

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0544.8
(1753 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
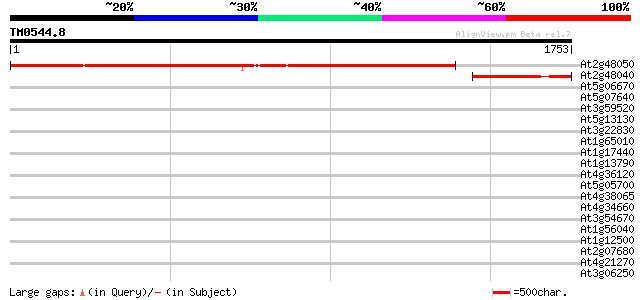
Score E
Sequences producing significant alignments: (bits) Value
At2g48050 hypothetical protein 1554 0.0
At2g48040 unknown protein 364 e-100
At5g06670 kinesin heavy chain-like protein 36 0.17
At5g07640 putative protein 36 0.23
At3g59520 unknown protein 35 0.30
At5g13130 putative protein 35 0.39
At3g22830 putative heat shock protein 34 0.66
At1g65010 hypothetical protein 34 0.66
At1g17440 unknown protein 34 0.86
At1g13790 hypothetical protein 34 0.86
At4g36120 myosin-like protein 33 1.9
At5g05700 arginine-tRNA-protein transferase 1 homolog 32 3.3
At4g38065 hypothetical protein 32 3.3
At4g34660 SH3 domain-containing protein 2 32 3.3
At3g54670 structural maintenance of chromosomes (SMC) - like pro... 32 3.3
At1g56040 hypothetical protein 32 3.3
At1g12500 phosphate/phosphoenolpyruvate translocator protein like 32 3.3
At2g07680 putative ABC transporter 32 4.3
At4g21270 kinesin-related protein katA 31 5.6
At3g06250 unknown protein 31 5.6
>At2g48050 hypothetical protein
Length = 1500
Score = 1554 bits (4024), Expect = 0.0
Identities = 805/1417 (56%), Positives = 1024/1417 (71%), Gaps = 36/1417 (2%)
Query: 1 MKIYAVFVFVSIYSIGVVSSSDMWFPMIIDLQTAFGYNPAASMFQNIWESLAILVVMQLY 60
+K YAV VF+ IY + S +W IDL GYN A + N+WESLA+L+VMQLY
Sbjct: 82 LKAYAVLVFMFIYCLSSFVSLQLWLSGFIDLYFYLGYNSKAPLLDNVWESLAVLIVMQLY 141
Query: 61 SYERRQSKSSGSSDFDLPEVGPFPFTRRLLVRHTEKILFLALFYASLSPISAFGFLYLLG 120
SYERRQS L G F F R L H +KILF ALFYASLSPIS FGF+YLLG
Sbjct: 142 SYERRQSGHYIPGQSSLLHPGVFGFFERFLAWHGQKILFAALFYASLSPISVFGFVYLLG 201
Query: 121 LINCSRLPKSSQIPAKVFLVYSGLLIMVEYLFQMLGDQAEMFPGQKYSQLSLIMGLQLYK 180
L+ C+ PKSS IP+K FL+Y+G L+ EYLFQ+ G QA+MFPGQKY++LS +GL++Y+
Sbjct: 202 LVICTTFPKSSSIPSKSFLIYTGFLVSAEYLFQLWGMQAQMFPGQKYAELSFYLGLRVYE 261
Query: 181 PGFKGLESGFRGKVVVIVACILQYNVFRWLEKMQYVDPNGGKWNEPCPLFNMVDIPNETT 240
PGF G+ESG RGKV+V+ AC LQYNVFRWLE+ + GK+ EPCPLF V + T
Sbjct: 262 PGFWGIESGLRGKVLVVAACTLQYNVFRWLERTSGLTVIKGKYEEPCPLF--VSAEDTTA 319
Query: 241 ACTPQNKQAETST---SATIKR-LARSRSCPTVNSALSQGTD------SGIEGDSTKKLR 290
+ + N + +ST S ++K+ A S S P + +QG G E S++K
Sbjct: 320 SVSSSNGENPSSTDHASISMKQGEATSNSWPFFSPRGNQGAGFLHPKTGGSESGSSRKFS 379
Query: 291 QLHFWESSKDSLKWNRKRLLFLRKERLEMQKTALKVSLKFWIENMFTLFGLEINMIALLL 350
HFW S K+S +WNR+R+L L+KER E QK LK+ LKFWIENMF L+GLEINMIALLL
Sbjct: 380 FGHFWGSIKESHRWNRRRILALKKERFETQKNLLKIYLKFWIENMFNLYGLEINMIALLL 439
Query: 351 ASFAVLNAISLLYIASLAACVLLHRLLIKKLWPIFVFLFASVVTVEYLAIWMR-LTFMNL 409
ASFA+LNAIS++YIA LAACVLL R +I+KLWP+ VFLFAS++ +EY+A W L
Sbjct: 440 ASFALLNAISMVYIALLAACVLLRRRVIQKLWPVVVFLFASILAIEYVATWNSFLPSDQA 499
Query: 410 EIEEHVPCRDCWRVSDIYFSYCRKCWLGIIVDDPRMLIGYYGVFMFSCFKFRADKSSSLT 469
E V C DCW ++ +YF +CR+CWLG+ VDDPR LI Y+ VFM +CFK RAD SS +
Sbjct: 500 PSETSVHCHDCWSIAALYFKFCRECWLGVRVDDPRTLISYFVVFMLACFKLRADHISSFS 559
Query: 470 GLEMYQKIHSQWKSASVLSNLSFETKGYWTFLDHLRLYGYCHLLDFVLSLILITGTLEYD 529
Y ++ SQ K++ V +LSFETK WT LD+LRLY Y HLLD VL LILITGTLEYD
Sbjct: 560 ESSTYHQMKSQRKNSFVWRDLSFETKSMWTVLDYLRLYCYVHLLDVVLILILITGTLEYD 619
Query: 530 FLHLGYLGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQSPFLGDFSEIKSGF 589
LHLGYL FALVF RMRL+ILK+ N IFR+LR+YNF +I+ SLAYQSPF+G+F++ K
Sbjct: 620 ILHLGYLAFALVFARMRLEILKKKNKIFRFLRVYNFVLIIFSLAYQSPFVGNFNDGKCET 679
Query: 590 IEYINELVGFQKYDYGFRITSRSALVEIIIYMLVSLQSYMFSFPEFDYVSTYLEKEQIGA 649
++YI E++GF KYDYGFRIT+RSALVEIII+MLVSLQSYMFS EFDYVS YLE EQIGA
Sbjct: 680 VDYIYEVIGFYKYDYGFRITARSALVEIIIFMLVSLQSYMFSSQEFDYVSRYLEAEQIGA 739
Query: 650 ILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNAS 709
I+R+QEKKAA KT QL+ IR+AEE K R+LQVEKMKSEMLNL+ QLH M++D+N+ AS
Sbjct: 740 IVREQEKKAARKTEQLQQIREAEEKKRQRNLQVEKMKSEMLNLRVQLHRMNSDSNFGVAS 799
Query: 710 LKNDGLRERRNSFLENE------------FRKQALDTNTESIGTLDVNQSLLSEKSESPL 757
+ +GLR R++ +L + RK+ + +S + ++ +S E+
Sbjct: 800 PRTEGLRRRKSPYLIPDSGAASPEIDGVVHRKEEQPIDEDSQYPFEAHEFPVSTTPEALD 859
Query: 758 VPEYMKHPMGSPHGIVEVKERTKNNDFLDLEIRNRYKLPGKRNALVSAVHFIGSGISQAQ 817
PEY SP I EV++ + D + +E + K GK N L+SAV IG G+SQ Q
Sbjct: 860 SPEYSFG--ASPCEITEVQQ---DLDVMSMERERKQKSEGKENPLISAVQLIGDGVSQVQ 914
Query: 818 SLGNMAVNNLVNYLKIEREGQEPTDDSSEDEEYY-EIENQNSGAEPLEPTFSTNSINEHT 876
+GN AVNNLVN+L I E + + SS D+E Y E+E+Q P E + S+
Sbjct: 915 FIGNQAVNNLVNFLNISPENSDTNEQSSVDDEVYDEMESQKRKHTPFE---RSTSLQSDR 971
Query: 877 GSDTSCLQIGILVRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNT 936
SD + QIG + RHIWSRM+SNN++VCYCCFI+ +LWNFSLLS+VYLAALFLYALC +T
Sbjct: 972 SSDGTSFQIGRIFRHIWSRMQSNNDIVCYCCFIIAFLWNFSLLSMVYLAALFLYALCVHT 1031
Query: 937 GPSYIFWVLILIYTEVCILLQYLYQIIIQHTEFEFPMSLLQELGFPAKKITSSFVTSNLP 996
GP++IFWV++L+YTE+ ILLQYLYQIIIQH LL ELGFP ++I SSFV S+LP
Sbjct: 1032 GPTHIFWVIMLMYTEIYILLQYLYQIIIQHCGLSIDAPLLHELGFPTQRIKSSFVVSSLP 1091
Query: 997 FFLIYIFTLLQISITMKGGGWTISVDPSFFRRRNQKSYLEDVKCSTYHERVQRLFLPLSN 1056
FLIYIFTL+Q SIT+K G W S D F RRN + +D+ +R+ +F L +
Sbjct: 1092 LFLIYIFTLIQSSITVKDGDWVPSAD--FTSRRNARGSQKDLTRIRLSQRILDVFKKLRD 1149
Query: 1057 ALKLLTRSLCRYWKSLTWGAETPPYFVQLSMEVNLWPKEGIQPKRIESKINKLLRILRHR 1116
+ KL+ RS+ RYW SLT GAE+PPYFVQ++M+V++WP++GIQP+R+E ++N+LLR++ +
Sbjct: 1150 SAKLVIRSIYRYWISLTRGAESPPYFVQVTMDVHMWPEDGIQPERVECRMNQLLRLVHNE 1209
Query: 1117 RCKEEKIFNLHSASRVRVQSIEKSEENENLCLVIFEVLYASPPNEFAAEEWYSSLTPAAD 1176
RC++ +SRV VQSIE+S E N LV+ EV YASP N ++ EWY SLTPA+D
Sbjct: 1210 RCEKGNPDLCPYSSRVHVQSIERSTETPNEALVVLEVEYASPTNGCSSAEWYKSLTPASD 1269
Query: 1177 VSYEIQKAQHAGILKEIGFPYRILSVIGGGKREVDLYAYIFGADLAVFFLIAIFYESIMK 1236
V+ EI+KAQH+G+ + GFPY ILSVIGGGKR+ DLYAYIFGADL VFFL+AIFY+S++K
Sbjct: 1270 VAKEIRKAQHSGLGEGTGFPYPILSVIGGGKRDTDLYAYIFGADLIVFFLVAIFYQSVIK 1329
Query: 1237 ANSEFLEVYQLEDQFPEDFVLVLMVVFFLIVLDRIIYLCSFATAKVIFYLFSLVLFTYSV 1296
SEF++VYQLEDQFP DFV++LMV+FFLIV+DR+IYLCSFAT KV++YLFSL+LFTY+V
Sbjct: 1330 NKSEFIDVYQLEDQFPFDFVIILMVIFFLIVVDRVIYLCSFATGKVVYYLFSLILFTYAV 1389
Query: 1297 TKYAWDMDPLRRYSGRLALRAIYFTKAISLVLQAIQFHFGMPHKSILYRQFLTSSVSRIN 1356
T+YAW + P ++++ LALR I+ KA+SL LQAIQ +G+PHKS LYRQFLTS VSRIN
Sbjct: 1390 TEYAWSIYPTQQHAAGLALRIIFLAKAMSLALQAIQIRYGLPHKSTLYRQFLTSEVSRIN 1449
Query: 1357 VMGFRLYRAIPFLYELRCVLDWSCTTTSLTMYDWLKL 1393
G+RLYRA+PFLYELRCVLDWSCT TSLTMYDWLK+
Sbjct: 1450 YYGYRLYRALPFLYELRCVLDWSCTATSLTMYDWLKV 1486
>At2g48040 unknown protein
Length = 294
Score = 364 bits (934), Expect = e-100
Identities = 180/312 (57%), Positives = 226/312 (71%), Gaps = 25/312 (8%)
Query: 1446 MYSSGNPTNIANPIKDASARVHIKTLSGRLTLFETTLCEIISWETLEARTSLDPKGYLST 1505
MYSSGNPTNIANPIKDAS ++ +KT+ G+LTL++TTLCE IS + ++ L + +L T
Sbjct: 1 MYSSGNPTNIANPIKDASVQIDLKTVGGKLTLYQTTLCERISGDNIDLGLDLGSQSFLPT 60
Query: 1506 YNEKDIQLICCQSDASTLWLVPPIVQARFMKSLSW--NMDLAFSWEFTRDRPRGKEVVKY 1563
YN+ DIQLICCQ+DAS LWLVP V RF++SL W +MD+ F+W RDRP+GKE VKY
Sbjct: 61 YNKNDIQLICCQADASVLWLVPDTVVTRFIQSLDWDTDMDITFTWVLNRDRPKGKETVKY 120
Query: 1564 ELTIQEQDLPKSSEVTEVFNGTSNSFSAFNIYPRYFRVTGSGDVRSLEQSVELVSGDLVL 1623
E ++ DLPK S++ V NG+ + F N+YP++FRVTGSGDVRS E + VS D+++
Sbjct: 121 ERSVDPLDLPKRSDIQMVLNGSMDGFRVHNLYPKFFRVTGSGDVRSFEDQTDEVSADILI 180
Query: 1624 NRGNPE-WWSFYDLDISD-AHGCGKFPGPMAIIVSEETPQGIIGDTLSKFSIWGLYITFV 1681
N N + WWSF++L S+ C GP+AII+SEETP
Sbjct: 181 NHANFKWWWSFHNLKASENISACEGMDGPVAIIMSEETPP-------------------- 220
Query: 1682 LAVGRFIRLQCSDLRMRIPFENLPSCDRLMALCEDIYAARAEGELEVEEVLYWTLVKIYR 1741
VGRFIRLQCSDLRMRIP+ENLPSCDRL+A+CED+YAARAEGEL VEEVLYWTLVKIYR
Sbjct: 221 -PVGRFIRLQCSDLRMRIPYENLPSCDRLIAICEDLYAARAEGELGVEEVLYWTLVKIYR 279
Query: 1742 TPHMLLEYTQAE 1753
+PHMLLEYT+ +
Sbjct: 280 SPHMLLEYTKLD 291
>At5g06670 kinesin heavy chain-like protein
Length = 997
Score = 36.2 bits (82), Expect = 0.17
Identities = 24/90 (26%), Positives = 48/90 (52%)
Query: 607 RITSRSALVEIIIYMLVSLQSYMFSFPEFDYVSTYLEKEQIGAILRQQEKKAAWKTAQLR 666
R++ SAL + II + + +++ SF E + T ++ ILR+Q+K + + AQ
Sbjct: 604 RLSEGSALADQIIETMENREAHEDSFHEIETPETRIKMIDQMEILREQQKTLSEEMAQQS 663
Query: 667 HIRKAEELKHLRSLQVEKMKSEMLNLQNQL 696
K + ++ Q E++K+E++NL +
Sbjct: 664 RSFKLLSEEAAKAPQNEEIKAEIINLNGDI 693
>At5g07640 putative protein
Length = 316
Score = 35.8 bits (81), Expect = 0.23
Identities = 18/68 (26%), Positives = 32/68 (46%)
Query: 396 EYLAIWMRLTFMNLEIEEHVPCRDCWRVSDIYFSYCRKCWLGIIVDDPRMLIGYYGVFMF 455
E L + +TF+ ++ + + WR D+Y C C+ + D+ + G + F F
Sbjct: 122 EALPVMKDITFVLKLAKDAIASQIRWREGDVYMETCPVCYEHVTSDEKFEVPGCFHRFCF 181
Query: 456 SCFKFRAD 463
C K +AD
Sbjct: 182 DCIKKQAD 189
>At3g59520 unknown protein
Length = 269
Score = 35.4 bits (80), Expect = 0.30
Identities = 45/191 (23%), Positives = 76/191 (39%), Gaps = 33/191 (17%)
Query: 474 YQKIHSQWKSASVLSNLSFETKGYWTF-----LDHLRLYGYCHLLDFVLSLILITGTLEY 528
++ I S SVL +L F W+ L H+ L G + L + L L++ +G L
Sbjct: 52 WRMITSALSHISVL-HLVFNMSALWSLGVVEQLGHVGL-GTAYYLHYTLVLVVFSGVLVI 109
Query: 529 DFLHLGYLGFALVFFR--------------MRLKILKQGN---NIFRYLRM------YNF 565
HL F + +FR M + +KQ + N+F L + +
Sbjct: 110 GIYHLLIARFKIDYFRRVTAVGYSCVVFGWMTILSVKQPSSKLNLFGLLSLPISFAPFES 169
Query: 566 AVIVLSLAYQSPFLGDFSEIKSGFIEYINELVGFQKYDYGFRITSRSALVEIIIYMLVSL 625
+ + Q+ FLG S I G+ + G Y + +T +V + ++ L
Sbjct: 170 LIFTSIIVPQASFLGHLSGILVGYAISWGLIGGMNNY---WALTMLGWIVVVFVFSLKKS 226
Query: 626 QSYMFSFPEFD 636
+Y FSF E +
Sbjct: 227 GAYDFSFLEIE 237
>At5g13130 putative protein
Length = 706
Score = 35.0 bits (79), Expect = 0.39
Identities = 26/85 (30%), Positives = 46/85 (53%), Gaps = 9/85 (10%)
Query: 650 ILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNA- 708
+L QE KA +++++ KA+ K + LQ+++ K+ + NL+N+ +ST A
Sbjct: 617 LLELQESKA-----KIQNLEKAQREKEVLELQLKESKARIQNLENRQEGVSTIFQQERAR 671
Query: 709 -SLKNDGLRE--RRNSFLENEFRKQ 730
+ DGLR+ R S + + RKQ
Sbjct: 672 RDVTEDGLRKKLREASDVIDGLRKQ 696
>At3g22830 putative heat shock protein
Length = 406
Score = 34.3 bits (77), Expect = 0.66
Identities = 45/211 (21%), Positives = 84/211 (39%), Gaps = 20/211 (9%)
Query: 665 LRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLKNDGLRERRNSFLE 724
L++IR+ + + +Q + SE +L N + + D LR + +
Sbjct: 145 LKNIRRRKTSNNSNQMQ-QPQSSEQQSLDN----FCIEVGRYGLDGEMDSLRRDKQVLMM 199
Query: 725 NEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMKHPMGSPHGIVEVKERTKNNDF 784
R + +T+ TL + + L +S+ + ++ M +P I ++ E+ +
Sbjct: 200 ELVRLRQQQQSTKMYLTL-IEEKLKKTESKQKQMMSFLARAMQNPDFIQQLVEQKEKRKE 258
Query: 785 LDLEIRNRYKLP---GKRNAL-----------VSAVHFIGSGISQAQSLGNMAVNNLVNY 830
++ I + + P GKRN V+A G+SQ + GNM+ +
Sbjct: 259 IEEAISKKRQRPIDQGKRNVEDYGDESGYGNDVAASSSALIGMSQEYTYGNMSEFEMSEL 318
Query: 831 LKIEREGQEPTDDSSEDEEYYEIENQNSGAE 861
K+ Q D+SS EE +E N E
Sbjct: 319 DKLAMHIQGLGDNSSAREEVLNVEKGNDEEE 349
>At1g65010 hypothetical protein
Length = 1318
Score = 34.3 bits (77), Expect = 0.66
Identities = 33/138 (23%), Positives = 60/138 (42%), Gaps = 17/138 (12%)
Query: 634 EFDYVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQ 693
E Y+ E ++ IL QE K + +++ E + ++ K++ ++LN +
Sbjct: 878 ETAYLKKIEELSKLHEILSDQETKLQISNHEKEELKERETAYLKKIEELSKVQEDLLNKE 937
Query: 694 NQLHSMSTDAN--YSNASLKNDGLRERRN---SFLENEFRKQAL-----DTNTESIGTL- 742
N+LH M + S SL + E N S L E QA+ + ++ + TL
Sbjct: 938 NELHGMVVEIEDLRSKDSLAQKKIEELSNFNASLLIKENELQAVVCENEELKSKQVSTLK 997
Query: 743 ------DVNQSLLSEKSE 754
D+ QSL+ ++ E
Sbjct: 998 TIDELSDLKQSLIHKEKE 1015
Score = 31.6 bits (70), Expect = 4.3
Identities = 54/278 (19%), Positives = 105/278 (37%), Gaps = 54/278 (19%)
Query: 643 EKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTD 702
EKE AI+ ++ KA + +++ EEL +L+ ++K LQ H
Sbjct: 1013 EKELQAAIVENEKLKAEAALS----LQRIEELTNLKQTLIDKQNE----LQGVFHENEEL 1064
Query: 703 ANYSNASLKN-DGLRERRNSFLENEFRKQALDTNTESIGTLD------------VNQSLL 749
+SLK D L S+LE E Q + + T D + +SLL
Sbjct: 1065 KAKEASSLKKIDELLHLEQSWLEKESEFQRVTQENLELKTQDALAAKKIEELSKLKESLL 1124
Query: 750 SEKSE----SPLVPEYMKHPMGSPHGIVEVKERTKNNDFLDLEIRN------------RY 793
+++E E M+ P S HG E+ K+ D + N Y
Sbjct: 1125 EKETELKCREAAALEKMEEP--SKHGNSELNSIGKDYDLVQFSEVNGASNGDEKTKTDHY 1182
Query: 794 KLPGKRNALV---------------SAVHFIGSGISQAQSLGNMAVNNLVNYLKIEREGQ 838
+ + + + +A+H + + +++ + + + KIE+
Sbjct: 1183 QQRSREHMIQESPMEAIDKHLMGERAAIHKVAHRVEGERNVEKESEFKMWDSYKIEKSEV 1242
Query: 839 EPTDDSSEDEEYYEIENQNSGAEPLEPTFSTNSINEHT 876
P ++ D E++++ +E ++ + S+ +HT
Sbjct: 1243 SPERETELDSVEEEVDSKAESSENMDQYSNGFSLTDHT 1280
>At1g17440 unknown protein
Length = 683
Score = 33.9 bits (76), Expect = 0.86
Identities = 37/164 (22%), Positives = 69/164 (41%), Gaps = 6/164 (3%)
Query: 626 QSYMFSFPEFDYVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKM 685
Q S P+ S L ++QI I++QQ++++ +Q+ ++L+ ++ ++
Sbjct: 403 QQQQLSSPQLHQSSMSLNQQQISQIIQQQQQQSQLGQSQMNQSHSQQQLQQMQQQLQQQP 462
Query: 686 KSEMLNLQNQLHSMSTDANY-SNASLKNDGLRERRNSFLENEFRKQALDT--NTESIGTL 742
+ +M Q Q M + S L + G + + + E + T + S GT
Sbjct: 463 QQQMQQQQQQQQQMQINQQQPSPRMLSHAGQKSVSLTGSQPEATQSGTTTPGGSSSQGTE 522
Query: 743 DVNQSLLSEKSESPLVPEYMKHPMGSPHGIVEVKERTKNNDFLD 786
NQ LL ++ LV + H P VE +DF+D
Sbjct: 523 ATNQ-LLGKRKIQDLVSQVDVHAKLDPD--VEDLLLEVADDFID 563
>At1g13790 hypothetical protein
Length = 736
Score = 33.9 bits (76), Expect = 0.86
Identities = 27/126 (21%), Positives = 56/126 (44%), Gaps = 8/126 (6%)
Query: 661 KTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLKNDGLRERRN 720
+TA H+RK +LK + + E + + N +++ T S+ + + + ++ +
Sbjct: 331 RTAIGDHLRKQGDLKSVSGKEAEDQRKTFTLVSNLENTLVTK---SDNLQQMESIYKQTS 387
Query: 721 SFLENEFRKQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMKHPMGSPHGIVEVKERTK 780
S LE +++ E I T + S++ + + L Y +H S H + KE
Sbjct: 388 SVLEKRMKEK-----DEMINTHNEKMSIMQQTARDYLASIYEEHEKASQHLEAQRKEYED 442
Query: 781 NNDFLD 786
++LD
Sbjct: 443 RENYLD 448
>At4g36120 myosin-like protein
Length = 981
Score = 32.7 bits (73), Expect = 1.9
Identities = 41/179 (22%), Positives = 79/179 (43%), Gaps = 13/179 (7%)
Query: 669 RKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDANYSNASLKNDGLRERRNSFLENEFR 728
+K +++ ++ Q +K+K+E+ ++L A NA+L L+ER +
Sbjct: 161 KKLQDVILAKTSQWDKIKAELEGKIDELSEGLHRAASDNAALTRS-LQERSEMIVRISEE 219
Query: 729 KQALDTNTESIGTLDVNQSLLSEKSESPLVPEYMKHPMGSPHGIVEVKERTKNNDFLDLE 788
+ + + E + T L+EK S Y+K+ + VE++ KN +
Sbjct: 220 RSKAEADVEKLKT----NLQLAEKEIS-----YLKYDLHVASKEVEIRNEEKNMSLKSAD 270
Query: 789 IRNRYKLPG-KRNALVSAVHFIGSGISQAQSLGNMAVNNLVNYLKIEREGQEPTDDSSE 846
I N+ L G K+ A + A G+ + + G A+ + L++E G E TD ++
Sbjct: 271 IANKQHLEGVKKIAKLEAECHRLRGLLRKKLPGPAAMAQM--KLEVEGLGHEFTDPRAQ 327
>At5g05700 arginine-tRNA-protein transferase 1 homolog
Length = 632
Score = 32.0 bits (71), Expect = 3.3
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 1/40 (2%)
Query: 834 EREGQEPTDDSSEDEEYYEIENQNSGAEPLEPTFSTNSIN 873
E E ++ DD +DEE YE E+++S E +P N IN
Sbjct: 542 EDEDEDDDDDDDDDEEMYETESEDSHIES-DPGSKDNDIN 580
>At4g38065 hypothetical protein
Length = 1050
Score = 32.0 bits (71), Expect = 3.3
Identities = 16/50 (32%), Positives = 30/50 (60%), Gaps = 1/50 (2%)
Query: 653 QQEKKAAWKTAQLRHIRKA-EELKHLRSLQVEKMKSEMLNLQNQLHSMST 701
Q E+K WK Q +H+ +A E+LK+L ++ + E L ++++S+ T
Sbjct: 181 QVEEKLKWKKEQFKHLEEAYEKLKNLFKDSKKEWEEEKSKLLDEIYSLQT 230
>At4g34660 SH3 domain-containing protein 2
Length = 368
Score = 32.0 bits (71), Expect = 3.3
Identities = 29/121 (23%), Positives = 58/121 (46%), Gaps = 7/121 (5%)
Query: 621 MLVSLQSYMFSFPEFDYVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSL 680
+L+ L+S + T L KE A+ ++++ +L + ++E H R L
Sbjct: 186 ILMKLESAEAKLHDLKSNMTILGKEAASALASVEDQQQKLTLERLLSMVESERAYHQRVL 245
Query: 681 QV-EKMKSEMLNLQNQLHSMSTDANYSNASLKNDGLRERRNSFLENEFRKQALDTNTESI 739
Q+ ++++ EM++ + ++ + ST + S S+ E N F Q DT+T+S+
Sbjct: 246 QILDQLEGEMVSERQRIEAPSTPS--SADSMPPPPSYEEANGV----FASQMHDTSTDSM 299
Query: 740 G 740
G
Sbjct: 300 G 300
>At3g54670 structural maintenance of chromosomes (SMC) - like
protein
Length = 1265
Score = 32.0 bits (71), Expect = 3.3
Identities = 22/75 (29%), Positives = 38/75 (50%), Gaps = 7/75 (9%)
Query: 649 AILRQQEKKAAWKTAQLRHIRKAEELKHLR------SLQVEKMKSEMLNLQNQLHSMSTD 702
A L Q+KK +L+ +K E KHLR +L+ E+ ++ N++N + + D
Sbjct: 192 AALIYQKKKTIGNEKKLKKAQKEEAEKHLRLQEELKALKRERFLWQLYNIENDIEKANED 251
Query: 703 ANYSNASLKNDGLRE 717
+ S S + D +RE
Sbjct: 252 VD-SEKSNRKDVMRE 265
>At1g56040 hypothetical protein
Length = 437
Score = 32.0 bits (71), Expect = 3.3
Identities = 25/109 (22%), Positives = 51/109 (45%), Gaps = 1/109 (0%)
Query: 644 KEQIGAILRQQEKKAAWKTAQLRHIRKAEELKHLRSLQVEKMKSEMLNLQNQLHSMSTDA 703
KE+ G + Q + A+LRH ++ E L + ++EK+K ++ ++N++ + A
Sbjct: 180 KEEQGKLQLQAKALEHKLEAELRHRKETETLLAIERDRIEKVKIQLETVENEIDNTRLKA 239
Query: 704 NYSNASLKNDGLRERRNSFLENEFRKQALDTNTESIGTLDVNQSLLSEK 752
+ + + RR S + E K+ L+ + T + Q LS +
Sbjct: 240 EEFERKYEGEMIL-RRESEIALEKEKKELEEVKLKLETYEREQENLSSE 287
>At1g12500 phosphate/phosphoenolpyruvate translocator protein like
Length = 361
Score = 32.0 bits (71), Expect = 3.3
Identities = 21/50 (42%), Positives = 28/50 (56%), Gaps = 12/50 (24%)
Query: 361 LLYIASLAACVLLH----------RLLIKKLW--PIFVFLFASVVTVEYL 398
LLY+A +AAC+LL R+LI+K P+ +FL A TV YL
Sbjct: 241 LLYMAPMAACILLPFTLYIEGNVLRVLIEKARTDPLIIFLLAGNATVAYL 290
>At2g07680 putative ABC transporter
Length = 1146
Score = 31.6 bits (70), Expect = 4.3
Identities = 25/102 (24%), Positives = 44/102 (42%), Gaps = 10/102 (9%)
Query: 500 FLDHLRLYGYCHLLDFVLSLILITGTLEYDFLHLGYLGFALVFFRMRLKILKQGNNIFRY 559
F++HL LY + + SL L L L LG +V F + +L G N
Sbjct: 791 FIEHLTLYQRTSYSEIIASLWLS--------LRLQLLGSMIVLFVAVMAVLGSGGNF--P 840
Query: 560 LRMYNFAVIVLSLAYQSPFLGDFSEIKSGFIEYINELVGFQK 601
+ ++ L+L+Y +P + + + F E E+V ++
Sbjct: 841 ISFGTPGLVGLALSYAAPLVSLLGSLLTSFTETEKEMVSVER 882
>At4g21270 kinesin-related protein katA
Length = 793
Score = 31.2 bits (69), Expect = 5.6
Identities = 37/144 (25%), Positives = 63/144 (43%), Gaps = 15/144 (10%)
Query: 621 MLVSLQSYMFSFPEFDYVSTYLEKEQIGAILRQQEKKAAWKTAQLRHIRK-----AEELK 675
M LQ Y S +++ + E + A L + EK+ + L +R ++L
Sbjct: 221 MYKRLQEYNTSLQQYNS-KLQTDLETVRAALTRAEKEKSSILENLSTLRGHSKSLQDQLS 279
Query: 676 HLRSLQVEKMK------SEMLNLQNQLHSMSTDANYSNASLKNDGLRERRNSFLENEFR- 728
R LQ + +K SE+ NL+N+L + D + +++ L E + EN +
Sbjct: 280 SSRVLQDDAIKQKDSLLSEVTNLRNELQQVRDDRD--RQVVQSQKLSEEIRKYQENVGKS 337
Query: 729 KQALDTNTESIGTLDVNQSLLSEK 752
Q LD T G+L+ SL E+
Sbjct: 338 SQELDILTAKSGSLEETCSLQKER 361
>At3g06250 unknown protein
Length = 764
Score = 31.2 bits (69), Expect = 5.6
Identities = 19/50 (38%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Query: 1545 AFSWEFTRDRPRGKEVVKYELTIQEQDLPKSSEVTEVFNGTSNSFSAFNI 1594
AFSW F + R + + +QDLP V +VF GT + FSA+ I
Sbjct: 418 AFSWLF-QTWLRAMSGRRPRSMVADQDLPIQQAVAQVFPGTHHRFSAWQI 466
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,164,947
Number of Sequences: 26719
Number of extensions: 1635538
Number of successful extensions: 5649
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 27
Number of HSP's that attempted gapping in prelim test: 5606
Number of HSP's gapped (non-prelim): 51
length of query: 1753
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1640
effective length of database: 8,299,349
effective search space: 13610932360
effective search space used: 13610932360
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0544.8


