

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.5
(191 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
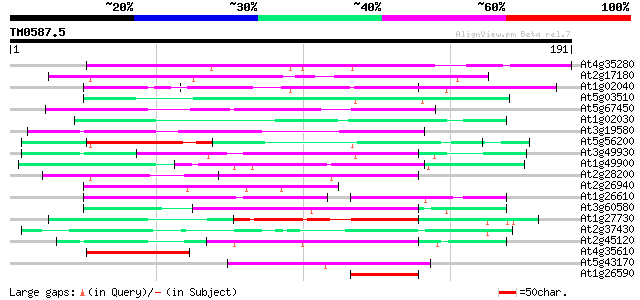
Score E
Sequences producing significant alignments: (bits) Value
At4g35280 putative zinc-finger protein 137 3e-33
At2g17180 putative C2H2-type zinc finger protein 114 3e-26
At1g02040 hypothetical protein 70 9e-13
At5g03510 unknown protein 59 1e-09
At5g67450 Cys2/His2-type zinc finger protein 1 (dbj|BAA85108.1) 58 3e-09
At1g02030 unknown protein 58 3e-09
At3g19580 zinc finger protein, putative 57 6e-09
At5g56200 unknown protein 56 1e-08
At3g49930 zinc-finger-like protein 55 2e-08
At1g49900 Cys2/His2-type zinc finger protein, putative 55 3e-08
At2g28200 putative zinc-finger protein 52 2e-07
At2g26940 putative C2H2-type zinc finger protein 51 4e-07
At1g26610 zinc-finger protein (ZPT4-4), putative 50 6e-07
At3g60580 zinc finger protein - like 49 2e-06
At1g27730 salt-tolerance zinc finger protein like 49 2e-06
At2g37430 putative C2H2-type zinc finger protein 48 3e-06
At2g45120 putative C2H2-type zinc finger protein 47 6e-06
At4g35610 putative protein 44 4e-05
At5g43170 Cys2/His2-type zinc finger protein 3 (dbj|BAA85109.1) 44 5e-05
At1g26590 C2H2 zinc finger protein, putative 44 5e-05
>At4g35280 putative zinc-finger protein
Length = 284
Score = 137 bits (346), Expect = 3e-33
Identities = 86/218 (39%), Positives = 106/218 (48%), Gaps = 65/218 (29%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQP------QPQPPQTTVITQE 80
C+ECG+KFWS KAL GHMRCHPER+WRGINPPP +R P+ Q P + +++E
Sbjct: 79 CTECGRKFWSWKALFGHMRCHPERQWRGINPPPNYRVPTAASSKQLNQILPNWVSFMSEE 138
Query: 81 EHEVAACLLLLANA---------------------------------NAKG----DTCSS 103
+HEVA+CLL+L+N N KG +
Sbjct: 139 DHEVASCLLMLSNGTPSSSSIERFECGGCKKVFGSHQALGGHRASHKNVKGCFAITNVTD 198
Query: 104 SKSEISCSSSHH----------HHKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPP 153
+S SS H HHKC+IC RVFSSGQALGGH R HWEK +
Sbjct: 199 DPMTVSTSSGHDHQGKILTFSGHHKCNICFRVFSSGQALGGHMRCHWEKEEE-------- 250
Query: 154 PAPAPAPVLDLNLPPPESGSDPITVPLDFAALDLSLRL 191
P + LDLN+PP + D T LDL L L
Sbjct: 251 --PMISGALDLNVPP--TIQDLSTSDTSGCCLDLRLGL 284
Score = 33.5 bits (75), Expect = 0.070
Identities = 14/26 (53%), Positives = 19/26 (72%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPERE 51
+C+ C + F S +AL GHMRCH E+E
Sbjct: 223 KCNICFRVFSSGQALGGHMRCHWEKE 248
>At2g17180 putative C2H2-type zinc finger protein
Length = 270
Score = 114 bits (285), Expect = 3e-26
Identities = 65/157 (41%), Positives = 87/157 (55%), Gaps = 17/157 (10%)
Query: 14 PSPSSSPSGKWSQ----CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKF-RRPSQPQ 68
P P + P SQ C+ECGK+F S KAL GHMRCHPER+WRGINPP F RR +
Sbjct: 50 PKPVTQPDPDASQIARPCTECGKQFGSLKALFGHMRCHPERQWRGINPPSNFKRRINSNA 109
Query: 69 PQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSS 128
+ ++EEH +A+CLL++AN GD + S S +C C +VF S
Sbjct: 110 ASSSSSWDPSEEEHNIASCLLMMAN----GDVPTRS------SEVEERFECDGCKKVFGS 159
Query: 129 GQALGGHKRRHWEKGGDLIHHHL--PPPAPAPAPVLD 163
QALGGH+ H + G + ++ PP P P ++D
Sbjct: 160 HQALGGHRATHKDVKGCFANKNITEDPPPPPPQEIVD 196
>At1g02040 hypothetical protein
Length = 324
Score = 69.7 bits (169), Expect = 9e-13
Identities = 52/169 (30%), Positives = 74/169 (43%), Gaps = 25/169 (14%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
QC C K F S +AL GH H ++ +G F + + + + + E
Sbjct: 151 QCKACKKVFTSHQALGGHRASH--KKVKGC-----FASQDKEEEEEEEYKEDDDDNDE-- 201
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGD 145
+ + + D S + I+ S+ H +C+IC RVFSSGQALGGHKR HW +
Sbjct: 202 -------DEDEEEDEEDKSTAHIARKRSNAH-ECTICHRVFSSGQALGGHKRCHWLTPSN 253
Query: 146 LI--------HHHLPPPAPAPAPVLDLNLPPPESGSDPITVPLDFAALD 186
+ HH + P P P LDLNL E DP + + D
Sbjct: 254 YLRMTSLHDHHHSVGRPQPLDQPSLDLNLACQEYSVDPTAMSVGMIERD 302
Score = 42.4 bits (98), Expect = 2e-04
Identities = 27/94 (28%), Positives = 41/94 (42%), Gaps = 20/94 (21%)
Query: 59 PKFRRPSQPQPQPPQTTVITQEEHEVAACLLLLANA-------------NAKGDTCSSSK 105
P ++R Q + E+ E+A CL+LL+N+ + KG T K
Sbjct: 86 PPWKRERSMSTQHHHLNLSPNEDEELANCLVLLSNSGDAHGGDQHKQHGHGKGKTVKKQK 145
Query: 106 SEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
+ +C C +VF+S QALGGH+ H
Sbjct: 146 TA-------QVFQCKACKKVFTSHQALGGHRASH 172
>At5g03510 unknown protein
Length = 292
Score = 59.3 bits (142), Expect = 1e-09
Identities = 50/160 (31%), Positives = 59/160 (36%), Gaps = 34/160 (21%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
QC C K F S +AL GH R S +P+ + E+ +
Sbjct: 119 QCKTCDKSFHSFQALGGH-------------------RASHKKPKLGASVFKCVEKKTAS 159
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHH-HKCSICLRVFSSGQALGGHKRRH----- 139
A + A A G S + S H+CSIC FSSGQALGGH RRH
Sbjct: 160 ASTVETVEAGAVGSFLSLQVTSSDGSKKPEKTHECSICKAEFSSGQALGGHMRRHRGLTI 219
Query: 140 ---------WEKGGDLIHHHLPPPAPAPAPVLDLNLPPPE 170
HHH P LDLNLP PE
Sbjct: 220 NANATSAIKTAISSSSHHHHEESIRPKNFLQLDLNLPAPE 259
Score = 33.9 bits (76), Expect = 0.054
Identities = 36/110 (32%), Positives = 42/110 (37%), Gaps = 45/110 (40%)
Query: 12 VVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQP 71
V S S K +CS C +F S +AL GHMR H RG
Sbjct: 179 VTSSDGSKKPEKTHECSICKAEFSSGQALGGHMRRH-----RG----------------- 216
Query: 72 PQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSI 121
L NANA S+ K+ IS SSSHHHH+ SI
Sbjct: 217 ------------------LTINANA----TSAIKTAIS-SSSHHHHEESI 243
Score = 33.1 bits (74), Expect = 0.092
Identities = 22/74 (29%), Positives = 34/74 (45%), Gaps = 12/74 (16%)
Query: 78 TQEEHEVAACLLLLANAN-AKGDTCSSSKSEISCSSSHHH-----------HKCSICLRV 125
T E+ ++A CL+LL+ + AK S + S+ ++C C +
Sbjct: 67 TDEDEDMANCLILLSQGHQAKSSDDHLSMQRMGFFSNKKPVASLGLGLDGVYQCKTCDKS 126
Query: 126 FSSGQALGGHKRRH 139
F S QALGGH+ H
Sbjct: 127 FHSFQALGGHRASH 140
>At5g67450 Cys2/His2-type zinc finger protein 1 (dbj|BAA85108.1)
Length = 245
Score = 58.2 bits (139), Expect = 3e-09
Identities = 38/133 (28%), Positives = 57/133 (42%), Gaps = 25/133 (18%)
Query: 13 VPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPP 72
+PS +S + +C+ CGK F S +AL GH H R+P+
Sbjct: 85 LPSRASPSDHRDYKCTVCGKSFSSYQALGGHKTSH--------------RKPTNTSITSG 130
Query: 73 QTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQAL 132
++ H + +++ N S K H CSIC + F+SGQAL
Sbjct: 131 NQE-LSNNSHSNSGSVVINVTVNTGNGVSQSGKI----------HTCSICFKSFASGQAL 179
Query: 133 GGHKRRHWEKGGD 145
GGHKR H++ G +
Sbjct: 180 GGHKRCHYDGGNN 192
>At1g02030 unknown protein
Length = 267
Score = 58.2 bits (139), Expect = 3e-09
Identities = 42/147 (28%), Positives = 56/147 (37%), Gaps = 48/147 (32%)
Query: 23 KWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEH 82
KW +C C K F S +AL GH H ++
Sbjct: 158 KWFECETCEKVFKSYQALGGHRASHKKK-------------------------------- 185
Query: 83 EVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEK 142
+A + G K + S SS HH+C IC +VF+SGQALGGHKR H
Sbjct: 186 --------IAETDQLGSDELKKKKKKSTSS---HHECPICAKVFTSGQALGGHKRSHASA 234
Query: 143 GGDLIHHHLPPPAPAPAPVLDLNLPPP 169
+ + ++DLNLP P
Sbjct: 235 NNEFTRR-----SGIIISLIDLNLPAP 256
Score = 37.7 bits (86), Expect = 0.004
Identities = 16/36 (44%), Positives = 23/36 (63%), Gaps = 6/36 (16%)
Query: 117 HKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLP 152
HKC +C + F++G+ALGGH R H ++ H LP
Sbjct: 5 HKCKLCWKSFANGRALGGHMRSH------MLIHPLP 34
Score = 35.8 bits (81), Expect = 0.014
Identities = 24/78 (30%), Positives = 38/78 (47%), Gaps = 2/78 (2%)
Query: 65 SQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLR 124
++ +P + T EE +VA L+LL+ + + S E + +C C +
Sbjct: 110 TEQEPMSSVSDAATTEE-DVALSLMLLSRDKWEKEE-EESDEERWKKKRNKWFECETCEK 167
Query: 125 VFSSGQALGGHKRRHWEK 142
VF S QALGGH+ H +K
Sbjct: 168 VFKSYQALGGHRASHKKK 185
Score = 27.7 bits (60), Expect = 3.9
Identities = 11/22 (50%), Positives = 14/22 (63%)
Query: 26 QCSECGKKFWSEKALHGHMRCH 47
+C C K F + +AL GHMR H
Sbjct: 6 KCKLCWKSFANGRALGGHMRSH 27
>At3g19580 zinc finger protein, putative
Length = 273
Score = 57.0 bits (136), Expect = 6e-09
Identities = 40/135 (29%), Positives = 57/135 (41%), Gaps = 34/135 (25%)
Query: 7 HTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQ 66
H Q + P S + + +C+ C K F S +AL GH H + PP +
Sbjct: 89 HQQSQSLTPPPESKNLPY-KCNVCEKAFPSYQALGGHKASHRIK-------PPTVISTTA 140
Query: 67 PQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVF 126
P +++ E+H +AA S H+CSIC +VF
Sbjct: 141 DDSTAPTISIVAGEKHPIAA--------------------------SGKIHECSICHKVF 174
Query: 127 SSGQALGGHKRRHWE 141
+GQALGGHKR H+E
Sbjct: 175 PTGQALGGHKRCHYE 189
>At5g56200 unknown protein
Length = 493
Score = 56.2 bits (134), Expect = 1e-08
Identities = 48/178 (26%), Positives = 66/178 (36%), Gaps = 37/178 (20%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQ----CSECGKKFWSEKALHGHMRCHPEREWRGINPPPK 60
H H VV + G ++ C C K F S +AL GH H
Sbjct: 318 HDHQLDQVVVVAAEEGGGAGAREKHVCVTCNKSFSSYQALGGHRASH------------- 364
Query: 61 FRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHH-HHKC 119
+ ++ + A LL S+ + S SSSH+ H C
Sbjct: 365 -----------NKVKILENHQARANAEASLLGTEAIITGLASAQGTNTSLSSSHNGDHVC 413
Query: 120 SICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPPAPAPAPVLDLNLPPPESGSDPIT 177
+IC + FS+GQALGGHKR HW + P P AP + P + S +T
Sbjct: 414 NICHKSFSTGQALGGHKRCHWTAPVSTV---APTTVPTAAPTV-----PATASSSQVT 463
Score = 53.5 bits (127), Expect = 7e-08
Identities = 24/43 (55%), Positives = 28/43 (64%), Gaps = 4/43 (9%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQP 69
C C KKF S KAL+GHMR HP+R W+G+ PPP P P P
Sbjct: 132 CCVCYKKFTSMKALYGHMRFHPDRGWKGVLPPP----PLLPHP 170
Score = 41.2 bits (95), Expect = 3e-04
Identities = 36/135 (26%), Positives = 50/135 (36%), Gaps = 44/135 (32%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVAA 86
C ECGK+F S KAL GH R H R+ S +P+ V E ++
Sbjct: 81 CCECGKRFVSGKALGGHKRIHVLET----------RKFSMMRPKMVSGMVGRSERGDLEV 130
Query: 87 CLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDL 146
C +C + F+S +AL GH R H ++G
Sbjct: 131 A-------------------------------CCVCYKKFTSMKALYGHMRFHPDRGWKG 159
Query: 147 IHHHLPPPAPAPAPV 161
+ LPPP P P+
Sbjct: 160 V---LPPPPLLPHPL 171
Score = 32.3 bits (72), Expect = 0.16
Identities = 14/23 (60%), Positives = 16/23 (68%)
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
H C C + F SG+ALGGHKR H
Sbjct: 79 HICCECGKRFVSGKALGGHKRIH 101
>At3g49930 zinc-finger-like protein
Length = 215
Score = 55.1 bits (131), Expect = 2e-08
Identities = 36/100 (36%), Positives = 50/100 (50%), Gaps = 9/100 (9%)
Query: 44 MRCHPEREWRGINPPPKFRRPSQ---PQPQPPQTTVITQEEHEVAACLLLLANANAKGDT 100
+RC E E + K +R + QP PP + EE +A CLL+LA ++ +
Sbjct: 22 LRCLDETEPENLESWTKRKRTKRHRIDQPNPPPS-----EEEYLALCLLMLARGSSDHHS 76
Query: 101 CSSSKSEISCSSSHHH-HKCSICLRVFSSGQALGGHKRRH 139
S +S S H +KCS+C + F S QALGGHK H
Sbjct: 77 PPSDHHSLSPLSDHQKDYKCSVCGKSFPSYQALGGHKTSH 116
Score = 53.9 bits (128), Expect = 5e-08
Identities = 52/177 (29%), Positives = 61/177 (34%), Gaps = 55/177 (31%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRP 64
HS H SP S + +CS CGK F S +AL GH H R+P
Sbjct: 75 HSPPSDHHSLSPLSDHQKDY-KCSVCGKSFPSYQALGGHKTSH--------------RKP 119
Query: 65 SQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLR 124
TV N N S H CSIC +
Sbjct: 120 VSVDVNNSNGTVTN--------------NGNISNGLVGQSGKT---------HNCSICFK 156
Query: 125 VFSSGQALGGHKRRHWEKG-----GDLIHHHLPPPAPAPAPVLDLNLPPPESGSDPI 176
F SGQALGGHKR H++ G GD H DLNLP + + I
Sbjct: 157 SFPSGQALGGHKRCHYDGGNGNSNGDNSHK------------FDLNLPADQVSDETI 201
>At1g49900 Cys2/His2-type zinc finger protein, putative
Length = 917
Score = 54.7 bits (130), Expect = 3e-08
Identities = 49/195 (25%), Positives = 68/195 (34%), Gaps = 67/195 (34%)
Query: 4 SHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRR 63
S + QK P P QC+ CG++ S +AL GH H + PP +
Sbjct: 729 SQTQLQKVTQPQTQMLPKSDSYQCNVCGRELPSYQALGGHKASHRTK------PPVE--- 779
Query: 64 PSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICL 123
NA G+ K + S HKCSIC
Sbjct: 780 -------------------------------NATGEKMRPKK----LAPSGKIHKCSICH 804
Query: 124 RVFSSGQALGGHKRRHWE-----------------------KGGDLIHHHLPPPAPAPAP 160
R FS+GQ+LGGHKR H+E G + H+P P +
Sbjct: 805 REFSTGQSLGGHKRLHYEGVLRGHKRSQEKEAVSQGDKLSPSGNGSVVTHVPDPKQSRKG 864
Query: 161 VLDLNLPPPESGSDP 175
++ +N P +DP
Sbjct: 865 LIVINKVPSPEFNDP 879
Score = 52.0 bits (123), Expect = 2e-07
Identities = 35/98 (35%), Positives = 50/98 (50%), Gaps = 16/98 (16%)
Query: 57 PPPKFRRPSQPQPQPPQTT---------VITQEE----HEVAACLLLLANANAKGDTCSS 103
PPP+ + SQ Q PP++ V T + H+ + + NA D
Sbjct: 175 PPPQLQ--SQTQTAPPKSDLFKCSICEKVFTSYQALGGHKASHSIKAAQLENAGADAGEK 232
Query: 104 SKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWE 141
++S++ S S HKC IC +F +GQALGGHKRRH+E
Sbjct: 233 TRSKM-LSPSGKIHKCDICHVLFPTGQALGGHKRRHYE 269
Score = 38.9 bits (89), Expect = 0.002
Identities = 18/29 (62%), Positives = 20/29 (68%)
Query: 118 KCSICLRVFSSGQALGGHKRRHWEKGGDL 146
KCSIC +VF+S QALGGHK H K L
Sbjct: 194 KCSICEKVFTSYQALGGHKASHSIKAAQL 222
Score = 29.6 bits (65), Expect = 1.0
Identities = 15/31 (48%), Positives = 18/31 (57%)
Query: 19 SPSGKWSQCSECGKKFWSEKALHGHMRCHPE 49
SPSGK +C C F + +AL GH R H E
Sbjct: 239 SPSGKIHKCDICHVLFPTGQALGGHKRRHYE 269
>At2g28200 putative zinc-finger protein
Length = 284
Score = 51.6 bits (122), Expect = 2e-07
Identities = 29/73 (39%), Positives = 40/73 (54%), Gaps = 5/73 (6%)
Query: 72 PQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEI-----SCSSSHHHHKCSICLRVF 126
P TT TQEE ++A CL++LA + +I S +SS + ++C C R F
Sbjct: 63 PVTTDCTQEEEDMAICLIMLARGTVLPSPDLKNSRKIHQKISSENSSFYVYECKTCNRTF 122
Query: 127 SSGQALGGHKRRH 139
SS QALGGH+ H
Sbjct: 123 SSFQALGGHRASH 135
Score = 49.7 bits (117), Expect = 9e-07
Identities = 42/141 (29%), Positives = 58/141 (40%), Gaps = 30/141 (21%)
Query: 12 VVPSPSSSPSGKWSQ-------------CSECGKKFWSEKALHGHMRCHPEREWRGINPP 58
V+PSP S K Q C C + F S +AL GH H
Sbjct: 87 VLPSPDLKNSRKIHQKISSENSSFYVYECKTCNRTFSSFQALGGHRASHK---------- 136
Query: 59 PKFRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHK 118
K R ++ + + P T + E ++ G +S S I + ++ H+
Sbjct: 137 -KPRTSTEEKTRLPLTQPKSSASEEGQN-----SHFKVSGSALASQASNI-INKANKVHE 189
Query: 119 CSICLRVFSSGQALGGHKRRH 139
CSIC F+SGQALGGH RRH
Sbjct: 190 CSICGSEFTSGQALGGHMRRH 210
>At2g26940 putative C2H2-type zinc finger protein
Length = 286
Score = 50.8 bits (120), Expect = 4e-07
Identities = 33/105 (31%), Positives = 50/105 (47%), Gaps = 18/105 (17%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPP-------KFRRPSQPQPQPPQTTVIT 78
+C CGK+F + +L GHMR HP+R W+GI PPP + V++
Sbjct: 120 RCCLCGKEFQTMHSLFGHMRRHPDRTWKGIRPPPPSEKFNLSYLDDDVDDEDEDDDDVMS 179
Query: 79 Q--------EEHEVAACLLLL---ANANAKGDTCSSSKSEISCSS 112
+ E+ + AAC+L++ A+ N S KSE+ SS
Sbjct: 180 RSMMMSDVPEDVQGAACILMMLSYASRNYWAGIEESPKSEVRSSS 224
Score = 34.3 bits (77), Expect = 0.041
Identities = 16/36 (44%), Positives = 19/36 (52%)
Query: 104 SKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
S+ E H C IC + FSSG+A GGH R H
Sbjct: 36 SEDEDESPQPQRKHFCVICEKQFSSGKAYGGHVRIH 71
Score = 33.1 bits (74), Expect = 0.092
Identities = 30/128 (23%), Positives = 53/128 (40%), Gaps = 25/128 (19%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVAA 86
C C K+F S KA GH+R H E+ K ++ + + + ++ +E+ +
Sbjct: 51 CVICEKQFSSGKAYGGHVRIH-SIEYNNKGKMKKMKKMKLKKKKKRKIGLVKKEKEKE-- 107
Query: 87 CLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDL 146
+ LA + +G +C +C + F + +L GH RRH ++
Sbjct: 108 --IDLARTDVEGKI-----------------RCCLCGKEFQTMHSLFGHMRRHPDRTWKG 148
Query: 147 IHHHLPPP 154
I PPP
Sbjct: 149 IR---PPP 153
>At1g26610 zinc-finger protein (ZPT4-4), putative
Length = 455
Score = 50.4 bits (119), Expect = 6e-07
Identities = 28/57 (49%), Positives = 34/57 (59%), Gaps = 7/57 (12%)
Query: 117 HKCSICLRVFSSGQALGGHKRRHW----EKGGDLIHHHLPPPAPAPAPVLDLNLPPP 169
H+C IC RVF SGQALGGHKR H+ E +I H + A ++DLNLP P
Sbjct: 399 HECPICFRVFKSGQALGGHKRSHFIGNQEHRTLVIQHQV---AHEMHTLIDLNLPAP 452
Score = 42.7 bits (99), Expect = 1e-04
Identities = 29/83 (34%), Positives = 40/83 (47%), Gaps = 2/83 (2%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C ECGK F S KAL GHM CH ERE ++ S P + + V+ + H A
Sbjct: 94 RCRECGKGFVSSKALCGHMACHSEREKIVMDSQSDTEASSSPIRRRSKRVVV--KPHHKA 151
Query: 86 ACLLLLANANAKGDTCSSSKSEI 108
A ++ + + SS SEI
Sbjct: 152 AFVVGGNGIMNQSISASSDASEI 174
Score = 27.7 bits (60), Expect = 3.9
Identities = 10/21 (47%), Positives = 13/21 (61%)
Query: 119 CSICLRVFSSGQALGGHKRRH 139
C C + F G++LGGH R H
Sbjct: 11 CKFCSKRFPCGKSLGGHIRTH 31
>At3g60580 zinc finger protein - like
Length = 288
Score = 48.9 bits (115), Expect = 2e-06
Identities = 41/147 (27%), Positives = 56/147 (37%), Gaps = 46/147 (31%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTTVITQEEHEVA 85
+C CGK F S +AL GH H +
Sbjct: 174 KCETCGKVFKSYQALGGHRASHKKNRV--------------------------------- 200
Query: 86 ACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGD 145
+N K + S ++ + + H+C ICLRVF+SGQALGGHKR H G+
Sbjct: 201 --------SNNKTEQRSETEYDNVVVVAKRIHECPICLRVFASGQALGGHKRSHGV--GN 250
Query: 146 LI---HHHLPPPAPAPAPVLDLNLPPP 169
L + ++DLNLP P
Sbjct: 251 LSVNQQRRVHRNESVKQRMIDLNLPAP 277
Score = 43.1 bits (100), Expect = 9e-05
Identities = 25/90 (27%), Positives = 42/90 (45%), Gaps = 13/90 (14%)
Query: 63 RPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTC-------------SSSKSEIS 109
+P Q P + T E ++A CL++L+ K + S ++I+
Sbjct: 106 KPESDQEPPHSSASDTTTEEDLAFCLMMLSRDKWKKNKSNKEVVEEIETEEESEGYNKIN 165
Query: 110 CSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
+++ +KC C +VF S QALGGH+ H
Sbjct: 166 RATTKGRYKCETCGKVFKSYQALGGHRASH 195
Score = 34.7 bits (78), Expect = 0.032
Identities = 12/23 (52%), Positives = 17/23 (73%)
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
+KC +C + F +G+ALGGH R H
Sbjct: 4 YKCRVCFKSFVNGKALGGHMRSH 26
Score = 29.6 bits (65), Expect = 1.0
Identities = 17/41 (41%), Positives = 21/41 (50%), Gaps = 6/41 (14%)
Query: 26 QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQ 66
+C C K F + KAL GHMR H N + +RPSQ
Sbjct: 5 KCRVCFKSFVNGKALGGHMRSHMS------NSHEEEQRPSQ 39
>At1g27730 salt-tolerance zinc finger protein like
Length = 227
Score = 48.9 bits (115), Expect = 2e-06
Identities = 47/180 (26%), Positives = 68/180 (37%), Gaps = 49/180 (27%)
Query: 14 PSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQ 73
P P + +CS C K F S +AL GH H + Q
Sbjct: 69 PPPPPAVEKLSYKCSVCDKTFSSYQALGGHKASH--------------------RKNLSQ 108
Query: 74 TTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALG 133
T ++H T S++ + + S H C+IC + F SGQALG
Sbjct: 109 TLSGGGDDHS----------------TSSATTTSAVTTGSGKSHVCTICNKSFPSGQALG 152
Query: 134 GHKRRHWEKGGDLIHHHLPPPAPAPAPV--------LDLNLPP-PE----SGSDPITVPL 180
GHKR H+E ++ + A + DLN+PP PE +G D + P+
Sbjct: 153 GHKRCHYEGNNNINTSSVSNSEGAGSTSHVSSSHRGFDLNIPPIPEFSMVNGDDEVMSPM 212
Score = 42.4 bits (98), Expect = 2e-04
Identities = 26/63 (41%), Positives = 38/63 (60%), Gaps = 9/63 (14%)
Query: 77 ITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHK 136
+T+EE+ +A CL+LLA N + + ++S +KCS+C + FSS QALGGHK
Sbjct: 49 LTEEEY-LAFCLMLLARDNRQPPP-PPAVEKLS-------YKCSVCDKTFSSYQALGGHK 99
Query: 137 RRH 139
H
Sbjct: 100 ASH 102
Score = 36.2 bits (82), Expect = 0.011
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 1/47 (2%)
Query: 3 NSHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPE 49
+ HS T S ++ SGK C+ C K F S +AL GH RCH E
Sbjct: 115 DDHS-TSSATTTSAVTTGSGKSHVCTICNKSFPSGQALGGHKRCHYE 160
>At2g37430 putative C2H2-type zinc finger protein
Length = 178
Score = 48.1 bits (113), Expect = 3e-06
Identities = 50/179 (27%), Positives = 68/179 (37%), Gaps = 68/179 (37%)
Query: 5 HSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRP 64
+ HT+ H + +C C K+F S +AL GH H + PK
Sbjct: 36 NQHTESH---------TSNQFECKTCNKRFSSFQALGGHRASHKK---------PK---- 73
Query: 65 SQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLR 124
+T E+ +V L+N + KG+ H HKCSIC +
Sbjct: 74 ------------LTVEQKDVKH----LSN-DYKGN---------------HFHKCSICSQ 101
Query: 125 VFSSGQALGGHKRRHWEKGGDLIHHHLPPPAPAPAPV------------LDLNLPPPES 171
F +GQALGGH RRH + + P PV LDLNL P E+
Sbjct: 102 SFGTGQALGGHMRRH--RSSMTVEPSFISPMIPSMPVLKRCGSSKRILSLDLNLTPLEN 158
Score = 37.0 bits (84), Expect = 0.006
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 3/57 (5%)
Query: 83 EVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
++A CL++LA + + +E S + + +C C + FSS QALGGH+ H
Sbjct: 16 DIAKCLMILAQTSMVKQIGLNQHTE---SHTSNQFECKTCNKRFSSFQALGGHRASH 69
>At2g45120 putative C2H2-type zinc finger protein
Length = 314
Score = 47.0 bits (110), Expect = 6e-06
Identities = 44/162 (27%), Positives = 62/162 (38%), Gaps = 56/162 (34%)
Query: 17 SSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPPQTT- 75
SS G++ +C CGK F S +AL GH H K + + + +T
Sbjct: 187 SSKSRGRF-KCETCGKVFKSYQALGGHRASH------------KKNKACMTKTEQVETEY 233
Query: 76 VITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGH 135
V+ +E +V H+C IC RVF+SGQALGGH
Sbjct: 234 VLGVKEKKV--------------------------------HECPICFRVFTSGQALGGH 261
Query: 136 KRRHWEKGG--------DLIHHHLPPPAPAPAPVLDLNLPPP 169
KR H G ++ + ++DLNLP P
Sbjct: 262 KRSHGSNIGAGRGLSVSQIV--QIEEEVSVKQRMIDLNLPAP 301
Score = 41.2 bits (95), Expect = 3e-04
Identities = 24/82 (29%), Positives = 35/82 (42%), Gaps = 10/82 (12%)
Query: 68 QPQPPQTTVITQEEHEVAACLLLLANANAKG----------DTCSSSKSEISCSSSHHHH 117
+P+ + T E ++A CL++L+ K D + S S
Sbjct: 135 EPEHHSSASDTTTEEDLAFCLIMLSRDKWKQQKKKKQRVEEDETDHDSEDYKSSKSRGRF 194
Query: 118 KCSICLRVFSSGQALGGHKRRH 139
KC C +VF S QALGGH+ H
Sbjct: 195 KCETCGKVFKSYQALGGHRASH 216
Score = 33.5 bits (75), Expect = 0.070
Identities = 12/23 (52%), Positives = 16/23 (69%)
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
+KC C + F +G+ALGGH R H
Sbjct: 4 YKCRFCFKSFINGRALGGHMRSH 26
Score = 26.9 bits (58), Expect = 6.6
Identities = 11/22 (50%), Positives = 14/22 (63%)
Query: 26 QCSECGKKFWSEKALHGHMRCH 47
+C C K F + +AL GHMR H
Sbjct: 5 KCRFCFKSFINGRALGGHMRSH 26
>At4g35610 putative protein
Length = 271
Score = 44.3 bits (103), Expect = 4e-05
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKF 61
C CG+ F S KA+ GHMR H +R ++G PPP F
Sbjct: 140 CHICGRGFGSWKAVFGHMRAHRDRNYQGFLPPPTF 174
Score = 28.9 bits (63), Expect = 1.7
Identities = 15/36 (41%), Positives = 19/36 (52%), Gaps = 3/36 (8%)
Query: 119 CSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPP 154
C IC R F S +A+ GH R H ++ LPPP
Sbjct: 140 CHICGRGFGSWKAVFGHMRAHRDRN---YQGFLPPP 172
>At5g43170 Cys2/His2-type zinc finger protein 3 (dbj|BAA85109.1)
Length = 193
Score = 43.9 bits (102), Expect = 5e-05
Identities = 24/77 (31%), Positives = 41/77 (53%), Gaps = 8/77 (10%)
Query: 75 TVITQEEHEVAACLLLLANANAKGDTCSSSKS--------EISCSSSHHHHKCSICLRVF 126
TV + ++ C ++ A G +S +S + + S++ H CS+C + F
Sbjct: 68 TVAEKPSYKCGVCYKTFSSYQALGGHKASHRSLYGGGENDKSTPSTAVKSHVCSVCGKSF 127
Query: 127 SSGQALGGHKRRHWEKG 143
++GQALGGHKR H++ G
Sbjct: 128 ATGQALGGHKRCHYDGG 144
Score = 33.5 bits (75), Expect = 0.070
Identities = 13/21 (61%), Positives = 15/21 (70%)
Query: 27 CSECGKKFWSEKALHGHMRCH 47
CS CGK F + +AL GH RCH
Sbjct: 120 CSVCGKSFATGQALGGHKRCH 140
>At1g26590 C2H2 zinc finger protein, putative
Length = 361
Score = 43.9 bits (102), Expect = 5e-05
Identities = 17/23 (73%), Positives = 19/23 (81%)
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
H+C IC R+F SGQALGGHKR H
Sbjct: 315 HECPICFRMFKSGQALGGHKRSH 337
Score = 36.6 bits (83), Expect = 0.008
Identities = 15/23 (65%), Positives = 16/23 (69%)
Query: 27 CSECGKKFWSEKALHGHMRCHPE 49
C ECG+ F S KAL GHM CH E
Sbjct: 66 CRECGRVFVSLKALRGHMACHGE 88
Score = 31.2 bits (69), Expect = 0.35
Identities = 12/21 (57%), Positives = 14/21 (66%)
Query: 119 CSICLRVFSSGQALGGHKRRH 139
C C + F SG+ALGGH R H
Sbjct: 11 CKYCYKTFPSGKALGGHIRIH 31
Score = 30.0 bits (66), Expect = 0.78
Identities = 17/40 (42%), Positives = 18/40 (44%)
Query: 27 CSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQ 66
C C K F S KAL GH+R H G N K R Q
Sbjct: 11 CKYCYKTFPSGKALGGHIRIHTNENSVGYNGNKKKRLVDQ 50
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,813,574
Number of Sequences: 26719
Number of extensions: 297661
Number of successful extensions: 3264
Number of sequences better than 10.0: 166
Number of HSP's better than 10.0 without gapping: 126
Number of HSP's successfully gapped in prelim test: 40
Number of HSP's that attempted gapping in prelim test: 2203
Number of HSP's gapped (non-prelim): 975
length of query: 191
length of database: 11,318,596
effective HSP length: 94
effective length of query: 97
effective length of database: 8,807,010
effective search space: 854279970
effective search space used: 854279970
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0587.5


