

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590a.3
(270 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
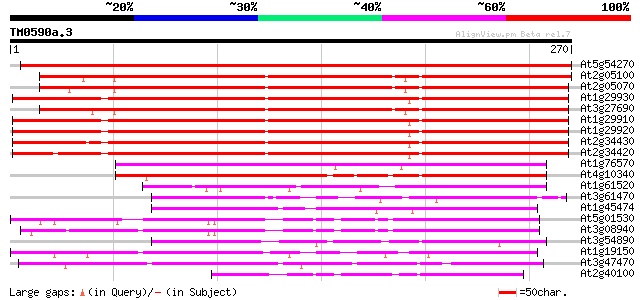
Score E
Sequences producing significant alignments: (bits) Value
At5g54270 Lhcb3 chlorophyll a/b binding protein (gb|AAD28773.1) 497 e-141
At2g05100 putative chlorophyll a/b binding protein 361 e-100
At2g05070 putative chlorophyll a/b binding protein 359 e-99
At1g29930 unknown protein 357 4e-99
At3g27690 putative chlorophyll A-B binding protein 356 8e-99
At1g29910 chlorophyll a/b-binding protein 355 1e-98
At1g29920 chlorophyll a/b-binding protein 355 1e-98
At2g34430 putative photosystem II type I chlorophyll a/b binding... 352 9e-98
At2g34420 photosystem II type I chlorophyll a/b binding protein 350 3e-97
At1g76570 putative chlorophyll A-B binding protein 194 5e-50
At4g10340 chlorophyll a/b-binding protein - like 191 5e-49
At1g61520 PSI type III chlorophyll a/b-binding protein 143 1e-34
At3g61470 Lhca2 protein 126 1e-29
At1g45474 light-harvesting complex protein 124 7e-29
At5g01530 chlorophyll a/b-binding protein CP29 120 1e-27
At3g08940 Lhcb4-protein (CP29, Lhcb4.2) 120 1e-27
At3g54890 chlorophyll a/b-binding protein 119 2e-27
At1g19150 unknown protein 116 1e-26
At3g47470 light-harvesting chlorophyll a/b-binding protein (Cab4) 111 4e-25
At2g40100 putative chlorophyll a/b binding protein 93 2e-19
>At5g54270 Lhcb3 chlorophyll a/b binding protein (gb|AAD28773.1)
Length = 265
Score = 497 bits (1280), Expect = e-141
Identities = 237/265 (89%), Positives = 250/265 (93%)
Query: 6 MAATTASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYLGPFS 65
MA+T SS +V+ P+ FLGQTK S+ N LRDVVS+G+ KYTMGNDLWYGPDRVKYLGPFS
Sbjct: 1 MASTFTSSSSVLTPTTFLGQTKASSFNPLRDVVSLGSPKYTMGNDLWYGPDRVKYLGPFS 60
Query: 66 AQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWV 125
QTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGA GCITPEVL+KWV
Sbjct: 61 VQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGAFGCITPEVLQKWV 120
Query: 126 RVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPG 185
RVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQV+LMGLVEGFRINGL G
Sbjct: 121 RVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVILMGLVEGFRINGLDG 180
Query: 186 VGEGNNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLE 245
VGEGN+LYPGGQYFDPLGLADDP TFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLE
Sbjct: 181 VGEGNDLYPGGQYFDPLGLADDPVTFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLE 240
Query: 246 NLLDHLDDPVANNAWVYATKFVPGS 270
NLLDHLD+PVANNAW +ATKF PG+
Sbjct: 241 NLLDHLDNPVANNAWAFATKFAPGA 265
>At2g05100 putative chlorophyll a/b binding protein
Length = 265
Score = 361 bits (926), Expect = e-100
Identities = 188/265 (70%), Positives = 206/265 (76%), Gaps = 12/265 (4%)
Query: 15 TVIKPSPFLGQTKGSNSNTL-RDVVSMGTGKYTMGN-------DLWYGPDRVKYLGPFSA 66
+ I+ S F GQT SN L R V G G+ TM +WYGPDR KYLGPFS
Sbjct: 4 SAIQQSSFAGQTALKPSNELLRKVGVSGGGRVTMRRTVKSTPQSIWYGPDRPKYLGPFSE 63
Query: 67 QTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWVR 126
TPSYLTGE+PGDYGWDTAGLSADPE FAKNR LEVIH RWAMLGALGC PE+L K
Sbjct: 64 NTPSYLTGEYPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCTFPEILSK-NG 122
Query: 127 VDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPGV 186
V F E VWFKAGSQIFSEGGLDYLGNPNL+HAQSILA+ QV+LMG +EG+RI G P +
Sbjct: 123 VKFGEAVWFKAGSQIFSEGGLDYLGNPNLIHAQSILAIWAVQVVLMGFIEGYRIGGGP-L 181
Query: 187 GEG-NNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLE 245
GEG + LYPGG FDPL LA+DP F+ELKVKE+KNGRLAMFSMFGFFVQAIVTGKGP+E
Sbjct: 182 GEGLDPLYPGGA-FDPLNLAEDPEAFSELKVKELKNGRLAMFSMFGFFVQAIVTGKGPIE 240
Query: 246 NLLDHLDDPVANNAWVYATKFVPGS 270
NL DHL DPVANNAW YAT FVPG+
Sbjct: 241 NLFDHLADPVANNAWSYATNFVPGN 265
>At2g05070 putative chlorophyll a/b binding protein
Length = 265
Score = 359 bits (921), Expect = e-99
Identities = 185/264 (70%), Positives = 207/264 (78%), Gaps = 12/264 (4%)
Query: 15 TVIKPSPFLGQTK-GSNSNTLRDVVSMGTGKYTMGN-------DLWYGPDRVKYLGPFSA 66
+ I+ S F GQT +S+ ++ V +G G+ TM +WYGPDR KYLGPFS
Sbjct: 4 SAIQQSSFAGQTALKPSSDLIQKVGVLGGGRVTMRRTVKSTPQSIWYGPDRPKYLGPFSE 63
Query: 67 QTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWVR 126
TPSYLTGE+PGDYGWDTAGLSADPE FAKNR LEVIH RWAMLGALGC PE+L K
Sbjct: 64 NTPSYLTGEYPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCTFPEILSK-NG 122
Query: 127 VDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPGV 186
V F E VWFKAGSQIFSEGGLDYLGNPNL+HAQSILA+ QV+LMG +EG+RI G P +
Sbjct: 123 VKFGEAVWFKAGSQIFSEGGLDYLGNPNLIHAQSILAIWAVQVVLMGFIEGYRIGGGP-L 181
Query: 187 GEG-NNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLE 245
GEG + LYPGG FDPL LA+DP F+ELKVKE+KNGRLAMFSMFGFFVQAIVTGKGP+E
Sbjct: 182 GEGLDPLYPGGA-FDPLNLAEDPEAFSELKVKELKNGRLAMFSMFGFFVQAIVTGKGPIE 240
Query: 246 NLLDHLDDPVANNAWVYATKFVPG 269
NL DHL DPVANNAW YAT FVPG
Sbjct: 241 NLFDHLADPVANNAWSYATNFVPG 264
>At1g29930 unknown protein
Length = 267
Score = 357 bits (916), Expect = 4e-99
Identities = 181/269 (67%), Positives = 207/269 (76%), Gaps = 6/269 (2%)
Query: 2 AATLMAATTASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYL 61
A+T+ ++ A +G +K SP + GS T+R V+ G WYG DRVKYL
Sbjct: 3 ASTMALSSPAFAGKAVKLSPAASEVLGSGRVTMRKTVAKPKGP---SGSPWYGSDRVKYL 59
Query: 62 GPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVL 121
GPFS ++PSYLTGEFPGDYGWDTAGLSADPE FA+NR LEVIH RWAMLGALGC+ PE+L
Sbjct: 60 GPFSGESPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 119
Query: 122 EKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRIN 181
+ V F E VWFKAGSQIFS+GGLDYLGNP+LVHAQSILA+ QV+LMG VEG+R+
Sbjct: 120 AR-NGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVA 178
Query: 182 GLPGVGEGNN-LYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTG 240
G +GE + LYPGG FDPLGLA DP FAELKVKE+KNGRLAMFSMFGFFVQAIVTG
Sbjct: 179 GNGPLGEAEDLLYPGGS-FDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTG 237
Query: 241 KGPLENLLDHLDDPVANNAWVYATKFVPG 269
KGP+ENL DHL DPV NNAW +AT FVPG
Sbjct: 238 KGPIENLADHLADPVNNNAWAFATNFVPG 266
>At3g27690 putative chlorophyll A-B binding protein
Length = 266
Score = 356 bits (913), Expect = 8e-99
Identities = 185/265 (69%), Positives = 204/265 (76%), Gaps = 13/265 (4%)
Query: 15 TVIKPSPFLGQTKGSNSNTLRDVV--SMGTGKYTMGN-------DLWYGPDRVKYLGPFS 65
+ I+ S F GQT SN L + S G G+ M +WYGPDR KYLGPFS
Sbjct: 4 SAIQHSSFAGQTTLKPSNDLLRKIGASNGGGRIIMRRTVKSTPQSIWYGPDRPKYLGPFS 63
Query: 66 AQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWV 125
TPSYLTGE+PGDYGWDTAGLSADPE FAKNR LEVIH RWAMLGALGC PE+L K
Sbjct: 64 ENTPSYLTGEYPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCTFPEILSK-N 122
Query: 126 RVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPG 185
V F E VWFKAGSQIFSEGGLDYLGNPNL+HAQSILA+ QV+LMG +EG+RI G P
Sbjct: 123 GVKFGEAVWFKAGSQIFSEGGLDYLGNPNLIHAQSILAIWACQVVLMGFIEGYRIGGGP- 181
Query: 186 VGEG-NNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPL 244
+GEG + LYPGG FDPL LA+DP F+ELKVKE+KNGRLAMFSMFGFFVQAIVTGKGP+
Sbjct: 182 LGEGLDPLYPGGA-FDPLNLAEDPEAFSELKVKELKNGRLAMFSMFGFFVQAIVTGKGPI 240
Query: 245 ENLLDHLDDPVANNAWVYATKFVPG 269
ENL DH+ DPVANNAW YAT FVPG
Sbjct: 241 ENLFDHIADPVANNAWAYATNFVPG 265
>At1g29910 chlorophyll a/b-binding protein
Length = 267
Score = 355 bits (911), Expect = 1e-98
Identities = 180/269 (66%), Positives = 206/269 (75%), Gaps = 6/269 (2%)
Query: 2 AATLMAATTASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYL 61
A+T+ ++ A +G + SP + GS T+R V+ G WYG DRVKYL
Sbjct: 3 ASTMALSSPAFAGKAVNLSPAASEVLGSGRVTMRKTVAKPKGP---SGSPWYGSDRVKYL 59
Query: 62 GPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVL 121
GPFS ++PSYLTGEFPGDYGWDTAGLSADPE FA+NR LEVIH RWAMLGALGC+ PE+L
Sbjct: 60 GPFSGESPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 119
Query: 122 EKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRIN 181
+ V F E VWFKAGSQIFS+GGLDYLGNP+LVHAQSILA+ QV+LMG VEG+R+
Sbjct: 120 AR-NGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVA 178
Query: 182 GLPGVGEGNN-LYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTG 240
G +GE + LYPGG FDPLGLA DP FAELKVKE+KNGRLAMFSMFGFFVQAIVTG
Sbjct: 179 GNGPLGEAEDLLYPGGS-FDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTG 237
Query: 241 KGPLENLLDHLDDPVANNAWVYATKFVPG 269
KGP+ENL DHL DPV NNAW +AT FVPG
Sbjct: 238 KGPIENLADHLADPVNNNAWAFATNFVPG 266
>At1g29920 chlorophyll a/b-binding protein
Length = 267
Score = 355 bits (911), Expect = 1e-98
Identities = 180/269 (66%), Positives = 206/269 (75%), Gaps = 6/269 (2%)
Query: 2 AATLMAATTASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYL 61
A+T+ ++ A +G + SP + GS T+R V+ G WYG DRVKYL
Sbjct: 3 ASTMALSSPAFAGKAVNLSPAASEVLGSGRVTMRKTVAKPKGP---SGSPWYGSDRVKYL 59
Query: 62 GPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVL 121
GPFS ++PSYLTGEFPGDYGWDTAGLSADPE FA+NR LEVIH RWAMLGALGC+ PE+L
Sbjct: 60 GPFSGESPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 119
Query: 122 EKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRIN 181
+ V F E VWFKAGSQIFS+GGLDYLGNP+LVHAQSILA+ QV+LMG VEG+R+
Sbjct: 120 AR-NGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVA 178
Query: 182 GLPGVGEGNN-LYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTG 240
G +GE + LYPGG FDPLGLA DP FAELKVKE+KNGRLAMFSMFGFFVQAIVTG
Sbjct: 179 GNGPLGEAEDLLYPGGS-FDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTG 237
Query: 241 KGPLENLLDHLDDPVANNAWVYATKFVPG 269
KGP+ENL DHL DPV NNAW +AT FVPG
Sbjct: 238 KGPIENLADHLADPVNNNAWAFATNFVPG 266
>At2g34430 putative photosystem II type I chlorophyll a/b binding
protein.
Length = 266
Score = 352 bits (904), Expect = 9e-98
Identities = 182/269 (67%), Positives = 206/269 (75%), Gaps = 7/269 (2%)
Query: 2 AATLMAATTASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYL 61
A+T+ ++ A +G +K SP + G+ T+R S TG WYG DRVKYL
Sbjct: 3 ASTMALSSPALTGKAVKLSPAASEVFGTGRITMRKA-SKPTGP---SGSPWYGSDRVKYL 58
Query: 62 GPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVL 121
GPFS + PSYLTGEFPGDYGWDTAGLSADPE FA+NR LEVIH RWAMLGALGC+ PE+L
Sbjct: 59 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 118
Query: 122 EKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRIN 181
+ V F E VWFKAGSQIFS+GGLDYLGNP+LVHAQSILA+ QV+LMG VEG+R+
Sbjct: 119 AR-NGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVA 177
Query: 182 GLPGVGEGNN-LYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTG 240
G +GE + LYPGG FDPLGLA DP FAELKVKE+KNGRLAMFSMFGFFVQAIVTG
Sbjct: 178 GDGPLGEAEDLLYPGGS-FDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTG 236
Query: 241 KGPLENLLDHLDDPVANNAWVYATKFVPG 269
KGPLENL DHL DPV NNAW +AT FVPG
Sbjct: 237 KGPLENLADHLADPVNNNAWAFATNFVPG 265
>At2g34420 photosystem II type I chlorophyll a/b binding protein
Length = 265
Score = 350 bits (899), Expect = 3e-97
Identities = 180/269 (66%), Positives = 205/269 (75%), Gaps = 8/269 (2%)
Query: 2 AATLMAATTASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYL 61
++T+ ++ A +G +KP+ GS T+R V+ G WYG DRVKYL
Sbjct: 3 SSTMALSSPAFAGKAVKPAA--SDVLGSGRVTMRKTVAKPKGP---SGSPWYGSDRVKYL 57
Query: 62 GPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVL 121
GPFS + PSYLTGEFPGDYGWDTAGLSADPE FA+NR LEVIH RWAMLGALGC+ PE+L
Sbjct: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 117
Query: 122 EKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRIN 181
+ V F E VWFKAGSQIFS+GGLDYLGNP+LVHAQSILA+ QV+LMG VEG+R+
Sbjct: 118 AR-NGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVA 176
Query: 182 GLPGVGEGNN-LYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTG 240
G +GE + LYPGG FDPLGLA DP FAELKVKE+KNGRLAMFSMFGFFVQAIVTG
Sbjct: 177 GDGPLGEAEDLLYPGGS-FDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTG 235
Query: 241 KGPLENLLDHLDDPVANNAWVYATKFVPG 269
KGPLENL DHL DPV NNAW +AT FVPG
Sbjct: 236 KGPLENLADHLADPVNNNAWAFATNFVPG 264
>At1g76570 putative chlorophyll A-B binding protein
Length = 327
Score = 194 bits (492), Expect = 5e-50
Identities = 101/215 (46%), Positives = 129/215 (59%), Gaps = 8/215 (3%)
Query: 52 WYGPDRVKYLGPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLG 111
WYG +R ++ GP P YLTGE PGDYG+D AGL D F K E++H RWAML
Sbjct: 106 WYGKERPRWFGPIPYDYPPYLTGELPGDYGFDIAGLGKDRLTFDKYFNFEILHARWAMLA 165
Query: 112 ALGCITPEVLEKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNL--VHAQSILAVLGFQV 169
ALG + PEV + F EPVW++ G L+YLG P L +Q ++ + QV
Sbjct: 166 ALGALIPEVFDLTGTFHFAEPVWWRVGYSKLQGETLEYLGIPGLHVAGSQGVIVIAICQV 225
Query: 170 LLMGLVEGFRINGLPGVG------EGNNLYPGGQYFDPLGLADDPATFAELKVKEIKNGR 223
LLM E R G+ + G+ YPGG FDPL L++DP F +LKVKEIKNGR
Sbjct: 226 LLMVGPEYARYCGIEALEPLGIYLPGDINYPGGTLFDPLNLSEDPVAFEDLKVKEIKNGR 285
Query: 224 LAMFSMFGFFVQAIVTGKGPLENLLDHLDDPVANN 258
LAM + GF+ QA TGKGP++NL+DH+ DP+ NN
Sbjct: 286 LAMVAWLGFYAQAAFTGKGPVQNLVDHVSDPLHNN 320
>At4g10340 chlorophyll a/b-binding protein - like
Length = 280
Score = 191 bits (484), Expect = 5e-49
Identities = 108/210 (51%), Positives = 132/210 (62%), Gaps = 9/210 (4%)
Query: 52 WYGPDRVKYLGPF---SAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWA 108
WYGPDR +L ++ P YL GE GDYG+D GL PE FAK +A E+IH RWA
Sbjct: 62 WYGPDRRIFLPDGLLDRSEIPEYLNGEVAGDYGYDPFGLGKKPENFAKYQAFELIHARWA 121
Query: 109 MLGALGCITPEVLEKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQ 168
MLGA G I PE L K+ E VWFK G+ + L+Y G + +LAV+ +
Sbjct: 122 MLGAAGFIIPEALNKYGANCGPEAVWFKTGALLLDGNTLNYFGKN--IPINLVLAVVA-E 178
Query: 169 VLLMGLVEGFRINGLPGVGEGNNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFS 228
V+L+G E +RI G+ + L+PGG FDPLGLA DP A LKVKEIKNGRLAMF+
Sbjct: 179 VVLLGGAEYYRITN--GLDFEDKLHPGGP-FDPLGLAKDPEQGALLKVKEIKNGRLAMFA 235
Query: 229 MFGFFVQAIVTGKGPLENLLDHLDDPVANN 258
M GFF+QA VTG+GP+ENL HL DP NN
Sbjct: 236 MLGFFIQAYVTGEGPVENLAKHLSDPFGNN 265
>At1g61520 PSI type III chlorophyll a/b-binding protein
Length = 273
Score = 143 bits (360), Expect = 1e-34
Identities = 90/217 (41%), Positives = 120/217 (54%), Gaps = 33/217 (15%)
Query: 65 SAQTPSYLTGEFPGDYGWDTAGLSADPEA---FAKNRAL---EVIHGRWAMLGALGCITP 118
S+Q+ SYL G PGDYG+D GLS DPE F + R L E+I+GR+AMLGA G I P
Sbjct: 59 SSQSLSYLDGSLPGDYGFDPLGLS-DPEGTGGFIEPRWLAYGEIINGRFAMLGAAGAIAP 117
Query: 119 EVLEKWVRVDFKEPV-WFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGF---------- 167
E+L K + + + WF+ G + G N +A++GF
Sbjct: 118 EILGKAGLIPAETALPWFQTG--VIPPAGTYTYWADNYTLFVLEMALMGFAEHRRLQDWY 175
Query: 168 ------QVLLMGLVEGFRINGLPGVGEGNNLYPGGQYFDPLGLADDPATFAELKVKEIKN 221
+ +GL +G G GN YPGG +F+PLG D + ELK+KE+KN
Sbjct: 176 NPGSMGKQYFLGLEKGL-------AGSGNPAYPGGPFFNPLGFGKDEKSLKELKLKEVKN 228
Query: 222 GRLAMFSMFGFFVQAIVTGKGPLENLLDHLDDPVANN 258
GRLAM ++ G+F+Q +VTG GP +NLLDHL DPV NN
Sbjct: 229 GRLAMLAILGYFIQGLVTGVGPYQNLLDHLADPVNNN 265
>At3g61470 Lhca2 protein
Length = 257
Score = 126 bits (317), Expect = 1e-29
Identities = 83/218 (38%), Positives = 112/218 (51%), Gaps = 41/218 (18%)
Query: 69 PSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWVRVD 128
P +L G PGD+G+D GLS+DP++ N E++H RWAMLGA G PE L K + +
Sbjct: 62 PEWLDGSLPGDFGFDPLGLSSDPDSLKWNVQAEIVHCRWAMLGAAGIFIPEFLTK-IGI- 119
Query: 129 FKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEG----------- 177
P W+ AG Q +Y + + +++L+G EG
Sbjct: 120 LNTPSWYTAGEQ-------EYFTDKTTLFV--------VELILIGWAEGRRWADIIKPGS 164
Query: 178 ------FRINGLPGVGEGNNLYPGGQYFDPLGL-ADDPATFAELKVKEIKNGRLAMFSMF 230
F N L G G YPGG +FDPLG + PA EL+ KEIKNGRLAM ++
Sbjct: 165 VNTDPVFPNNKLTGTDVG---YPGGLWFDPLGWGSGSPAKLKELRTKEIKNGRLAMLAVM 221
Query: 231 GFFVQAIVTGKGPLENLLDHLDDPVANNAWVYATKFVP 268
G + Q I TG GP++NL HL DP +A ++A F P
Sbjct: 222 GAWFQHIYTGTGPIDNLFAHLADP--GHATIFAA-FTP 256
>At1g45474 light-harvesting complex protein
Length = 256
Score = 124 bits (310), Expect = 7e-29
Identities = 78/194 (40%), Positives = 102/194 (52%), Gaps = 15/194 (7%)
Query: 69 PSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWVRVD 128
P YL G GDYG+D GL DPE+ E++H R+AMLG G + ++L +
Sbjct: 56 PPYLDGNLAGDYGFDPLGLGEDPESLKWYVQAELVHSRFAMLGVAGILFTDLLRTTGIRN 115
Query: 129 FKEPVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLV-------EGFRIN 181
PVW++AG+ F D+ L+ Q +L M V EG
Sbjct: 116 L--PVWYEAGAVKF-----DFASTKTLIVVQFLLMGFAETKRYMDFVSPGSQAKEGSFFF 168
Query: 182 GLPGVGEGNNL-YPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTG 240
GL EG YPGG +PLGLA D + K+KEIKNGRLAM +M GFFVQA VT
Sbjct: 169 GLEAALEGLEPGYPGGPLLNPLGLAKDVQNAHDWKLKEIKNGRLAMMAMLGFFVQASVTH 228
Query: 241 KGPLENLLDHLDDP 254
GP++NL++HL +P
Sbjct: 229 TGPIDNLVEHLSNP 242
>At5g01530 chlorophyll a/b-binding protein CP29
Length = 290
Score = 120 bits (300), Expect = 1e-27
Identities = 104/304 (34%), Positives = 132/304 (43%), Gaps = 74/304 (24%)
Query: 1 MAATLMAATTASS--GTVIKPS--PFLGQTKGSNSNTLRDVVSMGTGKYTMGND--LWYG 54
MAAT AA ASS GT + P P G+ + + K T+ D LWY
Sbjct: 1 MAATSAAAAAASSIMGTRVAPGIHPGSGRFTAVFGFGKKKAAPKKSAKKTVTTDRPLWYP 60
Query: 55 PDRVKYLGPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAF-----------AKN------ 97
A +P +L G GDYG+D GL E AKN
Sbjct: 61 ----------GAISPDWLDGSLVGDYGFDPFGLGKPAEYLQFDIDSLDQNLAKNLAGDVI 110
Query: 98 --------------------------RALEVIHGRWAMLGALGCITPEVLEKWVRVDFKE 131
R E+IHGRWAML LG ++ E L
Sbjct: 111 GTRTEAADAKSTPFQPYSEVFGIQRFRECELIHGRWAMLATLGALSVEWLTG-------- 162
Query: 132 PVWFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPGVGEGNN 191
W AG +G YLG P SI ++ +VL++G +E F+ N +
Sbjct: 163 VTWQDAGKVELVDGS-SYLGQPLPF---SISTLIWIEVLVIGYIE-FQRNA--ELDSEKR 215
Query: 192 LYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLENLLDHL 251
LYPGG++FDPLGLA DP A+L++ EIK+ RLAM + GF VQA TGKGPL N HL
Sbjct: 216 LYPGGKFFDPLGLAADPEKTAQLQLAEIKHARLAMVAFLGFAVQAAATGKGPLNNWATHL 275
Query: 252 DDPV 255
DP+
Sbjct: 276 SDPL 279
>At3g08940 Lhcb4-protein (CP29, Lhcb4.2)
Length = 287
Score = 120 bits (300), Expect = 1e-27
Identities = 102/294 (34%), Positives = 130/294 (43%), Gaps = 62/294 (21%)
Query: 6 MAAT-TASSGTVIKPSPFLGQTKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYLGPF 64
MAAT TA++ + I + + SNS+ GT K + DR +
Sbjct: 1 MAATSTAAAASSIMGTRVVSDIS-SNSSRFTARFGFGTKKASPKKAKTVISDRPLWFP-- 57
Query: 65 SAQTPSYLTGEFPGDYGWDTAGLSADPEAF-----------AKN---------------- 97
A++P YL G GDYG+D GL E AKN
Sbjct: 58 GAKSPEYLDGSLVGDYGFDPFGLGKPAEYLQFDLDSLDQNLAKNLYGEVIGTRTEAVDPK 117
Query: 98 ----------------RALEVIHGRWAMLGALGCITPEVLEKWVRVDFKEPVWFKAGSQI 141
R E+IHGRWAML LG IT E L W AG
Sbjct: 118 STPFQPYSEVFGLQRFRECELIHGRWAMLATLGAITVEWLTG--------VTWQDAGKVE 169
Query: 142 FSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPGVGEGNNLYPGGQYFDP 201
+G YLG P SI ++ +VL++G +E F+ N + LYPGG++FDP
Sbjct: 170 LVDGS-SYLGQPLPF---SISTLIWIEVLVIGYIE-FQRNA--ELDSEKRLYPGGKFFDP 222
Query: 202 LGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLENLLDHLDDPV 255
LGLA DP A+L++ EIK+ RLAM GF VQA TGKGPL N HL DP+
Sbjct: 223 LGLASDPVKKAQLQLAEIKHARLAMVGFLGFAVQAAATGKGPLNNWATHLSDPL 276
>At3g54890 chlorophyll a/b-binding protein
Length = 241
Score = 119 bits (298), Expect = 2e-27
Identities = 81/192 (42%), Positives = 97/192 (50%), Gaps = 16/192 (8%)
Query: 69 PSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEKWVRVD 128
P+YL G PGD+G+D GL P + + E+IH RWAML G + PE L
Sbjct: 55 PAYLDGSAPGDFGFDPLGLGEVPANLERYKESELIHCRWAMLAVPGILVPEAL------- 107
Query: 129 FKEPVWFKAGSQIFSEGG-LDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRINGLPGVG 187
W KA GG YLGNP V ++ +L + L + VE R
Sbjct: 108 -GYGNWVKAQEWAALPGGQATYLGNP--VPWGTLPTILAIEFLAIAFVEHQR---SMEKD 161
Query: 188 EGNNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFV-QAIVTGKGPLEN 246
YPGG FDPLG + DP ELKVKEIKNGRLA+ + GF V Q+ G GPLEN
Sbjct: 162 PEKKKYPGGA-FDPLGYSKDPKKLEELKVKEIKNGRLALLAFVGFCVQQSAYPGTGPLEN 220
Query: 247 LLDHLDDPVANN 258
L HL DP NN
Sbjct: 221 LATHLADPWHNN 232
>At1g19150 unknown protein
Length = 270
Score = 116 bits (290), Expect = 1e-26
Identities = 91/279 (32%), Positives = 126/279 (44%), Gaps = 50/279 (17%)
Query: 1 MAATLMAATTASSGTVIKPS---PFLGQTKGSNSNTLRD-VVSMGTGKYTMGNDLWYGPD 56
+A+ L + T S+ V P+ P + T + +R+ +V + GK PD
Sbjct: 5 IASALTSTLTLSTSRVQNPTQRRPHVASTSSTGGRLMRERLVVVRAGKEVSSVCEPLPPD 64
Query: 57 RVKYLGPFSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCI 116
R + + P +L G PGD+G+D GL +DP+ E+IH RWAML G I
Sbjct: 65 RPLWFP--GSSPPEWLDGSLPGDFGFDPLGLGSDPDTLKWFAQAELIHSRWAMLAVTGII 122
Query: 117 TPEVLEKWVRVDFKEPV-WFKAGSQIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLV 175
PE LE R+ F E W+ AGS+ +Y + + Q++LMG
Sbjct: 123 IPECLE---RLGFIENFSWYDAGSR-------EYFADSTTLFVA--------QMVLMGWA 164
Query: 176 EGFR-------------------INGLPGVGEGNNLYPGGQYFD-PLGLADDPATFAELK 215
EG R +N P VG YPGG +FD + P L+
Sbjct: 165 EGRRWADLIKPGSVDIEPKYPHKVNPKPDVG-----YPGGLWFDFMMWGRGSPEPVMVLR 219
Query: 216 VKEIKNGRLAMFSMFGFFVQAIVTGKGPLENLLDHLDDP 254
KEIKNGRLAM + GF QA T + P+ENL+ HL DP
Sbjct: 220 TKEIKNGRLAMLAFLGFCFQATYTSQDPIENLMAHLADP 258
>At3g47470 light-harvesting chlorophyll a/b-binding protein (Cab4)
Length = 251
Score = 111 bits (278), Expect = 4e-25
Identities = 87/255 (34%), Positives = 119/255 (46%), Gaps = 13/255 (5%)
Query: 5 LMAATTASSGTVIKPSPFLGQ-TKGSNSNTLRDVVSMGTGKYTMGNDLWYGPDRVKYLGP 63
+ TT +S ++ +P + GS+ RD+ G + + ++L
Sbjct: 1 MATVTTHASASIFRPCTSKPRFLTGSSGRLNRDLSFTSIGSSAKTSSFKVEAKKGEWLPG 60
Query: 64 FSAQTPSYLTGEFPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGALGCITPEVLEK 123
++ P YLTG GD G+D GL+ DPE E+++GRWAMLG G + PEV K
Sbjct: 61 LAS--PDYLTGSLAGDNGFDPLGLAEDPENLKWFVQAELVNGRWAMLGVAGMLLPEVFTK 118
Query: 124 WVRVDFKEPVWFKAGS-QIFSEGGLDYLGNPNLVHAQSILAVLGFQVLLMGLVEGFRING 182
++ P W+ AG Q F+ ++ L H I + G V I
Sbjct: 119 IGIINV--PEWYDAGKEQYFASSSTLFVIEFILFHYVEIRRWQ--DIKNPGSVNQDPIFK 174
Query: 183 LPGVGEGNNLYPGGQYFDPLGLADDPATFAELKVKEIKNGRLAMFSMFGFFVQAIVTGKG 242
+ +G YPGG F+PL A E K KE+ NGRLAM + GF VQ VTGKG
Sbjct: 175 QYSLPKGEVGYPGG-IFNPLNFAPTQ----EAKEKELANGRLAMLAFLGFVVQHNVTGKG 229
Query: 243 PLENLLDHLDDPVAN 257
P ENLL HL DP N
Sbjct: 230 PFENLLQHLSDPWHN 244
>At2g40100 putative chlorophyll a/b binding protein
Length = 276
Score = 92.8 bits (229), Expect = 2e-19
Identities = 64/150 (42%), Positives = 79/150 (52%), Gaps = 16/150 (10%)
Query: 98 RALEVIHGRWAMLGALGCITPEVLEKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPNLVH 157
R E+IHGRWAMLG LG I E L W AG EG YLG P
Sbjct: 138 RECELIHGRWAMLGTLGAIAVEALTGIA--------WQDAGKVELVEGS-SYLGQPLPF- 187
Query: 158 AQSILAVLGFQVLLMGLVEGFRINGLPGVGEGNNLYPGGQYFDPLGLADDPATFAELKVK 217
S+ ++ +VL++G +E R + L +YPGG YFDPLGLA DP LK+
Sbjct: 188 --SLTTLIWIEVLVVGYIEFQRNSELD---PEKRIYPGG-YFDPLGLAADPEKLDTLKLA 241
Query: 218 EIKNGRLAMFSMFGFFVQAIVTGKGPLENL 247
EIK+ RLAM + F +QA TGKGP+ L
Sbjct: 242 EIKHSRLAMVAFLIFALQAAFTGKGPVSFL 271
Score = 31.2 bits (69), Expect = 0.61
Identities = 16/38 (42%), Positives = 22/38 (57%), Gaps = 1/38 (2%)
Query: 76 FPGDYGWDTAGLSADPEAFAKNRALEVIHGRWAMLGAL 113
+PG Y +D GL+ADPE + E+ H R AM+ L
Sbjct: 218 YPGGY-FDPLGLAADPEKLDTLKLAEIKHSRLAMVAFL 254
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,588,599
Number of Sequences: 26719
Number of extensions: 300076
Number of successful extensions: 700
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 605
Number of HSP's gapped (non-prelim): 37
length of query: 270
length of database: 11,318,596
effective HSP length: 98
effective length of query: 172
effective length of database: 8,700,134
effective search space: 1496423048
effective search space used: 1496423048
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0590a.3


