

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.11
(269 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
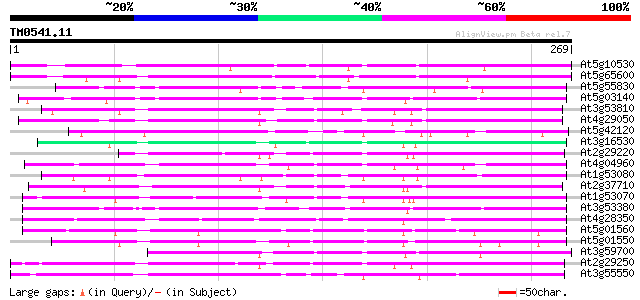
Sequences producing significant alignments: (bits) Value
At5g10530 lectin-like protein kinase - like 129 2e-30
At5g65600 receptor protein kinase-like protein 118 3e-27
At5g55830 serine/threonine-specific kinase like protein 96 2e-20
At5g03140 receptor like protein kinase 93 2e-19
At3g53810 serine/threonine-specific kinase like protein 86 2e-17
At4g29050 serine/threonine-specific kinase like protein 85 4e-17
At5g42120 receptor lectin kinase-like protein 83 1e-16
At3g16530 putative lectin 82 2e-16
At2g29220 putative protein kinase 82 3e-16
At4g04960 unknown protein 81 7e-16
At1g53080 protein kinase, putative 81 7e-16
At2g37710 putative receptor-like protein kinase 80 1e-15
At1g53070 protein kinase, putative 80 1e-15
At3g53380 receptor lectin kinase -like protein 80 1e-15
At4g28350 receptor protein kinase like protein 79 3e-15
At5g01560 receptor like protein kinase 76 2e-14
At5g01550 receptor like protein kinase 75 4e-14
At3g59700 serine/threonine-specific kinase lecRK1 precursor,lect... 75 5e-14
At2g29250 putative protein kinase 74 8e-14
At3g55550 probable serine/threonine-specific protein kinase 74 1e-13
>At5g10530 lectin-like protein kinase - like
Length = 651
Score = 129 bits (324), Expect = 2e-30
Identities = 101/273 (36%), Positives = 139/273 (49%), Gaps = 23/273 (8%)
Query: 1 MANSFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKK 60
MANS + +LPF+ S FN RF SD I GDA G V
Sbjct: 1 MANSILLFSFVLVLPFVC--------SVQFNISRFGSDVSEIAYQGDARANGAV------ 46
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPK--GSEPHGDGFTFYIA 118
+L I G T+ V L + T K +DF+T FSF ++ + G +G GF F++A
Sbjct: 47 ELTNIDYTCRAGWATYGKQVPLWNPGTSKPSDFSTRFSFRIDTRNVGYGNYGHGFAFFLA 106
Query: 119 GLDFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDI 177
P + GGFLGLF+ N +SA +V VEFD+FTN WDP + HVGI+
Sbjct: 107 PARIQLPPN-SAGGFLGLFNGTNN-QSSAFPLVYVEFDTFTNPEWDPLDVKS--HVGINN 162
Query: 178 NSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIE-TFVAYEVDLG 236
NS+ S W Q +IG+ LI Y SA + LSV Y + +E + ++Y +DL
Sbjct: 163 NSLVSSNYTSWNATSHNQ-DIGRVLIFYDSARRNLSVSWTYDLTSDPLENSSLSYIIDLS 221
Query: 237 AVLSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
VL V +GF+A +G V+E + +LSW F+S L
Sbjct: 222 KVLPSEVTIGFSATSGGVTEGNRLLSWEFSSSL 254
>At5g65600 receptor protein kinase-like protein
Length = 675
Score = 118 bits (296), Expect = 3e-27
Identities = 95/275 (34%), Positives = 133/275 (47%), Gaps = 23/275 (8%)
Query: 1 MANSFSISLLTTLLPFILLITTVKSDSFSFNFPRF-ESDTKTILSDGDANTT-GGVLQLT 58
M+NS L LPF++ DS FNF F + D I GDA G +
Sbjct: 16 MSNSILFLSLFLFLPFVV-------DSLYFNFTSFRQGDPGDIFYHGDATPDEDGTVNFN 68
Query: 59 KKDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA 118
+ + G T+ V + ++TGK +DF+T FSF ++ + G G F++A
Sbjct: 69 NAE-----QTSQVGWITYSKKVPIWSHKTGKASDFSTSFSFKIDARNLSADGHGICFFLA 123
Query: 119 GLDFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNV-WDPSYPSNSPHVGIDI 177
+ P GGFL LF KN +++S +V VEFD+F N WDP+ HVGI+
Sbjct: 124 PMGAQLPA-YSVGGFLNLFTRKNNYSSSF-PLVHVEFDTFNNPGWDPN--DVGSHVGINN 179
Query: 178 NSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVY---PNSPVKIETFVAYEVD 234
NS+ S W Q +I A ISY S +K LSV Y S K + ++Y +D
Sbjct: 180 NSLVSSNYTSWNASSHSQ-DICHAKISYDSVTKNLSVTWAYELTATSDPKESSSLSYIID 238
Query: 235 LGAVLSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
L VL V+ GF AA G+ +E H +LSW +S L
Sbjct: 239 LAKVLPSDVMFGFIAAAGTNTEEHRLLSWELSSSL 273
>At5g55830 serine/threonine-specific kinase like protein
Length = 681
Score = 95.9 bits (237), Expect = 2e-20
Identities = 77/256 (30%), Positives = 131/256 (51%), Gaps = 26/256 (10%)
Query: 23 VKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSKNSFGQTTFFGAVHL 82
V SD+ +F F F T L GD++ GV+ LT++ G+P +S G + +
Sbjct: 26 VSSDNMNFTFKSFTIRNLTFL--GDSHLRNGVVGLTRE--LGVPDTSS-GTVIYNNPIRF 80
Query: 83 SDNRTGKVADFTTEFSFVVNPKGSEPH--GDGFTFYIAGLDFDFPEDPKEGGFLGLFDPK 140
D + A F+T FSF V +P GDG F+++ D D P GG+LGL
Sbjct: 81 YDPDSNTTASFSTHFSFTVQNLNPDPTSAGDGLAFFLSH-DNDTLGSP--GGYLGLV--- 134
Query: 141 NAFNTSANQVVAVEFDSFTNVWDPSYPS-NSPHVGIDINSIRSVATAPWPIDLVPQGEIG 199
N+ N+ VA+EFD+ DP + N H+G+D++S+ S++T+ + + G
Sbjct: 135 NSSQPMKNRFVAIEFDTKL---DPHFNDPNGNHIGLDVDSLNSISTSDPLLSSQIDLKSG 191
Query: 200 KAL---ISYRSASKVLSVFVVYPNSPVKI-----ETFVAYEVDLGAVLSDWVLVGFTAAT 251
K++ I Y++ ++L+VF+ Y + PV + ++ +DL L+ + VGF+ +T
Sbjct: 192 KSITSWIDYKNDLRLLNVFLSYTD-PVTTTKKPEKPLLSVNIDLSPFLNGEMYVGFSGST 250
Query: 252 GSVSETHDILSWAFNS 267
+E H I +W+F +
Sbjct: 251 EGSTEIHLIENWSFKT 266
>At5g03140 receptor like protein kinase
Length = 711
Score = 92.8 bits (229), Expect = 2e-19
Identities = 87/270 (32%), Positives = 130/270 (47%), Gaps = 24/270 (8%)
Query: 5 FSI-SLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSD---GDANTTGGVLQLTKK 60
FS+ S L +L L + V+ D F RF+ T T+ S GDA+ G ++LT++
Sbjct: 9 FSVYSTLIHILASFLCSSDVRGD---FPATRFDLGTLTLSSLKLLGDAHLNNGTIKLTRE 65
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGL 120
+P+ + G+ + V T A FTT FSF V G G F I+
Sbjct: 66 --LSVPTSTA-GKALYGKPVKFRHPETKSPASFTTYFSFSVTNLNPSSIGGGLAFVISP- 121
Query: 121 DFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSI 180
D D+ GGFLGL + S + VAVEFD+ +V N HVG+D+N++
Sbjct: 122 DEDYLGST--GGFLGLTEETG----SGSGFVAVEFDTLMDVQFKDVNGN--HVGLDLNAV 173
Query: 181 RSVATAPW---PIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGA 237
S A A IDL G + I+Y + +VL+V+V Y N K ++ +DL
Sbjct: 174 VSAAVADLGNVDIDL-KSGNAVNSWITYDGSGRVLTVYVSYSNLKPK-SPILSVPLDLDR 231
Query: 238 VLSDWVLVGFTAATGSVSETHDILSWAFNS 267
+SD + VGF+ +T +E H + W+F+S
Sbjct: 232 YVSDSMFVGFSGSTQGSTEIHSVDWWSFSS 261
>At3g53810 serine/threonine-specific kinase like protein
Length = 677
Score = 86.3 bits (212), Expect = 2e-17
Identities = 81/268 (30%), Positives = 123/268 (45%), Gaps = 35/268 (13%)
Query: 16 FILL--ITTVKSDSFSFNFPRFESDTKTILSDGDANTT-GGVLQLTKKDLYGIPSKNSFG 72
F LL I S + +F + F I G A T G+L+LT S G
Sbjct: 11 FFLLCQIMISSSQNLNFTYNGFHPPLTDISLQGLATVTPNGLLKLTNT------SVQKTG 64
Query: 73 QTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA---GLDFDFPEDPK 129
+ D++ G V+ F+T F F ++ + G G F +A GL F P
Sbjct: 65 HAFCTERIRFKDSQNGNVSSFSTTFVFAIHSQIPTLSGHGIAFVVAPTLGLPFALPSQ-- 122
Query: 130 EGGFLGLFDPKNAFNTSANQVVAVEFDSF--TNVWDPSYPSNSPHVGIDINSIRSV--AT 185
++GLF+ N N + N + AVEFD+ + DP N HVGID+N +RS +T
Sbjct: 123 ---YIGLFNISNNGNDT-NHIFAVEFDTIQSSEFGDP----NDNHVGIDLNGLRSANYST 174
Query: 186 APWPID--------LVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGA 237
A + D L+ + I + I Y + S + V V +S + V+Y DL +
Sbjct: 175 AGYRDDHDKFQNLSLISRKRI-QVWIDYDNRSHRIDVTVAPFDSDKPRKPLVSYVRDLSS 233
Query: 238 VLSDWVLVGFTAATGSVSETHDILSWAF 265
+L + + VGF++ATGSV H ++ W+F
Sbjct: 234 ILLEDMYVGFSSATGSVLSEHFLVGWSF 261
>At4g29050 serine/threonine-specific kinase like protein
Length = 669
Score = 85.1 bits (209), Expect = 4e-17
Identities = 82/275 (29%), Positives = 125/275 (44%), Gaps = 33/275 (12%)
Query: 5 FSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYG 64
F + +L +L F +S+ F F + D I + G+++LT
Sbjct: 3 FFVLVLLLVLQFFSNKALSQSEEGEFGFNGYLYDNSGIA----ITNSKGLMKLTNS---- 54
Query: 65 IPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA---GLD 121
S+ S+G + V ++ G V+ F+T F F + + G G F I+ GL
Sbjct: 55 --SEFSYGHVFYNSPVRFKNSPNGTVSSFSTTFVFAIVSNVNALDGHGLAFVISPTKGLP 112
Query: 122 FDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIR 181
+ +LGLF+ N + S N +VAVEFD+F N +N HVGIDINS+
Sbjct: 113 YS-----SSSQYLGLFNLTNNGDPS-NHIVAVEFDTFQNQEFDDMDNN--HVGIDINSLS 164
Query: 182 S--VATAPWPID---------LVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVA 230
S +TA + D L+ Q I +A I Y S+ + L+V + + P ++
Sbjct: 165 SEKASTAGYYEDDDGTFKNIRLINQKPI-QAWIEYDSSRRQLNVTIHPIHLPKPKIPLLS 223
Query: 231 YEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAF 265
DL L D + VGFT+ATG + +H IL W F
Sbjct: 224 LTKDLSPYLFDSMYVGFTSATGRLRSSHYILGWTF 258
>At5g42120 receptor lectin kinase-like protein
Length = 691
Score = 83.2 bits (204), Expect = 1e-16
Identities = 78/264 (29%), Positives = 122/264 (45%), Gaps = 46/264 (17%)
Query: 29 SFNFP----RFESDTKTILSDGDANTTGGVLQLTKKDLY--------GIPSKNSFGQTTF 76
S +FP RF + + GDA + LT++ PS + G+ +
Sbjct: 18 SLDFPTLSHRFSPPLQNLTLYGDAFFRDRTISLTQQQPCFPSVTTPPSKPSSSGIGRALY 77
Query: 77 FGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGGFLGL 136
+ + T A F+ FSF + S P GDGF F I F GFLGL
Sbjct: 78 VYPIKFLEPSTNTTASFSCRFSFSIIASPSCPFGDGFAFLITSNADSF---VFSNGFLGL 134
Query: 137 FDPKNAFNTSANQVVAVEFDSFTNVWDPSYPS-NSPHVGIDINSIRSVATAPWPIDLVPQ 195
+P ++F +AVEFD+ +DP + N HVGID++SI SV++ +D + +
Sbjct: 135 PNPDDSF-------IAVEFDT---RFDPVHGDINDNHVGIDVSSIFSVSS----VDAISK 180
Query: 196 G---EIGK---ALISYRSASKVLSVFVVY----PNSPVKIETFVAYEVDLGAVLSDWVLV 245
G + GK A I Y K++ V+V Y P SPV ++ ++DL + +++ V
Sbjct: 181 GFDLKSGKKMMAWIEYSDVLKLIRVWVGYSRVKPTSPV-----LSTQIDLSGKVKEYMHV 235
Query: 246 GFTAATGSV-SETHDILSWAFNSF 268
GF+A+ + S H + W F +F
Sbjct: 236 GFSASNAGIGSALHIVERWKFRTF 259
>At3g16530 putative lectin
Length = 276
Score = 82.4 bits (202), Expect = 2e-16
Identities = 71/280 (25%), Positives = 110/280 (38%), Gaps = 40/280 (14%)
Query: 14 LPFILLITTVKSDSFSFNFPRFESDTKTILSDG------DANTTGGVLQLTKKDLYGIPS 67
L F++L + + FNF F+ L D D + G L +T+ + +
Sbjct: 6 LCFLVLFLANAAFAVKFNFDSFDGSNLLFLGDAELGPSSDGVSRSGALSMTRDE-----N 60
Query: 68 KNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPE- 126
S GQ + + + T F T F+F + P+ G GF F I PE
Sbjct: 61 PFSHGQGLYINQIPFKPSNTSSPFSFETSFTFSITPRTKPNSGQGFAFIIT------PEA 114
Query: 127 ---DPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSV 183
+GG+LG+ + N N ++A+EFD+F N N HVG++INS+ S+
Sbjct: 115 DNSGASDGGYLGILNKTND-GKPENHILAIEFDTFQNKEFLDISGN--HVGVNINSMTSL 171
Query: 184 ATAP--------------WPIDLV--PQGEIGKALISYRSASKVLSVFVVYPNSPVKIET 227
W V GE KA + +R+ ++V + N
Sbjct: 172 VAEKAGYWVQTRVGKRKVWSFKDVNLSSGERFKAWVEFRNKDSTITVTLAPENVKKPKRA 231
Query: 228 FVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNS 267
+ L VL + GF + G E HDI SW+F +
Sbjct: 232 LIEAPRVLNEVLLQNMYAGFAGSMGRAVERHDIWSWSFEN 271
>At2g29220 putative protein kinase
Length = 627
Score = 82.0 bits (201), Expect = 3e-16
Identities = 68/227 (29%), Positives = 112/227 (48%), Gaps = 28/227 (12%)
Query: 53 GVLQLTKKDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDG 112
G+L+LT S GQ + LS+ + F+T F F + +G+ G G
Sbjct: 51 GLLELTNT------SMRQIGQAFHGFPIPLSNPNSTNSVSFSTSFIFAIT-QGTGAPGHG 103
Query: 113 FTFYIA-GLDFD--FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSN 169
F I+ +DF FP + +LGLF+ N N S N+++A+EFD+ V N
Sbjct: 104 LAFVISPSMDFSGAFPSN-----YLGLFNTSNNGN-SLNRILAIEFDTVQAVELNDIDDN 157
Query: 170 SPHVGIDINSIRSVATAPWP---------IDL-VPQGEIGKALISYRSASKVLSVFVVYP 219
HVGID+N + S+A+AP I L + G+ + I Y + +L+V +
Sbjct: 158 --HVGIDLNGVISIASAPAAYFDDREAKNISLRLASGKPVRVWIEYNATETMLNVTLAPL 215
Query: 220 NSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFN 266
+ P ++ +++L + S VGF+A+TG+V+ +H +L W+FN
Sbjct: 216 DRPKPSIPLLSRKMNLSGIFSQEHHVGFSASTGTVASSHFVLGWSFN 262
>At4g04960 unknown protein
Length = 686
Score = 80.9 bits (198), Expect = 7e-16
Identities = 79/277 (28%), Positives = 120/277 (42%), Gaps = 39/277 (14%)
Query: 8 SLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPS 67
+LL L F++L + + F FN F + + G A +L LT + +
Sbjct: 3 ALLFLLTLFLILPNPISAIDFIFN--GFNDSSSNVSLFGIATIESKILTLTNQTSFAT-- 58
Query: 68 KNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPED 127
G+ + + D T V F+T F F + P + G G F A P
Sbjct: 59 ----GRALYNRTIRTKDPITSSVLPFSTSFIFTMAPYKNTLPGHGIVFLFA------PST 108
Query: 128 PKEGG----FLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSV 183
G LGLF+ N N S N + VEFD F N +N HVGID+NS+ SV
Sbjct: 109 GINGSSSAQHLGLFNLTNNGNPS-NHIFGVEFDVFANQEFSDIDAN--HVGIDVNSLHSV 165
Query: 184 ---ATAPWPIDLVP-------QGEIGKALISYRSASKVLSVFV---VYPNSPVKIETFVA 230
+ W D V G + I YR +++ V + P P+ ++
Sbjct: 166 YSNTSGYWSDDGVVFKPLKLNDGRNYQVWIDYRDFVVNVTMQVAGKIRPKIPL-----LS 220
Query: 231 YEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNS 267
++L V+ D + VGFTAATG + ++H IL+W+F++
Sbjct: 221 TSLNLSDVVEDEMFVGFTAATGRLVQSHKILAWSFSN 257
>At1g53080 protein kinase, putative
Length = 283
Score = 80.9 bits (198), Expect = 7e-16
Identities = 76/280 (27%), Positives = 125/280 (44%), Gaps = 38/280 (13%)
Query: 16 FILLITTVKSDSFS---FNFPRFESDTKTILSDG------DANTTGGVLQLTKKDLYGIP 66
F+L I+ + S + S FNF RF+ + D + G L +T+ + IP
Sbjct: 9 FVLFISLLSSKTISAVKFNFNRFDGTNLIFIGYAELGPATDGMSRSGALSMTRDN---IP 65
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPH-GDGFTFYIAG-LDFDF 124
+ G T S+N + V F T F+F + P+ S P+ G G F + + +++
Sbjct: 66 FSHGQGLYTDPIPFKSSNNTSSSVYSFKTSFTFSITPRRSNPNPGHGIAFIVVPTVAYEY 125
Query: 125 PEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPS-NSPHVGIDINSIRSV 183
+D G FLGL + N + N + AVEFD F D + N HVG++INS+ S
Sbjct: 126 DQDSTRG-FLGLVNLTTNGNPN-NHLFAVEFDVFQ---DKRFGDINDNHVGVNINSVNSK 180
Query: 184 ATAP--------------WPIDLVP--QGEIGKALISYRSASKVLSVFVVYPNSPVKIET 227
+ W V G+ KA I Y+++ ++ + + P +
Sbjct: 181 VSEKAGYWIQTRTRGKNQWLFKEVKLSSGDNYKAWIEYKNSKVIVWLAPAHLKKPKR--P 238
Query: 228 FVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNS 267
+ +VDL V+ + + GF+ + G E HDI SW+F +
Sbjct: 239 LIETQVDLSEVVLETMYTGFSGSMGRGVERHDIWSWSFEN 278
>At2g37710 putative receptor-like protein kinase
Length = 675
Score = 80.1 bits (196), Expect = 1e-15
Identities = 75/272 (27%), Positives = 117/272 (42%), Gaps = 31/272 (11%)
Query: 10 LTTLLPFILLITTVKSDSFSFNFPR---FESDTKTILSDGDANTTGGVLQLTKKDLYGIP 66
L T+ F +S S S NF F T + T G+L+LT +
Sbjct: 5 LLTIFFFFFFNLIFQSSSQSLNFAYNNGFNPPTDLSIQGITTVTPNGLLKLTNTTV---- 60
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA---GLDFD 123
G + + D+ G V+ F+T F F ++ + + G G F +A L +
Sbjct: 61 --QKTGHAFYTKPIRFKDSPNGTVSSFSTSFVFAIHSQIAILSGHGIAFVVAPNASLPYG 118
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSV 183
P ++GLF+ N N + N V AVE D+ + +N HVGIDINS++SV
Sbjct: 119 NPSQ-----YIGLFNLANNGNET-NHVFAVELDTILST--EFNDTNDNHVGIDINSLKSV 170
Query: 184 ATAP---WP-------IDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEV 233
++P W + L+ + + + + Y + + V + N V
Sbjct: 171 QSSPAGYWDEKGQFKNLTLISRKPM-QVWVDYDGRTNKIDVTMAPFNEDKPTRPLVTAVR 229
Query: 234 DLGAVLSDWVLVGFTAATGSVSETHDILSWAF 265
DL +VL + VGF++ATGSV H IL W+F
Sbjct: 230 DLSSVLLQDMYVGFSSATGSVLSEHYILGWSF 261
>At1g53070 protein kinase, putative
Length = 272
Score = 80.1 bits (196), Expect = 1e-15
Identities = 80/283 (28%), Positives = 122/283 (42%), Gaps = 37/283 (13%)
Query: 7 ISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDAN--------TTGGVLQLT 58
I TTL F+ + T+ + ++ F F F + T I GDA + G + LT
Sbjct: 5 ILCFTTL--FLAIFTSQVTTAYKFKFDYFGNGTDPISFHGDAEYGPDTDGKSRSGAIALT 62
Query: 59 KKDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA 118
+ ++ S G+ F + N + + F T F+F + PK + G G F +
Sbjct: 63 RDNI-----PFSHGRAIFTTPITFKPNASA-LYPFKTSFTFSITPKTNPNQGHGLAFIVV 116
Query: 119 GLDFDFPEDPKEG-GFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPS-NSPHVGID 176
+ D G G+L L + N N + N + AVEFD F D S N HVGID
Sbjct: 117 PSN---QNDAGSGLGYLSLLNRTNNGNPN-NHLFAVEFDVFQ---DKSLGDMNDNHVGID 169
Query: 177 INSIRSVATAP---WPI--------DL-VPQGEIGKALISYRSASKVLSVFVVYPNSPVK 224
INS+ SV + W + DL + G+ KA I Y + KV+SV + +
Sbjct: 170 INSVDSVVSVKSGYWVMTRSGWLFKDLKLSSGDRYKAWIEYNNNYKVVSVTIGLAHLKKP 229
Query: 225 IETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNS 267
+ + DL V+ + + GF + G E H+I W F +
Sbjct: 230 NRPLIEAKFDLSKVIHEVMYTGFAGSMGRGVERHEIWDWTFQN 272
>At3g53380 receptor lectin kinase -like protein
Length = 715
Score = 79.7 bits (195), Expect = 1e-15
Identities = 76/264 (28%), Positives = 121/264 (45%), Gaps = 19/264 (7%)
Query: 7 ISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIP 66
+SL + ILL + + F+F +L GDA + G++ LT+ DL +P
Sbjct: 1 MSLFLSFFISILLCFFNGATTTQFDFSTLAISNLKLL--GDARLSNGIVGLTR-DL-SVP 56
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPE 126
+ + G+ + + T F++ FSF + G G F I+ D
Sbjct: 57 NSGA-GKVLYSNPIRFRQPGTHFPTSFSSFFSFSITNVNPSSIGGGLAFVISP---DANS 112
Query: 127 DPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATA 186
GG LGL P N S ++ VAVEFD+ +V NS HVG D+N + S +
Sbjct: 113 IGIAGGSLGLTGP----NGSGSKFVAVEFDTLMDV--DFKDINSNHVGFDVNGVVSSVSG 166
Query: 187 PWP---IDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWV 243
IDL G + I Y ++V +V V Y N K+ +++ +DL ++D++
Sbjct: 167 DLGTVNIDL-KSGNTINSWIEYDGLTRVFNVSVSYSNLKPKVP-ILSFPLDLDRYVNDFM 224
Query: 244 LVGFTAATGSVSETHDILSWAFNS 267
VGF+ +T +E H I W+F+S
Sbjct: 225 FVGFSGSTQGSTEIHSIEWWSFSS 248
>At4g28350 receptor protein kinase like protein
Length = 649
Score = 79.0 bits (193), Expect = 3e-15
Identities = 72/272 (26%), Positives = 123/272 (44%), Gaps = 26/272 (9%)
Query: 7 ISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIP 66
+S+L L +LL + F +N F + +L + + +L LT + + I
Sbjct: 5 VSILLFSLASLLLFRSTTGIEFIYN-SNFTTTNTLLLGNATVKSPPSILTLTNQTTFSI- 62
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPE 126
G+ + ++ S + + F T F F + P G GF F L F
Sbjct: 63 -----GRGLYPSRINASSSSASPLP-FATSFIFSMAPFKHLSPGHGFAFVF--LPFSETS 114
Query: 127 DPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATA 186
LGLF+ N + ++ ++ AVEFD F N N HVG+D+NS+ SVA+
Sbjct: 115 AASSSQHLGLFNFTNNGDPNS-RIFAVEFDVFAN--QEFNDINDNHVGVDVNSLTSVASE 171
Query: 187 P-----------WPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDL 235
+ + GE +A I + ++ ++V + +S I ++ ++L
Sbjct: 172 TAGFYGGRDGQRFTELKLNSGENYQAWIEFNGSA--INVTMARASSRKPIRPLISIPLNL 229
Query: 236 GAVLSDWVLVGFTAATGSVSETHDILSWAFNS 267
VL D + VGFTA+TG + ++H ILSW+F++
Sbjct: 230 TGVLLDDMFVGFTASTGQLVQSHRILSWSFSN 261
>At5g01560 receptor like protein kinase
Length = 685
Score = 75.9 bits (185), Expect = 2e-14
Identities = 77/280 (27%), Positives = 126/280 (44%), Gaps = 32/280 (11%)
Query: 7 ISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDAN-TTGGVLQLTKKDLYGI 65
+SLL L + T ++ + F F F+ + I GD+ T+ G+L+LT ++
Sbjct: 2 VSLLLVLFLVRAHVATTETTT-EFIFHGFKGNQSEIHMQGDSTITSNGLLRLTDRN---- 56
Query: 66 PSKNSFGQTTFFGAVHLSDNRTGK--VADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
+ G + V L D+ + V F+T F F++ + G GFTF ++
Sbjct: 57 --SDVVGTAFYHKPVRLLDSNSTNTTVRSFSTSFIFIIPSSSTSNGGFGFTFTLSPTPNR 114
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSV 183
DP++ ++GL + +N N S+N V AVEFD+ D + + H+G++ NS+ S
Sbjct: 115 TDADPEQ--YMGLLNERNDGN-SSNHVFAVEFDTVQGFKDGTNRIGN-HIGLNFNSLSSD 170
Query: 184 ATAP-----------WPIDLVPQGEIGKALISYRSASKVLSVFVVYPNS---PVKIETFV 229
P LV GE + + Y +K L++ VYP +I
Sbjct: 171 VQEPVAYFNNNDSQKEEFQLV-SGEPIQVFLDYHGPTKTLNL-TVYPTRLGYKPRIPLIS 228
Query: 230 AYEVDLGAVLSDWVLVGFTAATG--SVSETHDILSWAFNS 267
L ++ D + VGFTAATG S H ++ W+F S
Sbjct: 229 REVPKLSDIVVDEMFVGFTAATGRHGQSSAHYVMGWSFAS 268
>At5g01550 receptor like protein kinase
Length = 688
Score = 75.1 bits (183), Expect = 4e-14
Identities = 73/273 (26%), Positives = 123/273 (44%), Gaps = 43/273 (15%)
Query: 21 TTVKSDSFSFNFPRFESDTKTILSDGDANTT-GGVLQLTKKDLYGIPSKNSFGQTTFFGA 79
TT ++ F F F + I++ G A G+L+LT ++ N G + +
Sbjct: 17 TTTETPKTEFIFRGFSGNQSNIVTTGAATIKLDGLLRLTDRN------SNVTGTSFYHKP 70
Query: 80 VHLSDNRTGK----VADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDPKEGG--- 132
V L + T + F+T F FV+ P S G GFTF ++ P + G
Sbjct: 71 VRLLETNTSSTNSTIRSFSTSFVFVIIPTSSSNGGFGFTFTLS------PTPDRTGAESA 124
Query: 133 -FLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSVATAP---- 187
+LGL + N N S N V AVEFD+ D + + + H+G++ NS+ S P
Sbjct: 125 QYLGLLNKANDGN-STNHVFAVEFDTVQGFKDGADRTGN-HIGLNFNSLTSDVQEPVVYY 182
Query: 188 ----------WPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVK-IETFVAYEV-DL 235
+P+ G+ +A++ Y ++ L++ V N + + ++ V L
Sbjct: 183 DNEDPNRKEDFPLQ---SGDPIRAILDYDGPTQTLNLTVYPANLKSRPVRPLISRPVPKL 239
Query: 236 GAVLSDWVLVGFTAATG-SVSETHDILSWAFNS 267
++ + + VGFTAATG S H ++ W+F+S
Sbjct: 240 SQIVQEEMYVGFTAATGRDQSSAHYVMGWSFSS 272
>At3g59700 serine/threonine-specific kinase lecRK1 precursor,lectin
receptor-like
Length = 661
Score = 74.7 bits (182), Expect = 5e-14
Identities = 61/218 (27%), Positives = 103/218 (46%), Gaps = 24/218 (11%)
Query: 67 SKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA---GLDFD 123
+K++FGQ V + ++ TG ++ F+ F F + P+ ++ G TF I+ GL
Sbjct: 47 TKHTFGQAFENEHVEIKNSSTGVISSFSVNFFFAIVPEHNQQGSHGMTFVISPTRGLP-- 104
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSV 183
+LG+F+ N S N V+A+E D + N HVGI+IN +RSV
Sbjct: 105 ---GASSDQYLGIFNKTNNGKAS-NNVIAIELDIHKDEEFGDIDDN--HVGININGLRSV 158
Query: 184 ATAPW-----------PIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVK-IETFVAY 231
A+A + L+ + E+ + I Y + L+V + PV ++ ++
Sbjct: 159 ASASAGYYDDKDGSFKKLSLISR-EVMRLSIVYSQPDQQLNVTLFPAEIPVPPLKPLLSL 217
Query: 232 EVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFNSFL 269
DL L + + +GFTA+TGSV H ++ W N +
Sbjct: 218 NRDLSPYLLEKMYLGFTASTGSVGAIHYLMGWLVNGVI 255
>At2g29250 putative protein kinase
Length = 623
Score = 73.9 bits (180), Expect = 8e-14
Identities = 73/277 (26%), Positives = 124/277 (44%), Gaps = 26/277 (9%)
Query: 1 MANSFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKK 60
MAN++ S+ +++ F+L + V S + + + S G L+LT
Sbjct: 1 MANTYK-SIAVSII-FLLYFSCVSSQQQTKFLNHGFLEANLLKSGSSKIHPSGHLELTNT 58
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA-G 119
S GQ + + + + F T F F + P P G G F I+
Sbjct: 59 ------SMRQIGQAFHGFPIPFLNPNSSNLVSFPTSFVFAITPGPGAP-GHGLAFVISPS 111
Query: 120 LDFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINS 179
LDF +LGLF+ N N S N ++AVEFD+ V N HVGID+N
Sbjct: 112 LDFS---GALPSNYLGLFNTSNNGN-SLNCILAVEFDTVQAVELNDIDDN--HVGIDLNG 165
Query: 180 IRSV-ATAPWPID---------LVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFV 229
+ S+ +T+ D + G+ + I Y + +L+V + + P +
Sbjct: 166 VISIESTSAEYFDDREAKNISLRLASGKPIRVWIEYNATETMLNVTLAPLDRPKPKLPLL 225
Query: 230 AYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFN 266
+ +++L ++S+ VGF+AATG+V+ +H +L W+F+
Sbjct: 226 SRKLNLSGIISEENYVGFSAATGTVTSSHFVLGWSFS 262
>At3g55550 probable serine/threonine-specific protein kinase
Length = 684
Score = 73.6 bits (179), Expect = 1e-13
Identities = 76/276 (27%), Positives = 120/276 (42%), Gaps = 27/276 (9%)
Query: 1 MANSFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKK 60
M+ +F++ LL LL F+ + + FSF + S T+ + TG + T+
Sbjct: 1 MSQTFAVILL--LLIFLTHLVSSLIQDFSFIGFKKASPNLTLNGVAEIAPTGAIRLTTE- 57
Query: 61 DLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGL 120
++ G + + + F+T F+ + P+ G G F I
Sbjct: 58 ------TQRVIGHAFYSLPIRFKPIGVNRALSFSTSFAIAMVPEFVTLGGHGLAFAITPT 111
Query: 121 DFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPS-NSPHVGIDINS 179
P + +LGL + ++ AVEFD+ V D + N HVGIDINS
Sbjct: 112 PDLRGSLPSQ--YLGLLNSSRV--NFSSHFFAVEFDT---VRDLEFEDINDNHVGIDINS 164
Query: 180 IRSVATAPWPIDLVPQ---------GEIGKALISYRSASKVLSVFVVYPNSPVKIETFVA 230
+ S + P L G + +A I Y S K L V + P S + ++
Sbjct: 165 MESSISTPAGYFLANSTKKELFLDGGRVIQAWIDYDSNKKRLDV-KLSPFSEKPKLSLLS 223
Query: 231 YEVDLGAVLSDWVLVGFTAATGSVSETHDILSWAFN 266
Y+VDL +VL D + VGF+A+TG ++ +H IL W FN
Sbjct: 224 YDVDLSSVLGDEMYVGFSASTGLLASSHYILGWNFN 259
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,283,294
Number of Sequences: 26719
Number of extensions: 287073
Number of successful extensions: 691
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 43
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 570
Number of HSP's gapped (non-prelim): 70
length of query: 269
length of database: 11,318,596
effective HSP length: 98
effective length of query: 171
effective length of database: 8,700,134
effective search space: 1487722914
effective search space used: 1487722914
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0541.11


