

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.15
(322 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
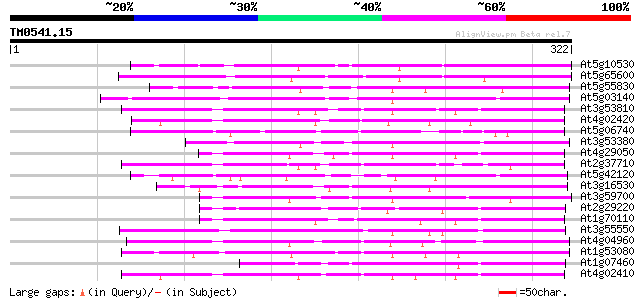
Score E
Sequences producing significant alignments: (bits) Value
At5g10530 lectin-like protein kinase - like 112 2e-25
At5g65600 receptor protein kinase-like protein 109 2e-24
At5g55830 serine/threonine-specific kinase like protein 97 1e-20
At5g03140 receptor like protein kinase 96 3e-20
At3g53810 serine/threonine-specific kinase like protein 91 8e-19
At4g02420 81 7e-16
At5g06740 lectin-like protein kinase 80 1e-15
At3g53380 receptor lectin kinase -like protein 79 3e-15
At4g29050 serine/threonine-specific kinase like protein 78 7e-15
At2g37710 putative receptor-like protein kinase 78 7e-15
At5g42120 receptor lectin kinase-like protein 77 1e-14
At3g16530 putative lectin 74 1e-13
At3g59700 serine/threonine-specific kinase lecRK1 precursor,lect... 72 4e-13
At2g29220 putative protein kinase 72 5e-13
At1g70110 hypothetical protein 71 7e-13
At3g55550 probable serine/threonine-specific protein kinase 70 1e-12
At4g04960 unknown protein 69 3e-12
At1g53080 protein kinase, putative 69 4e-12
At1g07460 putative lectin, predicted GPI-anchored protein 69 4e-12
At4g02410 unknown protein 68 6e-12
>At5g10530 lectin-like protein kinase - like
Length = 651
Score = 112 bits (281), Expect = 2e-25
Identities = 89/260 (34%), Positives = 130/260 (49%), Gaps = 18/260 (6%)
Query: 70 LIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSS 129
L+FS +L+ V S + F+ F DV I GDA NG ++LT ID +
Sbjct: 6 LLFSFVLVLPFVCS--VQFNISRFGSDVSEIAYQGDARA-NGAVELTNIDYTCR-----A 57
Query: 130 GISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP----HGDGFTFFLASLDFEFPEGGG 185
G + + V L + T + +DF T F+F ++ +G GF FFLA + P
Sbjct: 58 GWATYGKQVPLWNPGTSKPSDFSTRFSFRIDTRNVGYGNYGHGFAFFLAPARIQLPPNSA 117
Query: 186 GGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNF--PHIGIDINTIRSAVTIPWSV 242
GG FLGLF+ T +S +V VEFD+F+N +DP H+GI+ N++ S+ W+
Sbjct: 118 GG-FLGLFNG-TNNQSSAFPLVYVEFDTFTNPEWDPLDVKSHVGINNNSLVSSNYTSWNA 175
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSA 302
Q I I Y S R+L+V Y + ++ Y ID+ L V IGFSA
Sbjct: 176 TSHNQD-IGRVLIFYDSARRNLSVSWTYDLTSDPLENSSLSYIIDLSKVLPSEVTIGFSA 234
Query: 303 TTGEFAETHSILSWSFNSTL 322
T+G E + +LSW F+S+L
Sbjct: 235 TSGGVTEGNRLLSWEFSSSL 254
>At5g65600 receptor protein kinase-like protein
Length = 675
Score = 109 bits (272), Expect = 2e-24
Identities = 86/269 (31%), Positives = 135/269 (49%), Gaps = 17/269 (6%)
Query: 63 TSELLATLIFSLLLITTTVKSDSLSFSFPTF-DPDVRVILLNGDANTT-NGTLQLTKIDQ 120
+S + +++F L + DSL F+F +F D I +GDA +GT+ +Q
Sbjct: 13 SSSMSNSILFLSLFLFLPFVVDSLYFNFTSFRQGDPGDIFYHGDATPDEDGTVNFNNAEQ 72
Query: 121 YGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDF 178
G + V + KTG+ +DF T F+F ++ + G G FFLA +
Sbjct: 73 TSQV-----GWITYSKKVPIWSHKTGKASDFSTSFSFKIDARNLSADGHGICFFLAPMGA 127
Query: 179 EFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNF--PHIGIDINTIRSA 235
+ P GGFL LF + Y++S +V VEFD+F+N +DPN H+GI+ N++ S+
Sbjct: 128 QLP-AYSVGGFLNLFTRKNNYSSSF-PLVHVEFDTFNNPGWDPNDVGSHVGINNNSLVSS 185
Query: 236 VTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYP--PGQNTTNIDAIKYPIDIESYLS 293
W+ + Q I +AKISY SV+++L+V Y + ++ Y ID+ L
Sbjct: 186 NYTSWNASSHSQD-ICHAKISYDSVTKNLSVTWAYELTATSDPKESSSLSYIIDLAKVLP 244
Query: 294 EWVVIGFSATTGEFAETHSILSWSFNSTL 322
V+ GF A G E H +LSW +S+L
Sbjct: 245 SDVMFGFIAAAGTNTEEHRLLSWELSSSL 273
>At5g55830 serine/threonine-specific kinase like protein
Length = 681
Score = 97.1 bits (240), Expect = 1e-20
Identities = 78/253 (30%), Positives = 121/253 (46%), Gaps = 23/253 (9%)
Query: 81 VKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHL 140
V SD+++F+F +F +R + GD++ NG + LT+ + G P SSG + +
Sbjct: 26 VSSDNMNFTFKSFT--IRNLTFLGDSHLRNGVVGLTR--ELGVPDT-SSGTVIYNNPIRF 80
Query: 141 SDKKTGRVADFRTEFTFVVNHSQPH----GDGFTFFLASLDFEFPEGGGGGGFLGLFDEE 196
D + A F T F+F V + P GDG FFL+ + G GG+LGL +
Sbjct: 81 YDPDSNTTASFSTHFSFTVQNLNPDPTSAGDGLAFFLSHDNDTL---GSPGGYLGLVNSS 137
Query: 197 TAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVT---IPWSVNDQPQGAIQ 251
KN VA+EFD+ + DPN HIG+D++++ S T + S D G
Sbjct: 138 ---QPMKNRFVAIEFDTKLDPHFNDPNGNHIGLDVDSLNSISTSDPLLSSQIDLKSGKSI 194
Query: 252 NAKISYSSVSRDLTVVVHYPPGQNTTNIDA---IKYPIDIESYLSEWVVIGFSATTGEFA 308
+ I Y + R L V + Y TT + ID+ +L+ + +GFS +T
Sbjct: 195 TSWIDYKNDLRLLNVFLSYTDPVTTTKKPEKPLLSVNIDLSPFLNGEMYVGFSGSTEGST 254
Query: 309 ETHSILSWSFNST 321
E H I +WSF ++
Sbjct: 255 EIHLIENWSFKTS 267
>At5g03140 receptor like protein kinase
Length = 711
Score = 95.5 bits (236), Expect = 3e-20
Identities = 87/275 (31%), Positives = 134/275 (48%), Gaps = 20/275 (7%)
Query: 53 LNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGT 112
L P ++ S L+ I + L ++ V+ D + F + + L GDA+ NGT
Sbjct: 2 LKLPPRFFSVYSTLIH--ILASFLCSSDVRGDFPATRFDLGTLTLSSLKLLGDAHLNNGT 59
Query: 113 LQLTKIDQYGNPSPHSSGISAFYGA-VHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFT 170
++LT+ P S+ A YG V +T A F T F+F V + P G
Sbjct: 60 IKLTRELSV----PTSTAGKALYGKPVKFRHPETKSPASFTTYFSFSVTNLNPSSIGGGL 115
Query: 171 FFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGID 228
F+ S D ++ G GGFLGL +E S + VAVEFD+ + D N H+G+D
Sbjct: 116 AFVISPDEDYL--GSTGGFLGLTEE----TGSGSGFVAVEFDTLMDVQFKDVNGNHVGLD 169
Query: 229 INTIRSAVTIPW-SVN-DQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPI 286
+N + SA +V+ D G N+ I+Y R LTV V Y + + I + P+
Sbjct: 170 LNAVVSAAVADLGNVDIDLKSGNAVNSWITYDGSGRVLTVYVSYSNLKPKSPI--LSVPL 227
Query: 287 DIESYLSEWVVIGFSATTGEFAETHSILSWSFNST 321
D++ Y+S+ + +GFS +T E HS+ WSF+S+
Sbjct: 228 DLDRYVSDSMFVGFSGSTQGSTEIHSVDWWSFSSS 262
>At3g53810 serine/threonine-specific kinase like protein
Length = 677
Score = 90.9 bits (224), Expect = 8e-19
Identities = 83/272 (30%), Positives = 117/272 (42%), Gaps = 32/272 (11%)
Query: 65 ELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKIDQYGN 123
+L F LL S +L+F++ F P + I L G A T NG L+LT N
Sbjct: 4 KLKLIFFFFLLCQIMISSSQNLNFTYNGFHPPLTDISLQGLATVTPNGLLKLT------N 57
Query: 124 PSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP--HGDGFTFFLA---SLDF 178
S +G + + D + G V+ F T F F ++ P G G F +A L F
Sbjct: 58 TSVQKTGHAFCTERIRFKDSQNGNVSSFSTTFVFAIHSQIPTLSGHGIAFVVAPTLGLPF 117
Query: 179 EFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAV 236
P ++GLF+ N + N I AVEFD+ +S DPN H+GID+N +RSA
Sbjct: 118 ALPSQ-----YIGLFNISNNGNDT-NHIFAVEFDTIQSSEFGDPNDNHVGIDLNGLRSAN 171
Query: 237 TIPWSVNDQPQGAIQNAK----------ISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPI 286
D QN I Y + S + V V P + + Y
Sbjct: 172 YSTAGYRDD-HDKFQNLSLISRKRIQVWIDYDNRSHRIDVTVA-PFDSDKPRKPLVSYVR 229
Query: 287 DIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
D+ S L E + +GFS+ TG H ++ WSF
Sbjct: 230 DLSSILLEDMYVGFSSATGSVLSEHFLVGWSF 261
>At4g02420
Length = 669
Score = 81.3 bits (199), Expect = 7e-16
Identities = 79/264 (29%), Positives = 115/264 (42%), Gaps = 28/264 (10%)
Query: 71 IFSLLLITTTVKSDS--LSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKIDQYGNPSPH 127
IF L ++KS S + F++ F P I + G A T NG L+LT N +
Sbjct: 9 IFFLSFFWQSLKSSSQIIDFTYNGFRPPPTDISILGIATITPNGLLKLT------NTTMQ 62
Query: 128 SSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHGDGFTFFLA---SLDFEFPEGG 184
S+G + + + D G V+ F T F F ++ P G F +A L F G
Sbjct: 63 STGHAFYTKPIRFKDSPNGTVSSFSTTFVFAIHSQIPIAHGMAFVIAPNPRLPF-----G 117
Query: 185 GGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN--SFDPNFPHIGIDINTIRSAVTIP--- 239
+LGLF+ N +N + AVE D+ N D N H+GIDIN++ S + P
Sbjct: 118 SPLQYLGLFNVTNNGNV-RNHVFAVELDTIMNIEFNDTNNNHVGIDINSLNSVKSSPAGY 176
Query: 240 WSVNDQPQG-AIQNAKISYSSVSRD----LTVVVHYPPGQNTTNIDAIKYPIDIESYLSE 294
W NDQ + ++K V D L V P G+ + D+ S L +
Sbjct: 177 WDENDQFHNLTLISSKRMQVWVDFDGPTHLIDVTMAPFGEVKPRKPLVSIVRDLSSVLLQ 236
Query: 295 WVVIGFSATTGEFAETHSILSWSF 318
+ +GFS+ TG +L WSF
Sbjct: 237 DMFVGFSSATGNIVSEIFVLGWSF 260
>At5g06740 lectin-like protein kinase
Length = 652
Score = 80.1 bits (196), Expect = 1e-15
Identities = 77/266 (28%), Positives = 122/266 (44%), Gaps = 35/266 (13%)
Query: 70 LIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPS---P 126
L+F +L + L F FP F+ + L+ ++ G +Q+T D G P
Sbjct: 9 LLFLILTCKIETQVKCLKFDFPGFNVSNELELIRDNSYIVFGAIQVTP-DVTGGPGGTIA 67
Query: 127 HSSGISAFYGAVHLSDKKTGRVADFRTEFTF-VVNHSQPHGDGFTFFLASLDFEFPEGGG 185
+ +G + + L K + A F T F + N + P G+G F L E
Sbjct: 68 NQAGRALYKKPFRLWSKH--KSATFNTTFVINISNKTDPGGEGLAFVLTPE--ETAPQNS 123
Query: 186 GGGFLGLFDEETAYNTSKNSIVAVEFDSF-SNSFDPNFPHIGIDINTIRSAVTIPWSVND 244
G +LG+ +E T N +++ IV+VEFD+ S+S D + H+ +++N I S V
Sbjct: 124 SGMWLGMVNERTNRN-NESRIVSVEFDTRKSHSDDLDGNHVALNVNNINSVV-------- 174
Query: 245 QPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTT-----NIDAIKY-------PIDIESYL 292
Q ++ I S DLT V Y G+N + N+D + ID+ +YL
Sbjct: 175 --QESLSGRGIKIDS-GLDLTAHVRYD-GKNLSVYVSRNLDVFEQRNLVFSRAIDLSAYL 230
Query: 293 SEWVVIGFSATTGEFAETHSILSWSF 318
E V +GF+A+T F E + + SWSF
Sbjct: 231 PETVYVGFTASTSNFTELNCVRSWSF 256
>At3g53380 receptor lectin kinase -like protein
Length = 715
Score = 79.0 bits (193), Expect = 3e-15
Identities = 67/226 (29%), Positives = 107/226 (46%), Gaps = 18/226 (7%)
Query: 102 LNGDANTTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNH 161
L GDA +NG + LT+ N +G + + T F + F+F + +
Sbjct: 36 LLGDARLSNGIVGLTRDLSVPNSG---AGKVLYSNPIRFRQPGTHFPTSFSSFFSFSITN 92
Query: 162 SQPH--GDGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF- 218
P G G F ++ + G GG LGL T N S + VAVEFD+ +
Sbjct: 93 VNPSSIGGGLAFVISP---DANSIGIAGGSLGL----TGPNGSGSKFVAVEFDTLMDVDF 145
Query: 219 -DPNFPHIGIDINTIRSAVTIPW-SVN-DQPQGAIQNAKISYSSVSRDLTVVVHYPPGQN 275
D N H+G D+N + S+V+ +VN D G N+ I Y ++R V V Y
Sbjct: 146 KDINSNHVGFDVNGVVSSVSGDLGTVNIDLKSGNTINSWIEYDGLTRVFNVSVSY--SNL 203
Query: 276 TTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSFNST 321
+ + +P+D++ Y+++++ +GFS +T E HSI WSF+S+
Sbjct: 204 KPKVPILSFPLDLDRYVNDFMFVGFSGSTQGSTEIHSIEWWSFSSS 249
>At4g29050 serine/threonine-specific kinase like protein
Length = 669
Score = 77.8 bits (190), Expect = 7e-15
Identities = 66/228 (28%), Positives = 96/228 (41%), Gaps = 32/228 (14%)
Query: 109 TNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVV--NHSQPHG 166
+ G ++LT N S S G + V + G V+ F T F F + N + G
Sbjct: 45 SKGLMKLT------NSSEFSYGHVFYNSPVRFKNSPNGTVSSFSTTFVFAIVSNVNALDG 98
Query: 167 DGFTFFLASLDFEFPEGG----GGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DP 220
G F ++ P G +LGLF+ T N IVAVEFD+F N D
Sbjct: 99 HGLAFVIS------PTKGLPYSSSSQYLGLFNL-TNNGDPSNHIVAVEFDTFQNQEFDDM 151
Query: 221 NFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAK----------ISYSSVSRDLTVVVHY 270
+ H+GIDIN++ S + G +N + I Y S R L V +H
Sbjct: 152 DNNHVGIDINSLSSEKASTAGYYEDDDGTFKNIRLINQKPIQAWIEYDSSRRQLNVTIH- 210
Query: 271 PPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
P I + D+ YL + + +GF++ TG +H IL W+F
Sbjct: 211 PIHLPKPKIPLLSLTKDLSPYLFDSMYVGFTSATGRLRSSHYILGWTF 258
>At2g37710 putative receptor-like protein kinase
Length = 675
Score = 77.8 bits (190), Expect = 7e-15
Identities = 81/278 (29%), Positives = 121/278 (43%), Gaps = 44/278 (15%)
Query: 65 ELLATLIFSLLLITTTVKSDSLSFSFPT-FDPDVRVILLNGDANTTNGTLQLTKIDQYGN 123
+LL F + S SL+F++ F+P + + T NG L+LT N
Sbjct: 4 KLLTIFFFFFFNLIFQSSSQSLNFAYNNGFNPPTDLSIQGITTVTPNGLLKLT------N 57
Query: 124 PSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP---HGDGFTFFLA---SLD 177
+ +G + + + D G V+ F T F F + HSQ G G F +A SL
Sbjct: 58 TTVQKTGHAFYTKPIRFKDSPNGTVSSFSTSFVFAI-HSQIAILSGHGIAFVVAPNASLP 116
Query: 178 FEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSF-SNSF-DPNFPHIGIDINTIRSA 235
+ G ++GLF+ N + N + AVE D+ S F D N H+GIDIN+++S
Sbjct: 117 Y-----GNPSQYIGLFNLANNGNET-NHVFAVELDTILSTEFNDTNDNHVGIDINSLKSV 170
Query: 236 VTIPWSVNDQPQGAIQNAKISYSSVSRD-LTVVVHYPPGQNTTNIDAIKYPI-------- 286
+ P D+ +G +N + +SR + V V Y T ID P
Sbjct: 171 QSSPAGYWDE-KGQFKNLTL----ISRKPMQVWVDYDG--RTNKIDVTMAPFNEDKPTRP 223
Query: 287 ------DIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
D+ S L + + +GFS+ TG H IL WSF
Sbjct: 224 LVTAVRDLSSVLLQDMYVGFSSATGSVLSEHYILGWSF 261
>At5g42120 receptor lectin kinase-like protein
Length = 691
Score = 77.0 bits (188), Expect = 1e-14
Identities = 83/274 (30%), Positives = 125/274 (45%), Gaps = 47/274 (17%)
Query: 70 LIFSLLLITTTVKSDSLSFSFPT----FDPDVRVILLNGDANTTNGTLQLTKIDQYGNPS 125
+IF L+L LS FPT F P ++ + L GDA + T+ LT+ Q PS
Sbjct: 9 VIFHLILF--------LSLDFPTLSHRFSPPLQNLTLYGDAFFRDRTISLTQ-QQPCFPS 59
Query: 126 -------PHSSGI--SAFYGAVHLSDKKTGRVADFRTEFTF--VVNHSQPHGDGFTFFLA 174
P SSGI + + + + T A F F+F + + S P GDGF F +
Sbjct: 60 VTTPPSKPSSSGIGRALYVYPIKFLEPSTNTTASFSCRFSFSIIASPSCPFGDGFAFLIT 119
Query: 175 SLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDP-----NFPHIGIDI 229
S F GFLGL + + +S +AVEFD+ FDP N H+GID+
Sbjct: 120 SNADSFV---FSNGFLGLPNPD-------DSFIAVEFDT---RFDPVHGDINDNHVGIDV 166
Query: 230 NTIRSAVTIPWSVN--DQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPID 287
++I S ++ D G A I YS V + + V V Y + T+ + + ID
Sbjct: 167 SSIFSVSSVDAISKGFDLKSGKKMMAWIEYSDVLKLIRVWVGYSRVKPTSPV--LSTQID 224
Query: 288 IESYLSEWVVIGFSAT-TGEFAETHSILSWSFNS 320
+ + E++ +GFSA+ G + H + W F +
Sbjct: 225 LSGKVKEYMHVGFSASNAGIGSALHIVERWKFRT 258
>At3g16530 putative lectin
Length = 276
Score = 73.9 bits (180), Expect = 1e-13
Identities = 71/267 (26%), Positives = 115/267 (42%), Gaps = 45/267 (16%)
Query: 85 SLSFSFPTFDPDVRVILLNGDAN--------TTNGTLQLTKIDQYGNPSPHSSGISAFYG 136
++ F+F +FD + L GDA + +G L +T+ + NP H G+ +
Sbjct: 19 AVKFNFDSFDGSNLLFL--GDAELGPSSDGVSRSGALSMTRDE---NPFSHGQGL--YIN 71
Query: 137 AVHLSDKKTGRVADFRTEFTFVVN-HSQPH-GDGFTFFLASLDFEFPE----GGGGGGFL 190
+ T F T FTF + ++P+ G GF F + PE G GG+L
Sbjct: 72 QIPFKPSNTSSPFSFETSFTFSITPRTKPNSGQGFAFIIT------PEADNSGASDGGYL 125
Query: 191 GLFDEETAYNTSKNSIVAVEFDSFSNS--FDPNFPHIGIDINTIRSAVTIP--------- 239
G+ ++ T +N I+A+EFD+F N D + H+G++IN++ S V
Sbjct: 126 GILNK-TNDGKPENHILAIEFDTFQNKEFLDISGNHVGVNINSMTSLVAEKAGYWVQTRV 184
Query: 240 -----WSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDA-IKYPIDIESYLS 293
WS D + + K ++D T+ V P A I+ P + L
Sbjct: 185 GKRKVWSFKDVNLSSGERFKAWVEFRNKDSTITVTLAPENVKKPKRALIEAPRVLNEVLL 244
Query: 294 EWVVIGFSATTGEFAETHSILSWSFNS 320
+ + GF+ + G E H I SWSF +
Sbjct: 245 QNMYAGFAGSMGRAVERHDIWSWSFEN 271
>At3g59700 serine/threonine-specific kinase lecRK1 precursor,lectin
receptor-like
Length = 661
Score = 72.0 bits (175), Expect = 4e-13
Identities = 56/228 (24%), Positives = 98/228 (42%), Gaps = 25/228 (10%)
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVV--NHSQPHGD 167
NG LT N + H+ G + V + + TG ++ F F F + H+Q
Sbjct: 38 NGYFTLT------NTTKHTFGQAFENEHVEIKNSSTGVISSFSVNFFFAIVPEHNQQGSH 91
Query: 168 GFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHI 225
G TF ++ G +LG+F++ T + N+++A+E D + D + H+
Sbjct: 92 GMTFVISPT--RGLPGASSDQYLGIFNK-TNNGKASNNVIAIELDIHKDEEFGDIDDNHV 148
Query: 226 GIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYP 285
GI+IN +RS + D G+ + + V R L++V P Q + + P
Sbjct: 149 GININGLRSVASASAGYYDDKDGSFKKLSLISREVMR-LSIVYSQPDQQLNVTLFPAEIP 207
Query: 286 I-----------DIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
+ D+ YL E + +GF+A+TG H ++ W N +
Sbjct: 208 VPPLKPLLSLNRDLSPYLLEKMYLGFTASTGSVGAIHYLMGWLVNGVI 255
>At2g29220 putative protein kinase
Length = 627
Score = 71.6 bits (174), Expect = 5e-13
Identities = 67/227 (29%), Positives = 102/227 (44%), Gaps = 31/227 (13%)
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYG-AVHLSDKKTGRVADFRTEFTFVVNHSQ-PHGD 167
+G L+LT N S G AF+G + LS+ + F T F F + G
Sbjct: 50 SGLLELT------NTSMRQIG-QAFHGFPIPLSNPNSTNSVSFSTSFIFAITQGTGAPGH 102
Query: 168 GFTFFLA-SLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFS----NSFDPNF 222
G F ++ S+DF G +LGLF+ N S N I+A+EFD+ N D N
Sbjct: 103 GLAFVISPSMDFS---GAFPSNYLGLFNTSNNGN-SLNRILAIEFDTVQAVELNDIDDN- 157
Query: 223 PHIGIDINTIRSAVTIPWSVNDQPQ----------GAIQNAKISYSSVSRDLTVVVHYPP 272
H+GID+N + S + P + D + G I Y++ L V + P
Sbjct: 158 -HVGIDLNGVISIASAPAAYFDDREAKNISLRLASGKPVRVWIEYNATETMLNVTLA-PL 215
Query: 273 GQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSFN 319
+ +I + +++ S+ +GFSA+TG A +H +L WSFN
Sbjct: 216 DRPKPSIPLLSRKMNLSGIFSQEHHVGFSASTGTVASSHFVLGWSFN 262
>At1g70110 hypothetical protein
Length = 666
Score = 71.2 bits (173), Expect = 7e-13
Identities = 66/225 (29%), Positives = 102/225 (45%), Gaps = 29/225 (12%)
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN-HSQPHGD- 167
NG ++LT N +P ++G + + + G V+ F T F F + H+ +G
Sbjct: 45 NGLIRLT------NSTPQTTGQVFYNDQLRFKNSVNGTVSSFSTTFVFSIEFHNGIYGGY 98
Query: 168 GFTFFLA---SLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDS-FSNSF-DPNF 222
G F + L FP +LGLF+ KN IVAVE D+ F D +
Sbjct: 99 GIAFVICPTRDLSPTFPTT-----YLGLFNRSNM-GDPKNHIVAVELDTKVDQQFEDKDA 152
Query: 223 PHIGIDINTIRS---AVTIPWSVNDQPQGAIQNAK------ISYSSVSRDLTVVVHYPPG 273
H+GIDINT+ S A+ + N + + N+ I Y S + + V +H P
Sbjct: 153 NHVGIDINTLVSDTVALAGYYMDNGTFRSLLLNSGQPMQIWIEYDSKQKQINVTLH-PLY 211
Query: 274 QNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSF 318
I + D+ YL E + +GF++TTG+ +H IL W+F
Sbjct: 212 VPKPKIPLLSLEKDLSPYLLELMYVGFTSTTGDLTASHYILGWTF 256
>At3g55550 probable serine/threonine-specific protein kinase
Length = 684
Score = 70.5 bits (171), Expect = 1e-12
Identities = 71/268 (26%), Positives = 109/268 (40%), Gaps = 22/268 (8%)
Query: 64 SELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKIDQYG 122
S+ A ++ L+ +T V S FSF F + LNG A G ++LT Q
Sbjct: 2 SQTFAVILLLLIFLTHLVSSLIQDFSFIGFKKASPNLTLNGVAEIAPTGAIRLTTETQ-- 59
Query: 123 NPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHGDGFTFFLASLDFEFPE 182
G + + + R F T F + G A
Sbjct: 60 ----RVIGHAFYSLPIRFKPIGVNRALSFSTSFAIAMVPEFVTLGGHGLAFAITPTPDLR 115
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIP- 239
G +LGL + +S AVEFD+ + D N H+GIDIN++ S+++ P
Sbjct: 116 GSLPSQYLGLLNSSRVNFSSH--FFAVEFDTVRDLEFEDINDNHVGIDINSMESSISTPA 173
Query: 240 --WSVNDQPQ------GAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESY 291
+ N + G + A I Y S + L V + P + + Y +D+ S
Sbjct: 174 GYFLANSTKKELFLDGGRVIQAWIDYDSNKKRLDVKLS--PFSEKPKLSLLSYDVDLSSV 231
Query: 292 LSEWVVIGFSATTGEFAETHSILSWSFN 319
L + + +GFSA+TG A +H IL W+FN
Sbjct: 232 LGDEMYVGFSASTGLLASSHYILGWNFN 259
>At4g04960 unknown protein
Length = 686
Score = 68.9 bits (167), Expect = 3e-12
Identities = 70/273 (25%), Positives = 112/273 (40%), Gaps = 36/273 (13%)
Query: 68 ATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPH 127
A L L + ++ F F F+ + L G A + L LT N +
Sbjct: 3 ALLFLLTLFLILPNPISAIDFIFNGFNDSSSNVSLFGIATIESKILTLT------NQTSF 56
Query: 128 SSGISAFYGAVHLSDKKTGRVADFRTEFTFVV----NHSQPHGDGFTFFLASLDFEFPEG 183
++G + + + D T V F T F F + N HG F F ++ G
Sbjct: 57 ATGRALYNRTIRTKDPITSSVLPFSTSFIFTMAPYKNTLPGHGIVFLFAPST----GING 112
Query: 184 GGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNS--FDPNFPHIGIDINTIRSA------ 235
LGLF+ N S N I VEFD F+N D + H+GID+N++ S
Sbjct: 113 SSSAQHLGLFNLTNNGNPS-NHIFGVEFDVFANQEFSDIDANHVGIDVNSLHSVYSNTSG 171
Query: 236 -------VTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDI 288
V P +ND G I Y ++T+ V G+ I + +++
Sbjct: 172 YWSDDGVVFKPLKLND---GRNYQVWIDYRDFVVNVTMQV---AGKIRPKIPLLSTSLNL 225
Query: 289 ESYLSEWVVIGFSATTGEFAETHSILSWSFNST 321
+ + + +GF+A TG ++H IL+WSF+++
Sbjct: 226 SDVVEDEMFVGFTAATGRLVQSHKILAWSFSNS 258
>At1g53080 protein kinase, putative
Length = 283
Score = 68.6 bits (166), Expect = 4e-12
Identities = 72/282 (25%), Positives = 121/282 (42%), Gaps = 32/282 (11%)
Query: 65 ELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNG------DANTTNGTLQLTKI 118
+L ++F LL + T+ + + F+F FD + + D + +G L +T+
Sbjct: 5 KLCIFVLFISLLSSKTISA--VKFNFNRFDGTNLIFIGYAELGPATDGMSRSGALSMTRD 62
Query: 119 DQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPH-GDGFTFFLA- 174
+ P H G+ S+ + V F+T FTF + S P+ G G F +
Sbjct: 63 NI---PFSHGQGLYTDPIPFKSSNNTSSSVYSFKTSFTFSITPRRSNPNPGHGIAFIVVP 119
Query: 175 SLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTI 232
++ +E+ + G FLGL + T N + N + AVEFD F + D N H+G++IN++
Sbjct: 120 TVAYEYDQDSTRG-FLGLVNLTTNGNPN-NHLFAVEFDVFQDKRFGDINDNHVGVNINSV 177
Query: 233 RSAVT------IPWSVNDQPQGAIQNAKIS-------YSSVSRDLTVVVHYPPGQNTTNI 279
S V+ I + Q + K+S + +V P
Sbjct: 178 NSKVSEKAGYWIQTRTRGKNQWLFKEVKLSSGDNYKAWIEYKNSKVIVWLAPAHLKKPKR 237
Query: 280 DAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSFNST 321
I+ +D+ + E + GFS + G E H I SWSF +T
Sbjct: 238 PLIETQVDLSEVVLETMYTGFSGSMGRGVERHDIWSWSFENT 279
>At1g07460 putative lectin, predicted GPI-anchored protein
Length = 258
Score = 68.6 bits (166), Expect = 4e-12
Identities = 61/200 (30%), Positives = 90/200 (44%), Gaps = 19/200 (9%)
Query: 133 AFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHGDGFTFFLA-SLDFEFPEGGGGGGFLG 191
AFYG T F +FV + P G G TF ++ S+DF +LG
Sbjct: 4 AFYGFPIAFKNSTNSSNSFSFSTSFVFSIDAP-GHGLTFLISPSMDFT---QAMPSQYLG 59
Query: 192 LFDEETAYNTSKNSIVAVEFDSF-SNSF-DPNFPHIGIDINTIRSAVTIPWSVNDQPQGA 249
LF+ T S N I+AVEFD+ SN F D + H+GID+N + S + P + Q
Sbjct: 60 LFNT-TNNGNSTNRILAVEFDTVKSNEFLDIDDNHVGIDVNGLVSVESAPAAFFSNKQSK 118
Query: 250 IQNAKIS----------YSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIG 299
+ K+S Y+ V R L V + + N + +++ E + +G
Sbjct: 119 NISLKLSSKDPIRAWIEYNGVERLLNVTLA-TLDTSKPNFPLLSRQMNLSEIFMEKMYVG 177
Query: 300 FSATTGEFAETHSILSWSFN 319
FSA+TG H +L WSF+
Sbjct: 178 FSASTGNITSNHDVLGWSFS 197
>At4g02410 unknown protein
Length = 674
Score = 68.2 bits (165), Expect = 6e-12
Identities = 69/270 (25%), Positives = 116/270 (42%), Gaps = 26/270 (9%)
Query: 65 ELLATLIFSLLLITTTVKSDS--LSFSFPTFDPDVRVILLNGDAN-TTNGTLQLTKIDQY 121
+L F ++L++ + S S L+F++ +F I + G A T+NG L+LT
Sbjct: 4 KLFTIFFFFIILLSKPLNSSSQSLNFTYNSFHRPPTNISIQGIATVTSNGILKLT----- 58
Query: 122 GNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQP--HGDGFTFFLASLDFE 179
+ + S+G + + + D V+ F T F + P G G FF+A
Sbjct: 59 -DKTVISTGHAFYTEPIRFKDSPNDTVSSFSTTFVIGIYSGIPTISGHGMAFFIAPNPVL 117
Query: 180 FPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINT---IRS 234
+LGLF N + N I+AVEFD+ N D N H+GI+IN+ ++S
Sbjct: 118 --SSAMASQYLGLFSSTNNGNDT-NHILAVEFDTIMNPEFDDTNDNHVGININSLTSVKS 174
Query: 235 AVTIPWSVNDQPQGAIQNAK------ISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDI 288
++ W +Q ++ + Y + + V + P G+ + D+
Sbjct: 175 SLVGYWDEINQFNNLTLISRKRMQVWVDYDDRTNQIDVTMA-PFGEVKPRKALVSVVRDL 233
Query: 289 ESYLSEWVVIGFSATTGEFAETHSILSWSF 318
S + + +GFSA TG H + WSF
Sbjct: 234 SSVFLQDMYLGFSAATGYVLSEHFVFGWSF 263
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,474,676
Number of Sequences: 26719
Number of extensions: 328870
Number of successful extensions: 938
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 37
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 844
Number of HSP's gapped (non-prelim): 62
length of query: 322
length of database: 11,318,596
effective HSP length: 99
effective length of query: 223
effective length of database: 8,673,415
effective search space: 1934171545
effective search space used: 1934171545
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0541.15


