPNWEGPYRIIKVLGNGAYHLEELDGRRLPRSFNGLSLRYFY 993
KL +WEGPY I KV+ G Y L +DG +PRS+N +L+ +Y
Sbjct: 1731 KVFENTVEQDAGKLGAHWEGPYLISKVVKPGVYELLTMDGTPIPRSWNSANLKRYY 1786
>At1g20390 hypothetical protein
Length = 1791
Score = 367 bits (943), Expect = e-101
Identities = 210/545 (38%), Positives = 302/545 (54%), Gaps = 20/545 (3%)
Query: 468 ATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYL 527
ATNN AEYE LIAGL+LA ++I ++ TDSQL+ Q+ G ++ K+ + YL+ V+ +
Sbjct: 1247 ATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQLM 1306
Query: 528 MTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEG----ELMACV 583
F+ + +PR +N ADALA LA T + + E++ PSI+ E++ +
Sbjct: 1307 TKDFENFKLSKIPRGDNAPADALAALALTSDSDLRRIIPVESIDKPSIDSTDAVEIVNTI 1366
Query: 584 SRGRT--------WMDPIISILEGDPAEVEQCTKEQQR-EASHYTLIDGHLYRRGFSTPL 634
W I L ++ T + R +A+ YTL+ HL + +
Sbjct: 1367 RSSNAPDPADPTDWRVEIRDYLSDGTLPSDKWTARRLRIKAAKYTLMKEHLLKVSAFGAM 1426
Query: 635 LKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 694
L C+ + IM E HEG +H GGR+LA K+ + GFYWPT+ DC F +C++CQ
Sbjct: 1427 LNCLHGTEINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQR 1486
Query: 695 FADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAK 754
A P + L AP+PF W +D+VGP P +R Q +FILV DYFTKW+EAE A
Sbjct: 1487 HAPTIHQPTELLRAGVAPYPFMRWAMDIVGPMPASR-QKRFILVMTDYFTKWVEAESYAT 1545
Query: 755 ITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNG 814
I + + NF WK I+CR G+P I++DNG+QF S FC I++ ++ +PQ NG
Sbjct: 1546 IRANDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNG 1605
Query: 815 QVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPV 874
Q E+ N+ IL GL++RL E KGAW DEL VLWSY TT +S T +TPF YG++AM P
Sbjct: 1606 QAEATNKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPA 1665
Query: 875 EIDNFTWRTRPGFE--EENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVRE 932
E+ + R + E N M LD L E R+ A R + A +N +V R
Sbjct: 1666 EVGYSSLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRH 1725
Query: 933 MQVGDLVLK--WRSGAPGN--KLTPNWEGPYRIIKVLGNGAYHLEELDGRRLPRSFNGLS 988
VGDLVL+ + + A N KL NWEG Y++ K++ G Y L + G +PR++N +
Sbjct: 1726 FDVGDLVLRKVFENTAEINAGKLGANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSMH 1785
Query: 989 LRYFY 993
L+ +Y
Sbjct: 1786 LKRYY 1790
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 289 bits (739), Expect = 6e-78
Identities = 188/569 (33%), Positives = 282/569 (49%), Gaps = 42/569 (7%)
Query: 453 IKEIVKASGAMVAIKATNNQAEYEA----LIAGLKLAREVKIRSLLIR-TDSQLVENQVK 507
IK V++ A A EYEA + A L L REV R T EN
Sbjct: 774 IKTWVRSRNLSDAPTANRYSGEYEAKDACMEAYLNLVREVSGRFEQFELTRIPRAENSAA 833
Query: 508 GTFQVKDPNLIKYLERVRYLMTLFQEVV----VEYVP-RAENQRADALAKLASTRKPGNN 562
L RV + T+ Q + + +V RA +R DA + + G++
Sbjct: 834 NALAALASTFEVTLPRVIPVETISQPSIRLDEISFVTTRAMRRRLDAQSAENGLHQLGDD 893
Query: 563 KSVIQETLAYPSIEGELMACVSR-----------GRTWMDPIIS-ILEGDPAEVEQCTKE 610
+ + +E + + G W +PI IL G + ++
Sbjct: 894 EEISDAVHPTEIVENQSLPDNHNAPLPDQPPHDWGADWREPIRDYILNGTLPAEKWAARK 953
Query: 611 QQREASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLR 670
+ + + + + LYRR FS P C+ E+ +M EVH+G C +H GGRSLA KV +
Sbjct: 954 LKATCARFCIANDILYRRIFSAPDAVCIFGEQTRTVMKEVHDGTCGNHTGGRSLAFKVRK 1013
Query: 671 AGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTAR 730
G+YWPTL DC + ++C++CQ A L P + L T+SAP+PF W +D+VGP +
Sbjct: 1014 YGYYWPTLVADCEAYARKCEQCQKHAPLILQPAELLTTVSAPYPFMKWLMDIVGPLHVS- 1072
Query: 731 AQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQ 790
T+ +EA + IT ++ NF WK I+CR G+P IV+DNG+QF S Q
Sbjct: 1073 -------------TRGVEAAAYSNITHVQVWNFIWKDIICRHGLPYEIVTDNGSQFISEQ 1119
Query: 791 TREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYN 850
FC+E I++ ++ +PQ NGQ E+ N+ I+ L+++L KGAW EL VLW+
Sbjct: 1120 FEVFCEEWQIRLSHSTPRYPQGNGQAEAMNKTIISNLKKKLNAYKGAWFGELQNVLWAVR 1179
Query: 851 TTEQSTTRETPFRMTYGVDAMLPVEIDNFTWR--TRPGFEEENQANMAVELDLLSETRDE 908
TT + T ETPF + YG++A++P EI + R P E EN + +D + E R+
Sbjct: 1180 TTPRRATDETPFSLIYGMEAVIPAEIKVPSARRIRNPQNETENNEMIIDVIDTIDERRNR 1239
Query: 909 AHIRETAMKQRVAAKFNSRVRVREMQVGDLVLKW----RSGAPGNKLTPNWEGPYRIIKV 964
A R A +NS VR R +VG LVL+ ++ KL +WEGPY+I V
Sbjct: 1240 ALARMQNYHNAGARYYNSNVRNRSFEVGTLVLRRVQQNKAEKGAGKLGISWEGPYKITHV 1299
Query: 965 LGNGAYHLEELDGRRLPRSFNGLSLRYFY 993
+ NG Y L ++G+ + R++N + L+ FY
Sbjct: 1300 VRNGVYRLINMEGKTVRRAWNSMHLKRFY 1328
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 278 bits (711), Expect = 1e-74
Identities = 178/548 (32%), Positives = 274/548 (49%), Gaps = 94/548 (17%)
Query: 490 IRSLLIRTDS-------QLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRA 542
+ +LIR D ++ NQ G ++ KD + YL V+ L F++ + +PR
Sbjct: 772 VSGVLIREDRGEQKPIYYIIVNQFNGDYEAKDSRMEAYLHVVKDLAKNFRKFELIRIPRG 831
Query: 543 ENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELMACV---------SRGRTWMD-- 591
+N ADALA LAST P N + E ++ SI+ E A V RG T ++
Sbjct: 832 QNTTADALAALASTSDPEVNSIIPVECISERSIKEEKEAFVVTRSRAAARDRGDTAVELP 891
Query: 592 ------------PIISILEGDPAE-VEQCTKEQQREASHYTLID---------------G 623
P ++ E E +E+ + + EA +D
Sbjct: 892 PVKRRKSKEATVPKPAVQENIGIELIEEVISDAKIEAETEPWVDEDILAPNEHPEPEALN 951
Query: 624 HLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCM 683
LYRRG S P L + E +MSEVHEG+C SH GR++A K+ + +YWPT+ DC+
Sbjct: 952 SLYRRGVSDPYLLSIFGPVVEIVMSEVHEGLCGSHSSGRAMAFKIKKMCYYWPTMITDCV 1011
Query: 684 DFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYF 743
+ ++CK CQ+ A L P + ++SAP+PF W +D++GP + ++++LV DYF
Sbjct: 1012 KYAQRCKRCQLHAPLIHQPSELFSSISAPYPFMRWSMDIIGPLHRSTRGVQYLLVLTDYF 1071
Query: 744 TKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMR 803
+KWIEAE + ++N+ GI++
Sbjct: 1072 SKWIEAEAI-------LLNW-----------------------------------GIKVS 1089
Query: 804 FASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFR 863
++++ +PQ NGQ E+AN+ IL L++RL KG W DEL VLW+Y TT + +T ETPF
Sbjct: 1090 YSTLRYPQGNGQAEAANKTILSNLKKRLNHLKGGWYDELQPVLWAYRTTPRRSTGETPFS 1149
Query: 864 MTYGVDAMLPVEID--NFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVA 921
+ YG+ A++P E++ P E+EN A + LD ++E RD+A I+ + A
Sbjct: 1150 LVYGMKAVVPAELNVPGLRRTEAPLNEKENSAMLDDSLDTINERRDQALIQIQNYQHAAA 1209
Query: 922 AKFNSRVRVREMQVGDLVLKW----RSGAPGNKLTPNWEGPYRIIKVLGNGAYHLEELDG 977
+NS+V+ R VGD VLK + KL NWEGPY + +V+ NG Y L++L+
Sbjct: 1210 RYYNSKVKSRPFFVGDYVLKRVFDNKKEEGAGKLGINWEGPYIVTEVVRNGVYRLKDLED 1269
Query: 978 RRLPRSFN 985
R + R +N
Sbjct: 1270 RPVQRPWN 1277
>At3g31530 hypothetical protein
Length = 831
Score = 234 bits (598), Expect = 1e-61
Identities = 158/454 (34%), Positives = 225/454 (48%), Gaps = 77/454 (16%)
Query: 467 KATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRY 526
+ATNN AEYEAL+AGL LA +KI DSQL+ NQ G + +D + YL V+
Sbjct: 439 EATNNVAEYEALVAGLNLAWGLKIGKTRAFCDSQLIANQFNGEYTTQDKKMEAYLIHVQN 498
Query: 527 LMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIE-----GELMA 581
L F E + +PR EN ADALA LAST + + E + PSIE L
Sbjct: 499 LAKNFDEFELTRIPRGENTSADALAALASTSDTSLKRVIPVEFIEKPSIELGKEEHVLPI 558
Query: 582 CVSRGRT--------WMDPIIS-ILEGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFST 632
+S + WM+PIIS I EG + ++ + +A+ + L+D LY+ S
Sbjct: 559 QISADQDDPDDCNSEWMEPIISYISEGKLPSDKWKARKLKAQAARFVLVDTKLYKWRLSG 618
Query: 633 PLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKEC 692
PL+ CV E IM E+H G SH PT+ +
Sbjct: 619 PLMTCVEAEAICKIMKEIHSG---SHA----------------PTIHQ------------ 647
Query: 693 QVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPL 752
P + L ++++P+PF W +D++GP ++ Q K +LV DYF+KWIEAE
Sbjct: 648 ---------PAELLSSIASPYPFMRWSMDIIGPMHPSK-QKKLVLVLTDYFSKWIEAESY 697
Query: 753 AKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQT 812
A I A++ NF K I+CR GIP IV+DNG+QF S++ + FC + GI++ ++ +PQ
Sbjct: 698 ASIKDAQVENFVLKHILCRHGIPYEIVTDNGSQFISTRFQGFCDKWGIRLSKSTPRYPQG 757
Query: 813 NGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAML 872
NGQ E+AN EL VLW + TT + TRETPF YG + ++
Sbjct: 758 NGQAEAAN--------------------ELEGVLWLHRTTPRRATRETPFASIYGTECVI 797
Query: 873 PVE--IDNFTWRTRPGFEEENQANMAVELDLLSE 904
P E + + P + +N ELDL+ E
Sbjct: 798 PAEMIVPSLRRSLSPENDPDNTQTPLDELDLIDE 831
>At2g13230 pseudogene
Length = 353
Score = 231 bits (588), Expect = 2e-60
Identities = 128/361 (35%), Positives = 200/361 (54%), Gaps = 34/361 (9%)
Query: 520 YLERVRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIE--- 576
YL V+ F E + +PR EN ADALA LAST + + E + PSIE
Sbjct: 4 YLALVQDHAKQFHEFKLTRIPRGENTSADALAALASTSDLNMRRVIPVEFIEKPSIELDK 63
Query: 577 -------------------GELMACVSRGRTWMDPIISILEGDPAEVEQC-TKEQQREAS 616
+ C S W++PI + + +++ ++ + EA+
Sbjct: 64 VEHAFPIQVVAGYEDASDNSPSLGCDSE---WIEPIRNFISKRKVRLDKWEARKLKAEAA 120
Query: 617 HYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWP 676
+ L++ L++ S PL+ CV E +M EVH G C ++ GGR+LA K+ R G++WP
Sbjct: 121 RFVLVEEKLFKWRLSGPLMTCVEKEAARKVMKEVHGGSCGNNSGGRALAIKIKRLGYFWP 180
Query: 677 TLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFI 736
T+ KDC ++CQ A P + L ++++P+P W +D++GP ++ Q K +
Sbjct: 181 TMIKDC-------EKCQRNAPTIHLPAELLSSIASPYPLMRWSMDIIGPMHASK-QKKLV 232
Query: 737 LVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCK 796
LV DYF+K EAE A I A++ +F WK I+CR G+P IV+DNG+QF S++ + FC
Sbjct: 233 LVLTDYFSKLREAESYASIKEAQVESFMWKHILCRHGVPYEIVTDNGSQFISTRFQGFCD 292
Query: 797 EMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQST 856
+ GI++ ++ + Q NGQ ++AN+ IL GL++RL KG+W+DEL VLWS+ TT +
Sbjct: 293 KWGIRLSKSTPRYLQGNGQADAANKTILDGLKKRLTAKKGSWVDELEGVLWSHRTTPRRA 352
Query: 857 T 857
T
Sbjct: 353 T 353
>At2g11940 putative retroelement gag/pol polyprotein
Length = 1212
Score = 191 bits (486), Expect = 1e-48
Identities = 149/558 (26%), Positives = 235/558 (41%), Gaps = 130/558 (23%)
Query: 490 IRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVPRAENQRADA 549
+ LI + G ++ KD + YL+ +R L F++ + VPRAEN ADA
Sbjct: 730 LADFLIELPLATTASDYSGDYEAKDNRMEAYLDLLRELAEKFEKFELIKVPRAENSAADA 789
Query: 550 LAKLASTRKPGNNKSVIQETLAYPSIEGELMACVSR------------------------ 585
LA LAST + + + ET+A P+I E + V+R
Sbjct: 790 LAALASTPEITVRRIISVETIAQPNIRIEEINFVTRAMRRQLEVQANNPELQQLHDEDEI 849
Query: 586 ---------------------GRTWMDPIISILEGDPAEVEQC-TKEQQREASHYTLIDG 623
G W +PI + + ++ ++ + + + L D
Sbjct: 850 PEAVPLVADEDSTEYNLPHDWGADWREPIRNYIFNGVLPTDKWEARKLKATCARFCLADD 909
Query: 624 HLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCM 683
LYRR S P C+ E+ ++ E+H+G C +H GGRSLA K+ + G++WPT+ DC
Sbjct: 910 ILYRRIISAPDAICIFGEQTRTVIKEIHDGTCGNHSGGRSLAFKLKKYGYFWPTMVADCE 969
Query: 684 DFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYF 743
+ + + F +P PF W +D+VGP + ++F+L+
Sbjct: 970 AYAHRSRRTFHFG-------------ISPLPFMKWSMDIVGPLHVSTRGVRFLLL----- 1011
Query: 744 TKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMR 803
+W+EA + IT ++ +F WK I+CR G+P IV+DNG QF S Q + F E I++
Sbjct: 1012 -EWVEAAAYSYITQVQVRHFIWKEIICRHGLPYEIVTDNGPQFISEQFKAFYAEWQIRLN 1070
Query: 804 FASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFR 863
++ +PQ G VE+
Sbjct: 1071 RSTPRYPQ--GNVEA--------------------------------------------- 1083
Query: 864 MTYGVDAMLPVEIDNFTWRTR----PGFEEENQANMAVELDLLSETRDEAHIRETAMKQR 919
++P EI RTR P E N M +D + E R+ A +R
Sbjct: 1084 -------VIPAEIK--VPRTRRIRNPKNEAANNKMMVDVIDTIDERRNRALVRMQNYHNA 1134
Query: 920 VAAKFNSRVRVREMQVGDLVLKW----RSGAPGNKLTPNWEGPYRIIKVLGNGAYHLEEL 975
A +NS VR R VG LVL+ + KL +WEGPY+I ++ NG L +
Sbjct: 1135 AAQYYNSNVRSRSFDVGTLVLRRIHQNTAEKGAGKLGISWEGPYKITHIVRNGDC-LINM 1193
Query: 976 DGRRLPRSFNGLSLRYFY 993
+G+ + R++N L+ FY
Sbjct: 1194 EGKTVRRAWNAAHLKRFY 1211
>At1g37200 hypothetical protein
Length = 1564
Score = 185 bits (470), Expect = 1e-46
Identities = 115/383 (30%), Positives = 186/383 (48%), Gaps = 62/383 (16%)
Query: 464 VAIKATNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLER 523
+ A+NN++EYEALI G+KLA+ ++IR + +DSQLV +Q G ++ KD + YLE
Sbjct: 1074 LGFNASNNESEYEALIDGIKLAQGMRIRDIHAHSDSQLVTSQFHGEYEAKDERMEAYLEL 1133
Query: 524 VRYLMTLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIE------- 576
V+ L F+ + +PR EN DALA LAST P + + E + +PSI+
Sbjct: 1134 VKTLTQQFESFELTRIPRGENTSTDALAALASTLDPFVKRIIPVEGIEHPSIDLTVKHAG 1193
Query: 577 --------------------GELMACVSRGRT----------------------WMDPII 594
L A G W PII
Sbjct: 1194 METSATCNFTRVTRSTTAAARALAAAAEAGAAEEEAKLRTPEQLASYEPEPYNDWRIPII 1253
Query: 595 SILEGDPAEVEQC-TKEQQREASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEG 653
+E ++ T++ + +++ Y +++G L +R + P + C ++ + +M +HEG
Sbjct: 1254 DYIERGITPPDKWETRKLKAQSARYCIMEGRLMKRSVAGPYMVCTYGQQTKDLMKSMHEG 1313
Query: 654 VCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPW 713
C SH GR+L C +L DC+ +C +CQ A PP+E+ ++S+P+
Sbjct: 1314 QCGSHYSGRTLRCIMLA----------DCIAHSLRCDKCQRHAPTLHQPPEEMSSISSPY 1363
Query: 714 PFAMWGVDLVGPFPTA--RAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCR 771
PF W +D+V P + + ++K +LV DY TKWIE + ++T ++ F + IV R
Sbjct: 1364 PFMKWSMDVVRPMEASGGKKRLKNLLVLTDYSTKWIEGKAFQQVTEKQVEVFLLENIVYR 1423
Query: 772 FGIPRAIVSDNGTQFSSSQTREF 794
GIP IV+DNG +++ F
Sbjct: 1424 HGIPYEIVTDNGLTSPREKSKHF 1446
Score = 56.6 bits (135), Expect = 7e-08
Identities = 33/99 (33%), Positives = 54/99 (54%), Gaps = 5/99 (5%)
Query: 900 DLLSETRDEAHIRETAMKQRVAAKFNSRVRVREMQVGDLVLKW----RSGAPGNKLTPNW 955
D + E RD A I+ +Q +NS +++R +VG+LVL+ KL NW
Sbjct: 1465 DFIDERRDHAMIQVQNYQQPAVRYYNSNIKIRRFEVGELVLRKVFSNTRELNAGKLGTNW 1524
Query: 956 EGPYRIIKVLGNGAYHLEEL-DGRRLPRSFNGLSLRYFY 993
EGPYRI +V+ +G Y L ++ +G R +N + L+ ++
Sbjct: 1525 EGPYRITEVVRDGIYKLVKVFNGVPELRPWNAMHLKKYH 1563
>At2g13060 pseudogene
Length = 930
Score = 181 bits (460), Expect = 1e-45
Identities = 116/384 (30%), Positives = 183/384 (47%), Gaps = 17/384 (4%)
Query: 567 QETLAYPSIEGELMACVSRGRTWMDPIISILEGDPAEVEQCT----KEQQREASHYTLID 622
+ T + S E + V + W I++ L D E + T K RE Y +
Sbjct: 116 ESTRKWSSTENCAVTAVEKDYPWYADIVNYLAAD-VEPDNFTDYNKKRFLREIRRYQWDE 174
Query: 623 GHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDC 682
+LY+ + +C+ + +I+S H H KVL+A F+WPT+ +D
Sbjct: 175 PYLYKHSYDGIYRRCIAATEVPSILSHCHSSSYGGHFATFKTVSKVLQADFWWPTMFRDT 234
Query: 683 MDFVKQCKECQVFADLSKA---PPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVA 739
F+ QC CQ +SK PP + + F WG+D +GPFP + + +ILV
Sbjct: 235 QKFISQCDPCQRRGKISKRNEMPPNFRLEVEV---FDRWGIDFMGPFPPSNKNL-YILVD 290
Query: 740 VDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMG 799
VDY +KW+EA K SA ++ + I RFG+PR ++SD F + + + G
Sbjct: 291 VDYVSKWVEAIASLKNDSAVVMKLFKSIIFPRFGVPRIVISDGDKHFINKILEKLLLQYG 350
Query: 800 IQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRE 859
+Q R A+ HPQT+GQVE +NR I L + + +AK W +L LW+Y T ++
Sbjct: 351 VQHRVATPYHPQTSGQVEVSNRQIKEILEKTVGKAKKEWSYKLYDALWAYKTAFKTPLGT 410
Query: 860 TPFRMTYGVDAMLPVEIDN-FTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQ 918
TPF + YG LPVE+++ W + + A + ELD E R A+ K+
Sbjct: 411 TPFHLLYGKACHLPVELEHKAAWAVKMMNFDIKSAGES-ELD---EIRIHAYDNSKLYKE 466
Query: 919 RVAAKFNSRVRVREMQVGDLVLKW 942
R A + ++ R + D +L++
Sbjct: 467 RTKAYHDKKILTRTFEPNDQILRF 490
>At2g06320 putative retroelement pol polyprotein
Length = 466
Score = 172 bits (437), Expect = 6e-43
Identities = 117/376 (31%), Positives = 186/376 (49%), Gaps = 18/376 (4%)
Query: 613 REASHYTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAG 672
R+ HY + +LY V ++ E I+ H+ H K+L+AG
Sbjct: 66 RDIHHYYWDEPYLYTLCKDKIYRSYVSEDEVEGILLHCHDSAYGGHFATFKTVSKILQAG 125
Query: 673 FYWPTLRKDCMDFVKQCKECQVFADLSKA---PPKELVTMSAPWPFAMWGVDLVGPFPTA 729
F+WPT+ KD +FV +C CQ ++S+ P ++ + F +WG+D +GPFP++
Sbjct: 126 FWWPTMFKDAQEFVSKCDSCQRKGNISRRNEMPQNPILEVDI---FDVWGIDFMGPFPSS 182
Query: 730 RAQMKFILVAVDYFTKWIEAEPLAKITS-AKIVNFYWKRIVC-RFGIPRAIVSDNGTQFS 787
K+ILV VDY +KW+EA +A T+ AK+V +K I+ RFG+ ++SD G F
Sbjct: 183 YGN-KYILVVVDYVSKWVEA--IASPTNDAKVVLKLFKTIIFPRFGVSWVVISDGGKHFI 239
Query: 788 SSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLW 847
+ K+ G++ + A+ HPQT+GQVE +NR I L + + + W +L LW
Sbjct: 240 NKVFENLLKKHGVKHKVATPYHPQTSGQVEISNREIKTILEKTVGITRKDWSTKLDDALW 299
Query: 848 SYNTTEQSTTRETPFRMTYGVDAMLPVEID-NFTWRTR-PGFE-EENQANMAVELDLLSE 904
+Y T ++ TPF + L VE++ W + F+ + + ++L L E
Sbjct: 300 AYKTAFKTPIGTTPFNLLCVKSCHLHVELEYKAMWAVKLLNFDIKTAEEKRLIQLSELDE 359
Query: 905 TRDEAHIRETAMKQRVAAKFNSRVRVREMQVGDLVLKWRSGA---PGNKLTPNWEGPYRI 961
R EA+ K+R + ++ ++ QVGD VL + S PG KL W GP+ I
Sbjct: 360 IRLEAYESSKIYKERTKLFHDKKIITKDFQVGDQVLLFNSRLKIFPG-KLKSRWSGPFCI 418
Query: 962 IKVLGNGAYHLEELDG 977
+V GA L G
Sbjct: 419 TEVRPYGAVTLAGKSG 434
>At1g36830 hypothetical protein
Length = 243
Score = 169 bits (429), Expect = 5e-42
Identities = 82/226 (36%), Positives = 137/226 (60%), Gaps = 4/226 (1%)
Query: 720 VDLVGPFPTA--RAQMKFILVAVDYFTKWIEAEPLAKITSAKIVNFYWKRIVCRFGIPRA 777
+D+VGP + + ++K +LV DY TKWIEA+ ++T ++ +F W+ IVC+ GIP
Sbjct: 1 MDVVGPMEASGGKKKLKNLLVLTDYSTKWIEAKAFQQVTEKQVEDFLWENIVCQHGIPYE 60
Query: 778 IVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGA 837
I++DNGT +S + + FC + I++ ++ +PQ NGQ E+AN+ IL +++ L K
Sbjct: 61 IITDNGTNLTSRKIKAFCDKWKIRLTTSTPHYPQGNGQAEAANKAILSNIKKILDSKKSM 120
Query: 838 WLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVE--IDNFTWRTRPGFEEENQANM 895
W D L VLW+Y TT + +T+ETPF + YG++A++P+ I + + N +
Sbjct: 121 WSDVLHGVLWAYRTTPRKSTQETPFSLAYGLEAVIPIVTIIPSVRRTASHANSDMNTQML 180
Query: 896 AVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVREMQVGDLVLK 941
+D + E RD+A IR +Q V +NS +++R +VG+LVL+
Sbjct: 181 RDNMDFIDERRDQAMIRVQNYQQAVTRYYNSNIKIRRFEVGELVLR 226
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 159 bits (403), Expect = 6e-39
Identities = 110/368 (29%), Positives = 179/368 (47%), Gaps = 48/368 (13%)
Query: 618 YTLIDGHLYRRGFSTPLLKCVPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPT 677
YTL +YR CV ++ E I+ H H K+L+AGF+WP+
Sbjct: 1375 YTLCKDKIYRT--------CVSEDEIEGILLHCHGFAYGGHFATFKTMSKILQAGFWWPS 1426
Query: 678 LRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFIL 737
+ KD +F+ +C + F +WG+D +GPFP++ K+IL
Sbjct: 1427 MFKDAQEFISKCDSFE--------------------NFDVWGIDFMGPFPSSYGN-KYIL 1465
Query: 738 VAVDYFTKWIEAEPLAKITS-AKIVNFYWKRIVC-RFGIPRAIVSDNGTQFSSSQTREFC 795
VA+DY +KW+EA +A T+ A++V +K I+ RFG+PR ++SD G F +
Sbjct: 1466 VAIDYVSKWVEA--IASHTNDARVVLKLFKTIIFPRFGVPRIVISDGGKHFINKGFENLL 1523
Query: 796 KEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQS 855
K+ G++ + T+GQVE +NR I L + + + W +L LW+Y T ++
Sbjct: 1524 KKHGVKHK--------TSGQVEISNREIKAILEKTVGSTRKDWSAKLNDTLWAYRTAFKT 1575
Query: 856 TTRETPFRMTYGVDAMLPVEID-NFTWRTR-PGFE-EENQANMAVELDLLSETRDEAHIR 912
TPF + YG LPVE++ W + F+ + + ++L+ L++ R EA+
Sbjct: 1576 PIGTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRLIQLNDLNKIRLEAYES 1635
Query: 913 ETAMKQRVAAKFNSRVRVREMQVGDLVLKWRSGA---PGNKLTPNWEGPYRIIKVLGNGA 969
K+R + + ++ R+ +VGD VL + S PG KL W GP+ + V GA
Sbjct: 1636 SKIYKERTKSFHDKKIVSRDFKVGDQVLLFNSRLRLFPG-KLKSRWSGPFSVTAVRPYGA 1694
Query: 970 YHLEELDG 977
L +G
Sbjct: 1695 ITLAGKNG 1702
>At4g07850 putative polyprotein
Length = 1138
Score = 119 bits (299), Expect = 6e-27
Identities = 92/339 (27%), Positives = 151/339 (44%), Gaps = 35/339 (10%)
Query: 569 TLAYPSIEGELMACVSRGRTWMD---PIISILEGDPAEVEQCTKEQQREASHYTLIDGH- 624
TL YP+ + EL A V +TW P ++ D ++ +Q+ H ++
Sbjct: 658 TLNYPTYDKELYALVRALQTWQHYLWPKEFVIHTDHESLKHLKGQQKLNKRHARWVEFIE 717
Query: 625 ------LYRRGFSTPLLK----------------CVPPEKY-EAIMSEVHEGVCASHIGG 661
Y++G + C+P E + E H G H G
Sbjct: 718 TFPYVIKYKKGKDNVVADALSRRHEGFLFYDNRLCIPNSSLRELFIREAHGGGLMGHFGV 777
Query: 662 RSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFADLSKAPPKELVTMSAPWPFAMWG-- 719
S KV++ F+WP +++D ++C C+ +K+ P L T P P W
Sbjct: 778 -SKTLKVMQDHFHWPHMKRDVERMCERCTTCKQAK--AKSQPHGLCT-PLPIPLHPWNDI 833
Query: 720 -VDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITSAK-IVNFYWKRIVCRFGIPRA 777
+D V P R I V VD F+K P K A I N +++ +V G+P+
Sbjct: 834 SMDFVVGLPRTRTGKDSIFVVVDRFSKMAHFIPCHKTDDAMHIANLFFREVVRLHGMPKT 893
Query: 778 IVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGA 837
IVSD T+F S + ++G ++ F++ HPQT+GQ E NR + LR + +
Sbjct: 894 IVSDRDTKFLSYFWKTLWSKLGTKLLFSTTCHPQTDGQTEVVNRTLSTLLRALIKKNLKT 953
Query: 838 WLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEI 876
W D LP V ++YN + S T+ +PF++ YG + + P+++
Sbjct: 954 WEDCLPHVEFAYNHSVHSATKFSPFQIVYGFNPITPLDL 992
>At3g31970 hypothetical protein
Length = 1329
Score = 117 bits (292), Expect = 4e-26
Identities = 102/357 (28%), Positives = 165/357 (45%), Gaps = 28/357 (7%)
Query: 637 CVPPEKY--EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 694
CVP ++ I+SE H + + H + + L+ + W +++D ++V +C CQ+
Sbjct: 895 CVPKDEELRREILSEAHASMFSIHPRATKMY-RDLKRYYQWVGMKRDVANWVTECDVCQL 953
Query: 695 FADLSKAPPKELVTMSA-PWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLA 753
+ P L ++ W + +D V P +R + I V VD TK +
Sbjct: 954 VKAEHQVPGGLLQSLPILEWKWDFITMDFVVGLPVSRTK-DAIWVIVDRLTKSAHFLAIR 1012
Query: 754 KITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQT 812
K A ++ Y IV G+P +IVSD ++F+S+ R F EMG +++ ++ HPQT
Sbjct: 1013 KTDGAVLLAKKYVSEIVELHGVPVSIVSDRDSKFTSAFWRAFQGEMGTKVQMSTAYHPQT 1072
Query: 813 NGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAML 872
+GQ E + + LR + + G W D L V ++YN + Q++ R PF YG
Sbjct: 1073 DGQSERTIQTLEDMLRMCVLDRGGHWADHLSLVEFAYNNSYQASIRMAPFEALYGRPCRT 1132
Query: 873 PVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQ---RVAAKFNSRVR 929
P+ W T+ G A D + ET + + + MK+ R + + R R
Sbjct: 1133 PL-----CW-TQVGERSIYGA------DYVLETTERIRVLKLNMKEAQDRQRSYADKRRR 1180
Query: 930 VREMQVGDLVLKWRSGAPG-------NKLTPNWEGPYRIIKVLGNGAYHLEELDGRR 979
E +VGD V + G KLTP + GP+RI++ +G AY LE D R
Sbjct: 1181 ELEFEVGDRVYLKMAMLRGPNRSISETKLTPRYMGPFRIVERVGPVAYRLELPDVMR 1237
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 115 bits (288), Expect = 1e-25
Identities = 108/414 (26%), Positives = 185/414 (44%), Gaps = 29/414 (7%)
Query: 572 YPSIEGELMACVSRGRTWMDPIISILEGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFS 631
YP+ + E+ A V + W +I + + E++ T + E + T +G + G
Sbjct: 486 YPTHDLEMAAVVFALKIWRSYLIRLAQERDEEIKGWTLNNKTE--YQTSNNGTIVVNG-- 541
Query: 632 TPLLKCVPPEKY--EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQC 689
CVP ++ E I+ E H+ + H G + + L+ ++W +RKD +V +C
Sbjct: 542 ---RVCVPNDRALKEEILREAHQSKFSIHPGSNKMY-RDLKRYYHWVGMRKDVARWVAKC 597
Query: 690 KECQVFADLSKAPPKELVTMS-APWPFAMWGVDLVGPFPTA-RAQMKFILVAVDYFTKWI 747
CQ+ + P L + + W + +D V PT +++ + V VD TK
Sbjct: 598 PTCQLVKAEHQVPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSA 657
Query: 748 EAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFAS 806
++ A+I+ Y I+ GIP +IVSD T+F+S F K +G ++ ++
Sbjct: 658 HFMAISDKDGAEIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLST 717
Query: 807 VEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTY 866
HPQT+GQ E + + LR + + G W L + ++YN + Q++ +P+ Y
Sbjct: 718 AYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALY 777
Query: 867 GVDAMLPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNS 926
G P+ W T G E + +D +E I+ + R + N
Sbjct: 778 GRACRTPL-----CW-TPVG---ERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANK 828
Query: 927 RVRVREMQVGDLVLKWRSGAPG-------NKLTPNWEGPYRIIKVLGNGAYHLE 973
R + E QVGDLV G KL+P + GPY++I+ +G AY L+
Sbjct: 829 RRKELEFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLD 882
>At2g10780 pseudogene
Length = 1611
Score = 109 bits (273), Expect = 7e-24
Identities = 96/349 (27%), Positives = 159/349 (45%), Gaps = 22/349 (6%)
Query: 637 CVPPEKY--EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 694
CVP ++ E I+ E H+ + H G + + L+ ++W ++KD +V +C CQ+
Sbjct: 1152 CVPNDRALKEEILREAHQSKFSIHPGSNKMY-RDLKRYYHWVGMKKDVARWVAKCPTCQL 1210
Query: 695 FADLSKAPPKELVTMSAP-WPFAMWGVDLVGPFPTA-RAQMKFILVAVDYFTKWIEAEPL 752
+ P L + P W + +D V PT +++ + V VD TK +
Sbjct: 1211 VKAEHQVPSGLLQNLPIPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAI 1270
Query: 753 AKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQ 811
+ A+I+ Y IV GIP +IVSD T+F+S + F K +G ++ ++ HPQ
Sbjct: 1271 SDKDGAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQ 1330
Query: 812 TNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAM 871
T+ Q E + + LR + + G W L V ++YN + Q++ +P+ YG
Sbjct: 1331 TDEQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACR 1390
Query: 872 LPVEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAKFNSRVRVR 931
P+ W T G E + +D +E I+ + R + N R +
Sbjct: 1391 TPL-----CW-TPVG---ERRLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKEL 1441
Query: 932 EMQVGDLVLKWRSGAPG-------NKLTPNWEGPYRIIKVLGNGAYHLE 973
E QVGDLV G KL+P + GPY++I+ +G AY L+
Sbjct: 1442 EFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLD 1490
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 103 bits (258), Expect = 4e-22
Identities = 97/368 (26%), Positives = 158/368 (42%), Gaps = 58/368 (15%)
Query: 637 CVPPEKY--EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQV 694
CVP ++ I+SE H + + H G + + L+ + W +++D ++V +C CQ
Sbjct: 1008 CVPKDEELRREILSEAHASMFSIHPGATKMY-RDLKRYYQWVGMKRDVANWVAECDVCQ- 1065
Query: 695 FADLSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAK 754
LV P A+W V +D TK + K
Sbjct: 1066 -----------LVKAEHQVPDAIW-------------------VIMDRLTKSAHFLAIRK 1095
Query: 755 ITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTN 813
A ++ Y IV G+P +IVSD ++F+ + R F +MG +++ ++ HPQT+
Sbjct: 1096 TDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTFAFWRAFQAKMGTKVQMSTAYHPQTD 1155
Query: 814 GQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLP 873
GQ E + + LR + + G W D L V ++YN + Q++ PF YG P
Sbjct: 1156 GQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYQASIGMAPFEALYGRPCWTP 1215
Query: 874 VEIDNFTWRTRPGFEEENQANMAVELDLLSETRDEAHIRETAMKQ---RVAAKFNSRVRV 930
+ R+ G D + ET + + + MK+ R + + R R
Sbjct: 1216 LRWTQVEERSIYG------------ADYVQETTERIRVLKLNMKEAQARQRSYADKRRRE 1263
Query: 931 REMQVGDLVLKWRSGAPG-------NKLTPNWEGPYRIIKVLGNGAYHLEELD-GRRLPR 982
E +VGD V + G KL+P + GP+RI++ +G AY LE D R +
Sbjct: 1264 LEFEVGDRVYLKMAMLRGPNRSILETKLSPRYMGPFRIVERVGPVAYRLELPDVMRAFHK 1323
Query: 983 SFNGLSLR 990
F+ L LR
Sbjct: 1324 VFHVLMLR 1331
>At4g07610
Length = 276
Score = 103 bits (256), Expect = 6e-22
Identities = 56/165 (33%), Positives = 88/165 (52%), Gaps = 3/165 (1%)
Query: 638 VPPEKYEAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQVFAD 697
V ++ E I+ H + K+L+AGF+WPT+ K+ +FV +C C +
Sbjct: 76 VSEDEVEGILLHCHCSTYGGNFTTFKTVSKILQAGFWWPTMFKEAQEFVLKCDSCHRKGN 135
Query: 698 LSKAPPKELVTMSAPWPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPLAKITS 757
+SK + F +WG+D +GPFP++ K+ILVAVDY +KWIEA +
Sbjct: 136 ISKRNEMPQNPILEVELFDVWGIDFIGPFPSSYGN-KYILVAVDYVSKWIEA-TASPTND 193
Query: 758 AKIVNFYWKRIV-CRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQ 801
AK+V +K I+ RFG+PR ++SD G F + K+ G++
Sbjct: 194 AKVVLKLFKTIIFSRFGVPRVVISDGGKHFINKVFENLLKKHGVK 238
>At4g07830 putative reverse transcriptase
Length = 611
Score = 102 bits (254), Expect = 1e-21
Identities = 96/356 (26%), Positives = 165/356 (45%), Gaps = 22/356 (6%)
Query: 523 RVRYLMTLFQEVVVE--YVPRAENQRADALAKLASTRKPGNNKSVIQETLAYPSIEGELM 580
R+R M L + +E Y P N DAL++ PG + ETL G L
Sbjct: 50 RLRRWMKLVADYDLEIAYHPGKANVVTDALSRKRVGAAPGQSV----ETLVIEI--GALR 103
Query: 581 AC-VSRGRTWMDPIISILEGDPAEVEQCTKEQQREASHYTLIDGHLYRRGFSTPLLK--- 636
C V+R ++ ++ + D Q +E+ + +G Y+ + +L
Sbjct: 104 LCAVAREPLGLE---AVDQTDLLSRVQLAQEKDEGLIAASKAEGFEYQFAANGTILVHGR 160
Query: 637 -CVPPEKY--EAIMSEVHEGVCASHIGGRSLACKVLRAGFYWPTLRKDCMDFVKQCKECQ 693
CVP +K + I+SE H + + H G + + L+ + W +++D ++V++C CQ
Sbjct: 161 VCVPKDKELRQEILSEAHASMFSIHPGATKMY-RDLKRYYQWVGMKRDVGNWVEECDVCQ 219
Query: 694 VFADLSKAPPKELVTMSAP-WPFAMWGVDLVGPFPTARAQMKFILVAVDYFTKWIEAEPL 752
+ + L ++ P W + +D V P +R + I V VD TK +
Sbjct: 220 LVKIEHQVSGSLLQSLPIPEWKWDFITMDFVVGLPVSRTK-DAIWVIVDRLTKSAHFLAI 278
Query: 753 AKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQ 811
K A ++ Y IV G+P +IVSD ++F+S+ R F EMG +++ ++ HPQ
Sbjct: 279 RKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMSTAYHPQ 338
Query: 812 TNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYG 867
T+GQ E + + LR + + +G W D L V ++YN + Q++ PF + YG
Sbjct: 339 TDGQSERTIQTLEDMLRMCVLDWRGHWADHLSLVEFAYNNSYQASIGMAPFEVLYG 394
>At1g35370 hypothetical protein
Length = 1447
Score = 102 bits (253), Expect = 1e-21
Identities = 91/347 (26%), Positives = 158/347 (45%), Gaps = 30/347 (8%)
Query: 636 KCVPPEKYEAIMSEVHEGVCASHIGGRS---LACKVLRAGFYWPTLRKDCMDFVKQCKEC 692
K V P E I +++ + + S +GGRS + + +++ FYW + KD F++ C C
Sbjct: 1042 KIVVPNDVE-ITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFIRSCGTC 1100
Query: 693 QVFADLSKAPPKELVTMSAPWPFAMW---GVDLVGPFPTARAQMKFILVAVDYFTKWIEA 749
Q + A P L + P P +W +D + P + + I+V VD +K
Sbjct: 1101 QQCKSDNAAYPGLLQPL--PIPDKIWCDVSMDFIEGLPNSGGK-SVIMVVVDRLSKAAHF 1157
Query: 750 EPLAKITSA-KIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVE 808
LA SA + + + G P +IVSD F+S +EF K G+++R +S
Sbjct: 1158 VALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKLQGVELRMSSAY 1217
Query: 809 HPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGV 868
HPQ++GQ E NR + LR W LP + YNT S+++ TPF + YG
Sbjct: 1218 HPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSSQMTPFELVYG- 1276
Query: 869 DAMLPVEIDNFTWRTR-----PGFEEENQANMAVELDLLSETRDEAHIRETAMKQRVAAK 923
P+ + +++ +E + ++ L+ R + +++ A + R
Sbjct: 1277 -QAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLM---RAQHRMKQFADQHRTERT 1332
Query: 924 FN----SRVRVREMQVGDLVLKWRSGAPGNKLTPNWEGPYRIIKVLG 966
F+ V+++ + +VL+ KL+P + GPY+II+ G
Sbjct: 1333 FDIGDFVYVKLQPYRQQSVVLR-----VNQKLSPKYFGPYKIIEKCG 1374
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.336 0.143 0.468
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,295,113
Number of Sequences: 26719
Number of extensions: 801661
Number of successful extensions: 3044
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 85
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 2802
Number of HSP's gapped (non-prelim): 195
length of query: 993
length of database: 11,318,596
effective HSP length: 109
effective length of query: 884
effective length of database: 8,406,225
effective search space: 7431102900
effective search space used: 7431102900
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0590c.10


