

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0546.13
(1118 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
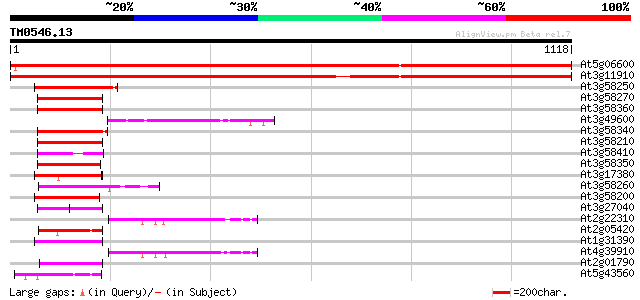
Score E
Sequences producing significant alignments: (bits) Value
At5g06600 ubiquitin carboxyl-terminal hydrolase 1915 0.0
At3g11910 ubiquitin carboxyl-terminal hydrolase like protein 1857 0.0
At3g58250 putative protein 132 8e-31
At3g58270 unknown protein 126 8e-29
At3g58360 putative protein 122 1e-27
At3g49600 putative protein 119 7e-27
At3g58340 putative protein 116 6e-26
At3g58210 putative protein 115 1e-25
At3g58410 putative protein 104 2e-22
At3g58350 putative protein 103 4e-22
At3g17380 unknown protein 103 5e-22
At3g58260 putative protein 101 3e-21
At3g58200 unknown protein 101 3e-21
At3g27040 hypothetical protein 100 4e-21
At2g22310 putative ubiquitin carboxyl terminal hydrolase 100 5e-21
At2g05420 hypothetical protein 99 1e-20
At1g31390 hypothetical protein 98 3e-20
At4g39910 ubiquitin-specific protease (AtUBP3) 96 1e-19
At2g01790 hypothetical protein 95 3e-19
At5g43560 unknown protein 94 3e-19
>At5g06600 ubiquitin carboxyl-terminal hydrolase
Length = 1126
Score = 1915 bits (4962), Expect = 0.0
Identities = 925/1128 (82%), Positives = 1014/1128 (89%), Gaps = 12/1128 (1%)
Query: 1 MTVMTPAPID---------QQEDEEMLVPHTDLPENNHQPMEVVAQPEAAPTVESQPVEE 51
MT+MTP P+D Q EDEEMLVP++DL + QPMEV AA TVE+QP E+
Sbjct: 1 MTMMTPPPVDVISDFYVLQQPEDEEMLVPNSDLVDGPAQPMEVTQPETAASTVENQPAED 60
Query: 52 PPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLP 111
PP +FTW I NFSR N +K YS+VFVVGGYKWR+LIFPKGNNVD+LSMYLDV+D+ +LP
Sbjct: 61 PPTLKFTWTIPNFSRQNTRKHYSDVFVVGGYKWRILIFPKGNNVDHLSMYLDVSDAASLP 120
Query: 112 YGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLND 171
YGWSRYAQFSLAVVNQI +YTVRK+TQHQFNARESDWGFTSFMPL ELYDPSRGYL+ND
Sbjct: 121 YGWSRYAQFSLAVVNQIHTRYTVRKETQHQFNARESDWGFTSFMPLSELYDPSRGYLVND 180
Query: 172 TLVVEAEVLVRRIVDYWTYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMP 231
T++VEAEV VR+++DYW+YDSKKETG+VGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMP
Sbjct: 181 TVLVEAEVAVRKVLDYWSYDSKKETGFVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMP 240
Query: 232 TTENDMPAGSIPLALQSLFYKLQYSDTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCE 291
TTEND P SIPLALQSLFYKLQY+DTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCE
Sbjct: 241 TTENDAPTASIPLALQSLFYKLQYNDTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCE 300
Query: 292 KLEDKMKGTVVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFD 351
KLEDKMKGTVVEGTIQ+LFEGHHMNYIECINVD+KSTRKESFYDLQLDVKGC DVYASFD
Sbjct: 301 KLEDKMKGTVVEGTIQQLFEGHHMNYIECINVDFKSTRKESFYDLQLDVKGCKDVYASFD 360
Query: 352 KYVEVEPLEGDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRY 411
KYVEVE LEGDNKYHAE +GLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRY
Sbjct: 361 KYVEVERLEGDNKYHAEGHGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRY 420
Query: 412 EFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKF 471
EFPLELDLDR+DGKYLSPDADR+VRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKF
Sbjct: 421 EFPLELDLDREDGKYLSPDADRSVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKF 480
Query: 472 DDERVTKEDTKRALEEQYGGEEELPQTNPGFNNT-PFKFTKYSNAYMLVYIREADKDKVI 530
DDERVTKED KRALEEQYGGEEELPQTNPGFNN PFKFTKYSNAYMLVYIRE+DKDK+I
Sbjct: 481 DDERVTKEDLKRALEEQYGGEEELPQTNPGFNNNPPFKFTKYSNAYMLVYIRESDKDKII 540
Query: 531 CNVDEKDIAEHLRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLV 590
CNVDEKDIAEHLR RLKKEQEEKE K++ KA+AHLYTIIKVAR+EDLKEQIGKDIYFDLV
Sbjct: 541 CNVDEKDIAEHLRVRLKKEQEEKEDKRRYKAQAHLYTIIKVARDEDLKEQIGKDIYFDLV 600
Query: 591 DHDKVRSFRVQKQMSFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQ 650
DHDKVRSFR+QKQ F FKEEVAKEFG+PVQ QRFW+WAKRQNHTYRPNRPLTP EE Q
Sbjct: 601 DHDKVRSFRIQKQTPFQQFKEEVAKEFGVPVQLQRFWIWAKRQNHTYRPNRPLTPQEELQ 660
Query: 651 SVGQVREVSNKVHNAELKLFLEVELGPDLRPIAPSDKTKDDILLFFKLYDPEKEELRYVG 710
VGQ+RE SNK + AELKLFLEVE DLRPI P +K+K+DILLFFKLYDPEK LRY G
Sbjct: 661 PVGQIREASNKANTAELKLFLEVEHLQDLRPIPPPEKSKEDILLFFKLYDPEKAVLRYAG 720
Query: 711 RLFVKSTGKPSEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDG 770
RL VKS+ KP +I +LNEM G+ PDEEI L+EEIKFEP VMCE +DKK +FR Q+EDG
Sbjct: 721 RLMVKSSSKPMDITGKLNEMVGFAPDEEIELFEEIKFEPCVMCEHLDKKTSFRLCQIEDG 780
Query: 771 DIICFQKAPAMDSEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTYDD 830
DIICFQK P ++ E YP VPS+LEYV NRQ+V FR+L+KPKED+F LE+S+ +TYDD
Sbjct: 781 DIICFQK-PLVNKEIECLYPAVPSFLEYVQNRQLVRFRALEKPKEDEFVLELSKQHTYDD 839
Query: 831 VVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEI 890
VVEKVA++L LDDPSK+RLT HNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYE+
Sbjct: 840 VVEKVAEKLGLDDPSKLRLTSHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEV 899
Query: 891 LDIPLPELQGLKTLKVAFYHATKDEVVSHTIRLPKQSTVGDVLDDLKTKVELSHPNAELR 950
LDIPLPELQGLKTLKVAF+HATK+EVV H IRLPKQSTVGDV+++LKTKVELSHP+AELR
Sbjct: 900 LDIPLPELQGLKTLKVAFHHATKEEVVIHNIRLPKQSTVGDVINELKTKVELSHPDAELR 959
Query: 951 LLEVFYHKIYKVFPPNEKIETINDQYWTLRAEEVPEEEKNLGPHDRLIHVYHFTKDTSQN 1010
LLEVFYHKIYK+FP E+IE INDQYWTLRAEE+PEEEKN+GP+DRLI VYHF K+T QN
Sbjct: 960 LLEVFYHKIYKIFPSTERIENINDQYWTLRAEEIPEEEKNIGPNDRLILVYHFAKETGQN 1019
Query: 1011 QMQIQNFGEPFFLVIREGETLTEIKVRIQKKLQVPDDEFEKWKFAFFALGRPEYLQDSDI 1070
Q Q+QNFGEPFFLVI EGETL EIK RIQKKL V D++F KWKFAF ++GRPEYLQD+D+
Sbjct: 1020 Q-QVQNFGEPFFLVIHEGETLEEIKNRIQKKLHVSDEDFAKWKFAFMSMGRPEYLQDTDV 1078
Query: 1071 VSNRFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1118
V NRFQRRDVYGA+EQYLGLEH D PKR+YA NQNRH +EKPVKIYN
Sbjct: 1079 VYNRFQRRDVYGAFEQYLGLEHADTTPKRAYAANQNRHAYEKPVKIYN 1126
>At3g11910 ubiquitin carboxyl-terminal hydrolase like protein
Length = 1089
Score = 1857 bits (4810), Expect = 0.0
Identities = 889/1118 (79%), Positives = 991/1118 (88%), Gaps = 29/1118 (2%)
Query: 1 MTVMTPAPIDQQEDEEMLVPHTDLPENNHQPMEVVAQPEAAPTVESQPVEEPPQSRFTWR 60
MT+MTP P+DQQEDEEMLVP+ DL E QPMEV AA VE+ P E+PP +FTW
Sbjct: 1 MTMMTPPPLDQQEDEEMLVPNPDLVEGP-QPMEVAQTDPAATAVENPPPEDPPSLKFTWT 59
Query: 61 IDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWSRYAQF 120
I F+R+N +K YS+VFVVGGYKWR+LIFPKGNNVD+LSMYLDVAD+ NLPYGWSRY+QF
Sbjct: 60 IPMFTRLNTRKHYSDVFVVGGYKWRILIFPKGNNVDHLSMYLDVADAANLPYGWSRYSQF 119
Query: 121 SLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVVEAEVL 180
SLAVVNQ+ N+Y++RK+TQHQFNARESDWGFTSFMPL ELY+P+RGYL+NDT+++EAEV
Sbjct: 120 SLAVVNQVNNRYSIRKETQHQFNARESDWGFTSFMPLSELYEPTRGYLVNDTVLIEAEVA 179
Query: 181 VRRIVDYWTYDSKKETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAG 240
VR+++DYW+YDSKKETG+VGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTEND P
Sbjct: 180 VRKVLDYWSYDSKKETGFVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDAPTA 239
Query: 241 SIPLALQSLFYKLQYSDTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGT 300
SIPLALQSLFYKLQY+DTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGT
Sbjct: 240 SIPLALQSLFYKLQYNDTSVATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMKGT 299
Query: 301 VVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLE 360
VVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGC DVYASFDKYVEVE LE
Sbjct: 300 VVEGTIQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDVKGCKDVYASFDKYVEVERLE 359
Query: 361 GDNKYHAEQYGLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLD 420
GDNKYHAE + LQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPL+LDLD
Sbjct: 360 GDNKYHAEGHDLQDAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLQLDLD 419
Query: 421 RDDGKYLSPDADRNVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKED 480
R+DG+YLSPDAD++VRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKED
Sbjct: 420 REDGRYLSPDADKSVRNLYTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKED 479
Query: 481 TKRALEEQYGGEEELPQTNPGFNNTPFKFTKYSNAYMLVYIREADKDKVICNVDEKDIAE 540
KRALEEQYGGEEELPQ NPGFNN PFKFTKYSNAYMLVYIRE+DKDK+ICNVDEKDIAE
Sbjct: 480 VKRALEEQYGGEEELPQNNPGFNNPPFKFTKYSNAYMLVYIRESDKDKIICNVDEKDIAE 539
Query: 541 HLRERLKKEQEEKEHKKKEKAEAHLYTIIKVARNEDLKEQIGKDIYFDLVDHDKVRSFRV 600
HLR RLKKEQEEKE K+K KA+AHL+T IKVAR++D+ EQIGK+IYFDLVDH+KVRSFR+
Sbjct: 540 HLRVRLKKEQEEKEDKRKYKAQAHLFTTIKVARDDDITEQIGKNIYFDLVDHEKVRSFRI 599
Query: 601 QKQMSFNLFKEEVAKEFGIPVQFQRFWLWAKRQNHTYRPNRPLTPAEEAQSVGQVREVSN 660
QKQ F FKEEVAKEFG+PVQ QRFW+WAKRQNHTYRPNRPL+P EE Q+
Sbjct: 600 QKQTPFQQFKEEVAKEFGVPVQLQRFWIWAKRQNHTYRPNRPLSPNEELQT--------- 650
Query: 661 KVHNAELKLFLEVELGPDLRPIAPSDKTKDDILLFFKLYDPEKEELRYVGRLFVKSTGKP 720
D PI P +KT +DILLFFKLYDPE LRYVGRL VKS+ KP
Sbjct: 651 -----------------DDLPIPPPEKTSEDILLFFKLYDPENAVLRYVGRLMVKSSSKP 693
Query: 721 SEILTRLNEMAGYDPDEEIGLYEEIKFEPNVMCEPIDKKLTFRASQLEDGDIICFQKAPA 780
+I+ +LN+MAG+ PDEEI L+EEIKFEP VMCE IDKK +FR Q+EDGDIIC+QK P
Sbjct: 694 MDIVGQLNKMAGFAPDEEIELFEEIKFEPCVMCEQIDKKTSFRLCQIEDGDIICYQK-PL 752
Query: 781 MDSEEHVRYPDVPSYLEYVHNRQVVHFRSLDKPKEDDFCLEMSRLYTYDDVVEKVAQQLN 840
E RYPDVPS+LEYV NR++V FR+L+KPKED+F +E+S+L+TYDDVVE+VA++L
Sbjct: 753 SIEESEFRYPDVPSFLEYVQNRELVRFRTLEKPKEDEFTMELSKLHTYDDVVERVAEKLG 812
Query: 841 LDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEILDIPLPELQG 900
LDDPSK+RLT HNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYE+LDIPLPELQG
Sbjct: 813 LDDPSKLRLTSHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQTSDILYYEVLDIPLPELQG 872
Query: 901 LKTLKVAFYHATKDEVVSHTIRLPKQSTVGDVLDDLKTKVELSHPNAELRLLEVFYHKIY 960
LKTLKVAF+ ATKDEV+ H IRLPKQSTVGDV+++LKTKVELSH +AELRLLEVF+HKIY
Sbjct: 873 LKTLKVAFHSATKDEVIIHNIRLPKQSTVGDVINELKTKVELSHQDAELRLLEVFFHKIY 932
Query: 961 KVFPPNEKIETINDQYWTLRAEEVPEEEKNLGPHDRLIHVYHFTKDTSQNQMQIQNFGEP 1020
K+FP E+IE INDQYWTLRAEE+PEEEKN+GP+DRLIHVYHFTK+ QNQ Q+QNFGEP
Sbjct: 933 KIFPSTERIENINDQYWTLRAEEIPEEEKNIGPNDRLIHVYHFTKEAGQNQ-QVQNFGEP 991
Query: 1021 FFLVIREGETLTEIKVRIQKKLQVPDDEFEKWKFAFFALGRPEYLQDSDIVSNRFQRRDV 1080
FFLVI EGETL EIK RIQKKL VPD++F KWKFA F++GRP+YL D+D+V NRFQRRDV
Sbjct: 992 FFLVIHEGETLEEIKTRIQKKLHVPDEDFAKWKFASFSMGRPDYLLDTDVVYNRFQRRDV 1051
Query: 1081 YGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1118
YGAWEQYLGLEH DNAPKR+YA NQNRH +EKPVKIYN
Sbjct: 1052 YGAWEQYLGLEHIDNAPKRAYAANQNRHAYEKPVKIYN 1089
>At3g58250 putative protein
Length = 317
Score = 132 bits (333), Expect = 8e-31
Identities = 74/171 (43%), Positives = 108/171 (62%), Gaps = 7/171 (4%)
Query: 50 EEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNV--DYLSMYLDVADS 107
++ + +F+W I NFS + +K+YS+ FV+ G +WR+L FPKGN+ D+LS+YLDVA+S
Sbjct: 4 DQADKKKFSWVIKNFSSLQSEKIYSDQFVIDGCRWRLLAFPKGNDTKSDHLSLYLDVAES 63
Query: 108 TNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGY 167
+LP GW R+AQFS +VN I K + RK+T H F + SDWGFT+ +PL EL G+
Sbjct: 64 ESLPCGWRRHAQFSFTIVNHIPEKCSQRKETIHWFCEKVSDWGFTNLVPLIELKAEDSGF 123
Query: 168 LLNDTL--VVEAEVL-VRRIVDYWTYDSKKETGYVGLKNQGATCYMNSLLQ 215
L+ L VVE EVL V +++ +S G+ L +Q Y+ SL +
Sbjct: 124 LVKGELKIVVEIEVLEVIGLLNVSESESMDVNGFHVLPSQAK--YVKSLFE 172
>At3g58270 unknown protein
Length = 343
Score = 126 bits (316), Expect = 8e-29
Identities = 60/131 (45%), Positives = 87/131 (65%)
Query: 55 SRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGW 114
++FTW I NFS +K YS+ F V G KWR+L FPKGN V+ LS+YL VA S LP GW
Sbjct: 7 NKFTWVIKNFSSQQSRKNYSDEFFVDGCKWRLLAFPKGNGVEKLSLYLAVAGSEFLPDGW 66
Query: 115 SRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLV 174
R+A F +VVNQ+ ++ + ++T++ F+A SDWGFTS + L +L+D G+L+N L
Sbjct: 67 RRHAYFHFSVVNQLSDELSQARETKNWFDASTSDWGFTSMLSLKKLHDKDGGFLVNGELK 126
Query: 175 VEAEVLVRRIV 185
+ +V V ++
Sbjct: 127 IVVDVSVLEVI 137
>At3g58360 putative protein
Length = 298
Score = 122 bits (305), Expect = 1e-27
Identities = 57/130 (43%), Positives = 84/130 (63%)
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWS 115
+ TW I+NFS ++ KK+YS+ F+VGG KWR L++PKGNNVDYL +YL+VAD +L W
Sbjct: 8 KITWAIENFSSLHSKKIYSDPFIVGGCKWRFLVYPKGNNVDYLFLYLEVADYESLSPEWR 67
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
R+A++ L VVNQ K + + + Q F+ + WG S PL E+ G+L+N L +
Sbjct: 68 RHARYLLNVVNQNSVKRSKQNEEQKWFDVQSPRWGRLSMFPLNEINAKDSGFLVNGELKI 127
Query: 176 EAEVLVRRIV 185
AE+ V ++
Sbjct: 128 VAEIEVLEVI 137
>At3g49600 putative protein
Length = 1672
Score = 119 bits (299), Expect = 7e-27
Identities = 103/357 (28%), Positives = 158/357 (43%), Gaps = 34/357 (9%)
Query: 195 ETGYVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQ 254
E+ GL N GATCY NS+LQ LY FR+ V+ + + + + + LF +L
Sbjct: 707 ESTPAGLTNLGATCYANSILQCLYMNTAFREGVFSVEV--HVLKQNPVLDQIARLFAQLH 764
Query: 255 YSDTS-VATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKM--KGTVVEGTIQKLFE 311
S S V + K+ D+ +Q D E +L LE + G + +Q LF
Sbjct: 765 ASQKSFVDSDAFVKTL---ELDNGVQQDTHEFLTLLLSLLERCLLHSGVKAKTIVQDLFS 821
Query: 312 G---HHMNYIECINVDYKSTRKESFYDLQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAE 368
G H +C S++ E FY L+L+VKG + AS + Y+ +E L GDN+Y
Sbjct: 822 GSVSHVTTCSKCGRDSEASSKMEDFYALELNVKGLKSLDASLNDYLSLEQLNGDNQYFCG 881
Query: 369 QYGLQ-DAKKGVLFIDFPPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYL 427
+ DA + + PPV+ QLKR + KI + FP LD+ G L
Sbjct: 882 SCNARVDATRCIKLRTLPPVITFQLKRCIFLPKTTAKKKITSSFSFPQVLDM----GSRL 937
Query: 428 SPDADRNVRNLYTLHSVLVHSG-GVHGGHYYAFIRPTLSDQWYKFDDERVTK-------E 479
+ + + Y L +VL+H G V+ GHY A I+ + W++FDDE V++ E
Sbjct: 938 AESSQNKL--TYDLSAVLIHKGSAVNSGHYVAHIKDEKTGLWWEFDDEHVSELGKRPCNE 995
Query: 480 DTKRALEEQYGGEEELPQTNPGFNN--------TPFKFTKYSNAYMLVYIREADKDK 528
+ + + G G + + S+AYML+Y DK +
Sbjct: 996 ASSSTPQSESNGTASSGNITDGIQSGSSDCRSAIKSEVFSSSDAYMLMYSLRCDKQE 1052
>At3g58340 putative protein
Length = 325
Score = 116 bits (291), Expect = 6e-26
Identities = 55/140 (39%), Positives = 85/140 (60%), Gaps = 2/140 (1%)
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWS 115
+F W I NFS +N ++ +S V+G KWR++ FPKG DYLS+YL+VAD +LP GW
Sbjct: 8 KFCWEIKNFSSLNSERCHSVPVVIGDCKWRLVAFPKGYKADYLSLYLEVADFKSLPSGWR 67
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
RY +F +VNQ+ + +V+++TQ F+ WGF + + L EL G+L+N +++
Sbjct: 68 RYVKFRACIVNQLSQELSVQQETQRWFDQNAPGWGFENMLLLTELNAKDGGFLVNGQVMI 127
Query: 176 EAEVLVRRIVDYWTYDSKKE 195
AEV ++ T D +E
Sbjct: 128 VAEVEFLEVIG--TLDESEE 145
>At3g58210 putative protein
Length = 330
Score = 115 bits (288), Expect = 1e-25
Identities = 55/133 (41%), Positives = 87/133 (65%), Gaps = 2/133 (1%)
Query: 55 SRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNV--DYLSMYLDVADSTNLPY 112
++FTW I NFS + + S FV+GG KWR+L++P+G N D+LS++L+VAD +LP
Sbjct: 7 NKFTWVIQNFSSSQSRVVPSNQFVIGGCKWRLLVYPEGFNKSGDHLSLFLEVADPRSLPP 66
Query: 113 GWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDT 172
GWSR+A++ L +VNQ +K + R + FN + WG ++ +PL +L+ G+L+ND
Sbjct: 67 GWSRHARYLLTIVNQHSDKISKRNEATKWFNQKIPGWGLSAMIPLTKLHAKDGGFLVNDE 126
Query: 173 LVVEAEVLVRRIV 185
L + AEV V ++
Sbjct: 127 LKIVAEVNVLEVI 139
>At3g58410 putative protein
Length = 328
Score = 104 bits (260), Expect = 2e-22
Identities = 53/131 (40%), Positives = 72/131 (54%), Gaps = 17/131 (12%)
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWS 115
+F W I NFS + KK YS F +G KWR+ I+PKGNN DYLS++L+VAD +LP GW
Sbjct: 29 KFAWVIKNFSSLQCKKFYSVPFQIGDCKWRLSIYPKGNNCDYLSLFLEVADFKSLPSGWR 88
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
RY + L +V Q WGF +PL +L+D G+L+N L++
Sbjct: 89 RYVKLRLYIVKQ-----------------EMWGWGFLYMLPLTKLHDEKEGFLVNGELMI 131
Query: 176 EAEVLVRRIVD 186
AEV +D
Sbjct: 132 VAEVDALGFID 142
>At3g58350 putative protein
Length = 355
Score = 103 bits (258), Expect = 4e-22
Identities = 51/129 (39%), Positives = 80/129 (61%), Gaps = 3/129 (2%)
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKG-NNVDYLSMYLDVADSTNLPYGW 114
+ TW I NF+ + +YS+ FVVGG KW + +PKG NN + LS++L VA T+LP GW
Sbjct: 62 KITWTIKNFASLLSDLIYSDHFVVGGCKWHLRAYPKGYNNANSLSLFLGVAVPTSLPSGW 121
Query: 115 SRYAQFSLAVVNQIQNKYTVRK--DTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDT 172
R+ +F L +VNQ+ +K + K + + F+ + ++WG +S PL E++ G+LLN
Sbjct: 122 RRHTKFRLTLVNQLSDKLSQSKLNELEQWFDEKTTNWGLSSMCPLNEIHAKDSGFLLNGE 181
Query: 173 LVVEAEVLV 181
L + E+ V
Sbjct: 182 LKIVVEIKV 190
>At3g17380 unknown protein
Length = 309
Score = 103 bits (257), Expect = 5e-22
Identities = 47/142 (33%), Positives = 88/142 (61%), Gaps = 7/142 (4%)
Query: 49 VEEPPQSRFTWRIDNFSRMN---VKKLYSEVFVVGGYKWRVLIFPKGNNV----DYLSMY 101
+ + P + + +I++FS + +++ +E F GGYKW+++++P GN D++S+Y
Sbjct: 14 ISDAPPTHYMVKIESFSLLTKHAIERYETESFEAGGYKWKLVLYPNGNKSKNTKDHVSVY 73
Query: 102 LDVADSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELY 161
L +ADS++L GW YA F L +++Q ++ Y + + + +F++ + +WGF F+P G
Sbjct: 74 LSLADSSSLSPGWEVYAVFRLYLLDQNKDNYLILQGNERRFHSVKREWGFDKFIPTGTFS 133
Query: 162 DPSRGYLLNDTLVVEAEVLVRR 183
D S GYL+ DT + A+V V +
Sbjct: 134 DASNGYLMEDTCMFGADVFVSK 155
Score = 75.5 bits (184), Expect = 2e-13
Identities = 38/139 (27%), Positives = 74/139 (52%), Gaps = 4/139 (2%)
Query: 49 VEEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNV---DYLSMYLDVA 105
+++ S+ W+I+NFS+++ + S F G KW++ +P G +LS+YL +
Sbjct: 168 IKDATSSKHVWKIENFSKLDKESYDSNAFFAGDRKWKIEFYPTGTKQGTGTHLSIYLTLV 227
Query: 106 DSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSR 165
D + G + +F++ + +Q+Q ++ K T+ F+ S+ G+ ++ + P+
Sbjct: 228 DPETISDGTKIFVEFTIRIFDQLQGRHIAGKVTK-WFSRSSSEHGWVKYVSMVYFTQPNS 286
Query: 166 GYLLNDTLVVEAEVLVRRI 184
G LL D +VEA+V V I
Sbjct: 287 GLLLKDVCLVEADVCVHGI 305
>At3g58260 putative protein
Length = 321
Score = 101 bits (251), Expect = 3e-21
Identities = 77/278 (27%), Positives = 129/278 (45%), Gaps = 58/278 (20%)
Query: 57 FTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVD---YLSMYLDVADST-NLPY 112
FTW I N S + ++ S++FVVGG KWR++ +P+ N+ D LS+YL V D +LP
Sbjct: 9 FTWVIKNLSTLQGLEVRSKIFVVGGCKWRLIAYPEVNDADGYLSLSVYLGVPDCCESLPS 68
Query: 113 GWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDT 172
GW R+A+FSL +VNQ+ + ++TQ F+ WGF + L ++ D G+L+ND
Sbjct: 69 GWKRHAKFSLTIVNQLSEGLSQVQETQAWFDENAPGWGFPPMLNLKDVSDKYGGFLVNDE 128
Query: 173 LVVEAEVLVRRIVDYWTYDSKKET---------------------------GYVGLKNQG 205
++V V V +V E+ +KNQ
Sbjct: 129 VMVAVAVDVIEVVGSLDAPEMSESMDIKGFKVLPSQVKSVNRLFESHPDIASKFSIKNQS 188
Query: 206 -ATCYMNSLL---QTLYHIPYFRKAVYHMPTTENDMPAGSIPLA-LQSLFYKLQYSDTSV 260
T YMN LL +TL+ P M +E+D+ LA ++S+ +KL + + +
Sbjct: 189 LKTAYMNVLLCLAETLHQSP--------MEISEDDLSDAKTTLAYMKSVGFKLDWLEKKL 240
Query: 261 ATKELTKSFGWDTYDSFMQHDVQELNRVLCEKLEDKMK 298
D + +E +++ + +E+++K
Sbjct: 241 --------------DELFEKKKEEADKIRMQNIEEELK 264
>At3g58200 unknown protein
Length = 319
Score = 101 bits (251), Expect = 3e-21
Identities = 47/132 (35%), Positives = 81/132 (60%), Gaps = 1/132 (0%)
Query: 49 VEEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKG-NNVDYLSMYLDVADS 107
+E+ ++F W I NFS + ++++S++FVVG KWR++ +PKG + S++L V D
Sbjct: 1 MEKEADNKFRWVIKNFSSLGSERVFSDIFVVGSCKWRLMAYPKGVRDNRCFSLFLVVTDF 60
Query: 108 TNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGY 167
LP W R+ + L VVNQ+ + ++ K+TQ F+ + WGF + +PL EL + G+
Sbjct: 61 KTLPCDWKRHTRLRLNVVNQLSEELSILKETQMWFDQKTPAWGFLAMLPLTELKAENGGF 120
Query: 168 LLNDTLVVEAEV 179
L+N+ + + EV
Sbjct: 121 LVNEEVKIVVEV 132
>At3g27040 hypothetical protein
Length = 358
Score = 100 bits (250), Expect = 4e-21
Identities = 48/131 (36%), Positives = 79/131 (59%), Gaps = 1/131 (0%)
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNV-DYLSMYLDVADSTNLPYGW 114
+FTW I N++ + +YS+ F G KWR+L FPKGNN+ DY +Y+ V +S +LP GW
Sbjct: 96 KFTWVIKNYNSLGSGSVYSDTFKAGRCKWRLLAFPKGNNIYDYFFLYICVPNSESLPSGW 155
Query: 115 SRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLV 174
R A+ S +VNQI + +++ + F+ +++ GF S L E+ +G+L+N +
Sbjct: 156 RRRAKVSFTMVNQIPGGLSQQREAVYWFDEKDTTHGFESMFLLSEIQSSDKGFLVNGEVK 215
Query: 175 VEAEVLVRRIV 185
+ AEV V ++
Sbjct: 216 IVAEVDVLEVI 226
Score = 55.1 bits (131), Expect = 2e-07
Identities = 25/66 (37%), Positives = 40/66 (59%), Gaps = 2/66 (3%)
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNN--VDYLSMYLDVADSTNLPYG 113
+F W I NF+ ++ ++YS+ F G KWR++ +PK + S++L V DS +LP G
Sbjct: 9 KFAWVIKNFNSLDTTRVYSDTFKAGRCKWRLVAYPKRRDRYTTSFSLFLCVPDSESLPSG 68
Query: 114 WSRYAQ 119
W R A+
Sbjct: 69 WRRRAK 74
>At2g22310 putative ubiquitin carboxyl terminal hydrolase
Length = 365
Score = 100 bits (249), Expect = 5e-21
Identities = 86/336 (25%), Positives = 146/336 (42%), Gaps = 50/336 (14%)
Query: 198 YVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKL--QY 255
Y G +N G TCY NS+LQ LY FR+ + ++ L LF ++ Q
Sbjct: 22 YFGFENFGNTCYCNSVLQALYFCAPFREQLLEHYANNKADAEENLLTCLADLFSQISSQK 81
Query: 256 SDTSVAT-----------KELTKSFGWDTYDSFMQHDVQELNRVL--------------- 289
T V EL +S+ F+ + + EL +L
Sbjct: 82 KKTGVIAPKRFVQRLKKQNELFRSYMHQDAHEFLNYLLNELVEILEKETQATKADNETSS 141
Query: 290 -CEKLEDKMKGTVVEGT--------IQKLFEGHHMNYIECINVDYKSTRKESFYDLQLDV 340
EK+ + +K + G + K+F+G N C+ + + R E+F DL LD+
Sbjct: 142 SPEKIANVLKAPLANGVHKEPIVTWVHKIFQGILTNETRCLRCETVTARDETFLDLSLDI 201
Query: 341 KGCHDVYASFDKYVEVEPLEGDNKYHAEQ-YGLQDAKKGVLFIDFPPVLQLQLKRFEYDF 399
+ + + + E L ++K+ ++ LQ+A+K + P +L + LKRF+Y
Sbjct: 202 EQNSSITSCLKNFSSTETLHAEDKFFCDKCCSLQEAQKRMKIKKPPHILVIHLKRFKYME 261
Query: 400 MRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSG-GVHGGHYYA 458
K++ R FPLEL L +Y+ + Y+L +V+VH G G + GHY +
Sbjct: 262 QLGRYKKLSYRVVFPLELKLSNTVDEYVDIE--------YSLFAVVVHVGSGPNHGHYVS 313
Query: 459 FIRPTLSDQWYKFDDERVTKEDTKRALEEQYGGEEE 494
++ + W FDDE V + + A++ +G +E
Sbjct: 314 LVKS--HNHWLFFDDESVEIIE-ESAVQTFFGSSQE 346
>At2g05420 hypothetical protein
Length = 297
Score = 99.4 bits (246), Expect = 1e-20
Identities = 51/136 (37%), Positives = 83/136 (60%), Gaps = 9/136 (6%)
Query: 58 TWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNN--------VDYLSMYLDVADSTN 109
TW I+NFS + ++S+ FVVG KWR+ +PKGN + L++YL+VA+S +
Sbjct: 11 TWVIENFSSLQSASIHSDQFVVGDCKWRLKAYPKGNEKATYLAYRANNLALYLNVANSKS 70
Query: 110 LPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLL 169
P GW+R+ +FSL +VNQ K + ++QH F+ + + GF + +PL L+ + G+L+
Sbjct: 71 FPIGWTRHTKFSLTLVNQKSEKLSKLTESQHWFDHKSTSRGFPAMIPLTNLH-TNEGFLV 129
Query: 170 NDTLVVEAEVLVRRIV 185
N L + A+V V +V
Sbjct: 130 NGELTLVAKVEVLEVV 145
>At1g31390 hypothetical protein
Length = 268
Score = 97.8 bits (242), Expect = 3e-20
Identities = 51/140 (36%), Positives = 82/140 (58%), Gaps = 3/140 (2%)
Query: 49 VEEPPQSRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDY---LSMYLDVA 105
+E+ + + TW I NFS + + + S++FVVG KW ++ +PKGN LS+YL+VA
Sbjct: 1 MEDQYEKKITWTIKNFSFVQSQAIDSDIFVVGDSKWHLVAYPKGNGESTNKCLSLYLNVA 60
Query: 106 DSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSR 165
D +LP GW R+ ++ L VVNQ+ K + ++ Q F GF + +PL +L D +
Sbjct: 61 DFQSLPNGWKRHIKYRLTVVNQMSEKLSEQEVIQGWFYKNFHISGFQTMLPLSKLLDKNG 120
Query: 166 GYLLNDTLVVEAEVLVRRIV 185
G+L+N + + EV V +V
Sbjct: 121 GFLVNGDVKIVVEVGVLEVV 140
>At4g39910 ubiquitin-specific protease (AtUBP3)
Length = 371
Score = 95.5 bits (236), Expect = 1e-19
Identities = 88/341 (25%), Positives = 148/341 (42%), Gaps = 55/341 (16%)
Query: 198 YVGLKNQGATCYMNSLLQTLYHIPYFRKAVYHMPTTENDMPAG--SIPLALQSLFYKL-- 253
Y G +N G TCY NS+LQ LY FR+ + T+ + ++ L LF ++
Sbjct: 22 YFGFENFGNTCYCNSVLQALYFCVPFREQLLEYYTSNKSVADAEENLMTCLADLFSQISS 81
Query: 254 QYSDTSVAT-----------KELTKSFGWDTYDSFMQHDVQELNRVL------------- 289
Q T V EL +S+ F+ + + E+ +L
Sbjct: 82 QKKKTGVIAPKRFVQRLKKQNELFRSYMHQDAHEFLNYLLNEVVDILEKEAKATKTEHET 141
Query: 290 -----CEKLEDKMKGTVVEGTIQK---------LFEGHHMNYIECINVDYKSTRKESFYD 335
EK+ + +K G + K +F+G N C+ + + R E+F D
Sbjct: 142 SSSSSPEKIANGLKVPQANGVVHKEPIVTWVHNIFQGILTNETRCLRCETVTARDETFLD 201
Query: 336 LQLDVKGCHDVYASFDKYVEVEPLEGDNKYHAEQ-YGLQDAKKGVLFIDFPPVLQLQLKR 394
L LD++ + + + E L ++K+ ++ LQ+A+K + P +L + LKR
Sbjct: 202 LSLDIEQNSSITSCLKNFSSTETLHAEDKFFCDKCCSLQEAQKRMKIKKPPHILVIHLKR 261
Query: 395 FEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVHSG-GVHG 453
F+Y K++ R FPLEL L + P AD Y+L +V+VH G G +
Sbjct: 262 FKYIEQLGRYKKLSYRVVFPLELKLSNT----VEPYADVE----YSLFAVVVHVGSGPNH 313
Query: 454 GHYYAFIRPTLSDQWYKFDDERVTKEDTKRALEEQYGGEEE 494
GHY + ++ + W FDDE V + + A++ +G +E
Sbjct: 314 GHYVSLVKS--HNHWLFFDDENVEMIE-ESAVQTFFGSSQE 351
>At2g01790 hypothetical protein
Length = 269
Score = 94.7 bits (234), Expect = 3e-19
Identities = 49/130 (37%), Positives = 78/130 (59%), Gaps = 3/130 (2%)
Query: 59 WRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNN---VDYLSMYLDVADSTNLPYGWS 115
W I+NFS ++ ++YS++FVVGG KW +L P+GNN DY S+YL V DS LP GW
Sbjct: 11 WVINNFSFLDSDRVYSDIFVVGGCKWCLLALPEGNNNYIYDYFSLYLCVPDSEYLPSGWR 70
Query: 116 RYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVV 175
R A+ S +VNQ+ + + +++ + F+ + + GF S L +G+L+N + +
Sbjct: 71 RRAKVSFTMVNQVTGELSQQQEGVYWFDEKNTTQGFGSMFRLLVFQSSYKGFLVNGEVDI 130
Query: 176 EAEVLVRRIV 185
AEV V ++
Sbjct: 131 VAEVDVVEVI 140
>At5g43560 unknown protein
Length = 1055
Score = 94.4 bits (233), Expect = 3e-19
Identities = 65/199 (32%), Positives = 107/199 (53%), Gaps = 28/199 (14%)
Query: 9 IDQQEDEEMLVPHTDLPENNH----QPMEVVAQPEAAPTVES-QPVEEPPQ--------- 54
+ + +E+ + L EN++ Q E +A+ ++ VE+ P PP
Sbjct: 1 MSESTNEDSGAGRSSLEENSNGQRSQSEEAIAEWRSSEQVENGTPSTSPPYWDIDDDDDF 60
Query: 55 --------SRFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNV-DYLSMYLDVA 105
+ TW I+ FS +N ++L +VF VGGYKW +LI+P+G +V ++LS++L VA
Sbjct: 61 GSKPSQLFGKNTWTIEKFSDINKRELRGDVFEVGGYKWYILIYPQGCDVCNHLSLFLCVA 120
Query: 106 DSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSR 165
L GWS +AQF++AV N+ K + DT H+F +E DWG+ F+ L +L
Sbjct: 121 HHEKLLPGWSHFAQFTIAVSNK-DPKKSKHSDTLHRFWKKEHDWGWKKFIELPKL---KE 176
Query: 166 GYLLND-TLVVEAEVLVRR 183
G++ + L ++A+V V R
Sbjct: 177 GFIDDSGCLTIKAQVQVIR 195
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,092,022
Number of Sequences: 26719
Number of extensions: 1298303
Number of successful extensions: 5125
Number of sequences better than 10.0: 134
Number of HSP's better than 10.0 without gapping: 96
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 4693
Number of HSP's gapped (non-prelim): 273
length of query: 1118
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1008
effective length of database: 8,379,506
effective search space: 8446542048
effective search space used: 8446542048
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0546.13


