

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
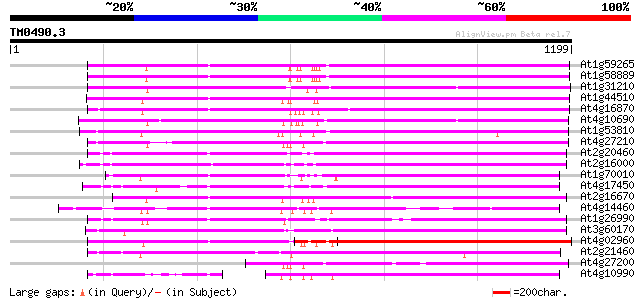
Query= TM0490.3
(1199 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g59265 polyprotein, putative 794 0.0
At1g58889 polyprotein, putative 792 0.0
At1g31210 putative reverse transcriptase 789 0.0
At1g44510 polyprotein, putative 783 0.0
At4g16870 retrotransposon like protein 764 0.0
At4g10690 retrotransposon like protein 761 0.0
At1g53810 745 0.0
At4g27210 putative protein 690 0.0
At2g20460 putative retroelement pol polyprotein 684 0.0
At2g16000 putative retroelement pol polyprotein 627 e-179
At1g70010 hypothetical protein 615 e-176
At4g17450 retrotransposon like protein 602 e-172
At2g16670 putative retroelement pol polyprotein 575 e-164
At4g14460 retrovirus-related like polyprotein 572 e-163
At1g26990 polyprotein, putative 550 e-156
At3g60170 putative protein 535 e-152
At4g02960 putative polyprotein of LTR transposon 523 e-148
At2g21460 putative retroelement pol polyprotein 497 e-140
At4g27200 putative protein 482 e-136
At4g10990 putative retrotransposon polyprotein 444 e-124
>At1g59265 polyprotein, putative
Length = 1466
Score = 794 bits (2050), Expect = 0.0
Identities = 466/1142 (40%), Positives = 624/1142 (53%), Gaps = 118/1142 (10%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GA+ H + L+ + + ++V G IPI +G TS+ T +PL L++
Sbjct: 331 WLLDSGATHHITSDFNNLSLHQPYTG-GDDVMVADGSTIPISHTGSTSLSTKSRPLNLHN 389
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY--PVTRS 283
+L+ P I KNLISV +L N VSV F P F V D TG+PLL+ + +LY P+ S
Sbjct: 390 ILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWPIASS 449
Query: 284 SPFAGLAS-------SVWHNRLGHPASSALNHLRNN-KLIFCEPSRSSSVCDSCVLGKHV 335
P + AS S WH RLGHPA S LN + +N L PS C C++ K
Sbjct: 450 QPVSLFASPSSKATHSSWHARLGHPAPSILNSVISNYSLSVLNPSHKFLSCSDCLINKSN 509
Query: 336 RLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYE 395
++PFS S I + RP + ++SD+W+SP+LS +RYYV+F+D T + W +P+ +KSQV E
Sbjct: 510 KVPFSQSTINSTRPLEYIYSDVWSSPILSHDNYRYYVIFVDHFTRYTWLYPLKQKSQVKE 569
Query: 396 TFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAER 455
TF T L++ +F I DNG E+ + Y +G+ S PHT NG +ER
Sbjct: 570 TFITFKNLLENRFQTRIGTFYSDNGGEFV--ALWEYFSQHGISHLTSPPHTPEHNGLSER 627
Query: 456 KIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSH 515
K R I TLL+H+S+P ++W +A +A YL+N LP LQ +SP Q L+ P+Y
Sbjct: 628 KHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLFGTSPNYDK 687
Query: 516 LRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDET 575
LRVFGC C+P +KL +S CVFLGY + Y C L ++ ISRHV FDE
Sbjct: 688 LRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYISRHVRFDEN 747
Query: 576 RFPFADLSLTPAPSYECFTED---------LP--------PSLI--HHWQTVSSRP---- 612
FPF++ T +P E E LP PS HH T S P
Sbjct: 748 CFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATPPSSPSAPF 807
Query: 613 -------------------------------PDPPVQP---------------SSPTDSS 626
P P QP ++PT+ S
Sbjct: 808 RNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTSQNNPTNES 867
Query: 627 PP--LSALVSPTASPSPLPLP----------PVPPA------PPV---------RTMTTR 659
P +L +P S S P P P PP+ PP+ + T
Sbjct: 868 PSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTH 927
Query: 660 SM-----HGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNT 714
SM GI KP +SL+VS+ + P QAL D W++AM SE NA I ++T
Sbjct: 928 SMGTRAKAGIIKPNPKYSLAVSL---AAESEPRTAIQALKDERWRNAMGSEINAQIGNHT 984
Query: 715 WELV-PRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKP 773
W+LV P P V ++ C WIF K S+G RYKARLV G +Q G+D ETFSPV+K
Sbjct: 985 WDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKS 1044
Query: 774 ATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLY 833
+IR VL +A+ RSWPI QLDV NAFL G L + +YM P GF D P+YVC+L+K+LY
Sbjct: 1045 TSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALY 1104
Query: 834 GLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLR 893
GLKQAPRAWY +Y+ +IGF +S SD SLF+ ++G I Y+L+YVDDI++ + L
Sbjct: 1105 GLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILITGNDPTLL 1164
Query: 894 KSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATP 953
+ + L+ F++KD L YFLGI R GL LSQ Y +++AR M + P TP
Sbjct: 1165 HNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTGLHLSQRRYILDLLARTNMITAKPVTTP 1224
Query: 954 VDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLAL 1013
+ KLS SGT D + YR + G+LQYL FTRPDISYAV ++ MH P +H+ AL
Sbjct: 1225 MAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMPTEEHLQAL 1284
Query: 1014 KRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSK 1073
KR+LRY+ GT +G+ L +L +Y+DADW G D ST+GY V+LG + ISWSSK
Sbjct: 1285 KRILRYLAGTPNHGIFLKKGNTLSLHAYSDADWAGDKDDYVSTNGYIVYLGHHPISWSSK 1344
Query: 1074 RQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVH 1133
+Q + RSS EAEYR VAN SE WI +LL EL L++ +++CDNV A YL NPV
Sbjct: 1345 KQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNVGATYLCANPVF 1404
Query: 1134 HQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPP 1193
H R KHI +D HF+R +V G RV+HV + Q+AD TK L R F +F S + V P
Sbjct: 1405 HSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQNFASKIGVTRVP 1464
Query: 1194 AS 1195
S
Sbjct: 1465 PS 1466
>At1g58889 polyprotein, putative
Length = 1466
Score = 792 bits (2046), Expect = 0.0
Identities = 465/1142 (40%), Positives = 623/1142 (53%), Gaps = 118/1142 (10%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GA+ H + L+ + + ++V G IPI +G TS+ T +PL L++
Sbjct: 331 WLLDSGATHHITSDFNNLSLHQPYTG-GDDVMVADGSTIPISHTGSTSLSTKSRPLNLHN 389
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY--PVTRS 283
+L+ P I KNLISV +L N VSV F P F V D TG+PLL+ + +LY P+ S
Sbjct: 390 ILYVPNIHKNLISVYRLCNANGVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWPIASS 449
Query: 284 SPFAGLAS-------SVWHNRLGHPASSALNHLRNN-KLIFCEPSRSSSVCDSCVLGKHV 335
P + AS S WH RLGHPA S LN + +N L PS C C++ K
Sbjct: 450 QPVSLFASPSSKATHSSWHARLGHPAPSILNSVISNYSLSVLNPSHKFLSCSDCLINKSN 509
Query: 336 RLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYE 395
++PFS S I + RP + ++SD+W+SP+LS +RYYV+F+D T + W +P+ +KSQV E
Sbjct: 510 KVPFSQSTINSTRPLEYIYSDVWSSPILSHDNYRYYVIFVDHFTRYTWLYPLKQKSQVKE 569
Query: 396 TFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAER 455
TF T L++ +F I DNG E+ + Y +G+ S PHT NG +ER
Sbjct: 570 TFITFKNLLENRFQTRIGTFYSDNGGEFV--ALWEYFSQHGISHLTSPPHTPEHNGLSER 627
Query: 456 KIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSH 515
K R I TLL+H+S+P ++W +A +A YL+N LP LQ +SP Q L+ P+Y
Sbjct: 628 KHRHIVETGLTLLSHASIPKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLFGTSPNYDK 687
Query: 516 LRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDET 575
LRVFGC C+P +KL +S CVFLGY + Y C L ++ ISRHV FDE
Sbjct: 688 LRVFGCACYPWLRPYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYISRHVRFDEN 747
Query: 576 RFPFADLSLTPAPSYECFTED---------LP--------PSLI--HHWQTVSSRP---- 612
FPF++ T +P E E LP PS HH T S P
Sbjct: 748 CFPFSNYLATLSPVQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAATPPSSPSAPF 807
Query: 613 -------------------------------PDPPVQP---------------SSPTDSS 626
P P QP ++PT+ S
Sbjct: 808 RNSQVSSSNLDSSFSSSFPSSPEPTAPRQNGPQPTTQPTQTQTQTHSSQNTSQNNPTNES 867
Query: 627 PP--LSALVSPTASPSPLPLP----------PVPPA------PPV---------RTMTTR 659
P +L +P S S P P P PP+ PP+ + T
Sbjct: 868 PSQLAQSLSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTH 927
Query: 660 SM-----HGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNT 714
SM GI KP +SL+VS+ + P QAL D W++AM SE NA I ++T
Sbjct: 928 SMGTRAKAGIIKPNPKYSLAVSL---AAESEPRTAIQALKDERWRNAMGSEINAQIGNHT 984
Query: 715 WELV-PRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKP 773
W+LV P P V ++ C WIF K S+G RYKAR V G +Q G+D ETFSPV+K
Sbjct: 985 WDLVPPPPSHVTIVGCRWIFTKKYNSDGSLNRYKARFVAKGYNQRPGLDYAETFSPVIKS 1044
Query: 774 ATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLY 833
+IR VL +A+ RSWPI QLDV NAFL G L + +YM P GF D P+YVC+L+K+LY
Sbjct: 1045 TSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALY 1104
Query: 834 GLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLR 893
GLKQAPRAWY +Y+ +IGF +S SD SLF+ ++G I Y+L+YVDDI++ + L
Sbjct: 1105 GLKQAPRAWYVELRNYLLTIGFVNSVSDTSLFVLQRGKSIVYMLVYVDDILITGNDPTLL 1164
Query: 894 KSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATP 953
+ + L+ F++KD L YFLGI R GL LSQ Y +++AR M + P TP
Sbjct: 1165 HNTLDNLSQRFSVKDHEELHYFLGIEAKRVPTGLHLSQRRYILDLLARTNMITAKPVTTP 1224
Query: 954 VDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLAL 1013
+ KLS SGT D + YR + G+LQYL FTRPDISYAV ++ MH P +H+ AL
Sbjct: 1225 MAPSPKLSLYSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMPTEEHLQAL 1284
Query: 1014 KRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSK 1073
KR+LRY+ GT +G+ L +L +Y+DADW G D ST+GY V+LG + ISWSSK
Sbjct: 1285 KRILRYLAGTPNHGIFLKKGNTLSLHAYSDADWAGDKDDYVSTNGYIVYLGHHPISWSSK 1344
Query: 1074 RQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVH 1133
+Q + RSS EAEYR VAN SE WI +LL EL L++ +++CDNV A YL NPV
Sbjct: 1345 KQKGVVRSSTEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNVGATYLCANPVF 1404
Query: 1134 HQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPP 1193
H R KHI +D HF+R +V G RV+HV + Q+AD TK L R F +F S + V P
Sbjct: 1405 HSRMKHIAIDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQNFASKIGVTRVP 1464
Query: 1194 AS 1195
S
Sbjct: 1465 PS 1466
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 789 bits (2037), Expect = 0.0
Identities = 435/1064 (40%), Positives = 599/1064 (55%), Gaps = 45/1064 (4%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W+ D+ A++H +S L S + + ++VG G +PI +G T+I + + + LN
Sbjct: 322 WHPDSAATAHVTSSTNGLQSATEYEG-DDAVLVGDGTYLPITHTGSTTIKSSNGKIPLNE 380
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSP 285
VL P I K+L+SV +L D V FD + D +T + LY V +
Sbjct: 381 VLVVPNIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLQTQKVVTTGPRRNGLY-VLENQE 439
Query: 286 FAGLASS--------VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRL 337
F L S+ VWH+RLGH S AL HL+N+K I SR+S VC+ C +GK RL
Sbjct: 440 FVALYSNRQCAATEEVWHHRLGHANSKALQHLQNSKAIQINKSRTSPVCEPCQMGKSSRL 499
Query: 338 PFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYET 396
PF S L P D +H DLW SPV+S G +YY +F+DD++ + W +P+ KS+
Sbjct: 500 PFLISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWFYPLHNKSEFLSV 559
Query: 397 FTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERK 456
F + L++ Q + IK Q D G E+ ++ + +G+ R SCP+T QNG AERK
Sbjct: 560 FISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCPYTPQQNGLAERK 619
Query: 457 IRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHL 516
R + + ++L HS P FW + A Y++N LP L+N SP + L+ P YS L
Sbjct: 620 HRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPYEALFGEKPDYSSL 679
Query: 517 RVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETR 576
RVFG C+P NK PRS CVFLGY ++GY+C+ K+ ISR+VIF+E+
Sbjct: 680 RVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKVYISRNVIFNESE 739
Query: 577 FPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQP----SSPTDSSPPLSAL 632
PF + + P Y L+ WQ P P S P D + +
Sbjct: 740 LPFKEKYQSLVPQYST-------PLLQAWQHNKISEISVPAAPVQLFSKPIDLNTYAGSQ 792
Query: 633 VS-------PTASPSPLPLPPVPPAPPV----------RTMTTRSMHGISKPKKPFSLSV 675
V+ PT++ P A + MTTRS GI KP ++L
Sbjct: 793 VTEQLTDPEPTSNNEGSDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQKPNTRYALIT 852
Query: 676 SIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRH 735
S + + P A+ P W A+ E N + +TW LVP D+N++ W+F+
Sbjct: 853 SRMNTAE---PKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNILSSKWVFKT 909
Query: 736 KKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDV 795
K +G ++ KARLV G Q GVD ETFSPVV+ ATIR VL ++ S+ WPI QLDV
Sbjct: 910 KLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKGWPIKQLDV 969
Query: 796 QNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGF 855
NAFLHG+L E ++M+ P GF DP P +VCRL K++YGLKQAPRAW+ F++++ GF
Sbjct: 970 SNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGF 1029
Query: 856 RHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYF 915
S SD SLF+ Q I Y+LLYVDDI+L S L + + L + F+MKDLGP YF
Sbjct: 1030 VCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYF 1089
Query: 916 LGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYR 975
LGI + +A GLFL Q+ YAT+I+ +AGM+ CNP TP+ Q+L + + + + +R
Sbjct: 1090 LGIQIEDYANGLFLHQTAYATDILQQAGMSDCNPMPTPL--PQQLDNLNSELFAEPTYFR 1147
Query: 976 SLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPV 1035
SL G LQYLT TRPDI +AV +C MH+P T LKR+LRY++GT+ GL + +
Sbjct: 1148 SLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMGLPIKRNST 1207
Query: 1036 ETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVS 1095
TL +Y+D+D GC +TRRST+G+C+ LG NLISWS+KRQPT+S SS EAEYR +
Sbjct: 1208 LTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAR 1267
Query: 1096 ESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQ 1155
E WI LL +L P T V+CDN+SA+YLS NP H R+KH + D H++RE+V G
Sbjct: 1268 EITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGL 1327
Query: 1156 ARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV-GEPPASTAG 1198
H+ + Q+AD+FTK LPR F D RS L V G P S G
Sbjct: 1328 IETQHISATFQLADVFTKSLPRRAFVDLRSKLGVSGSPTPSLRG 1371
>At1g44510 polyprotein, putative
Length = 1459
Score = 783 bits (2022), Expect = 0.0
Identities = 450/1128 (39%), Positives = 625/1128 (54%), Gaps = 101/1128 (8%)
Query: 165 PWYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALN 224
PW +D+GA+ H + L+ + + + +++ G G+ I+ +G T +P+ ++ LAL+
Sbjct: 336 PWLLDSGATHHITSDLNALSLHQPYNG-GEYVMIADGTGLTIKQTGSTFLPSQNRDLALH 394
Query: 225 HVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY--PVTR 282
VL+ P I KNLISV +L N VSV F P F V D TG LL+ + DLY PVT
Sbjct: 395 KVLYVPDIRKNLISVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDDLYEWPVTN 454
Query: 283 -------SSPFAGLASSVWHNRLGHPASSALNHLRNN-KLIFCEPSRSSSVCDSCVLGKH 334
+SP S WH+RLGHP++S LN L + L S + + C C++ K
Sbjct: 455 PPATALFTSPSPKTTLSSWHSRLGHPSASILNTLLSKFSLPVSVASSNKTSCSDCLINKS 514
Query: 335 VRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVY 394
+LPF++S+I + P + + +D+WTSP++S ++YY++ +D +T + W +P+ +KSQV
Sbjct: 515 HKLPFATSSIHSSSPLEYIFTDVWTSPIISHDNYKYYLVLVDHYTRYTWLYPLQQKSQVK 574
Query: 395 ETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAE 454
TF L++ +F A I+ L DNG E+ + + +NG+ S PHT NG +E
Sbjct: 575 ATFIAFKALVENRFQAKIRTLYSDNGGEFI--ALRDFLVSNGISHLTSPPHTPEHNGLSE 632
Query: 455 RKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYS 514
RK R I TLL +SVP +W +A A YL+N +P L SP Q L+ P+Y
Sbjct: 633 RKHRHIVETGLTLLTQASVPREYWTYAFATAVYLINRMPTPVLCLQSPFQKLFGSSPNYQ 692
Query: 515 HLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDE 574
LRVFGCLCFP T NKL+ RS CVFLGY + Y C D+ + ++ SRHV+FDE
Sbjct: 693 RLRVFGCLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDVDNNRLYTSRHVMFDE 752
Query: 575 TRFPFA-----------------DLSLTPAPS-YECFT--------------------ED 596
+ +PFA S +PA S + C D
Sbjct: 753 STYPFAASIREQSQSSLVTPPESSSSSSPANSGFPCSVLRLQSPPASSPETPSPPQQQND 812
Query: 597 LP--------PSLIHHWQ----TVSSRPPDPPVQPSSPTDSSPPLSALVSPTA---SPSP 641
P P+ HH Q T+S P +P++P ++ P A +P + P P
Sbjct: 813 SPVSPRQTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHENGPEPEAQSNPNSPFIGPLP 872
Query: 642 LPLPPVPPA----------------PPVRT------------------MTTRSMHGISKP 667
P P P+ PP +T M TRS + I+KP
Sbjct: 873 NPNPETNPSSSIEQRPVDKSTTTALPPNQTTIAATSNSRSQPPKNNHQMKTRSKNNITKP 932
Query: 668 KKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVI 727
K SL+V++ P +S P+ QAL D W+ AM EF+A R++TW+LVP +++
Sbjct: 933 KTKTSLTVALTQPHLSE-PNTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPPNPTQHLV 991
Query: 728 RCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRS 787
C W+F+ K NGL ++YKARLV G +Q GVD ETFSPV+K TIR VL +A+ ++
Sbjct: 992 GCRWVFKLKYLPNGLIDKYKARLVAKGFNQQYGVDYAETFSPVIKATTIRVVLDVAVKKN 1051
Query: 788 WPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFA 847
WP+ QLDV NAFL G L E +YM P GF D P +VCRL+K++YGLKQAPRAWY
Sbjct: 1052 WPLKQLDVNNAFLQGTLTEEVYMAQPPGFVDKDRPSHVCRLRKAIYGLKQAPRAWYMELK 1111
Query: 848 DYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMK 907
++ +IGF +S +D SLFI+ G+ + Y+L+YVDDII+ S H + ++ LA F++K
Sbjct: 1112 QHLLNIGFVNSLADTSLFIYSHGTTLLYLLVYVDDIIVTGSDHKSVSAVLSSLAERFSIK 1171
Query: 908 DLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTP 967
D L YFLGI TR GL L Q Y T+++A+ M P ATP+ T KL+ GT
Sbjct: 1172 DPTDLHYFLGIEATRTNTGLHLMQRKYMTDLLAKHNMLDAKPVATPLPTSPKLTLHGGTK 1231
Query: 968 CEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYG 1027
DAS YRS+ G+LQYL FTRPDI++AV ++ MH P + H A KRVLRY+ GT T+G
Sbjct: 1232 LNDASEYRSVVGSLQYLAFTRPDIAFAVNRLSQFMHQPTSDHWQAAKRVLRYLAGTTTHG 1291
Query: 1028 LHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEY 1087
+ L S L +++DADW G ST+ Y ++LG N ISWSSK+Q +SRSS E+EY
Sbjct: 1292 IFLNSSSPIHLHAFSDADWAGDSADYVSTNAYVIYLGRNPISWSSKKQRGVSRSSTESEY 1351
Query: 1088 RGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFV 1147
R VAN SE W+ +LL ELH L + CDN+ A Y+ NPV H R KHI +D HFV
Sbjct: 1352 RAVANAASEIRWLCSLLTELHIRLPHGPTIFCDNIGATYICANPVFHSRMKHIALDYHFV 1411
Query: 1148 REKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPAS 1195
R + RV HV + Q+AD TK L R F RS + V + P S
Sbjct: 1412 RGMIQSRALRVSHVSTNDQLADALTKSLSRPHFLSARSKIGVRQLPPS 1459
Score = 32.3 bits (72), Expect = 1.7
Identities = 42/178 (23%), Positives = 69/178 (38%), Gaps = 26/178 (14%)
Query: 19 PPASVDSSQQRPTYRSDQGHSNHRYGSGQNRNYQGGSGKPRKKGGSRYPGSPGSSGSSAA 78
PPAS + P ++D S + GS ++ + S SP S S
Sbjct: 796 PPASSPETPSPPQQQNDSPVSPRQTGSPTPSHHS-------QVRDSTLSPSPSVSNSEPT 848
Query: 79 SPPPWRPPPQASWNPWGWDPPPGWAPSPWGMPPCPYPTSQWSRPMGVPQQPGVLGPRPQA 138
+P P P+A NP SP+ + P P P + + + Q+ P ++
Sbjct: 849 APHENGPEPEAQSNP----------NSPF-IGPLPNPNPETNPSSSIEQR-----PVDKS 892
Query: 139 YTATASPAPTDIAAAMHTMSLTPPDTPWYMDTGASSHTA--ASQGTLTSYSNLSHLNQ 194
T P T IAA ++ S PP M T + ++ ++ +LT HL++
Sbjct: 893 TTTALPPNQTTIAATSNSRS-QPPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSE 949
>At4g16870 retrotransposon like protein
Length = 1474
Score = 764 bits (1974), Expect = 0.0
Identities = 448/1148 (39%), Positives = 614/1148 (53%), Gaps = 124/1148 (10%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GA+ H + L + + +++ G + I +G T +P+ + L LN
Sbjct: 333 WLLDSGATHHITSDLNALALHQPYN--GDDVMIADGTSLKITKTGSTFLPSNARDLTLNK 390
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY--PVTR- 282
VL+ P I KNL+SV +L N VSV F P F V D TG LL+ + +LY PVT
Sbjct: 391 VLYVPDIQKNLVSVYRLCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDELYEWPVTNP 450
Query: 283 ------SSPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSV-CDSCVLGKHV 335
++P S WH+RLGHP+SS LN L + + S S+ + C C + K
Sbjct: 451 KATALFTTPSPKTTLSSWHSRLGHPSSSILNTLISKFSLPVSVSASNKLACSDCFINKSH 510
Query: 336 RLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYE 395
+LPFS S+I + P + + SD+W SP+LS ++YY++ +D HT + W +P+ +KSQV
Sbjct: 511 KLPFSISSIKSTSPLEYIFSDVWMSPILSPDNYKYYLVLVDHHTRYTWLYPLQQKSQVKS 570
Query: 396 TFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAER 455
TF L++ +F A I+ L DNG E+ + + +NG+ S PHT NG +ER
Sbjct: 571 TFIAFKALVENRFQAKIRTLYSDNGGEFI--ALREFLVSNGISHLTSPPHTPEHNGLSER 628
Query: 456 KIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSH 515
K R I TLL +SVP +W +A A YL+N +P L +SP Q L+ P+Y
Sbjct: 629 KHRHIVETGLTLLTQASVPREYWPYAFAAAVYLINRMPTPVLSMESPFQKLFGSKPNYER 688
Query: 516 LRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDET 575
LRVFGCLCFP T NKL+ RS CVFLGY + Y C+D+ H+++ SRHV+FDE
Sbjct: 689 LRVFGCLCFPWLRPYTHNKLEERSRRCVFLGYSLTQTAYLCFDVEHKRLYTSRHVVFDEA 748
Query: 576 RFPFADL---SLTPAPSYECFTEDL-----------------PPSLIHHWQTV------- 608
FPF++L + P ++E + L P +++H Q
Sbjct: 749 SFPFSNLTSQNSLPTVTFEQSSSPLVTPILSSSSVLPSCLSSPCTVLHQQQPPVTTPNSP 808
Query: 609 -SSRPPDPPV--------------------------------QPSSPTDS-------SPP 628
SS+P P +P++P ++ SPP
Sbjct: 809 HSSQPTTSPAPLSPHRSTTMDFQVPQVRSSSPLLSSSSSLNSEPTAPNENGPEPEAQSPP 868
Query: 629 LSALVSPT--ASPSPLPLP-----------PVPPAPPVRTMTT----------------- 658
+ L +PT A PLP P P P PV+ TT
Sbjct: 869 IGPLSNPTHEAFIGPLPNPNRNPTNEIEPTPAPHPKPVKPTTTTTTPNRTTVSDASHQPT 928
Query: 659 -----------RSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFN 707
R+ + I KP FSL+ ++ + S S P N QAL D W+ AM EF+
Sbjct: 929 APQQNQHNMKTRAKNNIKKPNTKFSLTATLPNRSPSE-PTNVTQALKDKKWRFAMSDEFD 987
Query: 708 ALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETF 767
A R++TW+LVP + ++ C W+F+ K NG ++YKARLV G +Q GVD ETF
Sbjct: 988 AQQRNHTWDLVPHESQL-LVGCKWVFKLKYLPNGAIDKYKARLVAKGFNQQYGVDYAETF 1046
Query: 768 SPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCR 827
SPV+K TIR VL +A+ + W I QLDV NAFL G L E +YM P GF D P +VCR
Sbjct: 1047 SPVIKSTTIRLVLDVAVKKDWEIKQLDVNNAFLQGTLTEEVYMAQPPGFIDKDRPTHVCR 1106
Query: 828 LKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVA 887
L+K++YGLKQAPRAWY ++ +IGF +S SD SLFI+ G+ Y+L+YVDDII+
Sbjct: 1107 LRKAIYGLKQAPRAWYMELKQHLFNIGFVNSLSDASLFIYCHGTTFVYVLVYVDDIIVTG 1166
Query: 888 SSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASC 947
S + + LA F++KD L YFLGI TR GL L Q Y +++A+ MA
Sbjct: 1167 SDKSSIDAVLTSLAERFSIKDPTDLHYFLGIEATRTKQGLHLMQRKYIKDLLAKHNMADA 1226
Query: 948 NPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHT 1007
P TP+ T KL+ GT DAS YRS+ G+LQYL FTRPDI+YAV ++ M P
Sbjct: 1227 KPVLTPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLAFTRPDIAYAVNRLSQLMPQPTE 1286
Query: 1008 KHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNL 1067
H A KRVLRY+ GT T+G+ L + L +++DADW G D ST+ Y ++LG N
Sbjct: 1287 DHWQAAKRVLRYLAGTSTHGIFLDTTSPLNLHAFSDADWAGDSDDYVSTNAYVIYLGKNP 1346
Query: 1068 ISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYL 1127
ISWSSK+Q ++RSS E+EYR VAN SE W+ +LL +LH L + CDN+ A YL
Sbjct: 1347 ISWSSKKQRGVARSSTESEYRAVANAASEVKWLCSLLSKLHIRLPIRPSIFCDNIGATYL 1406
Query: 1128 SGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSL 1187
NPV H R KHI +D HFVR + G RV HV +R Q+AD TK L R F R +
Sbjct: 1407 CANPVFHSRMKHIAIDYHFVRNMIQSGALRVSHVSTRDQLADALTKPLSRAHFQSARFKI 1466
Query: 1188 SVGEPPAS 1195
V + P S
Sbjct: 1467 GVRQLPPS 1474
>At4g10690 retrotransposon like protein
Length = 1515
Score = 761 bits (1965), Expect = 0.0
Identities = 433/1133 (38%), Positives = 617/1133 (54%), Gaps = 94/1133 (8%)
Query: 147 PTDIAAAMHTMSLTPPDTP----WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQ 202
P D+ A M ++ + W D+ A++H + L + S + +IVG+G
Sbjct: 301 PEDLPNAFAAMRVSDQNQASSHEWLPDSAATAHITNTTDGLQNSQTYSG-DDSVIVGNGD 359
Query: 203 GIPIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDF 262
+PI G + L L VL P I K+L+SV +LT D S +FD + D
Sbjct: 360 FLPITHIGTIPLNISQGTLPLEDVLVCPGITKSLLSVSKLTDDYPCSFTFDSDSVVIKDK 419
Query: 263 KTGMPLLRCNSLGDLYPVTRSSPFAGLASS--------VWHNRLGHPASSALNHLRNNKL 314
+T L + N LY V + PF S+ VWH RLGHP L HL K
Sbjct: 420 RTQQLLTQGNKHKGLY-VLKDVPFQTYYSTRQQSSDDEVWHQRLGHPNKEVLQHLIKTKA 478
Query: 315 IFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVL 373
I + SS++C++C +GK RLPF +S ++ RP + +H DLW +PV S G +YYV+
Sbjct: 479 IVVNKT-SSNMCEACQMGKVCRLPFVASEFVSSRPLERIHCDLWGPAPVTSAQGFQYYVI 537
Query: 374 FLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCD 433
F+D+++ F W +P+ KS + F L++ Q+ I QCD G E+ + F +
Sbjct: 538 FIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKIAMFQCDGGGEFVSYKFVAHLA 597
Query: 434 ANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILP 493
+ G+ SCPHT QNG AER+ R + + +L+ HS VP W A + +L N+LP
Sbjct: 598 SCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSKVPHKLWVEAFFTSNFLSNLLP 657
Query: 494 RKTLQ-NDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHR 552
TL N SP ++L+ P Y+ LRVFG C+P NK P+S CVFLGY ++
Sbjct: 658 SSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYAKNKFDPKSLLCVFLGYNNKYK 717
Query: 553 GYKCYDLSHRKILISRHVIFDETRFPFADL-----SLTPAPSYECFTEDLPPS------- 600
GY+C K+ I RHV+FDE +FP++D+ +++ +P + + + +
Sbjct: 718 GYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISGSPLFTAWQKGFSSTALSRETP 777
Query: 601 ------LIHHWQTVSSR----------------------------PPDP------PVQPS 620
+I TVSS PP P P QP
Sbjct: 778 STNVEDIIFPSATVSSSVPTGCAPNIAETATAPDVDVAAAHDMVVPPSPITSTSLPTQPE 837
Query: 621 SPT--------DSSPPLSALVSPTASPSPL----PLPPVPPAPPVRT--------MTTRS 660
T DS +S+ ++P + L PP+ T M TR+
Sbjct: 838 ESTSDQNHYSTDSETAISSAMTPQSINVSLFEDSDFPPLQSVISSTTAAPETSHPMITRA 897
Query: 661 MHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPR 720
GI+KP ++L S P P + K+AL D W +AM E + ++TW+LVP
Sbjct: 898 KSGITKPNPKYAL---FSVKSNYPEPKSVKEALKDEGWTNAMGEEMGTMHETDTWDLVPP 954
Query: 721 PCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVL 780
++ C W+F+ K S+G +R KARLV G Q GVD ET+SPVV+ AT+R++L
Sbjct: 955 EMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEEGVDYVETYSPVVRSATVRSIL 1014
Query: 781 SIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPR 840
+A W + QLDV+NAFLH +L ET++M P GF DP PDYVC+LKK++Y LKQAPR
Sbjct: 1015 HVATINKWSLKQLDVKNAFLHDELKETVFMTQPPGFEDPSRPDYVCKLKKAIYDLKQAPR 1074
Query: 841 AWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALL 900
AW+ +F+ Y+ GF S SD SLF++ +G D+ ++LLYVDD+IL ++ L + + +L
Sbjct: 1075 AWFDKFSSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLYVDDMILTGNNDVLLQQLLNIL 1134
Query: 901 ASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKL 960
++EF MKD+G L YFLGI H GLFLSQ Y ++++ AGM+ C+ TP+ + L
Sbjct: 1135 STEFRMKDMGALHYFLGIQAHYHNDGLFLSQEKYTSDLLVNAGMSDCSSMPTPL--QLDL 1192
Query: 961 SSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYV 1020
+ P + + +R L G LQYLT TRPDI +AV VC MHAP LKR+L Y+
Sbjct: 1193 LQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFVCQKMHAPTMSDFHLLKRILHYL 1252
Query: 1021 RGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSR 1080
+GT+T G++L + L Y+D+DW GC DTRRST G+C FLG N+ISWS+KR PT+S+
Sbjct: 1253 KGTMTMGINLSSNTDSVLRCYSDSDWAGCKDTRRSTGGFCTFLGYNIISWSAKRHPTVSK 1312
Query: 1081 SSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHI 1140
SS EAEYR ++ SE WI LL E+ P Q ++CDN+SA+YLS NP H R+KH
Sbjct: 1313 SSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIPEMYCDNLSAVYLSANPALHSRSKHF 1372
Query: 1141 EMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPP 1193
++D ++VRE+V G V H+P+ Q+ADIFTK LP+ F D R L V PP
Sbjct: 1373 QVDYYYVRERVALGALTVKHIPASQQLADIFTKSLPQAPFCDLRFKLGVVLPP 1425
>At1g53810
Length = 1522
Score = 745 bits (1923), Expect = 0.0
Identities = 432/1136 (38%), Positives = 601/1136 (52%), Gaps = 99/1136 (8%)
Query: 149 DIAAAMHTMSLTPPDT----PWYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGI 204
D+ A+ TM +T W D+ AS+H ++ L S H + ++V G +
Sbjct: 305 DLPMALATMRITDVTDHHGHEWIPDSAASAHVTNNRHVLQQ-SQPYHGSDSIMVADGNFL 363
Query: 205 PIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKT 264
PI +G SI + + L VL P I+K+L+SV +LT+D SV FD ++D T
Sbjct: 364 PITHTGSGSIASSSGKIPLKEVLVCPDIVKSLLSVSKLTSDYPCSVEFDADSVRINDKAT 423
Query: 265 GMPLLRCNSLGDLYP-------VTRSSPFAGLASSVWHNRLGHPASSALNHLRNNKLIFC 317
L+ + LY V S+ +S VWH RLGH + L+ L ++K I
Sbjct: 424 KKLLVMGRNRDGLYSLEEPKLQVLYSTRQNSASSEVWHRRLGHANAEVLHQLASSKSIII 483
Query: 318 EPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLD 376
+VC++C LGK RLPF S RP + +H DLW SP S G RYYV+F+D
Sbjct: 484 INKVVKTVCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWGPSPTSSVQGFRYYVVFID 543
Query: 377 DHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANG 436
++ F W +P+ KS + TF L++ Q IK QCD G E+ + F ++ +G
Sbjct: 544 HYSRFTWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDHG 603
Query: 437 LIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKT 496
+ SCP+T QNG AERK R I + +++ S +P +W + A +++N+LP +
Sbjct: 604 IQQNMSCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTSS 663
Query: 497 LQN-DSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYK 555
L N +SP Q LY + P YS LRVFGC C+P K PRS CVFLGY ++GY+
Sbjct: 664 LDNNESPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYNEKYKGYR 723
Query: 556 CYDLSHRKILISRHVIF--------------------------------------DETRF 577
C +I ISRHV+F D++R+
Sbjct: 724 CLYPPTGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVTPTQPDQSRY 783
Query: 578 PFA--------DLSLTPAP-SYECFTEDLPPSLIHHWQTVSSRPPDPPVQPS-------- 620
P + DLS PA + E + +T+S P
Sbjct: 784 PVSSIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVSGSPERTTGLDSASIGD 843
Query: 621 ---SPT-DSSPPLSALVSPTASPSPLPLPPVP------PAPPVRTMTTRSMHGISKPKKP 670
SPT DSS P A SP +SP P+ P P M TR GISKP K
Sbjct: 844 SYHSPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNKR 903
Query: 671 FSL---SVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVI 727
+ L VSI P P +AL P W +AMQ E + TW LVP ++NV+
Sbjct: 904 YVLLTHKVSI------PEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVL 957
Query: 728 RCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRS 787
MW+FR K ++G ++ KARLV G Q G+D ET+SPVV+ T+R +L +A
Sbjct: 958 GSMWVFRTKLHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLK 1017
Query: 788 WPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFA 847
W + Q+DV+NAFLHGDL ET+YM P GF D PD+VC L KSLYGLKQ+PRAW+ RF+
Sbjct: 1018 WELKQMDVKNAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFS 1077
Query: 848 DYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMK 907
+++ GF S D SLF++ +D+ +LLYVDD+++ ++ +A L EF MK
Sbjct: 1078 NFLLEFGFICSLFDPSLFVYSSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMK 1137
Query: 908 DLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTP 967
D+G + YFLGI + + GGLF+SQ YA +++ A MA+C+P TP+ + S+
Sbjct: 1138 DMGQVHYFLGIQIQTYDGGLFMSQQKYAEDLLITASMANCSPMPTPLPLQLDRVSNQDEV 1197
Query: 968 CEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYG 1027
D + +RSL G LQYLT TRPDI +AV VC MH P LKR+LRY++GT++ G
Sbjct: 1198 FSDPTYFRSLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVSMG 1257
Query: 1028 LHLYPSPVETLVS----------YTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPT 1077
+ Y S ++VS Y+D+D+ C +TRRS GYC F+G N+ISWSSK+QPT
Sbjct: 1258 IQ-YNSNSSSVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQPT 1316
Query: 1078 LSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRT 1137
+SRSS EAEYR ++ SE W+ ++L E+ L + CDN+SA+YL+ NP H RT
Sbjct: 1317 VSRSSTEAEYRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHART 1376
Query: 1138 KHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPP 1193
KH ++D H++RE+V V H+P Q+ADIFTK LP F R L V PP
Sbjct: 1377 KHFDVDHHYIRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKLGVDFPP 1432
>At4g27210 putative protein
Length = 1318
Score = 690 bits (1780), Expect = 0.0
Identities = 403/1090 (36%), Positives = 556/1090 (50%), Gaps = 107/1090 (9%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W D+ A++H S +L S H ++V G +PI +G T++ + + L
Sbjct: 177 WLPDSAATAHVTNSPRSLQQ-SQPYHGTDAIMVDDGNYLPITHTGSTNLASSSGTVPLTD 235
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSP 285
VL P I K+L+S+ +LT D +V F+ G V+D T LL ++ LY +
Sbjct: 236 VLVCPSITKSLLSMSKLTQDFPCTVEFEYDGVRVNDKATKKLLLMGSNRDGLYCLKDDKQ 295
Query: 286 FAGLASS--------VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRL 337
F S+ VWH RLGHP H ++
Sbjct: 296 FQAFFSTRQRSASDEVWHRRLGHP--------------------------------HPQI 323
Query: 338 PFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYET 396
L+P + +H DLW + + S G RYY +F+D ++ F W +P+ KS Y
Sbjct: 324 ---------LQPLERVHCDLWGPTTITSVQGFRYYAVFIDHYSRFSWIYPLKLKSDFYNI 374
Query: 397 FTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERK 456
F L++ Q S I QCD G E+ + F ++ ++G+ + SCPHT QNG AERK
Sbjct: 375 FLAFHKLVENQLSQKISVFQCDGGGEFVSHKFLQHLQSHGIQQQLSCPHTPQQNGLAERK 434
Query: 457 IRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQND-SPTQLLYHRDPSYSH 515
R + + ++L S VP FW A A +L+N+LP L+ SP + LY + P Y+
Sbjct: 435 HRHLVELGLSMLFQSHVPHKFWVEAFFTANFLINLLPTSALKESISPYEKLYDKKPDYTS 494
Query: 516 LRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDET 575
LR FG CFP NK P S CVFLGY ++GY+C ++ ISRHVIFDE+
Sbjct: 495 LRSFGSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRCLYPPTGRLYISRHVIFDES 554
Query: 576 RFPFADL-------------------SLTPAPSYEC--------FTE-DLPP-------- 599
+PF+ S +PAPS FT D PP
Sbjct: 555 VYPFSHTYKHLHPQPRTPLLAAWLRSSDSPAPSTSTSPSSRSPLFTSADFPPLPQRKTPL 614
Query: 600 --------SLIHHWQTVSSRPPDPPVQPSSPTDS--------SPPLSALVSPTASPSPLP 643
S+ H + + PD + ++ DS S + T + +
Sbjct: 615 LPTLVPISSVSHASNITTQQSPDFDSERTTDFDSASIGDSSHSSQAGSDSEETIQQASVN 674
Query: 644 LPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQ 703
+ + V M TR+ GISKP + V + P P AL P W AM
Sbjct: 675 VHQTHASTNVHPMVTRAKVGISKPNPRY---VFLSHKVSYPEPKTVTAALKHPGWTGAMT 731
Query: 704 SEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDC 763
E + TW LVP D++V+ W+FR K ++G + KAR+V G Q G+D
Sbjct: 732 EEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHADGTLNKLKARIVAKGFLQEEGIDY 791
Query: 764 DETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPD 823
ET+SPVV+ T+R VL +A + +W I Q+DV+NAFLHGDL ET+YM P GF DP PD
Sbjct: 792 LETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFLHGDLKETVYMTQPAGFVDPSKPD 851
Query: 824 YVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDI 883
+VC L KS+YGLKQ+PRAW+ +F+ ++ GF S SD SLFI+ +++ +LLYVDD+
Sbjct: 852 HVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCSKSDPSLFIYAHNNNLILLLLYVDDM 911
Query: 884 ILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAG 943
++ +S S +A L EF M D+G L YFLGI V R GLF+SQ YA +++ A
Sbjct: 912 VITGNSSQTLTSLLAALNKEFRMTDMGQLHYFLGIQVQRQQNGLFMSQQKYAEDLLIAAS 971
Query: 944 MASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMH 1003
M C P TP+ + D + +RS+ G LQYLT TRPDI +AV VC MH
Sbjct: 972 MEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAGKLQYLTLTRPDIQFAVNFVCQKMH 1031
Query: 1004 APHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFL 1063
P LKR+LRY++GT+T G+ L +Y+D+DWG C TRRS G C F+
Sbjct: 1032 QPTISDFHLLKRILRYIKGTITMGISYSRDSPTLLQAYSDSDWGNCKQTRRSVGGLCTFM 1091
Query: 1064 GDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVS 1123
G NL+SWSSK+ PT+SRSS EAEY+ +++ SE W+ LL EL PL + CDN+S
Sbjct: 1092 GTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASEILWLSTLLRELRIPLPDTPELFCDNLS 1151
Query: 1124 AIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDF 1183
A+YL+ NP H RTKH ++D HFVRE+V V H+P QIADIFTK LP F
Sbjct: 1152 AVYLTANPAFHARTKHFDIDFHFVRERVALKALVVKHIPGSEQIADIFTKSLPYEAFIHL 1211
Query: 1184 RSSLSVGEPP 1193
R L V P
Sbjct: 1212 RGKLGVTLSP 1221
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 684 bits (1766), Expect = 0.0
Identities = 387/1039 (37%), Positives = 572/1039 (54%), Gaps = 42/1039 (4%)
Query: 166 WYMDTGASSHTAASQGTLTSY--SNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLAL 223
W +D+GA+ H + + + S +S +N + +G + I G G I +K + L
Sbjct: 442 WVIDSGATHHVSHDRKLFQTLDTSIVSFVN----LPTGPNVRISGVGTVLI---NKDIIL 494
Query: 224 NHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRS 283
+VL P NLIS+ LTTD V FDP + D G+ L +G+LY +
Sbjct: 495 QNVLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGLTLGEGKRIGNLYVLDTQ 554
Query: 284 SPFAGLAS----SVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPF 339
SP + + SVWH RLGHP+ S L+ L ++ S+ C C L K +L F
Sbjct: 555 SPAISVNAVVDVSVWHKRLGHPSFSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSF 614
Query: 340 SSSAIITLRPFDILHSDLWTSPVLSTA-GHRYYVLFLDDHTDFLWTFPISKKSQVYETFT 398
S+ I F++LH D+W + T G++Y++ +DDH+ W + + KS V F
Sbjct: 615 PSANNICNSTFELLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFP 674
Query: 399 TLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERKIR 458
L++ Q+ +K ++ DN +E +F + A G++ SCP T QN ERK +
Sbjct: 675 AFIDLVENQYDTRVKSVRSDNAKEL---AFTEFYKAKGIVSFHSCPETPEQNSVVERKHQ 731
Query: 459 TINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHLRV 518
I N+ R L+ S++ +W + A +L+N P L N +P ++L + P YS L+
Sbjct: 732 HILNVARALMFQSNMSLPYWGDCVLTAVFLINRTPSALLSNKTPFEVLTGKLPDYSQLKT 791
Query: 519 FGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETRFP 578
FGCLC+ S +K PRS CVFLGYP +GYK DL + ISR+V F E FP
Sbjct: 792 FGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKGYKLLDLESNVVHISRNVEFHEELFP 851
Query: 579 FADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPPLSALVSPTAS 638
A + + + FT P + +++S P P + PS+ T
Sbjct: 852 LASSQQSATTASDVFT---PMDPLSSGNSITSHLPSPQISPSTQISKR-------RITKF 901
Query: 639 PSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVS----IDDPSISPLPHNPKQALS 694
P+ L + H IS +S S I++ S P+P + +A
Sbjct: 902 PAHLQ------DYHCYFVNKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKD 955
Query: 695 DPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDG 754
W A+ E A+ R++TWE+ P + C W+F K ++G ER+KAR+V G
Sbjct: 956 SKEWCGAIDQEIGAMERTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKG 1015
Query: 755 RSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPL 814
+Q G+D ETFSPV K AT++ +L ++ S+ W ++QLD+ NAFL+GDL ETIYM P
Sbjct: 1016 YTQKEGLDYTETFSPVAKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPD 1075
Query: 815 GFRDPHH----PDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQG 870
G+ D P+ VCRLKKS+YGLKQA R W+ +F++ + ++GF DH+LF+ G
Sbjct: 1076 GYADIKGTSLPPNVVCRLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVRCIG 1135
Query: 871 SDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLS 930
S+ +L+YVDDI++ +++ +S L + F +++LGPL YFLG+ V R + G+ LS
Sbjct: 1136 SEFIVLLVYVDDIVIASTTEQAAQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLS 1195
Query: 931 QSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPD 990
Q YA E++ A M C PS+ P+ +LS + G ED +YR L G L YLT TRPD
Sbjct: 1196 QRKYALELLTSADMLDCKPSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPD 1255
Query: 991 ISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCP 1050
I++AV ++C AP T H+ A+ +VL+Y++GT+ GL TL YTDADWG CP
Sbjct: 1256 ITFAVNKLCQFSSAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTCP 1315
Query: 1051 DTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFP 1110
D+RRST+G+ +F+G +LISW SK+QPT+SRSSAEAEYR +A E W+ LLL L
Sbjct: 1316 DSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVH 1375
Query: 1111 LSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADI 1170
S +++ D+ +A+Y++ NPV H+RTKHIE+D H VREK+ GQ ++LHV ++ Q+ADI
Sbjct: 1376 -SGVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADI 1434
Query: 1171 FTKGLPRLLFDDFRSSLSV 1189
TK L F S +S+
Sbjct: 1435 LTKPLFPYQFAHLLSKMSI 1453
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 627 bits (1616), Expect = e-179
Identities = 371/1055 (35%), Positives = 557/1055 (52%), Gaps = 42/1055 (3%)
Query: 150 IAAAMHTMSLTPPDTPWYMDTGASSHTAASQGTLTSY--SNLSHLNQKLIVGSGQGIPIQ 207
+ A HT+S W +D+GA+ H + + +S S LS +N + +G + I
Sbjct: 419 LTVARHTLS----SATWVIDSGATHHVSHDRSLFSSLDTSVLSAVN----LPTGPTVKIS 470
Query: 208 GSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMP 267
G G + + + L +VL P NLIS+ LT D V FD + D G
Sbjct: 471 GVGTLKL---NDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRM 527
Query: 268 LLRCNSLGDLYPVTRS----SPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSS 323
L + + +LY + S A + S+WH RLGH + L+ + ++ ++ S
Sbjct: 528 LGQGRRVANLYLLDVGDQSISVNAVVDISMWHRRLGHASLQRLDAISDSLGTTRHKNKGS 587
Query: 324 SVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTA-GHRYYVLFLDDHTDFL 382
C C L K +L F +S + FD+LH D+W + T G++Y++ +DDH+
Sbjct: 588 DFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSRAT 647
Query: 383 WTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFS 442
W + + KS+V F ++ Q+ +K ++ DN E SF+ G++ S
Sbjct: 648 WMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELKFTSFYA---EKGIVSFHS 704
Query: 443 CPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSP 502
CP T QN ERK + I N+ R L+ S VP S W + A +L+N P + L N +P
Sbjct: 705 CPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTP 764
Query: 503 TQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHR 562
++L P Y LR FGCLC+ +K QPRS C+FLGYP ++GYK DL
Sbjct: 765 YEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLESN 824
Query: 563 KILISRHVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSP 622
+ ISR+V F E FP A + + S + FT +P S +S P PS
Sbjct: 825 TVFISRNVQFHEEVFPLAKNPGSES-SLKLFTPMVPVSS----GIISDTTHSPSSLPSQI 879
Query: 623 TDSSPPLSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVS----ID 678
+D P +S S P L TM + + IS +S S I+
Sbjct: 880 SDLPPQIS---SQRVRKPPAHLNDYH----CNTMQSDHKYPISSTISYSKISPSHMCYIN 932
Query: 679 DPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQ 738
+ + P+P N +A W A+ +E A+ ++NTWE+ P + C W+F K
Sbjct: 933 NITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCKWVFTLKFL 992
Query: 739 SNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNA 798
++G ERYKARLV G +Q G+D +TFSPV K TI+ +L ++ S+ W + QLDV NA
Sbjct: 993 ADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWFLKQLDVSNA 1052
Query: 799 FLHGDLHETIYMHHPLGFRDPHH----PDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIG 854
FL+G+L E I+M P G+ + + V RLK+S+YGLKQA R W+++F+ + S+G
Sbjct: 1053 FLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKKFSSSLLSLG 1112
Query: 855 FRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSY 914
F+ + DH+LF+ + +L+YVDDI++ ++S L F ++DLG L Y
Sbjct: 1113 FKKTHGDHTLFLKMYDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQRFKLRDLGDLKY 1172
Query: 915 FLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLY 974
FLG+ V R G+ + Q YA E++ GM +C P + P+ K+ G ED Y
Sbjct: 1173 FLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNLKMRKDDGDLIEDIEQY 1232
Query: 975 RSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSP 1034
R + G L YLT TRPDI++AV ++C AP T H+ A RVL+Y++GT+ GL S
Sbjct: 1233 RRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVGQGLFYSASS 1292
Query: 1035 VETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVV 1094
TL + D+DW C D+RRST+ + +F+GD+LISW SK+Q T+SRSSAEAEYR +A
Sbjct: 1293 DLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALAT 1352
Query: 1095 SESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRG 1154
E W+ LL+ L +++ D+ +AIY++ NPV H+RTKHI++D H VRE++ G
Sbjct: 1353 CEMVWLFTLLVSLQ-ASPPVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNG 1411
Query: 1155 QARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
+ ++LHV + Q+ADI TK L F+ +S +S+
Sbjct: 1412 ELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSI 1446
>At1g70010 hypothetical protein
Length = 1315
Score = 615 bits (1586), Expect = e-176
Identities = 368/1021 (36%), Positives = 533/1021 (52%), Gaps = 88/1021 (8%)
Query: 205 PIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKT 264
PI GS + + L LN VL P+ NL+SV LT + FD + D
Sbjct: 312 PISGSVHLG-----RHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATR 366
Query: 265 GMPLLRCNSLGDLY---------PVTRSSPFAGLASS--VWHNRLGHPASSALNHLRNNK 313
+ + + +LY P T SS +S +WH RLGHP+ L + +
Sbjct: 367 ELMVGMGKQVANLYIVDLDSLSHPGTDSSITVASVTSHDLWHKRLGHPSVQKLQPMSSLL 426
Query: 314 LIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTA-GHRYYV 372
+ + + C C + K LPF S + RPFD++H D W + T G+RY++
Sbjct: 427 SFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYFL 486
Query: 373 LFLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYC 432
+DD++ W + + KS V T T+++ QF IK ++ DN E + F ++
Sbjct: 487 TIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELN---FTQFY 543
Query: 433 DANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNIL 492
+ G++ SCP T QN ERK + I N+ R+L S +P S+W + A YL+N L
Sbjct: 544 HSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRL 603
Query: 493 PRKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHR 552
P L++ P ++L P+Y H++VFGCLC+ +K PR+ C F+GYP +
Sbjct: 604 PAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGFK 663
Query: 553 GYKCYDLSHRKILISRHVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRP 612
GYK DL I++SRHV+F E FPF L+ + F DL P+
Sbjct: 664 GYKLLDLETHSIIVSRHVVFHEELFPFLGSDLSQEE--QNFFPDLNPT------------ 709
Query: 613 PDPPVQPSS-----PTDSSPPLSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKP 667
PP+Q S P+DSS + L P+A+P+ P P V+T K
Sbjct: 710 --PPMQRQSSDHVNPSDSSSSVEIL--PSANPTNNV-----PEPSVQTSHR-------KA 753
Query: 668 KKPFSLSVSIDDPSISPLPHNPKQALS-----DPN------------------------W 698
KKP L +S PH ++ LS DP W
Sbjct: 754 KKPAYLQDYYCHSVVSSTPHEIRKFLSYDRINDPYLTFLACLDKTKEPSNYTEAEKLQVW 813
Query: 699 KSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQI 758
+ AM +EF+ L ++TWE+ P D I C WIF+ K S+G ERYKARLV G +Q
Sbjct: 814 RDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQK 873
Query: 759 AGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFR- 817
G+D +ETFSPV K +++ +L +A + QLD+ NAFL+GDL E IYM P G+
Sbjct: 874 EGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYAS 933
Query: 818 ---DPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIA 874
D P+ VCRLKKSLYGLKQA R WY +F+ + +GF S DH+ F+
Sbjct: 934 RQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFL 993
Query: 875 YILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTY 934
+L+Y+DDII+ +++ + + S F ++DLG L YFLG+ + R G+ +SQ Y
Sbjct: 994 CVLVYIDDIIIASNNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQRKY 1053
Query: 935 ATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYA 994
A +++ G C PS+ P+D + SG + YR L G L YL TRPDI++A
Sbjct: 1054 ALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFA 1113
Query: 995 VQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRR 1054
V ++ AP H+ A+ ++L+Y++GT+ GL + L Y +AD+ C D+RR
Sbjct: 1114 VNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRDSRR 1173
Query: 1055 STSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQA 1114
STSGYC+FLGD+LI W S++Q +S+SSAEAEYR ++ E W+ N L EL PLS+
Sbjct: 1174 STSGYCMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKP 1233
Query: 1115 TLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKG 1174
TL+ CDN +AI+++ N V H+RTKHIE D H VRE++++G + H+ + QIAD FTK
Sbjct: 1234 TLLFCDNEAAIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKP 1293
Query: 1175 L 1175
L
Sbjct: 1294 L 1294
>At4g17450 retrotransposon like protein
Length = 1433
Score = 602 bits (1551), Expect = e-172
Identities = 354/1031 (34%), Positives = 526/1031 (50%), Gaps = 51/1031 (4%)
Query: 156 TMSLTPPDTP--WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTS 213
++S P +P W +D+GA+ H ++ ++ +L + +L + + I G G+
Sbjct: 420 SVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENTFVRL--PNDCTVKIAGIGFIQ 477
Query: 214 IPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNS 273
+ ++L++VL+ P NLIS +LT + + V DF +
Sbjct: 478 LSDA---ISLHNVLYIPEFKFNLIS--ELTKELMIGRGSQVGNLYVLDFNENNHTVSLKG 532
Query: 274 LGDLYPVTRSSPFAGLASSVWHNRLGHPASSALNHLRNN-----KLIFCEPSRSSSVCDS 328
+ P + S WH RLGHPA S ++ L + K I E S VC
Sbjct: 533 TTSMCPEFSVCSSVVVDSVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHV 592
Query: 329 CVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPIS 388
C L K L F S + FD++H D W + T D W + +
Sbjct: 593 CHLSKQKHLSFQSRQNMCSAAFDLVHIDTWGPFSVPT-------------NDATWIYLLK 639
Query: 389 KKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSS 448
KS V F ++ TQ+ +K ++ DN E F A+G++ SCP T
Sbjct: 640 NKSDVLHVFPAFINMVHTQYQTKLKSVRSDNAHEL---KFTDLFAAHGIVAYHSCPETPE 696
Query: 449 QNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYH 508
QN ERK + I N+ R LL S++P FW + A +L+N LP L N SP + L +
Sbjct: 697 QNSVVERKHQHILNVARALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKN 756
Query: 509 RDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISR 568
P+Y L+ FGCLC+ +K +PR+ CVFLGYP+ ++GYK D+ + ISR
Sbjct: 757 IPPAYESLKTFGCLCYSSTSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISR 816
Query: 569 HVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPP 628
HVIF E FPF ++ +D P L +R D P++ +S D+ P
Sbjct: 817 HVIFHEDIFPFISSTIKDD------IKDFFPLL-----QFPARTDDLPLEQTSIIDTHPH 865
Query: 629 LSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPLPHN 688
S P PL PP + ++P F I++ + + +P
Sbjct: 866 QDVSSSKALVPFD-PLSKRQKKPPKHLQDFHCYNNTTEPFHAF-----INNITNAVIPQR 919
Query: 689 PKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKA 748
+A W AM+ E A++R+NTW +V P + I C W+F K ++G ERYKA
Sbjct: 920 YSEAKDFKAWCDAMKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKA 979
Query: 749 RLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETI 808
RLV G +Q G+D +ETFSPV K ++R +L +A W +HQLD+ NAFL+GDL E I
Sbjct: 980 RLVAKGYTQEEGLDYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEI 1039
Query: 809 YMHHPLGFRD----PHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSL 864
YM P G+ D P +CRL KS+YGLKQA R WY + ++ + +GF+ S +DH+L
Sbjct: 1040 YMKIPPGYADLVGEALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTL 1099
Query: 865 FIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHA 924
FI + +L+YVDDI++V++S D F A L S F ++DLG YFLGI + R
Sbjct: 1100 FIKYANGVLMGVLVYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSE 1159
Query: 925 GGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYL 984
G+ + Q Y E+++ G PS+ P+D KL+ G P D++ YR L G L YL
Sbjct: 1160 KGISICQRKYILELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYL 1219
Query: 985 TFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDA 1044
TRPDI+YAV +C HAP + H+ A+ +VLRY++GT+ GL L YTD+
Sbjct: 1220 QITRPDIAYAVNTLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDS 1279
Query: 1045 DWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLL 1104
D+G C D+RR + YC+F+GD L+SW SK+Q T+S S+AEAE+R ++ E W+ L
Sbjct: 1280 DFGSCTDSRRCVAAYCMFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLF 1339
Query: 1105 LELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSR 1164
+ P ++CDN +A+++ N V H+RTK +E+D + RE V G + + V +
Sbjct: 1340 DDFKVPFIPPAYLYCDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETG 1399
Query: 1165 HQIADIFTKGL 1175
Q+AD TK +
Sbjct: 1400 EQVADPLTKAI 1410
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 575 bits (1483), Expect = e-164
Identities = 355/1020 (34%), Positives = 524/1020 (50%), Gaps = 57/1020 (5%)
Query: 221 LALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPV 280
L + V + +LIS+ QL +N + V D + M + +G +
Sbjct: 313 LVMISVFYVEEFQSDLISIGQLMDENRCVLQMSDRFLVVQDRTSRMVMGAGRRVGGTFHF 372
Query: 281 TRSSPFAGLAS-------SVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGK 333
RS+ A + +WH+R+GHPA+ ++ + + + + + CD C K
Sbjct: 373 -RSTEIAASVTVKEEKNYELWHSRMGHPAARVVSLIPESS-VSVSSTHLNKACDVCHRAK 430
Query: 334 HVRLPFSSSAIITLRPFDILHSDLWTS-PVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQ 392
R F S TLR F++++ DLW S G RY++ +DD++ +W + ++ KS+
Sbjct: 431 QTRNSFPLSINKTLRIFELIYCDLWGPYRTPSHTGARYFLTIIDDYSRGVWLYLLNDKSE 490
Query: 393 VYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGK 452
+ QF+ IK ++ DNG E+ ++ G+I SC T +N +
Sbjct: 491 APCHLKNFFAMTDRQFNVKIKTVRSDNGTEFL--CLTKFFQEQGVIHERSCVATPERNDR 548
Query: 453 AERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPS 512
ERK R + N+ R L +++P FW + A YL+N P L + +P + L+ + P
Sbjct: 549 VERKHRHLLNVARALRFQANLPIQFWGECVLTAAYLINRTPSSVLNDSTPYERLHKKQPR 608
Query: 513 YSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIF 572
+ HLRVFG LC+ + +K RS CVF+GYP +G++ +DL + +SR V+F
Sbjct: 609 FDHLRVFGSLCYAHNRNRGGDKFAERSRRCVFVGYPHGQKGWRLFDLEQNEFFVSRDVVF 668
Query: 573 DETRFPFAD------LSLTPAPSYECFTEDLPPSLIHHWQTVSSRPP---DPPVQPSSPT 623
E FPF + + + L +H Q PP P+ PS+ +
Sbjct: 669 SELEFPFRISHEQNVIEEEEEALWAPIVDGLIEEEVHLGQNAGPTPPICVSSPISPSATS 728
Query: 624 D-----SSPPLSALVSPT-----------ASPSPLPLPPV------------PPA-PPVR 654
+S PL V PT +SP+ L P+ PPA PP R
Sbjct: 729 SRSEHSTSSPLDTEVVPTPATSTTSASSPSSPTNLQFLPLSRAKPTTAQAVAPPAVPPPR 788
Query: 655 TMTTRSMHGISKPKKPFSLSVSIDDPSISPLPH--NPKQALSDPNWKSAMQSEFNALI-- 710
+TR+ K F ++ ++ S S L Q D SA + + A+
Sbjct: 789 RQSTRNK-APPVTLKDFVVNTTVCQESPSKLNSILYQLQKRDDTRRFSASHTTYVAIDAQ 847
Query: 711 -RSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSP 769
++TW + P I W+++ K S+G ERYKARLV G Q G D ETF+P
Sbjct: 848 EENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARLVALGNKQKEGEDYGETFAP 907
Query: 770 VVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLK 829
V K AT+R L +A+ R+W IHQ+DV NAFLHGDL E +YM P GF + HP+ VCRL+
Sbjct: 908 VAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYMKLPPGF-EASHPNKVCRLR 966
Query: 830 KSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASS 889
K+LYGLKQAPR W+++ + GF+ S +D+SLF +GS IL+YVDD+I+ +S
Sbjct: 967 KALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGSVRIKILIYVDDLIITGNS 1026
Query: 890 HDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNP 949
+ F LAS F MKDLGPL YFLGI V R G+++ Q YA +II+ G+ P
Sbjct: 1027 QRATQQFKEYLASCFHMKDLGPLKYFLGIEVARSTTGIYICQRKYALDIISETGLLGVKP 1086
Query: 950 SATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKH 1009
+ P++ KL S+ D YR L G L YL TR D++++V + M P H
Sbjct: 1087 ANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTRLDLAFSVHILARFMQEPREDH 1146
Query: 1010 MLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLIS 1069
A RV+RY++ G+ L S + + D+DW G P +RRS +GY V GD+ IS
Sbjct: 1147 WAAALRVVRYLKADPGQGVFLRRSGDFQITGWCDSDWAGDPMSRRSVTGYFVQFGDSPIS 1206
Query: 1070 WSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSG 1129
W +K+Q T+S+SSAEAEYR ++ + SE W++ LL L Q ++ CD+ SAIY++
Sbjct: 1207 WKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLGVSHVQPMIMCCDSKSAIYIAT 1266
Query: 1130 NPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
NPV H+RTKHIE+D HFVR++ V+G HV + Q+ADIFTK L R F FR L +
Sbjct: 1267 NPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLADIFTKPLGRDCFSAFRIKLGI 1326
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 572 bits (1475), Expect = e-163
Identities = 385/1157 (33%), Positives = 545/1157 (46%), Gaps = 198/1157 (17%)
Query: 104 PSPWGMPPCPYPTSQWSRPMGVPQQPGVLGPRPQAYTATASPAPTDIAAAMHTMSLTP-- 161
P+ P P T M + G + P P + + D+ HT+S
Sbjct: 420 PAASSSNPSPLATVSEHGFMALTSTSGTIIPFP---STSLKYENNDLKFQNHTLSALQKF 476
Query: 162 -PDTPWYMDTGASSHTAASQGT---LTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTP 217
P W +D+GASSH + L S S H+ QKLI
Sbjct: 477 LPSDAWIIDSGASSHVCSDLAMFRELKSVSGTVHITQKLI-------------------- 516
Query: 218 HKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDL 277
L++VLH P NL+SV L + S F + + G+ + R +L
Sbjct: 517 -----LHNVLHVPDFKFNLMSVSSLVKTISCSAHFYVDCCLIQELSQGLMIGRGRLYHNL 571
Query: 278 YPV-------TRSSPFAGLASS-------VWHNRLGHPASSALNHLRNNKLIFCEPSRSS 323
Y + + S+P A L + +WH RLGHP+S L L+
Sbjct: 572 YILETENTSPSTSTPAACLFTGSVLNDGHLWHQRLGHPSSVVLQKLK------------- 618
Query: 324 SVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTS-PVLSTAGHRYYVLFLDDHTDFL 382
RL + S + PFD++H D+W + S G RY++ +DD T
Sbjct: 619 ------------RLAYISHNNLASNPFDLVHLDIWGPFSIESIEGFRYFLTVVDDCTRTT 666
Query: 383 WTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFS 442
W + + K V F L+ TQF+A IK ++ DN E F +G++ FS
Sbjct: 667 WVYMLRNKKDVSSVFPEFIKLVSTQFNAKIKAIRSDNAPEL---GFTEIVKEHGMLHHFS 723
Query: 443 CPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSP 502
C +T QN ERK + I N+ R LL S++P +W + A +L+N LP L N SP
Sbjct: 724 CAYTPQQNSVVERKHQHILNVARALLFQSNIPMQYWSDCVTTAVFLINRLPSPLLNNKSP 783
Query: 503 TQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHR 562
+L+ ++ P YS L+ FGCLCF + K PR+ CVFLGYP ++GYK DL
Sbjct: 784 YELILNKQPDYSLLKNFGCLCFVSTNAHERTKFTPRARACVFLGYPSGYKGYKVLDLESH 843
Query: 563 KILISRHVIFDETRFPF---------ADL---SLTPAPSYECFTEDLP----PSLI---- 602
+ +SR+V+F E FPF D+ S+ P P+ F E +P SLI
Sbjct: 844 SVTVSRNVVFKEHVFPFKTSELLNKAVDMFPNSILPLPAPLHFVETMPLIDEDSLIPTTT 903
Query: 603 ------HHWQTVSSRPPDPPVQPSSPTDSSP------PLSALVSPTASPSPLP------- 643
+H + SS P + PSS T++ P++ T +PS L
Sbjct: 904 DSRTADNHASSSSSALPSI-IPPSSNTETQDIDSNAVPITRSKRTTRAPSYLSEYHCSLV 962
Query: 644 -----LPPVPPAPPVRTMTTRSMHGISKPKK--PFSLSVSIDDPSISPL----------- 685
LPP + P+ + + S PKK P+ +S + +PL
Sbjct: 963 PSISTLPPTDSSIPIHPLP--EIFTASSPKKTTPYPISTVVSYDKYTPLCQSYIFAYNTE 1020
Query: 686 --PHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLF 743
P QA+ W E A+ + TW + P D NV+ C W+F K +G
Sbjct: 1021 TEPKTFSQAMKSEKWIRVAVEELQAMELNKTWSVESLPPDKNVVGCKWVFTIKYNPDGTV 1080
Query: 744 ERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGD 803
ERYKARLV G +Q G+D +TFSPV K + + +L +A W + Q+DV +AFLHGD
Sbjct: 1081 ERYKARLVAQGFTQQEGIDFLDTFSPVAKLTSAKMMLGLAAITGWTLTQMDVSDAFLHGD 1140
Query: 804 LHETIYMHHPLGFRDPHH----PDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHST 859
L E I+M P G+ P P+ VCRL KS+YGLKQA R WY+RF
Sbjct: 1141 LDEEIFMSLPQGYTPPAGTILPPNPVCRLLKSIYGLKQASRQWYKRFVA----------- 1189
Query: 860 SDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIA 919
L+Y+DDI++ +++ ++ ALL SEF +KDLGP +FLG+
Sbjct: 1190 ----------------ALVYIDDIMIASNNDAEVENLKALLRSEFKIKDLGPARFFLGLL 1233
Query: 920 VTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDG 979
C PS+ P+D L GTP + + YR L G
Sbjct: 1234 --------------------------GCKPSSIPMDPTLHLVRDMGTPLPNPTAYRKLIG 1267
Query: 980 ALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLV 1039
L YLT TRPDI+YAV Q+ + AP H+ A +VLRY++ GL +Y + E +
Sbjct: 1268 RLLYLTITRPDITYAVHQLSQFISAPSDIHLQAAHKVLRYIKANPGQGL-MYSADYEICL 1326
Query: 1040 S-YTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESC 1098
+ ++DADW C DTRRS SG+C++LG +LISW SK+Q SRSS E+EYR +A E
Sbjct: 1327 NGFSDADWAACKDTRRSISGFCIYLGTSLISWKSKKQAVASRSSTESEYRSMAQATCEII 1386
Query: 1099 WIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARV 1158
W++ LL +LH PL+ + CDN SA++ S NPV H+RTKHIE+D H VR+++ G +
Sbjct: 1387 WLQQLLKDLHIPLTCPAKLFCDNKSALHSSLNPVFHERTKHIEIDCHTVRDQIKAGNLKA 1446
Query: 1159 LHVPSRHQIADIFTKGL 1175
LHVP+ +Q ADI TK L
Sbjct: 1447 LHVPTENQHADILTKAL 1463
>At1g26990 polyprotein, putative
Length = 1436
Score = 550 bits (1417), Expect = e-156
Identities = 341/1058 (32%), Positives = 526/1058 (49%), Gaps = 84/1058 (7%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GAS H + +Y L +L +G + I+G+G+ + L+L++
Sbjct: 421 WVIDSGASHHVTHERNLYHTYKALDRTFVRL--PNGHTVKIEGTGFIQLTDA---LSLHN 475
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVT---- 281
VL P NL+SV LT VSF + + L + + +G+LY +
Sbjct: 476 VLFIPEFKFNLLSVSVLTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKS 535
Query: 282 ----RSSPFAGLASSV------WHNRLGHPASSALNHLRNNKLIFCEP-SRSSSVCDSCV 330
S P + SSV WH RLGHP+ + ++ L + ++ + ++ SS C C
Sbjct: 536 LVDVSSFPGKSVCSSVKNESEMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCH 595
Query: 331 LGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTA-GHRYYVLFLDDHTDFLWTFPISK 389
L K LPF S I + F+++H D W + T +RY++ +DD + W + + +
Sbjct: 596 LSKQKHLPFKSVNHIREKAFELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQ 655
Query: 390 KSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQ 449
KS V F + +++TQ+ + ++ DN E F+ G+ CP T Q
Sbjct: 656 KSDVLTVFPSFLKMVETQYHTKVCSVRSDNAHEL---KFNELFAKEGIKADHPCPETPEQ 712
Query: 450 NGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHR 509
N ERK + + N+ R L+ S +P +W + A +L+N L + N++P + L
Sbjct: 713 NFVVERKHQHLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKG 772
Query: 510 DPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRH 569
P YS L+ FGCLC+ + K PR+ C+FLGYPM ++GYK D+ + ISRH
Sbjct: 773 KPDYSSLKAFGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRH 832
Query: 570 VIFDETRFPFADLSLT------------PAPSYECFTEDLP----PSLIHHWQTVSSRPP 613
VIF E FPFA ++T PAP+ + E LP S H SS
Sbjct: 833 VIFYEDIFPFASSNITDAAKDFFPHIYLPAPNND---EHLPLVQSSSDAPHNHDESSSMI 889
Query: 614 DPPVQPSSPTDSSPP--LSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPF 671
P +P S P L +P+ P P + S +PF
Sbjct: 890 FVPSEPKSTRQRKLPSHLQDFHCYNNTPTTTKTSPYPLTNYI---------SYSYLSEPF 940
Query: 672 SLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMW 731
++I + + LP +A D W AM E +A +R+ TW + P + C W
Sbjct: 941 GAFINII--TATKLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKW 998
Query: 732 IFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIH 791
I K ++G ER+KARLV G +Q G+D TFSPV K T++ +LS+A W +H
Sbjct: 999 IITIKFLADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLAPKMKWYLH 1058
Query: 792 QLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVS 851
QLD+ NA L+GDL E IYM P G+ + + G + +P A
Sbjct: 1059 QLDISNALLNGDLEEEIYMKLPPGYSE-------------IQGQEVSPNA---------- 1095
Query: 852 SIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGP 911
DH+LF+ Q +L+YVDDI++ +++ + L+S F ++DLG
Sbjct: 1096 -----KCHGDHTLFVKAQDGFFLVVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGE 1150
Query: 912 LSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDA 971
+FLGI + R+A G+ L Q Y +++A + + C PS+ P++ QKLS +GT ED
Sbjct: 1151 PKFFLGIEIARNADGISLCQRKYVLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDG 1210
Query: 972 SLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLY 1031
YR + G LQYL TRPDI++AV ++ + AP H+ AL ++LRY++GT+ GL
Sbjct: 1211 KQYRRILGKLQYLCLTRPDINFAVSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFYG 1270
Query: 1032 PSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVA 1091
L ++D+DW CPDTRR +G+ +F+G++L+SW SK+Q +S SSAEAEYR ++
Sbjct: 1271 ADTNFDLRGFSDSDWQTCPDTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMS 1330
Query: 1092 NVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKV 1151
E W+ +L P + ++CDN +A++++ N V H+RTKHIE D H VRE +
Sbjct: 1331 VATKELIWLGYILTAFKIPFTHPAYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECI 1390
Query: 1152 VRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
G + + V + +Q+AD TK L F + S L +
Sbjct: 1391 EAGILKTIFVRTDNQLADTLTKPLYPKPFRENNSKLGL 1428
>At3g60170 putative protein
Length = 1339
Score = 535 bits (1379), Expect = e-152
Identities = 325/1052 (30%), Positives = 532/1052 (49%), Gaps = 41/1052 (3%)
Query: 163 DTPWYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLA 222
D W++D+G S+H S+ + N+ + +G+ + + G G +
Sbjct: 297 DEVWFLDSGCSNHMTGSKEWFSELEE--GFNRTVKLGNDTRMSVVGKGSVKVKVNGVTQV 354
Query: 223 LNHVLHTPRIIKNLISVRQLTT--------DNNVSVSFDPFGFSVSDFKTG--MPLLRCN 272
+ V + P + NL+S+ QL D V G + +G M L +
Sbjct: 355 IPEVYYVPELRNNLLSLGQLQERGLAILIRDGTCKVYHPSKGAIMETNMSGNRMFFLLAS 414
Query: 273 SLGDLYPVTRSSPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEP--SRSSSVCDSCV 330
++ + +WH R GH L L + K++ P + +C C+
Sbjct: 415 KPQKNSLCLQTEEVMDKENHLWHCRFGHLNQEGLKLLAHKKMVIGLPILKATKEICAICL 474
Query: 331 LGKHVRLPFSS-SAIITLRPFDILHSDLWTSPV--LSTAGHRYYVLFLDDHTDFLWTFPI 387
GK R S ++ + ++HSD+ P+ +S +G RY + F+DD T W + +
Sbjct: 475 TGKQHRESMSKKTSWKSSTQLQLVHSDI-CGPITPISHSGKRYILSFIDDFTRKTWVYFL 533
Query: 388 SKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTS 447
+KS+ + TF ++ + A + CL+ D G E+ ++ F +C ++G+ + + T
Sbjct: 534 HEKSEAFATFKIFKASVEKEIGAFLTCLRTDRGGEFTSNEFGEFCRSHGISRQLTAAFTP 593
Query: 448 SQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLY 507
QNG AERK RTI N +R++L+ VP FW A + + ++ N P ++ +P +
Sbjct: 594 QQNGVAERKNRTIMNAVRSMLSERQVPKMFWSEATKWSVHIQNRSPTAAVEGMTPEEAWS 653
Query: 508 HRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILIS 567
R P + RVFGC+ + P +KL +S CVFLG + ++ YD +KI+IS
Sbjct: 654 GRKPVVEYFRVFGCIGYVHIPDQKRSKLDDKSKKCVFLGVSEESKAWRLYDPVMKKIVIS 713
Query: 568 RHVIFDETR---FPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSP-- 622
+ V+FDE + + AD+ + EC ED + P+ +SP
Sbjct: 714 KDVVFDEDKSWDWDQADVEAKEV-TLECGDED----------DEKNSEVVEPIAVASPNH 762
Query: 623 --TDSSPPLSALVSPTA-SPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDD 679
+D++ S +++P++ +PSP+ P M + ++ S+ + +
Sbjct: 763 VGSDNNVSSSPILAPSSPAPSPVAAKVTRERRPPGWMADYETGEGEEIEENLSVMLLMMM 822
Query: 680 PSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQS 739
P+ + A+ D W+ AM+ E +++++NTWEL P I W+++ K
Sbjct: 823 TEADPIQFD--DAVKDKIWREAMEHEIESIVKNNTWELTTLPKGFTPIGVKWVYKTKLNE 880
Query: 740 NGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAF 799
+G ++YKARLV G +Q G+D E F+PV + T+RT+L+I+ +W I QLDV++AF
Sbjct: 881 DGEVDKYKARLVAKGYAQCYGIDYTEVFAPVARLDTVRTILAISSQFNWEIFQLDVKSAF 940
Query: 800 LHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHST 859
LHG+L E +Y+ P GF + V +L+K+LYGLKQAPRAWY R Y F
Sbjct: 941 LHGELKEEVYVRQPEGFIREGEEEKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEFERCP 1000
Query: 860 SDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIA 919
S+H+LF + +I + LYVDD+I S + F + EF M DLG + +FLGI
Sbjct: 1001 SEHTLFTKTRVGNILIVSLYVDDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIE 1060
Query: 920 VTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDG 979
V + GG+F+ Q YA E++AR GM N P+ KL+ D ++++ L G
Sbjct: 1061 VKQSDGGIFICQRRYAREVLARFGMDESNAVKNPIVPGTKLTKDENGEKVDETMFKQLVG 1120
Query: 980 ALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVET-- 1037
+L YLT TRPD+ Y V + M P H LA KR+LRY++GT+ G+ +
Sbjct: 1121 SLMYLTVTRPDLMYGVCLISRFMSNPRMSHWLAAKRILRYLKGTVELGIFYRRRKNRSLK 1180
Query: 1038 LVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSES 1097
L+++TD+D+ G + RRSTSG+ + I W+SK+QP ++ S+ EAEY A +
Sbjct: 1181 LMAFTDSDYAGDLNDRRSTSGFVFLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQC 1240
Query: 1098 CWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQAR 1157
W+R +L +L AT+++CDN S I LS +PV H ++KHIE+ H++R+ V +
Sbjct: 1241 VWLRKVLEKLGAEEKSATVINCDNSSTIQLSKHPVLHGKSKHIEVRFHYLRDLVNGDVVK 1300
Query: 1158 VLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
+ + P+ Q+ADIFTK L F+ R+ L +
Sbjct: 1301 LEYCPTEDQVADIFTKPLKLEQFEKLRALLGM 1332
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 523 bits (1346), Expect = e-148
Identities = 284/606 (46%), Positives = 369/606 (60%), Gaps = 21/606 (3%)
Query: 609 SSRPPDPPVQPSSPTDS---SPPLSALVSPTASPSPLPLPPVPPAPPV-----------R 654
S P P P PT S S P S S T++P PLPPV PAPP+
Sbjct: 854 SPLPQSPISSPHIPTPSTSISEPNSPSSSSTSTP---PLPPVLPAPPIIQVNAQAPVNTH 910
Query: 655 TMTTRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNT 714
+M TR+ GI KP + +S + S+ + + P QA+ D W+ AM SE NA I ++T
Sbjct: 911 SMATRAKDGIRKPNQKYSYATSL---AANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHT 967
Query: 715 WELVPRPC-DVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKP 773
W+LVP P V ++ C WIF K S+G RYKARLV G +Q G+D ETFSPV+K
Sbjct: 968 WDLVPPPPPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKS 1027
Query: 774 ATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLY 833
+IR VL +A+ RSWPI QLDV NAFL G L + +YM P GF D PDYVCRL+K++Y
Sbjct: 1028 TSIRIVLGVAVDRSWPIRQLDVNNAFLQGTLTDEVYMSQPPGFVDKDRPDYVCRLRKAIY 1087
Query: 834 GLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLR 893
GLKQAPRAWY Y+ ++GF +S SD SLF+ ++G I Y+L+YVDDI++ + L
Sbjct: 1088 GLKQAPRAWYVELRTYLLTVGFVNSISDTSLFVLQRGRSIIYMLVYVDDILITGNDTVLL 1147
Query: 894 KSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATP 953
K + L+ F++K+ L YFLGI R GL LSQ Y +++AR M + P ATP
Sbjct: 1148 KHTLDALSQRFSVKEHEDLHYFLGIEAKRVPQGLHLSQRRYTLDLLARTNMLTAKPVATP 1207
Query: 954 VDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLAL 1013
+ T KL+ SGT D + YR + G+LQYL FTRPD+SYAV ++ +MH P H AL
Sbjct: 1208 MATSPKLTLHSGTKLPDPTEYRGIVGSLQYLAFTRPDLSYAVNRLSQYMHMPTDDHWNAL 1267
Query: 1014 KRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSK 1073
KRVLRY+ GT +G+ L +L +Y+DADW G D ST+GY V+LG + ISWSSK
Sbjct: 1268 KRVLRYLAGTPDHGIFLKKGNTLSLHAYSDADWAGDTDDYVSTNGYIVYLGHHPISWSSK 1327
Query: 1074 RQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVH 1133
+Q + RSS EAEYR VAN SE WI +LL EL LS +++CDNV A YL NPV
Sbjct: 1328 KQKGVVRSSTEAEYRSVANTSSELQWICSLLTELGIQLSHPPVIYCDNVGATYLCANPVF 1387
Query: 1134 HQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPP 1193
H R KHI +D HF+R +V G RV+HV + Q+AD TK L R+ F +F + V + P
Sbjct: 1388 HSRMKHIALDYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRVAFQNFSRKIGVIKVP 1447
Query: 1194 ASTAGV 1199
S GV
Sbjct: 1448 PSCGGV 1453
Score = 329 bits (843), Expect = 7e-90
Identities = 201/581 (34%), Positives = 300/581 (51%), Gaps = 54/581 (9%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GA+ H + L+ + + +++ G IPI +G S+PT + L LN
Sbjct: 310 WLLDSGATHHITSDFNNLSFHQPYTG-GDDVMIADGSTIPITHTGSASLPTSSRSLDLNK 368
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY--PVTRS 283
VL+ P I KNLISV +L N VSV F P F V D TG+PLL+ + +LY P+ S
Sbjct: 369 VLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWPIASS 428
Query: 284 -------SPFAGLASSVWHNRLGHPASSALNH-LRNNKLIFCEPSRSSSVCDSCVLGKHV 335
SP + S WH+RLGHP+ + LN + N+ L PS C C + K
Sbjct: 429 QAVSMFASPCSKATHSSWHSRLGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFINKSH 488
Query: 336 RLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYE 395
++PFS+S I + +P + ++SD+W+SP+LS +RYYV+F+D T + W +P+ +KSQV +
Sbjct: 489 KVPFSNSTITSSKPLEYIYSDVWSSPILSIDNYRYYVIFVDHFTRYTWLYPLKQKSQVKD 548
Query: 396 TFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAER 455
TF +L++ +F I L DNG E+ Y +G+ S PHT NG +ER
Sbjct: 549 TFIIFKSLVENRFQTRIGTLYSDNGGEFV--VLRDYLSQHGISHFTSPPHTPEHNGLSER 606
Query: 456 KIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSH 515
K R I M TLL+H+SVP ++W +A +A YL+N LP LQ SP Q L+ + P+Y
Sbjct: 607 KHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPPNYEK 666
Query: 516 LRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDET 575
L+VFGC C+P +KL+ +S C F+GY + Y C + ++ SRHV FDE
Sbjct: 667 LKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQFDER 726
Query: 576 RFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSP-----TDSSP--- 627
FPF+ + + S E ++ P +W + ++ P P V P+ P D+SP
Sbjct: 727 CFPFSTTNFGVSTSQEQRSDSAP-----NWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPP 781
Query: 628 ----------------PLSALVSPTASPSPLPL--PPVPPAPPVRTMTTRSMHGISKPKK 669
P S++ SP++S P P P A P +T + S I
Sbjct: 782 SSPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNNPN 841
Query: 670 PFSLSVSIDDPSISPLPHN---------PKQALSDPNWKSA 701
P S S + + + SPLP + P ++S+PN S+
Sbjct: 842 PNSPSPNSPNQN-SPLPQSPISSPHIPTPSTSISEPNSPSS 881
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 497 bits (1279), Expect = e-140
Identities = 332/1044 (31%), Positives = 515/1044 (48%), Gaps = 46/1044 (4%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLA-LN 224
W MDTG S H + + + +G+ ++G G + + L
Sbjct: 287 WVMDTGCSYHMTYKREWFEDLNE--DAGGSVRMGNKTVSKVRGIGTIRVKNEAGMVVRLT 344
Query: 225 HVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY-----P 279
+V + P + +NL+S L T SF ++S LL LY P
Sbjct: 345 NVRYIPEMDRNLLS---LGTFEKSGYSFKLENGTLSIIAGDSVLLTVRRCYTLYLLQWRP 401
Query: 280 VTRSSPFAGLASS---VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVR 336
VT S +WH RLGH + ++ L L+ + C+ C+ GK R
Sbjct: 402 VTEESLSVVKRQDDTILWHRRLGHMSQKNMDLLLKKGLLDKKKVSKLETCEDCIYGKAKR 461
Query: 337 LPFSSSAIITLRPFDILHSDLWTSPVL--STAGHRYYVLFLDDHTDFLWTFPISKKSQVY 394
+ F+ + T + +HSDLW +P + S +Y++ F+DD+T + + + K + +
Sbjct: 462 IGFNLAQHDTREKLEYVHSDLWGAPSVPFSLGKCQYFISFIDDYTRKVRIYFLKTKDEAF 521
Query: 395 ETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAE 454
+ F A L++ Q IK L+ DNG E+ N SF +C G+++ +C +T QNG AE
Sbjct: 522 DKFVEWANLVENQTDKRIKTLRTDNGLEFCNRSFDEFCSQKGILWHRTCAYTPQQNGVAE 581
Query: 455 RKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYS 514
R RT+ +R++L+ S +P FW A L+N P L + P + + P YS
Sbjct: 582 RMNRTLMEKVRSMLSDSGLPKKFWAEATHTTAILINKTPSSALNYEVPDKRWSGKSPIYS 641
Query: 515 HLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDE 574
+LR FGC+ F KL PR+ + +GYP+ +GYK + L +K ++SR+VIF E
Sbjct: 642 YLRRFGCIAFV---HTDDGKLNPRAKKGILVGYPIGVKGYKIWLLEEKKCVVSRNVIFQE 698
Query: 575 TRFPFADLSLTPAPSYECFTEDLPPSL-----IHHWQTVSSRPPDPPVQPSSPTDSSPPL 629
+ + + E + PPS + H + ++S DP V+ SP + SP
Sbjct: 699 NA---SYKDMMQSKDAEKDENEAPPSSYLDLDLDHEEVITSGGDDPIVEAQSPFNPSPAT 755
Query: 630 SALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSI----DDPSISPL 685
+ S + + VR R++ + L+ ++ D I P
Sbjct: 756 TQTYSEGVNSETDIIQSPLSYQLVRDRDRRTIRAPVRFDDEDYLAEALYTTEDSGEIEPA 815
Query: 686 PHNP-KQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFE 744
++ K++++ WK AM E + I+++TW +V RP VI WI++ K G+ E
Sbjct: 816 DYSEAKRSMNWNKWKLAMNEEMESQIKNHTWTVVKRPQHQKVIGSRWIYKFKLGIPGVEE 875
Query: 745 -RYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGD 803
R+KARLV G +Q G+D E F+PVVK +IR ++SI + QLDV+ AFLHG+
Sbjct: 876 GRFKARLVAKGYAQRKGIDYHEIFAPVVKHVSIRILMSIVAQEDLELEQLDVKTAFLHGE 935
Query: 804 LHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHS 863
L E IYM P G+ + D VC L KSLYGLKQAP+ W ++F Y+S IGF S D
Sbjct: 936 LKEKIYMVPPEGYEEMFKEDEVCLLNKSLYGLKQAPKQWNEKFNAYMSEIGFIRSLYDSC 995
Query: 864 LFI--FRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAV- 920
+I GS + Y+LLYVDD+++ A + + L+ F MKDLG LG+ +
Sbjct: 996 AYIKELSDGSRV-YLLLYVDDMLVAAKNKEDISQLKEELSQRFDMKDLGAAKRILGMEII 1054
Query: 921 -TRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCED------ASL 973
R L+LSQ+ Y +I+ MA TP+ K+ +++ E +
Sbjct: 1055 RNREENTLWLSQNGYLNKILETYNMAESKHVVTPLGAHLKMRAATVEKQEQDEDYMKSIP 1114
Query: 974 YRSLDGALQY-LTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYP 1032
Y S G++ Y + TRPD++Y V + +M P +H L +K VLRY++G+L L
Sbjct: 1115 YSSAVGSIMYAMIGTRPDLAYPVGIISRYMSQPAREHWLGVKWVLRYIKGSLGTKLQYKR 1174
Query: 1033 SPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVAN 1092
S +V Y DAD C D RRS +G LG + ISW S +Q ++ S+ EAEY +
Sbjct: 1175 SSDFKVVGYCDADHAACKDRRRSITGLVFTLGGSTISWKSGQQRVVALSTTEAEYMSLTE 1234
Query: 1093 VVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVV 1152
V E+ W++ LL E + ++ + CD+ SAI LS N VHH+RTKHI++ ++R+ +
Sbjct: 1235 AVKEAVWMKGLLKEFGYE-QKSVEIFCDSQSAIALSKNNVHHERTKHIDVRYQYIRDIIA 1293
Query: 1153 RGQARVLHVPSRHQIADIFTKGLP 1176
G V+ + + ADIFTK +P
Sbjct: 1294 NGDGDVVKIDTEKNPADIFTKIVP 1317
>At4g27200 putative protein
Length = 819
Score = 482 bits (1241), Expect = e-136
Identities = 286/742 (38%), Positives = 381/742 (50%), Gaps = 73/742 (9%)
Query: 505 LLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKI 564
+L + S S R+ G CFP NK P S CVFLGY ++GY+C ++
Sbjct: 1 MLVRKRHSTSSWRI-GSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRCLYPPTGRL 59
Query: 565 LISRHVIFDETRFPFADL-------------------SLTPAPSYEC--------FTE-D 596
ISRHVIFDE+ +PF+ S +PAPS FT D
Sbjct: 60 YISRHVIFDESVYPFSHTYKHLHPQPRTPLLAAWLRSSDSPAPSTSTSPSSRSPLFTSAD 119
Query: 597 LPP----------------SLIHHWQTVSSRPPDPPVQPSSPTDS--------SPPLSAL 632
PP S+ H + + PD + ++ DS S +
Sbjct: 120 FPPLPQRKTPLLPTLVPISSVSHASNITTQQSPDFDSERTTDFDSASIGDSSHSSQAGSD 179
Query: 633 VSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQA 692
T + + + P + V M TR+ GISKP + V + P P A
Sbjct: 180 SEETIQQASVNVHQTPASTNVHPMVTRAKVGISKPNPRY---VFLSHKVSYPEPKTVTAA 236
Query: 693 LSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVG 752
L P W AM E + TW LVP D++V+ W+FR K ++G + KAR+V
Sbjct: 237 LKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVFRTKLHADGTLNKLKARIVA 296
Query: 753 DGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHH 812
G Q G+D ET+SPVV+ T+R VL +A + +W I Q+DV+NAFLHGDL ET+YM
Sbjct: 297 KGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQMDVKNAFLHGDLKETVYMTQ 356
Query: 813 PLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSD 872
P RD +VC L KS+YGLKQ+PRAW+ +F+ ++ GF SD SLFI+ ++
Sbjct: 357 PAANRD-----HVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCRKSDPSLFIYAHNNN 411
Query: 873 IAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSY-FLGIAVTRHAGGLFLSQ 931
+ +LL S +A L EF M D+G S FLGI V R GLF+SQ
Sbjct: 412 LILLLL-----------SQTLTSLLAALNKEFRMTDMGQHSLTFLGIQVQRQQNGLFMSQ 460
Query: 932 STYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDI 991
YA +++ A M C P TP+ + D + +RS+ G LQYLT TRPDI
Sbjct: 461 QKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTYFRSIAGKLQYLTLTRPDI 520
Query: 992 SYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPD 1051
+AV VC MH P LKR+LRY++GT+T G+ L +Y+D+DWG C
Sbjct: 521 QFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISYSRDSPTLLQAYSDSDWGNCKQ 580
Query: 1052 TRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPL 1111
TRRS G C F+G NL+SWSSK+ PT+SRSS EAEY+ +++ SE W+ LL EL PL
Sbjct: 581 TRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLSDAASEILWLSTLLRELRIPL 640
Query: 1112 SQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIF 1171
+ CDN+SA+YL+ NP H RTKH ++D HFVRE+V V H+P QIADIF
Sbjct: 641 PDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERVALKALVVKHIPGSEQIADIF 700
Query: 1172 TKGLPRLLFDDFRSSLSVGEPP 1193
TK LP F R L V PP
Sbjct: 701 TKSLPYEAFIHLRGKLGVTLPP 722
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 444 bits (1143), Expect = e-124
Identities = 254/684 (37%), Positives = 375/684 (54%), Gaps = 55/684 (8%)
Query: 547 YPMNHRGYKCYDLSHRKILISRHVIFDETRFPF---------ADL---SLTPAPSYECFT 594
YP ++GYK DL I I+R+V+F ET+FPF D+ S+ P P+ F
Sbjct: 335 YPSGYKGYKVLDLESHSISITRNVVFHETKFPFKTSKFLKESVDMFPNSILPLPAPLHFV 394
Query: 595 EDLP-----------PSLIHHWQTVSSRPPDPPVQPSSPTDS------SPPLSALVSPTA 637
E +P S + + SS PP P + TD+ S P++
Sbjct: 395 ESMPLDDDLRADDNNASTSNSASSASSIPPLPSTVNTQNTDALDIDTNSVPIARPKRNAK 454
Query: 638 SPSPLP------LPPVPPAPPVRTMTTRSMHGISKPKK---PFSLSVSIDDPSISPLPHN 688
+P+ L +P + P + + + PKK P+ +S +I ++PL H+
Sbjct: 455 APAYLSEYHCNSVPFLSSLSPTTSTSIETPSSSIPPKKITTPYPMSTAISYDKLTPLFHS 514
Query: 689 ----------PK---QALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRH 735
PK QA+ W A E +AL ++ TW + NV+ C W+F
Sbjct: 515 YICAYNVETEPKAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNVVGCKWVFTI 574
Query: 736 KKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDV 795
K +G ERYKARLV G +Q G+D ETFSPV K +++ +L +A + W + Q+DV
Sbjct: 575 KYNPDGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAATGWSLTQMDV 634
Query: 796 QNAFLHGDLHETIYMHHPLGFRDPHHPDY----VCRLKKSLYGLKQAPRAWYQRFADYVS 851
NAFLHG+L E IYM P G+ P VCRL KSLYGLKQA R WY+R +
Sbjct: 635 SNAFLHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQASRQWYKRLSSVFL 694
Query: 852 SIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGP 911
F S +D+++F+ + I +L+YVDD+++ ++ ++ LL SEF +KDLGP
Sbjct: 695 GANFIQSPADNTMFVKVSCTSIIVVLVYVDDLMIASNDSSAVENLKELLRSEFKIKDLGP 754
Query: 912 LSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDA 971
+FLG+ + R + G+ + Q YA ++ G++ C PS+ P+D L+ GT +A
Sbjct: 755 ARFFLGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLTKEMGTLLPNA 814
Query: 972 SLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLY 1031
+ YR L G L YL TRPDI++AV + + AP HM A +VLRY++G GL
Sbjct: 815 TSYRELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLKGNPGQGLMYS 874
Query: 1032 PSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVA 1091
S L ++DADWG C D+RRS +G+C++LG +LI+W SK+Q +SRSS E+EYR +A
Sbjct: 875 ASSELCLNGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLA 934
Query: 1092 NVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKV 1151
E W++ LL +LH ++ + CDN SA++L+ NPV H+RTKHIE+D H VR+++
Sbjct: 935 QATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQI 994
Query: 1152 VRGQARVLHVPSRHQIADIFTKGL 1175
G+ + LHVP+ +Q+ADI TK L
Sbjct: 995 KAGKLKTLHVPTGNQLADILTKPL 1018
Score = 71.2 bits (173), Expect = 3e-12
Identities = 76/292 (26%), Positives = 124/292 (42%), Gaps = 58/292 (19%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLI-VGSGQGIPIQGSGYTSIPTPHKPLALN 224
W +D+GASSH + LT + L H++ + + +G + I +G I + L L+
Sbjct: 99 WIIDSGASSHVCSD---LTMFRELIHVSGVTVTLPNGTRVAITHTGTICITST---LILH 152
Query: 225 HVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPV-TRS 283
+VL P NLISV L + S F + + G+ + R + +LY + T+
Sbjct: 153 NVLLVPDFKFNLISVCCLVKTLSYSAHFFADCCYIQELTRGLMIGRGKTYNNLYILETQR 212
Query: 284 SPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFSSSA 343
+ F+ PA+S+ + +V D C+L H RL
Sbjct: 213 TSFSPSL----------PAASSF---------------TGTVQDDCLLW-HQRLGIRH-- 244
Query: 344 IITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYETFTTLATL 403
LH + Y++ +DD T W + + KS+V F L
Sbjct: 245 --------YLHYR-----------NLYFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKL 285
Query: 404 IKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAER 455
I TQ++A IK ++ DN +E +F ++ G+I +FSC +T QN ER
Sbjct: 286 IFTQYNAKIKAIRSDNVKEL---AFTKFVKEQGMIHQFSCAYTPQQNSVVER 334
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.134 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,698,926
Number of Sequences: 26719
Number of extensions: 1473546
Number of successful extensions: 14097
Number of sequences better than 10.0: 473
Number of HSP's better than 10.0 without gapping: 246
Number of HSP's successfully gapped in prelim test: 237
Number of HSP's that attempted gapping in prelim test: 5722
Number of HSP's gapped (non-prelim): 3044
length of query: 1199
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1089
effective length of database: 8,379,506
effective search space: 9125282034
effective search space used: 9125282034
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0490.3


