

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0487.10
(289 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
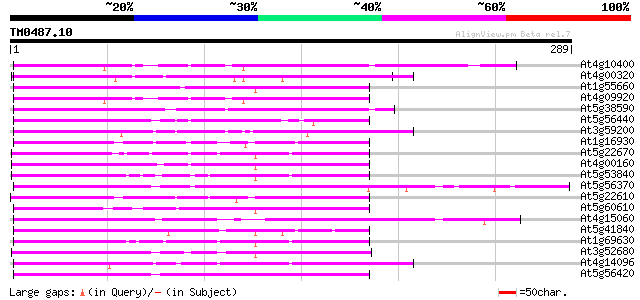
Score E
Sequences producing significant alignments: (bits) Value
At4g10400 putative protein 107 7e-24
At4g00320 106 2e-23
At1g55660 hypothetical protein 105 2e-23
At4g09920 hypothetical protein (probably remnant) 104 6e-23
At5g38590 unknown protein 102 3e-22
At5g56440 putative protein 101 5e-22
At3g59200 unknown protein 100 7e-22
At1g16930 hypothetical protein 100 7e-22
At5g22670 unknown protein 100 9e-22
At4g00160 unknown protein 100 9e-22
At5g53840 putative protein 99 2e-21
At5g56370 unknown protein 99 3e-21
At5g22610 unknown protein 98 6e-21
At5g60610 unknown protein 96 2e-20
At4g15060 hypothetical protein 96 3e-20
At5g41840 putative protein 94 7e-20
At1g69630 hypothetical protein 94 9e-20
At3g52680 unknown protein 93 1e-19
At4g14096 F-box protein - like 93 2e-19
At5g56420 unknown protein 92 4e-19
>At4g10400 putative protein
Length = 326
Score = 107 bits (267), Expect = 7e-24
Identities = 89/269 (33%), Positives = 135/269 (50%), Gaps = 31/269 (11%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDF-----DEANFRRTKG 57
DRIS LPDE+L ILSF+PTK AV+TSILSKRW+ LW +T L F E+ F+R +
Sbjct: 2 DRISGLPDEVLVKILSFVPTKVAVSTSILSKRWEFLWMWLTKLKFGSKRYSESEFKRLQC 61
Query: 58 WHSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICI 117
+ R + +HR I+ R+ H + P ++ +WV AV R E + I
Sbjct: 62 FLDRNL-PLHRA-------PVIESFRLVLSDSHFK-PEDIRMWVVVAVSRYIRE---LKI 109
Query: 118 YS-----LSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCL 172
YS + SS+++CK+LV+LKL+G L V LP LK L L+ ++ ++ L
Sbjct: 110 YSSHYGEKQNILPSSLYTCKSLVILKLDGGVLLDVPRMVCLPSLKTLELKGVRYFKQGSL 169
Query: 173 AQLLNGCPVLEDLKAKAIYFKKYKETGFEYKTLSKLVRADISGTSTDLFRMEVINNVDFL 232
+LL CPVLEDL + + G + L R +S S+ F ++ + + F
Sbjct: 170 QRLLCNCPVLEDL---VVNLSHHDNMGKLTVIVPSLQRLSLSTPSSREFVIDTPSLLSFQ 226
Query: 233 RIHEIYIRIPHDDQFTDMFHNLTHIELGY 261
+ +D+ T + N+ + Y
Sbjct: 227 LVDR------NDNSHTFLIENMPKLREAY 249
>At4g00320
Length = 773
Score = 106 bits (264), Expect = 2e-23
Identities = 70/210 (33%), Positives = 114/210 (53%), Gaps = 6/210 (2%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWH-SR 61
D+ S LPDE+L +LSFLPTK AV TS+LS RWK LW + L++D ++ ++G +R
Sbjct: 2 DKTSQLPDELLVKVLSFLPTKDAVRTSLLSMRWKSLWMWLPKLEYDFRHYSVSEGQGLAR 61
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVD--ICIYS 119
F+ ++ + H ++ + L++ + + S P ++ +WV+ AV + +C ++
Sbjct: 62 FI-TLSLLGHKAPAIESL-SLKLRYGAIGSIKPEDIYLWVSLAVHDSNVRELSLKLCTFA 119
Query: 120 LSPLNL-SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNG 178
P L S++ CK++V+LKL+ L V LP LK L L + S L +LL+
Sbjct: 120 ERPTKLPKSLYKCKSIVILKLKDEILVDVPRKVCLPSLKTLFLGRVTYSDEDSLHRLLSN 179
Query: 179 CPVLEDLKAKAIYFKKYKETGFEYKTLSKL 208
CPVLEDL + + K+L +L
Sbjct: 180 CPVLEDLVVERDRIDNLGKLSVVVKSLQRL 209
Score = 95.5 bits (236), Expect = 3e-20
Identities = 79/217 (36%), Positives = 114/217 (52%), Gaps = 29/217 (13%)
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFR-------- 53
+D IS LPD ++C+ILSFLPTK A +T++L+KRWKPL + LDFDE +FR
Sbjct: 278 KDGISGLPDAMICHILSFLPTKVAASTTVLAKRWKPLLAFMPNLDFDE-SFRFDPRMTCE 336
Query: 54 -RTKGWHSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEH 112
R KG S F+ V V+ + L +V ++V W+ ++RG +
Sbjct: 337 ERRKGSES-FMLVVDSVLALQ--AEANATLNKFYVKCEGVEQNSVLEWIPKVLKRGVLD- 392
Query: 113 VDICIYS---------LSPLNLSSIFSCKTLVVLKL---EGLDLNTTVSSVDLPFLKVLH 160
+D+ I S PL S IF KTLV LK+ +G ++N V LP LK LH
Sbjct: 393 IDLQIPSSRGFGSNSTFYPLP-SEIFVSKTLVRLKIQFQDGANVNVE-GDVSLPMLKTLH 450
Query: 161 LQVLQLSQRRCLAQLLNGCPVLEDLKAKAIYFKKYKE 197
L +++ R L +LL+GC LE+L + +K+ E
Sbjct: 451 LDYVKM-DTRMLQKLLSGCHTLEELLLMNLIWKESSE 486
>At1g55660 hypothetical protein
Length = 492
Score = 105 bits (263), Expect = 2e-23
Identities = 69/187 (36%), Positives = 102/187 (53%), Gaps = 6/187 (3%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWH-SR 61
D+IS LPDE+L +LSFL TK AV+TSILS RWK LW + L+++ ++ ++G +R
Sbjct: 53 DKISQLPDELLVKVLSFLSTKDAVSTSILSMRWKSLWMWLPKLEYNFRHYSVSEGQGLAR 112
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLS 121
F+ S RV P K R + S P ++ +WV+ AV + + +++ +
Sbjct: 113 FITSSLRVHKAPAIESLSLKFRYGAIG--SIKPKDIYLWVSLAVHVSNVRELSLKLFNFA 170
Query: 122 PLNL---SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNG 178
L S+ CK++V+LKL+ L V LP LK L L + S L +LL+
Sbjct: 171 ELPTKLPKSLCKCKSIVILKLKDEILVDVPRKVCLPSLKTLFLGRVTYSDANSLHRLLSN 230
Query: 179 CPVLEDL 185
CPVLEDL
Sbjct: 231 CPVLEDL 237
>At4g09920 hypothetical protein (probably remnant)
Length = 414
Score = 104 bits (259), Expect = 6e-23
Identities = 77/193 (39%), Positives = 108/193 (55%), Gaps = 22/193 (11%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDF-----DEANFRRTKG 57
DRI LPDE+L ILSF+PTK AV+TSILSKRW+ LW +T L F E+ F+R +
Sbjct: 2 DRIIGLPDEVLVKILSFVPTKVAVSTSILSKRWEFLWMWLTKLKFGSKRYSESEFKRLQC 61
Query: 58 WHSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICI 117
+ R + +HR I+ R+ H + P ++ +WV AV R E + I
Sbjct: 62 FLDRNL-PLHRA-------PVIESFRLVLSDSHFK-PEDIRMWVVVAVSRYIRE---LKI 109
Query: 118 YS-----LSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCL 172
YS + SS+++CK+LV+LKL+G L V LP LK L L+ ++ ++ L
Sbjct: 110 YSSHYGEKQNILPSSLYTCKSLVILKLDGGVLLDVPRMVCLPSLKTLELKGVRYFKQGSL 169
Query: 173 AQLLNGCPVLEDL 185
+LL CPVLEDL
Sbjct: 170 QRLLCNCPVLEDL 182
>At5g38590 unknown protein
Length = 410
Score = 102 bits (253), Expect = 3e-22
Identities = 71/198 (35%), Positives = 109/198 (54%), Gaps = 11/198 (5%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
D+I+ LPD++L ILS++PT AV+TSILSKRW+ LW + LD+ R+ R
Sbjct: 2 DKINGLPDDLLVKILSYVPTDIAVSTSILSKRWEFLWMWLPNLDYTSRWCRKPGDVGLRD 61
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEV-PSNVNVWVNAAVQRGGFEHVDICIYSLS 121
+ +H ++ ++ F S ++ P ++ W+ AV R + +DI +S +
Sbjct: 62 FIHKNLPLHRAPVIESLR-----FHSNSPDIKPEDIRRWIEIAVSRHVHD-LDIDHFSEN 115
Query: 122 P-LNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCP 180
+ LSS F+CK+LV LKL + L S V LP LK L L + + + L +LL+ CP
Sbjct: 116 ENIFLSSFFACKSLVTLKLRSVTLRDIPSMVCLPSLKTLLLDNVSFVEGKSLQELLSICP 175
Query: 181 VLEDLKAKAIYFKKYKET 198
VLEDL ++Y Y+ T
Sbjct: 176 VLEDL---SVYCDDYENT 190
>At5g56440 putative protein
Length = 430
Score = 101 bits (251), Expect = 5e-22
Identities = 75/192 (39%), Positives = 110/192 (57%), Gaps = 20/192 (10%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWH-SR 61
DRIS LPD+++ ILSF+PTK V+T++LSKRW+ LW+ V LD+ + + T+ W SR
Sbjct: 2 DRISLLPDDVVFKILSFVPTKVVVSTNLLSKRWRYLWKHVPKLDYRDPSLVDTEHWRASR 61
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFE-HVDICIYSL 120
FV + P ++ L +S +SR+ PS++ W++ A+ R H+ I S
Sbjct: 62 FVDKFLLLHEAP----VLETLHLS-LSRNCP-PSDIETWISVAISRRVRNLHIYRYIPST 115
Query: 121 SPLNL-SSIFSCKTLVVLKLEGLDLNTTVSSVDLPF------LKVLHLQVLQLSQRRCLA 173
P+ L S+++C+TLV L L L+ TV D PF LKVL L + S +
Sbjct: 116 GPIRLPRSLYTCETLVSLYLL---LDFTVD--DAPFMFCFRSLKVLVLLFAKFSSDEIVN 170
Query: 174 QLLNGCPVLEDL 185
+LL+GCPVLE L
Sbjct: 171 RLLSGCPVLEGL 182
>At3g59200 unknown protein
Length = 520
Score = 100 bits (250), Expect = 7e-22
Identities = 80/215 (37%), Positives = 119/215 (55%), Gaps = 14/215 (6%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTK-----G 57
DRISSLP+ ++ +ILSFLPTK+A +TS+LSK+W+ L+ VT LDFD+++++ K
Sbjct: 7 DRISSLPNPVVSHILSFLPTKEAASTSVLSKKWRYLFAYVTNLDFDDSDYQDGKPKSDVE 66
Query: 58 WHSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICI 117
F++ V RV+ + K + S + + V W+ + RG E +D+ I
Sbjct: 67 LSRSFMEFVDRVLALQG-NGSVNKFSLE-CSNYDVDLARVTGWILNVLGRGVSE-LDLSI 123
Query: 118 YSLSPLNLSSIFSCKTLVVLKL-EGLDLNTTVSSVD--LPFLKVLHLQVLQLSQRRC-LA 173
PL S IF KTLV LKL DL T+ D LP LK L++ + + +R
Sbjct: 124 LEY-PLP-SEIFVSKTLVRLKLGPANDLTLTLDRKDVFLPKLKTLYIDCVDVQERGFGFV 181
Query: 174 QLLNGCPVLEDLKAKAIYFKKYKETGFEYKTLSKL 208
+LL+GCPVLE+L I ++ +K KTL +L
Sbjct: 182 KLLSGCPVLEELVLMNIGWENWKFCSVSVKTLKRL 216
>At1g16930 hypothetical protein
Length = 449
Score = 100 bits (250), Expect = 7e-22
Identities = 70/196 (35%), Positives = 109/196 (54%), Gaps = 30/196 (15%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRIS+LPD +LC ILS L TK++V TS+LSKRW+ LW V LD D NF F
Sbjct: 15 DRISNLPDSLLCQILSDLSTKESVCTSVLSKRWRNLWLHVPVLDLDSNNFPD----DDVF 70
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSF-VSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSL- 120
V V+R + + + Q +++ ++ + V+ H S W+NA ++R +C +++
Sbjct: 71 VSFVNRFLGSEN-EQHLERFKLIYEVNEHD--ASRFKSWINAVIKR------RVCHFNVH 121
Query: 121 -----------SPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQR 169
PL S++SC+ LV L+L + L+ SV LP +K++HL +++
Sbjct: 122 NEVDDDDELVKMPL---SLYSCERLVNLQLYRVALDHP-ESVSLPCVKIMHLDMVKYDAD 177
Query: 170 RCLAQLLNGCPVLEDL 185
L L++GCPVLE+L
Sbjct: 178 STLEILISGCPVLEEL 193
>At5g22670 unknown protein
Length = 443
Score = 100 bits (249), Expect = 9e-22
Identities = 73/186 (39%), Positives = 105/186 (56%), Gaps = 10/186 (5%)
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
+D IS LPD++LC ILS LPTK AV TS+LSKRWK SV L+F+ + F G++
Sbjct: 10 QDSISLLPDDLLCRILSNLPTKVAVRTSVLSKRWKRFSLSVPLLEFNVSEFH---GYY-E 65
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLS 121
F + VH + T I KL+++F + + S + W++ AV+R +H+DI +S
Sbjct: 66 FARVVHGFLDT-SRETCIHKLKLAFEKKQHD-RSYLTQWIHNAVKR-KVQHLDIGRWSYL 122
Query: 122 PLNL--SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGC 179
L S+++C+TLV L+L + L V LP LK +HL L L++ C
Sbjct: 123 GQELIPHSLYTCETLVSLRLHNVSL-PDFDHVSLPRLKTMHLIDNIYPNDALLENLISSC 181
Query: 180 PVLEDL 185
PVLEDL
Sbjct: 182 PVLEDL 187
>At4g00160 unknown protein
Length = 453
Score = 100 bits (249), Expect = 9e-22
Identities = 65/187 (34%), Positives = 99/187 (52%), Gaps = 7/187 (3%)
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
+DRIS LPD +L ILSFLPTK VATS+ SK+W+PLW+ V L+FD ++ + +
Sbjct: 15 KDRISELPDALLIKILSFLPTKIVVATSVFSKQWRPLWKLVPNLEFDSEDYDDKEQYTFS 74
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLS 121
+ + H L + R+ F S + P ++ +WV A R E V + + +
Sbjct: 75 EIVCKSFLSHKAPVL---ESFRLEFESEKVD-PVDIGLWVGIAFSRHLRELVLVAADTGT 130
Query: 122 PLNL---SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNG 178
SS+ +C TL L+L L L S V + L+ LHL+++ + LL+G
Sbjct: 131 GTAFKFPSSLCTCNTLETLRLVLLILVDISSPVVMKSLRTLHLELVSYKDESSIRNLLSG 190
Query: 179 CPVLEDL 185
CP+LE+L
Sbjct: 191 CPILEEL 197
>At5g53840 putative protein
Length = 444
Score = 99.4 bits (246), Expect = 2e-21
Identities = 68/186 (36%), Positives = 100/186 (53%), Gaps = 11/186 (5%)
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
E+R+S LPD ++C ILS L TK AV TSILS RW+ LW+ V LDFD R + S
Sbjct: 17 EERLSQLPDHLICVILSHLSTKDAVRTSILSTRWRNLWQLVPVLDFDSRELRSFSEFVS- 75
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLS 121
F S +H + IQKLR+ + W++ V R +H+DI +++ S
Sbjct: 76 FAGSFF-YLHKDSY---IQKLRVCIYDLAGNY--YLTSWID-LVTRHRIQHIDISVFTCS 128
Query: 122 PLNL--SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGC 179
+ S+++C TLV LKL + + V V LP LK+L L + + L ++++
Sbjct: 129 GFGVIPLSLYTCDTLVHLKLSRVTM-VNVEFVSLPCLKILDLDFVNFTNETTLDKIISCS 187
Query: 180 PVLEDL 185
PVLE+L
Sbjct: 188 PVLEEL 193
>At5g56370 unknown protein
Length = 421
Score = 98.6 bits (244), Expect = 3e-21
Identities = 91/330 (27%), Positives = 143/330 (42%), Gaps = 58/330 (17%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
D IS LPD+ L ILS LPTK + TS+LSKRW+ LW+ V L + + G RF
Sbjct: 2 DSISLLPDDFLLRILSLLPTKDVLNTSVLSKRWRYLWKLVPKLQYSLIDKNADHGTFVRF 61
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFE-HVDICIYSLS 121
V + P ++ L + + SEV ++ WV AV++G E D Y
Sbjct: 62 VDRSLLLSMAP----VLESLHLKLGRQCSEV--DIGFWVRIAVEKGLCELDFDYEHYKTE 115
Query: 122 PLNL-SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCP 180
P L S+F+C TL VLKL+ + L V LK LHL+ + + +LL+ CP
Sbjct: 116 PCRLPQSLFTCGTLTVLKLKNVSLKDVQFPVCFKLLKTLHLEYVIFLDKETPQKLLSSCP 175
Query: 181 VLE-----------------------------------DLKAKAIYFKKYKETGFEYK-- 203
+LE +L K K +G Y+
Sbjct: 176 ILEVFDLTRDDDDVDNVMSFSVMVPSLQRFIYCGGSGAELVMNTPSLKYLKLSGCGYECM 235
Query: 204 --TLSKLVRADIS-GTSTDLFRMEVINNVDFLRIHEIYIRIPHDDQFT--DMFHNLTHIE 258
L ++V A + STD +++ ++ + + + +P + + +FH L H+E
Sbjct: 236 IGNLPEIVEAHVEVACSTD----DILTSL--ASVKRLLLCLPTEPELPTGTIFHQLEHLE 289
Query: 259 LGYYDYNTNWIEVVEFLNYCPKLQVLVINQ 288
+ T W ++ L + PKL+ L +N+
Sbjct: 290 --FCSCCTEWDILMFMLKHSPKLRSLKLNE 317
>At5g22610 unknown protein
Length = 502
Score = 97.8 bits (242), Expect = 6e-21
Identities = 70/196 (35%), Positives = 104/196 (52%), Gaps = 20/196 (10%)
Query: 1 MEDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHS 60
+ED IS LP+ +L ILS+LPTK V TS+LSKRWK +W + LD D + F +
Sbjct: 16 IEDLISKLPEVLLSQILSYLPTKDIVRTSVLSKRWKSVWLLIPGLDLDSSEFPH----YD 71
Query: 61 RFVQSVHRVIHTPDFLQP-IQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDI---- 115
FV ++ + P + KL++S + ++ PS V +W + V RG +H+D+
Sbjct: 72 TFVDFMNEFLFFSREENPCLHKLKLS-IQKNENDPSCVTLWTD-CVARGKLQHLDVEFGG 129
Query: 116 -----CIYSLSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVD-LPFLKVLHLQVLQLSQR 169
+ + PL S++ CKTL+ L+L + L SVD LP LK + L+ S
Sbjct: 130 RVMEREFWEMMPL---SLYICKTLLHLRLYRVLLGNFDQSVDSLPSLKSMCLEENVYSNE 186
Query: 170 RCLAQLLNGCPVLEDL 185
L L++ C VLEDL
Sbjct: 187 ASLESLISSCRVLEDL 202
>At5g60610 unknown protein
Length = 388
Score = 96.3 bits (238), Expect = 2e-20
Identities = 73/187 (39%), Positives = 104/187 (55%), Gaps = 15/187 (8%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRIS LPDE+L I+SF+PTK AV+TSILSKRW+ LW+ V L+ D T+ F
Sbjct: 2 DRISGLPDELLVKIISFVPTKVAVSTSILSKRWESLWKWVPKLECD-----CTEPALRDF 56
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFE-HVDICIYSLS 121
+ + + P + I+ L + F R S + ++ +W A+ E +D + +
Sbjct: 57 I-----LKNLPLQARIIESLYLRF-RRESFLFQDIKLWGGIAISHCLRELRIDFFSHYAN 110
Query: 122 PLNL--SSIFSCKTLVVLKLEGLDLNTTV-SSVDLPFLKVLHLQVLQLSQRRCLAQLLNG 178
P + S+++CK+LV LKL GL + V V LP LK L LQ + S+ L LL+
Sbjct: 111 PYVILPRSLYTCKSLVTLKLLGLGIRVDVPRDVCLPSLKTLLLQCVAYSEEDPLRLLLSC 170
Query: 179 CPVLEDL 185
CPVLEDL
Sbjct: 171 CPVLEDL 177
>At4g15060 hypothetical protein
Length = 555
Score = 95.5 bits (236), Expect = 3e-20
Identities = 76/267 (28%), Positives = 123/267 (45%), Gaps = 33/267 (12%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
D+IS LPD++L +L FLPTK AV+TSILSKRW+ LW + L++ N+ ++ R
Sbjct: 229 DKISRLPDDLLVKVLLFLPTKIAVSTSILSKRWEFLWMWLPKLEYHNTNYSASEEQRLRS 288
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLSP 122
+++ +H I+ LR+ F S P N+ W+ AV R C+
Sbjct: 289 FINLNLQLHRAPI---IESLRLKFSLGRSIKPQNIKQWIIIAVFR--------CVRD--- 334
Query: 123 LNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVL 182
K+LV+LKL+ L LP LK L L+ + L QLL+ CPVL
Sbjct: 335 ---------KSLVILKLKDQILVDVPRMAYLPSLKYLLLKRVTYKDSNSLHQLLSSCPVL 385
Query: 183 EDLKAKAIYFKKYKETGFEYKTLSKLVRADISGTSTDLFRMEVINNVDFLRIHEIYIRIP 242
++L + + + +L +L G S D E++ N L+ ++ +
Sbjct: 386 KNLVVERDEYNHDETLSITVSSLQRLTLKISRGGSFD----ELVINTPSLKYFKLTDYLG 441
Query: 243 H------DDQFTDMFHNLTHIELGYYD 263
DD ++ +F ++ +E + D
Sbjct: 442 ECETELDDDSYSYVFKDMPKLEEAHID 468
Score = 33.1 bits (74), Expect = 0.18
Identities = 24/92 (26%), Positives = 49/92 (53%), Gaps = 8/92 (8%)
Query: 200 FEYKTLSKLVRADISGTSTDLFRMEVINNVDFLRIHEIYIRIPHDDQFTD---MFHNLTH 256
F++K + L ADIS T D+++ + +V ++ + IR+ ++ +F+ L +
Sbjct: 22 FDFKDMPNLEEADISSTYPDIYKF--VRSVTSVKRLSLCIRVNAEEALYPDGIVFNQLEN 79
Query: 257 IELGYYDYNTNWIEV-VEFLNYCPKLQVLVIN 287
++L D +NW ++ V L Y P L+ L ++
Sbjct: 80 LKLCPCD--SNWSKLFVRLLKYSPNLRELEVS 109
>At5g41840 putative protein
Length = 540
Score = 94.4 bits (233), Expect = 7e-20
Identities = 69/202 (34%), Positives = 108/202 (53%), Gaps = 24/202 (11%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DRIS LPD ++C+ILSFLPTK+A +T++L+KRWKPL V L+FD++ + + +++
Sbjct: 14 DRISGLPDALICHILSFLPTKEAASTTVLAKRWKPLLAFVPNLNFDDSIYFHPRARRNKY 73
Query: 63 VQSVHRVIHTPDFLQPIQ-----KLRMSFVSRHSEVPSN-VNVWVNAAVQRGGFEHVDIC 116
+S + D + +Q L+ V V + V W+ ++RG +DI
Sbjct: 74 SKSYESFMSFVDSVLALQAKTKTPLKRFHVKCEDVVDQSWVLEWIPKVLKRG---VLDID 130
Query: 117 IYSLSPLNL----------SSIFSCKTLVVLKL---EGLDLNTTVSSVDLPFLKVLHLQV 163
++ S N S IF KTLV LK+ +G+ ++ V LP LK LHL
Sbjct: 131 LHITSSRNYCENSSFYSLPSKIFVSKTLVRLKIQFQDGVHIDVE-GGVSLPKLKTLHLDY 189
Query: 164 LQLSQRRCLAQLLNGCPVLEDL 185
++ + L +LL+GC LE+L
Sbjct: 190 FKI-ETSMLNKLLSGCHALEEL 210
>At1g69630 hypothetical protein
Length = 451
Score = 94.0 bits (232), Expect = 9e-20
Identities = 69/186 (37%), Positives = 98/186 (52%), Gaps = 8/186 (4%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
D IS LPD +LC +L LPTK V TS+LS+RW+ LW+ V LD D +F+ + S F
Sbjct: 18 DWISKLPDCLLCEVLLNLPTKDVVKTSVLSRRWRNLWKHVPGLDLDNTDFQEFNTFLS-F 76
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLSP 122
V S ++ FLQ L+ + + W+N V R +H+D+ S
Sbjct: 77 VDS-FLDFNSESFLQKF-ILKYDCDDEYDPDIFLIGRWINTIVTR-KVQHIDVLDDSYGS 133
Query: 123 LNL---SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGC 179
+ SSI++C++LV LKL GL L + V LP LKV+ L + + + L L+ C
Sbjct: 134 WEVQLPSSIYTCESLVSLKLCGLTL-ASPEFVSLPSLKVMDLIITKFADDMGLETLITKC 192
Query: 180 PVLEDL 185
PVLE L
Sbjct: 193 PVLESL 198
>At3g52680 unknown protein
Length = 456
Score = 93.2 bits (230), Expect = 1e-19
Identities = 65/194 (33%), Positives = 95/194 (48%), Gaps = 20/194 (10%)
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
+DRIS LPD +L ILS LPT VATS+LSK+W+ LW+ V L+FD ++ S
Sbjct: 20 KDRISELPDGLLLKILSSLPTNIVVATSVLSKQWRSLWKLVPNLEFDSDDYESEHYTFSE 79
Query: 62 FVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLS 121
V P ++ R+ FV+ + P ++ +WV A R H+ +
Sbjct: 80 IVCKSFLSHKAP----VLESFRLKFVNFN---PVDIGLWVGIAFSR----HLRELVLDFY 128
Query: 122 PLNL---------SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCL 172
P L SS+ +C TL LKL L S V + L+ LHL+ ++ +
Sbjct: 129 PAELGKGVTFTFPSSLCTCNTLETLKLVLCILVDIPSPVLMKSLRTLHLEFVRYKDESSV 188
Query: 173 AQLLNGCPVLEDLK 186
LL+GCP LE+L+
Sbjct: 189 RNLLSGCPGLEELR 202
>At4g14096 F-box protein - like
Length = 468
Score = 92.8 bits (229), Expect = 2e-19
Identities = 74/211 (35%), Positives = 105/211 (49%), Gaps = 10/211 (4%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEA---NFRRTKGWH 59
D ISSLP+ I C+ILSFLPTK+A +TS+LSK+W+ L+ V LD DE+ N
Sbjct: 8 DIISSLPEAISCHILSFLPTKEAASTSVLSKKWRYLFAFVPNLDLDESVYLNPENETEVS 67
Query: 60 SRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYS 119
S F+ V RV+ P+ K + P + W+N ++R G +D+ +Y
Sbjct: 68 SSFMDFVDRVLALQG-NSPLHKFSLKI--GDGVEPDRIIPWINNVLER-GVSDLDLHVYM 123
Query: 120 LSPLNL-SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRC-LAQLLN 177
+ S +F KTLV LKL L V LP LK L++ + L +LL+
Sbjct: 124 ETEFVFPSEMFLSKTLVRLKLMLYPL-LEFEDVYLPKLKTLYIDSCYFEKYGIGLTKLLS 182
Query: 178 GCPVLEDLKAKAIYFKKYKETGFEYKTLSKL 208
GCP+LEDL I + + TL +L
Sbjct: 183 GCPILEDLVLDDIPWCTWDFASVSVPTLKRL 213
>At5g56420 unknown protein
Length = 422
Score = 91.7 bits (226), Expect = 4e-19
Identities = 64/184 (34%), Positives = 89/184 (47%), Gaps = 5/184 (2%)
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DR+S LPD+ L ILS+LPTK + TS+LSKRW+ LW V L++D T S+F
Sbjct: 6 DRLSQLPDDFLLQILSWLPTKDVLVTSLLSKRWRFLWTLVPRLNYDLRLHDNTCPRFSQF 65
Query: 63 VQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIYSLSP 122
V + P ++ L + S +V VWV V R E P
Sbjct: 66 VDRSLLLHKAP----TLESLNIKIGSICFTAEKDVGVWVRIGVDRFVRELSVSYCSGEEP 121
Query: 123 LNL-SSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPV 181
+ L +F+C TL VLKLE + L V LK LHL ++ + L ++++ C
Sbjct: 122 IRLPKCLFTCSTLAVLKLENITLEDASCYVCFQSLKTLHLLDVKYLDDQSLPRIISSCSS 181
Query: 182 LEDL 185
LEDL
Sbjct: 182 LEDL 185
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,314,146
Number of Sequences: 26719
Number of extensions: 257253
Number of successful extensions: 1461
Number of sequences better than 10.0: 239
Number of HSP's better than 10.0 without gapping: 209
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 1007
Number of HSP's gapped (non-prelim): 313
length of query: 289
length of database: 11,318,596
effective HSP length: 98
effective length of query: 191
effective length of database: 8,700,134
effective search space: 1661725594
effective search space used: 1661725594
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0487.10


