

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.12
(583 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
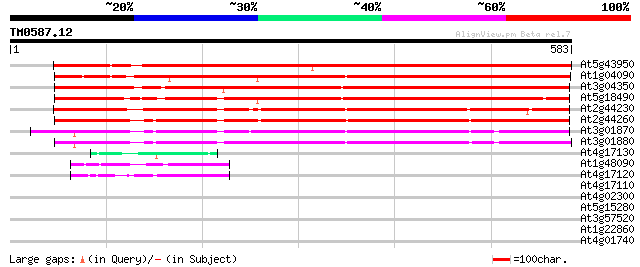
Score E
Sequences producing significant alignments: (bits) Value
At5g43950 unknown protein 665 0.0
At1g04090 unknown protein 651 0.0
At3g04350 unknown protein 644 0.0
At5g18490 Unknown protein 625 e-179
At2g44230 putative protein 467 e-132
At2g44260 unknown protein 466 e-131
At3g01870 hypothetical protein 423 e-118
At3g01880 hypothetical protein 404 e-113
At4g17130 hypothetical protein 48 2e-05
At1g48090 unknown protein 47 3e-05
At4g17120 hypothetical protein 47 4e-05
At4g17110 hypothetical protein 31 1.7
At4g02300 putative pectinesterase 31 1.7
At5g15280 putative protein 30 2.8
At3g57520 imbibition protein homolog 30 3.7
At1g22860 unknown protein 30 4.8
At4g01740 putative CHP-rich zinc finger protein 29 6.3
>At5g43950 unknown protein
Length = 566
Score = 665 bits (1716), Expect = 0.0
Identities = 326/544 (59%), Positives = 398/544 (72%), Gaps = 18/544 (3%)
Query: 46 TGQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQP 105
+GQGF G INLGELEV +TSFE VW + +K+V+FYKP +P+ FH L HYCQ
Sbjct: 35 SGQGFGLGRINLGELEVAEITSFEFVWRYCSRRDNKKSVSFYKPDKLPEDFHCLGHYCQ- 93
Query: 106 SYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSS-A 164
S + L GF+LVAR+V S PAL +PLDYTLVW SN S+E + S
Sbjct: 94 SDSHLLRGFLLVARQVNKSSE-----------PALVQPLDYTLVWSSNDLSEERQSESYG 142
Query: 165 YFWLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFS 224
YFWLPQPP+GYK +GYLVT P KP LD++ CVRADLTDKCE ++ I+ S+ P
Sbjct: 143 YFWLPQPPQGYKPIGYLVTTSPAKPELDQVRCVRADLTDKCEAHKVIITAISDSLSIPMF 202
Query: 225 VWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVV-CLKNLNPVLPAMPCLQQIHALIE 283
+W RP DRGM GKGVS GTFFC++ + L + CLKNL+ L AMP ++QIHA+I+
Sbjct: 203 IWKTRPSDRGMRGKGVSTGTFFCTTQSPEEDHLSTIACLKNLDSSLHAMPNIEQIHAMIQ 262
Query: 284 HYGPTVYFHPEEVYLPSSVDWFFSNRAMLF---RKGVSTGEAIDAGGSNLPIGGTNDGEF 340
HYGP VYFHP EVYLPSSV WFF N A+L V E ID GSNLP GGTND +
Sbjct: 263 HYGPRVYFHPNEVYLPSSVSWFFKNGALLCSNSNSSVINNEPIDETGSNLPHGGTNDKRY 322
Query: 341 WIDLPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITS 400
WIDLP +D +REF+K GDL+ +KLYVHVKPA GGTFTDLA W+FCPFNGP+TLK G+
Sbjct: 323 WIDLPINDQQRREFIKRGDLESSKLYVHVKPAFGGTFTDLAFWIFCPFNGPATLKLGLMD 382
Query: 401 IAFSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYING-NKAIVYSS 459
++ +K G+HV DWEHFT+RI NFSGEL+SIYFSQHSGG+W+ +LE++ G NKA+VYSS
Sbjct: 383 LSLAKTGQHVCDWEHFTVRISNFSGELYSIYFSQHSGGEWIKPENLEFVEGSNKAVVYSS 442
Query: 460 KSGHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGDVVREPQWLQ 519
K+GHASF G Y+QGS+ LGIGIRND+ S+L+VDSS++YEIVAAEYL V EP WL
Sbjct: 443 KNGHASFSKSGMYLQGSALLGIGIRNDSAKSDLFVDSSLKYEIVAAEYLRGAVVEPPWLG 502
Query: 520 YMREWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIG 579
YMREWGPKIVY S++E++K+ LP+RLR +++K+PVEL GEEGPTGPKEKNNW G
Sbjct: 503 YMREWGPKIVYNSRSEIEKLNERLPWRLRSWVDAVLRKIPVELSGEEGPTGPKEKNNWFG 562
Query: 580 DERW 583
DERW
Sbjct: 563 DERW 566
>At1g04090 unknown protein
Length = 572
Score = 651 bits (1680), Expect = 0.0
Identities = 318/546 (58%), Positives = 398/546 (72%), Gaps = 20/546 (3%)
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGI-PDGFHVLSHYCQP 105
GQGF SG INLG+L+V +T FE +W E +K ++FYKP G+ P FH L HYCQ
Sbjct: 36 GQGFGSGTINLGKLQVIKITDFEFIWRYR-STEKKKNISFYKPKGLLPKDFHCLGHYCQ- 93
Query: 106 SYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSA- 164
S + PL G+VL AR++ ++ +Q + PAL +P+D+TLVW SN ++ +S +
Sbjct: 94 SDSHPLRGYVLAARDL-------VDSLEQVEKPALVEPVDFTLVWSSNDSAENECSSKSE 146
Query: 165 --YFWLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFP 222
YFWLPQPPEGY+++G++VTK KP L+E+ CVRADLTD CEP+ I+ SE P
Sbjct: 147 CGYFWLPQPPEGYRSIGFVVTKTSVKPELNEVRCVRADLTDICEPHNVIVTAVSESLGVP 206
Query: 223 FSVWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQ---LPVVCLKNLNPVLPAMPCLQQIH 279
+W RP DRGM GKGVS GTFFC + + + + + CLKNL+ L AMP + QI
Sbjct: 207 LFIWRTRPSDRGMWGKGVSAGTFFCRTRLVAAREDLGIGIACLKNLDLSLHAMPNVDQIQ 266
Query: 280 ALIEHYGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEAIDAGGSNLPIGGTNDGE 339
ALI+HYGPT+ FHP E YLPSSV WFF N A+L KG E ID GSNLP GG+ND +
Sbjct: 267 ALIQHYGPTLVFHPGETYLPSSVSWFFKNGAVLCEKGNPIEEPIDENGSNLPQGGSNDKQ 326
Query: 340 FWIDLPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGIT 399
FWIDLPCDD QR+FVK G+L+ +KLY+H+KPALGGTFTDL W+FCPFNGP+TLK G+
Sbjct: 327 FWIDLPCDDQ-QRDFVKRGNLESSKLYIHIKPALGGTFTDLVFWIFCPFNGPATLKLGLV 385
Query: 400 SIAFSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYING-NKAIVYS 458
I+ +G+HV DWEHFTLRI NFSGEL+SIY SQHSGG+W+++YDLE I G NKA+VYS
Sbjct: 386 DISLISIGQHVCDWEHFTLRISNFSGELYSIYLSQHSGGEWIEAYDLEIIPGSNKAVVYS 445
Query: 459 SKSGHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYL--GDVVREPQ 516
SK GHASFP GTY+QGS+ LGIGIRND S L VDSS +YEI+AAEYL V+ EP
Sbjct: 446 SKHGHASFPRAGTYLQGSTMLGIGIRNDTARSELLVDSSSRYEIIAAEYLSGNSVLAEPP 505
Query: 517 WLQYMREWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNN 576
WLQYMREWGPK+VY S+ E+++++N P +R+S +++KLPVEL GEEGPTGPKEKNN
Sbjct: 506 WLQYMREWGPKVVYDSREEIERLVNRFPRTVRVSLATVLRKLPVELSGEEGPTGPKEKNN 565
Query: 577 WIGDER 582
W GDER
Sbjct: 566 WYGDER 571
>At3g04350 unknown protein
Length = 567
Score = 644 bits (1660), Expect = 0.0
Identities = 305/542 (56%), Positives = 392/542 (72%), Gaps = 21/542 (3%)
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
G+GFA+G I+LGE+EV +T F VW+S+ + K FY+ IP+GFH L HYCQP+
Sbjct: 38 GKGFATGRISLGEIEVVKITKFHRVWSSDSSHDKSKRATFYRADDIPEGFHCLGHYCQPT 97
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSAYF 166
++PL G+VL AR +A +++ P L+KP+ Y+LVW +++ YF
Sbjct: 98 -DQPLRGYVLAARTSKAVNADDF--------PPLKKPVSYSLVWSADSEKN----GGGYF 144
Query: 167 WLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPE----FP 222
WLP PP GY+A+G +VT +P +P +E+ CVR DLT+ CE IL+V S P
Sbjct: 145 WLPNPPVGYRAMGVIVTHEPGEPETEEVRCVREDLTESCETSEMILEVGSSKKSNGSSSP 204
Query: 223 FSVWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQL--PVVCLKNLNPVLPAMPCLQQIHA 280
FSVWS RPC+RGM+ +GV+VG+FFC + E+ + CLKNL+P L AMP L Q+HA
Sbjct: 205 FSVWSTRPCERGMLSQGVAVGSFFCCTYDLSSERTVPDIGCLKNLDPTLHAMPNLDQVHA 264
Query: 281 LIEHYGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEAIDAGGSNLPIGGTNDGEF 340
+IEH+GPTVYFHPEE Y+PSSV WFF N A+L+R G S G+ I++ GSNLP GG ND +F
Sbjct: 265 VIEHFGPTVYFHPEEAYMPSSVQWFFKNGALLYRSGKSEGQPINSTGSNLPAGGCNDMDF 324
Query: 341 WIDLPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITS 400
WIDLP +D++ + +K G+L+ ++LYVHVKPALGGTFTD+ MW+FCPFNGP+TLK G+ +
Sbjct: 325 WIDLP-EDEEAKSNLKKGNLESSELYVHVKPALGGTFTDIVMWIFCPFNGPATLKIGLFT 383
Query: 401 IAFSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYINGNKAIVYSSK 460
+ +++GEHVGDWEHFT RICNFSGELW ++FSQHSGG WVD+ D+E++ NK VYSSK
Sbjct: 384 LPMTRIGEHVGDWEHFTFRICNFSGELWQMFFSQHSGGGWVDASDIEFVKDNKPAVYSSK 443
Query: 461 SGHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLG-DVVREPQWLQ 519
GHASFPHPG Y+QGSSKLGIG+RND S VDSS +Y IVAAEYLG V EP WLQ
Sbjct: 444 HGHASFPHPGMYLQGSSKLGIGVRNDVAKSKYIVDSSQRYVIVAAEYLGKGAVIEPCWLQ 503
Query: 520 YMREWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIG 579
YMREWGP I Y S +E++KI+N LP +R S N+V P+ LYGEEGPTGPKEK+NW G
Sbjct: 504 YMREWGPTIAYDSGSEINKIMNLLPLVVRFSIENIVDLFPIALYGEEGPTGPKEKDNWEG 563
Query: 580 DE 581
DE
Sbjct: 564 DE 565
>At5g18490 Unknown protein
Length = 553
Score = 625 bits (1613), Expect = e-179
Identities = 305/540 (56%), Positives = 388/540 (71%), Gaps = 30/540 (5%)
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
G+GFA+G I+LGE++V VT F+ VW + +FYKPVGIP+GFH L HYCQP+
Sbjct: 37 GRGFATGRISLGEIQVVKVTEFDRVWKCGTSRGKLRCASFYKPVGIPEGFHCLGHYCQPN 96
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSAYF 166
N+PL GFVL AR +++ + D ++ P L+KPL+Y+LVW S+ S YF
Sbjct: 97 -NQPLRGFVLAAR-----ANKPGHLADDHR-PPLKKPLNYSLVWSSD--------SDCYF 141
Query: 167 WLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFSVW 226
WLP PP GY+A+G +VT ++P +DE+ CVR DLT+ CE ++L V S F+VW
Sbjct: 142 WLPNPPVGYRAVGVIVTDGSEEPEVDEVRCVREDLTESCETGEKVLGVGS------FNVW 195
Query: 227 SLRPCDRGMMGKGVSVGTFFCSSCVNMGEQ---LPVVCLKNLNPVLPAMPCLQQIHALIE 283
S +PC+RG+ +GV VG+F CS+ + + + CLKNL+P L MP L Q+HALI
Sbjct: 196 STKPCERGIWSRGVEVGSFVCSTNDLSSDNKAAMNIACLKNLDPSLQGMPNLDQVHALIH 255
Query: 284 HYGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEAIDAGGSNLPIGGTNDGEFWID 343
HYGP VYFHPEE Y+PSSV WFF N A+L R G S GE I++ GSNLP GG NDG FWID
Sbjct: 256 HYGPMVYFHPEETYMPSSVPWFFKNGALLHRFGKSQGEPINSAGSNLPAGGENDGSFWID 315
Query: 344 LPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAF 403
LP +D++ R +K G+++ ++LYVHVKPALGG FTD+ MW+FCPFNGP+TLK G+ ++
Sbjct: 316 LP-EDEEVRSNLKKGNIESSELYVHVKPALGGIFTDVVMWIFCPFNGPATLKIGLLTVPM 374
Query: 404 SKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYING-NKAIVYSSKSG 462
+++GEHVGDWEHFT RI NF+G+L ++FSQHSGG WVD DLE++ G NK +VYSSK G
Sbjct: 375 NRLGEHVGDWEHFTFRISNFNGDLTQMFFSQHSGGGWVDVSDLEFVKGSNKPVVYSSKHG 434
Query: 463 HASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGD-VVREPQWLQYM 521
HASFPHPG Y+QG SKLGIG+RND S VDSS +Y IVAAEYLG+ V EP WLQ+M
Sbjct: 435 HASFPHPGMYLQGPSKLGIGVRNDVAKSKYMVDSSQRYRIVAAEYLGEGAVSEPYWLQFM 494
Query: 522 REWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIGDE 581
REWGP IVY S E++KII+ LP LR SF +L P+ELYGEEGPTGPKEK+NW GDE
Sbjct: 495 REWGPTIVYDSAAEINKIIDLLPLILRNSFESL---FPIELYGEEGPTGPKEKDNWEGDE 551
>At2g44230 putative protein
Length = 542
Score = 467 bits (1202), Expect = e-132
Identities = 246/542 (45%), Positives = 339/542 (62%), Gaps = 37/542 (6%)
Query: 46 TGQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQP 105
+G+GFA G I+LG LEV V +F VW + F++P +P+GF +L Y QP
Sbjct: 31 SGEGFAKGRIDLGGLEVSQVDTFNKVWTVYEGGQDNLGATFFEPSSVPEGFSILGFYAQP 90
Query: 106 SYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEI-VTSSA 164
+ N+ L+G+ LV +++ GD +LR P+DY L+W + E +
Sbjct: 91 N-NRKLFGWTLVGKDLS---------GD-----SLRPPVDYLLLWSGKSTKVENNKVETG 135
Query: 165 YFWLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFS 224
YFW P PP+GY A+G +VT +KP LD++ CVR+DLTD+ EP I + + FS
Sbjct: 136 YFWQPVPPDGYNAVGLIVTTSDEKPPLDKIRCVRSDLTDQSEPDALIWETNG------FS 189
Query: 225 VWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVVCLKNLNPVLPAMPCLQQIHALIEH 284
V S +P +RG GVSVGTFF +S LP CLKN N MP QI AL +
Sbjct: 190 VSSSKPVNRGTQASGVSVGTFFSNS---PNPALP--CLKNNNFDFSCMPSKPQIDALFQT 244
Query: 285 YGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEA-IDAGGSNLPIGGTNDGEFWID 343
Y P +YFH +E YLPSSV+WFFSN A+L++KG + ++ G NLP G NDG +W+D
Sbjct: 245 YAPWIYFHKDEKYLPSSVNWFFSNGALLYKKGDESNPVPVEPNGLNLPQGEFNDGLYWLD 304
Query: 344 LPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAF 403
LP D R+ V+ GDL+ ++Y+H+KP GGTFTD+A+W+F PFNGPS K SI
Sbjct: 305 LPVASD-ARKRVQCGDLQSMEVYLHIKPVFGGTFTDIAVWMFYPFNGPSRAKLKAASIPL 363
Query: 404 SKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYI-NGNKAIVYSSKSG 462
++GEH+GDWEHFTLRI NFSG+L +Y SQHSGG W D+ ++E+ GNK + Y+S +G
Sbjct: 364 GRIGEHIGDWEHFTLRISNFSGKLHRMYLSQHSGGSWADASEIEFQGGGNKPVAYASLNG 423
Query: 463 HASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGDVVREPQWLQYMR 522
HA + PG +QG K +GIRND S +D+++++ +VAAEY+ + EP WL YMR
Sbjct: 424 HAMYSKPGLVLQG--KDNVGIRNDTGKSEKVIDTAVRFRVVAAEYMRGELEEPAWLNYMR 481
Query: 523 EWGPKIVYGSKTEL---DKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIG 579
WGPKI YG + E+ +KI+ + L+ +F + +K LP E++GEEGPTGPK K NW+G
Sbjct: 482 HWGPKIDYGHENEIRGVEKIM--VGESLKTTFRSAIKGLPNEVFGEEGPTGPKLKRNWLG 539
Query: 580 DE 581
DE
Sbjct: 540 DE 541
>At2g44260 unknown protein
Length = 553
Score = 466 bits (1200), Expect = e-131
Identities = 240/540 (44%), Positives = 337/540 (61%), Gaps = 33/540 (6%)
Query: 47 GQGFASGIINLGE-LEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQP 105
G GFA G I+LG LEV V++F VW++ F++P IP GF +L +Y QP
Sbjct: 41 GDGFAKGTIDLGGGLEVSQVSTFNKVWSTYEGGPDNLGATFFEPSSIPSGFSILGYYAQP 100
Query: 106 SYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTS-SA 164
+ N+ L+G+VL AR++ + + L+ P+DYTLV N S +I +
Sbjct: 101 N-NRNLFGWVLTARDLSSNT--------------LKPPVDYTLV--GNTESLKIKQDGTG 143
Query: 165 YFWLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLTDKCEPYRQILDVSSEIPEFPFS 224
YFW P PP+GY+A+G +VT KP LD++ C+R+DLT++CE I + +
Sbjct: 144 YFWQPVPPDGYQAVGLIVTNYSQKPPLDKLRCIRSDLTEQCEADTWIWGTNG------VN 197
Query: 225 VWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVVCLKNLNPVLPAMPCLQQIHALIEH 284
+ +L+P RG GV VGTF + + L CLKN MP QI L +
Sbjct: 198 ISNLKPTTRGTQATGVYVGTFTWQTQNSSPPSLS--CLKNTKLDFSTMPNGSQIEELFQT 255
Query: 285 YGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEA-IDAGGSNLPIGGTNDGEFWID 343
+ P +YFHP+E YLPSSV W+F+N A+L++KG + I++ GSNLP GG+NDG +W+D
Sbjct: 256 FSPCIYFHPDEEYLPSSVTWYFNNGALLYKKGEESKPIPIESNGSNLPQGGSNDGSYWLD 315
Query: 344 LPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKFGITSIAF 403
LP D + +E VK GDL+ K+Y+H+KP LG TFTD+++W+F PFNGP+ K ++
Sbjct: 316 LPIDKNG-KERVKKGDLQSTKVYLHIKPMLGATFTDISIWIFYPFNGPAKAKVKFVNLPL 374
Query: 404 SKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYING--NKAIVYSSKS 461
++GEH+GDWEH TLRI NF+GELW ++ SQHSGG W+D+ DLE+ +G NK + Y+S
Sbjct: 375 GRIGEHIGDWEHTTLRISNFTGELWRVFLSQHSGGIWIDACDLEFQDGGNNKFVAYASLH 434
Query: 462 GHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGDVVREPQWLQYM 521
GHA +P PG +QG G+GIRND +D+ + YE++AAEY G V EP W++Y
Sbjct: 435 GHAMYPKPGLVLQGDD--GVGIRNDTGKGKKVLDTGLGYEVIAAEYDGGGVVEPPWVKYF 492
Query: 522 REWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIGDE 581
R+WGPKI Y E+ + LP L+ +FV VKK+P E+YGE+GPTGPK K+NW GDE
Sbjct: 493 RKWGPKIDYNVDDEVKSVERILPGLLKKAFVKFVKKIPDEVYGEDGPTGPKLKSNWAGDE 552
>At3g01870 hypothetical protein
Length = 583
Score = 423 bits (1087), Expect = e-118
Identities = 237/569 (41%), Positives = 329/569 (57%), Gaps = 40/569 (7%)
Query: 22 FPSFSTTPMASRSVYSVIVHVCVITGQGFASGIINLGELEVCSVT----SFELVWNSNVM 77
F S ++ P+ + + + V G F G I+LG LEV V+ + + VW +
Sbjct: 46 FLSSTSLPVETAFTFPSALPVIPSGGGNFGKGRIDLGGLEVIQVSISTSTSQRVWRTYEG 105
Query: 78 VEMRKAVAFYKPVGIPDGFHVLSHYCQPSYNKPLWGFVLVAREVEACSSEKIEIGDQNKL 137
++ ++P+ +P F L Y QP+ N+ L+G+VL AR+V S
Sbjct: 106 GPDNMGLSIFQPINLPPSFSTLGFYGQPN-NRLLFGWVLAARDVSGNS------------ 152
Query: 138 PALRKPLDYTLVWCSNAGSKEI-VTSSAYFWLPQPPEGYKALGYLVTKKPDKPNLDE--M 194
LR P+DY V N S I +A+FW P P GY+A+G VT P KP+L + +
Sbjct: 153 --LRPPVDYIQV--INTTSMNINQEGAAFFWQPLCPNGYQAVGLYVTTSPIKPSLSQESI 208
Query: 195 SCVRADLTDKCEPYRQILDVSSEIPEFPFSVWSLRPCDRGMMGKGVSVGTFFCSSCVNMG 254
SCVR+DLT++ E + ++ SLRP +RG GV GTF C +N+
Sbjct: 209 SCVRSDLTEQSETDTWVWGTEE------MTLSSLRPANRGTEATGVHTGTFSCQP-LNIP 261
Query: 255 EQLPVVCLKNLNPVLPAMPCLQQIHALIEHYGPTVYFHPEEVYLPSSVDWFFSNRAMLFR 314
P+ CLKN L +MP Q L + Y P +Y HP+E ++ SSVDWFFSN A+LF+
Sbjct: 262 PPPPLFCLKNTKFDLSSMPSHNQTTVLFQSYSPWIYLHPDEDFISSSVDWFFSNGALLFQ 321
Query: 315 KGVSTGEA-IDAGGSNLPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKGAKLYVHVKPAL 373
KG + + GSNLP GG++DG FW+D P D + +E+VK GDL K+Y+H+KP
Sbjct: 322 KGNESNPVPVQPDGSNLPQGGSDDGLFWLDYPADKN-AKEWVKRGDLGHTKVYLHIKPMF 380
Query: 374 GGTFTDLAMWVFCPFNGPSTLKF-GITSIAFSKVGEHVGDWEHFTLRICNFSGELWSIYF 432
GGTFTD+ +W+F PFNG + LKF S++ +GEH+GDWEH TLRI NF+GELW YF
Sbjct: 381 GGTFTDIVVWIFYPFNGNARLKFLFFKSLSLGDIGEHIGDWEHVTLRISNFNGELWRAYF 440
Query: 433 SQHSGGKWVDSYDLEYINGNKAIVYSSKSGHASFPHPGTYIQGSSKLGIGIRNDACSSNL 492
S+HSGG V++ DLE+ GNK + YSS GHA F PG +QG G GIRND SN
Sbjct: 441 SEHSGGTLVEACDLEFQGGNKLVSYSSLHGHAMFSKPGLVLQGDD--GNGIRNDMARSNK 498
Query: 493 YVDSSIQYEIVAAEYLGDVVREPQWLQYMREWGPKIVYGSKTELDKIINALPFRLRISFV 552
+ D+ + YE+VA G ++EP WL Y R+WGP + + + L+ I +LP LR F
Sbjct: 499 FFDAGVAYELVA----GPGIQEPPWLNYFRKWGPLVPHDIQKNLEGIAKSLPGLLRKKFR 554
Query: 553 NLVKKLPVELYGEEGPTGPKEKNNWIGDE 581
NL+ K+P E+ E+GPTGPK K +W GD+
Sbjct: 555 NLINKIPREVLEEDGPTGPKVKRSWTGDD 583
>At3g01880 hypothetical protein
Length = 592
Score = 404 bits (1038), Expect = e-113
Identities = 231/547 (42%), Positives = 311/547 (56%), Gaps = 40/547 (7%)
Query: 47 GQGFASGIINLGELEVCSVT----SFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHY 102
G F I++G LEV ++ + VW + V+ ++P IP F L Y
Sbjct: 75 GGNFGKRSIDMGGLEVTQISISNSTSHRVWRTYEGGPDNMGVSIFEPTTIPRNFFKLGFY 134
Query: 103 CQPSYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTS 162
QP+ N+ L+G++LVA++V + LR P+DYT V N + I
Sbjct: 135 AQPN-NRQLFGWILVAKDVSGSN--------------LRPPVDYTEV--GNTTTLLIKQE 177
Query: 163 S-AYFWLPQPPEGYKALGYLVTKKPDKPNLDE--MSCVRADLTDKCEPYRQILDVSSEIP 219
AYFW P P GY A+G VT P KP+L + +SCVR+DLT++ E + +
Sbjct: 178 GPAYFWQPLCPNGYHAVGLYVTTSPMKPSLGQNSISCVRSDLTEQSEADTWVWRIKD--- 234
Query: 220 EFPFSVWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVVCLKNLNPVLPAMPCLQQIH 279
++ SLRP RG+ GV GTF C + P+ CLKN L +MP Q
Sbjct: 235 ---MTISSLRPATRGVEATGVFTGTFSCKQLNFLPHPPPLFCLKNTKFDLSSMPSENQTR 291
Query: 280 ALIEHYGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKG-VSTGEAIDAGGSNLPIGGTNDG 338
L + Y P +Y HP+E +LPSSV+W F+N A+L +KG S I GSNLP GG ND
Sbjct: 292 VLFKTYSPWIYLHPKEDFLPSSVNWVFANGALLHKKGNESIPVPIHPNGSNLPQGGCNDD 351
Query: 339 EFWIDLPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAMWVFCPFNGPSTLKF-G 397
FW+D D RE VK GDL+ K+Y+H+KP G TFTD+ +W+F P+NG + LKF
Sbjct: 352 LFWLDYLVDKK-AREKVKRGDLESTKVYLHIKPMFGATFTDIVVWLFFPYNGNAHLKFLF 410
Query: 398 ITSIAFSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYI-NGNKAIV 456
I S++ +GEHVGDWEH TLRI NF+GELW +YFS+HSGG VD+ DLE++ GNK +V
Sbjct: 411 IKSLSLGNIGEHVGDWEHVTLRISNFNGELWRVYFSEHSGGTLVDACDLEFMQGGNKPVV 470
Query: 457 YSSKSGHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYEIVAAEYLGDVVREPQ 516
YSS GHA F PG +QG K GIRND S+ D+ I YE++A G V EP
Sbjct: 471 YSSLHGHAMFSKPGVVLQGGGK--SGIRNDMARSDKCFDAGIGYEVIA----GPGVVEPP 524
Query: 517 WLQYMREWGPKIVYGSKTELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNN 576
WL Y R+WGP++ Y L+ + LP LR L+ K+P+E+ G++GPTGPK K
Sbjct: 525 WLNYFRKWGPRVHYRIDIFLNSVAKILPIFLRKGLRKLINKIPLEMRGQDGPTGPKVKVT 584
Query: 577 WIGDERW 583
W GDE++
Sbjct: 585 WTGDEQY 591
>At4g17130 hypothetical protein
Length = 747
Score = 47.8 bits (112), Expect = 2e-05
Identities = 37/141 (26%), Positives = 56/141 (39%), Gaps = 27/141 (19%)
Query: 85 AFYKPVGIPDGFHVLSHYCQPSYNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPL 144
AF++P P GF L Y P P G ++V L +++PL
Sbjct: 543 AFWRPHP-PPGFASLGDYLTPLDKPPTKGVLVV----------------NTNLMRVKRPL 585
Query: 145 DYTLVWC---------SNAGSKEIVTSSAYFWLPQPPEGYKALGYLVTKKPDKPNLDEMS 195
+ L+W S+ K+ SS W P+ P+GY AL +V+ P+L
Sbjct: 586 SFKLIWSPLASGGLGGSSMDDKDERDSSCSIWFPEAPKGYVALSCVVSSGSTPPSLASTF 645
Query: 196 CVRADLTDKCEPYRQILDVSS 216
C+ A C R + +SS
Sbjct: 646 CILASSVSPCS-LRDCVAISS 665
>At1g48090 unknown protein
Length = 4099
Score = 47.0 bits (110), Expect = 3e-05
Identities = 40/167 (23%), Positives = 69/167 (40%), Gaps = 25/167 (14%)
Query: 64 SVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPSYNKPLWGFVLVAREVEA 123
S +FE +W ++R+ V+ ++PV P GF +L P G + A + E
Sbjct: 2264 STPNFERIWWDKGG-DLRRPVSIWRPVPRP-GFAILGDSITEGLEPPALGILFKADDSEI 2321
Query: 124 CSSEKIEIGDQNKLPALRKPLDYTLV-WCSNAGSKEIVTSSAYFWLPQPPEGYKALGYLV 182
+ KP+ + V G E+ + W P P GY +LG ++
Sbjct: 2322 AA----------------KPVQFNKVAHIVGKGFDEV-----FCWFPVAPPGYVSLGCVL 2360
Query: 183 TKKPDKPNLDEMSCVRADLTDKCEPYR-QILDVSSEIPEFPFSVWSL 228
+K + P++D C R DL ++ Y + SS +S+W +
Sbjct: 2361 SKFDEAPHVDSFCCPRIDLVNQANIYEASVTRSSSSKSSQLWSIWKV 2407
>At4g17120 hypothetical protein
Length = 1661
Score = 46.6 bits (109), Expect = 4e-05
Identities = 40/168 (23%), Positives = 74/168 (43%), Gaps = 23/168 (13%)
Query: 64 SVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPSYNKPLWGFVLVAREVEA 123
+V +FEL+W + +K V+ ++P+ + +G Y P +
Sbjct: 113 AVATFELIWWNRGSGSQKK-VSIWRPI-VSEGMAYFGDIAVSGYEPP-----------NS 159
Query: 124 CSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEIVTSSAYFWLPQPPEGYKALGYLVT 183
C + + D + L+ +D+ LV K S FW+PQ P G+ +LG +
Sbjct: 160 C----VVLHDTSDQEILKAAVDFQLV---GRVKKHRGVESISFWMPQAPPGFVSLGCVAC 212
Query: 184 KKPDKP-NLDEMSCVRADLTDKCEPYRQILDVSSEIPE--FPFSVWSL 228
K KP + ++ C R+D+ + L +S++ + PFS+WS+
Sbjct: 213 KGSPKPYDFTKLRCARSDMVAGDHFADESLWDTSDVWQRVEPFSIWSI 260
>At4g17110 hypothetical protein
Length = 335
Score = 31.2 bits (69), Expect = 1.7
Identities = 17/64 (26%), Positives = 29/64 (44%), Gaps = 3/64 (4%)
Query: 143 PLDYTLVWCSNAGSKEIVTSSAYFWLPQPPEGYKALGYLVTKKPDKPNLDEMSCVRADLT 202
P+ Y LVW + S W P+ PEG+ + G + +P L+ + C+ L
Sbjct: 211 PVGYDLVW---RNCLDDYISPVSIWHPRAPEGFVSPGCVAVAGFIEPELNTVYCMPTSLA 267
Query: 203 DKCE 206
++ E
Sbjct: 268 EQTE 271
>At4g02300 putative pectinesterase
Length = 532
Score = 31.2 bits (69), Expect = 1.7
Identities = 24/84 (28%), Positives = 37/84 (43%), Gaps = 14/84 (16%)
Query: 414 EHFTLRICNFSGELWSIYFSQHSGGKWVDSYDL----EYINGNKAIVYSSKSGHASFPHP 469
+H C F G ++Y HS ++ D+ ++I GN A+V+ + S +A P+P
Sbjct: 335 DHSAFYRCEFDGYQDTLYV--HSAKQFYRECDIYGTIDFIFGNAAVVFQNSSLYARKPNP 392
Query: 470 GTYI--------QGSSKLGIGIRN 485
G I Q GI I N
Sbjct: 393 GHKIAFTAQSRNQSDQPTGISILN 416
>At5g15280 putative protein
Length = 1227
Score = 30.4 bits (67), Expect = 2.8
Identities = 26/83 (31%), Positives = 40/83 (47%), Gaps = 5/83 (6%)
Query: 491 NLYVDSSIQYE----IVAAEYLGDVVREPQWLQYMREWGPKIVYGSKTELDKIINALPFR 546
N+ +D S+ E IV A GD+ + L M WG K+ S L + + A
Sbjct: 520 NMVLDRSVLPEFNSLIVRASEDGDLQTALRLLDEMARWGQKLSRRSFAVLMRSLCASRAH 579
Query: 547 LRISFVNLVKKLPVELYGEEGPT 569
LR+S ++L++K P Y +G T
Sbjct: 580 LRVS-ISLLEKWPKLAYQLDGET 601
>At3g57520 imbibition protein homolog
Length = 773
Score = 30.0 bits (66), Expect = 3.7
Identities = 27/90 (30%), Positives = 39/90 (43%), Gaps = 12/90 (13%)
Query: 276 QQIHALIEHYGPTVYFHPEEVYLPSSVDWFFSNRAMLFRKGVSTGEAIDAGGSNLPIGGT 335
Q + A+ H + H E+ LPS +DWF F V T E +D G +L GGT
Sbjct: 173 QSVKAVERHM--QTFHHREKKKLPSFLDWFGWCTWDAFYTDV-TAEGVDEGLKSLSEGGT 229
Query: 336 -------NDGEFWIDLPCDDDDQREFVKHG 358
+DG W + + D+ V+ G
Sbjct: 230 PPKFLIIDDG--WQQIENKEKDENCVVQEG 257
>At1g22860 unknown protein
Length = 947
Score = 29.6 bits (65), Expect = 4.8
Identities = 14/40 (35%), Positives = 22/40 (55%), Gaps = 1/40 (2%)
Query: 427 LWSIYFSQHSGGKWVDSYDL-EYINGNKAIVYSSKSGHAS 465
+W ++ +S G W DS DL Y++ N+ I S K A+
Sbjct: 564 IWRLFTKNYSSGLWQDSDDLVPYLHDNELIRLSGKEAAAA 603
>At4g01740 putative CHP-rich zinc finger protein
Length = 652
Score = 29.3 bits (64), Expect = 6.3
Identities = 28/122 (22%), Positives = 47/122 (37%), Gaps = 18/122 (14%)
Query: 157 KEIVTSSAYFWLPQPPE---------GYKALGYLVTKKPDKPNLDEMSC------VRADL 201
KE++ W PPE Y +L L PN + C + L
Sbjct: 128 KELIQFDYCVW--HPPEVNHTLEVNHSYHSLHPLKLHTGQLPNYSDRKCRLCAKEIEVGL 185
Query: 202 TDKCEPYRQILDVSSEIPEFPFSVWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVVC 261
C LD+S + +W+L+ D + S+ +F CS+C G++ P +C
Sbjct: 186 FYHCSLCNFTLDMSCVLNPPQRYLWNLKAHDH-QLTLLPSLRSFLCSACGLNGDRSPYIC 244
Query: 262 LK 263
++
Sbjct: 245 VQ 246
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,983,470
Number of Sequences: 26719
Number of extensions: 703168
Number of successful extensions: 1431
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1348
Number of HSP's gapped (non-prelim): 19
length of query: 583
length of database: 11,318,596
effective HSP length: 105
effective length of query: 478
effective length of database: 8,513,101
effective search space: 4069262278
effective search space used: 4069262278
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0587.12


