

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0438.5
(434 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
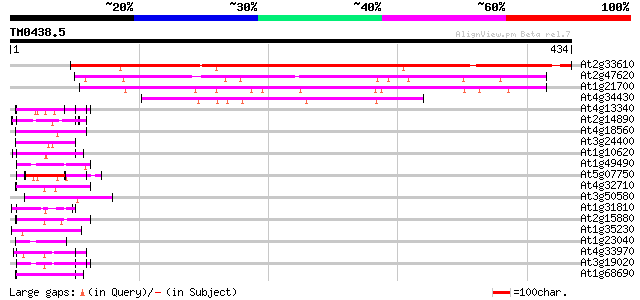
Score E
Sequences producing significant alignments: (bits) Value
At2g33610 putative SWI/SNF complex subunit SW13 416 e-116
At2g47620 SWI/SNF family like transcription activator 183 2e-46
At1g21700 putative transcriptional regulatory protein 135 6e-32
At4g34430 putative protein 107 1e-23
At4g13340 extensin-like protein 49 4e-06
At2g14890 arabinogalactan-protein AGP9 46 4e-05
At4g18560 pherophorin - like protein 46 5e-05
At3g24400 protein kinase, putative 45 6e-05
At1g10620 putative serine/threonine protein kinase emb|CAA18823.1 45 6e-05
At1g49490 hypothetical protein 45 8e-05
At5g07750 putative protein 45 1e-04
At4g32710 putative protein kinase 45 1e-04
At3g50580 proline-rich protein 45 1e-04
At1g31810 hypothetical protein 45 1e-04
At2g15880 unknown protein 44 2e-04
At1g35230 arabinogalactan-protein AGP5 44 2e-04
At1g23040 predicted GPI-anchored protein 44 2e-04
At4g33970 extensin-like protein 44 2e-04
At3g19020 hypothetical protein 44 2e-04
At1g68690 protein kinase, putative 44 2e-04
>At2g33610 putative SWI/SNF complex subunit SW13
Length = 469
Score = 416 bits (1070), Expect = e-116
Identities = 220/400 (55%), Positives = 292/400 (73%), Gaps = 23/400 (5%)
Query: 48 SATPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDS--SSKSPRVYKYYRNSIIKYFRY 105
S++ D + I VPS+S WFSW I+ CEVR LPEFFDS SSK+P+ Y Y RNSIIK +R
Sbjct: 40 SSSSDIDNIHVPSYSSWFSWTDINDCEVRSLPEFFDSRSSSKNPKFYLYLRNSIIKQYRD 99
Query: 106 NPNRKITFTDVRKMLVGDVGSIRRVFEFLEAWGLINYHPSSSFSKPFKWDDKD---TKVD 162
+ RKI+FTDVR+ LV DV SIRRVF+FL++WGLINY+ S+S +KP KW++K+ + D
Sbjct: 100 DHPRKISFTDVRRTLVSDVVSIRRVFDFLDSWGLINYNSSAS-AKPLKWEEKEAGKSAGD 158
Query: 163 SAEPPPPPVRETAKRVCSGCKALCTIACFVCDKYDLTLCARCYVRGNYRVGVSSSDFKRV 222
+A P V+ETAKR C+GCKA+C+IACF CDKYDLTLCARCYVR NYRVG++SS+FKRV
Sbjct: 159 AASEPATTVKETAKRNCNGCKAICSIACFACDKYDLTLCARCYVRSNYRVGINSSEFKRV 218
Query: 223 EISEETKTEWTEKETLNLVEAISHYGDDWKRVSHQVVGRTEKECVAHFLKLPFGEQFLGS 282
EISEE+K EW++KE L L+EA+ HYGDDWK+V+ V+GRTEK+CV+ F+KLPFGEQF+
Sbjct: 219 EISEESKPEWSDKEILLLLEAVMHYGDDWKKVASHVIGRTEKDCVSQFVKLPFGEQFVKE 278
Query: 283 AVSDDGCELKQHADESETVAS-------AESNKRMRLTPLADASNPIMAQAAFLSALAGP 335
+ S+DG E+ +S+ S + NKR++LTPLADASNPIMAQAAFLSALAG
Sbjct: 279 SDSEDGLEMFDQIKDSDIPESEGIDKDGSSPNKRIKLTPLADASNPIMAQAAFLSALAGT 338
Query: 336 EVAQAAAQAALTTLSDV-YKSTRINYRSLPKNTLQQDAGVASNGGNNSDSLQGARLNASL 394
VA+AAA+AA+ LSDV Y++ ++ ++ +QDA AS+G + + A +A
Sbjct: 339 NVAEAAARAAVRALSDVDYEAD----KNASRDPNRQDANAASSGETTRNESERAWADAKS 394
Query: 395 QLEKDESDVEKAISEIIEVQVCPILNCLMVFTLVKFKNLD 434
+EK+E +VE AI E +EV++ I + +V F+ LD
Sbjct: 395 LIEKEEHEVEGAIKETVEVEMKKIRD-----RIVHFEKLD 429
>At2g47620 SWI/SNF family like transcription activator
Length = 512
Score = 183 bits (464), Expect = 2e-46
Identities = 134/458 (29%), Positives = 211/458 (45%), Gaps = 101/458 (22%)
Query: 51 PDANVIL--VPSHSRWFSWDSIHQCEVRHLPEFFDSSS--KSPRVYKYYRNSIIKYFRYN 106
P A + L +P+ S WF WD IH+ E R EFF SS ++P+VYK YR+ II FR +
Sbjct: 6 PSAEIELYTIPAQSSWFLWDDIHEIERREFAEFFTESSITRTPKVYKEYRDFIINKFRED 65
Query: 107 PNRKITFTDVRKMLVGDVGSIRRVFEFLEAWGLINYHPSSSFSKPFKWDDKDTKVDSAE- 165
R++TFT VRK LVGDV +++VF FLE WGLIN FS K +D VD+A+
Sbjct: 66 TCRRLTFTSVRKFLVGDVNLLQKVFLFLEKWGLIN------FSSSLKKNDHLLSVDNAKI 119
Query: 166 --------------------PPPPPVRETAKR------------------------VCSG 181
PP V E + VC+
Sbjct: 120 EQGTPAGIRVTATPNSLRPITAPPLVEERVETGIKVPPLTSYSDVFSDLKKPDHVLVCAH 179
Query: 182 CKALCTIACFVCDKYDLTLCARCYVRGNYRVGVSSSDFKRVEISEETKTEWTEKETLNLV 241
C C + +K + +C +C+ GNY ++ DFK I WTE+E L L+
Sbjct: 180 CGERCDSPFYQHNKGIVNICEKCFKNGNYGENNTADDFKL--IGNSAAAVWTEEEILLLL 237
Query: 242 EAISHYGDDWKRVSHQVVGRTEKECVAHFLKLPFGEQFLGSA---------VSDDGCEL- 291
E++ +GDDW+ +S V ++ +C++ ++LPFGE +GSA D+ E
Sbjct: 238 ESVLKHGDDWELISQSVSTKSRLDCISKLIELPFGEFLMGSASGRLNPSILTEDENTEQV 297
Query: 292 ---KQHADESETVASAESN--------KRMRLTPLADASNPIMAQAAFLSALAGPEVAQA 340
Q +E+ET E KR R+ +++ + +M Q A +++ GP VA A
Sbjct: 298 QTDGQEHEETETREEKEDRVNEDEPPAKRKRVALISEGDSSLMKQVAAMASKVGPSVATA 357
Query: 341 AAQAALTTL------------SDVYKSTRINYRSLPKNTLQQDAGVASNG---------- 378
AA+AAL L +D Y + ++ + K+T ++ +G
Sbjct: 358 AAKAALAALCDEASCPKEIFDTDDYSNFTVDRANGEKDTDMEEQQEEKDGPQGLPVALRI 417
Query: 379 -GNNSDSLQGARLNASLQLEKDESDVEKAISEIIEVQV 415
+ + +L A A + +++E ++E+ + +IE Q+
Sbjct: 418 RASVATALGAAAAQAKILADQEEREMEQLAATVIEQQL 455
>At1g21700 putative transcriptional regulatory protein
Length = 807
Score = 135 bits (339), Expect = 6e-32
Identities = 113/443 (25%), Positives = 194/443 (43%), Gaps = 82/443 (18%)
Query: 55 VILVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKS--PRVYKYYRNSIIKYFRYNPNRKIT 112
V ++P HS WF+ +++ + E + +P+FF S + P Y +RN+I+ + NP + +T
Sbjct: 175 VHVLPMHSDWFAPNTVDRLERQVVPQFFSGKSPNHTPESYMEFRNAIVSKYVENPEKTLT 234
Query: 113 FTDVRKMLVG-DVGSIRRVFEFLEAWGLINY------HPSSSFSKPFKWDDKD------- 158
+D + ++ G D+ RVF FL+ WG+INY HP +D +
Sbjct: 235 ISDCQGLVDGVDIEDFARVFRFLDHWGIINYCATAQSHPGPLRDVSDVREDTNGEVNVPS 294
Query: 159 ---TKVDSA-EPPPPPVRETAKRVCSGCKAL---------------CTIACFVCD----- 194
T +DS + P R V S +L C C C
Sbjct: 295 AALTSIDSLIKFDKPNCRHKGGEVYSSLPSLDGDSPDLDIRIREHLCDSHCNHCSRPLPT 354
Query: 195 -------KYDLTLCARCYVRGNYRVGVSSSDFKRVE----ISEETKTEWTEKETLNLVEA 243
K D+ LC C+ G + VG S DF RV+ ++ WT++ETL L+EA
Sbjct: 355 VYFQSQKKGDILLCCDCFHHGRFVVGHSCLDFVRVDPMKFYGDQDGDNWTDQETLLLLEA 414
Query: 244 ISHYGDDWKRVSHQVVGRTEKECVAHFLKLPFGEQFLG----SAVSD-----DGCELKQH 294
+ Y ++W +++ V +++ +C+ HFL+LP + L S V++ +G + K
Sbjct: 415 VELYNENWVQIADHVGSKSKAQCILHFLRLPVEDGLLDNVEVSGVTNTENPTNGYDHKGT 474
Query: 295 ADESETVASAESNKRMRL-TPLADASNPIMAQAAFLSALAGPEVAQAAAQAALTTLS--D 351
+ +E + P + NP+MA AFL++ GP VA + A +L+ LS D
Sbjct: 475 DSNGDLPGYSEQGSDTEIKLPFVKSPNPVMALVAFLASAVGPRVAASCAHESLSVLSEDD 534
Query: 352 VYKSTRINYRS---LPKNTLQQDAGVASNGGNNSDS----------------LQGARLNA 392
KS + + L QQD ++ N +++ L A A
Sbjct: 535 RMKSEGMQGKEASLLDGENQQQDGAHKTSSQNGAEAQTPLPQDKVMAAFRAGLSAAATKA 594
Query: 393 SLQLEKDESDVEKAISEIIEVQV 415
L + +E ++++ + I+ Q+
Sbjct: 595 KLFADHEEREIQRLSANIVNHQL 617
>At4g34430 putative protein
Length = 827
Score = 107 bits (267), Expect = 1e-23
Identities = 80/277 (28%), Positives = 125/277 (44%), Gaps = 59/277 (21%)
Query: 103 FRYNPNRKITFTDVRKMLVGDVGSIRRVFEFLEAWGLINYHP------SSSFSKPFKWDD 156
F NPN +I D+ ++ VGD + + V EFL+ WGLIN+HP S+ S D
Sbjct: 4 FHSNPNIQIELKDLTELEVGDSEAKQEVMEFLDYWGLINFHPFPPTDTGSTASDHDDLGD 63
Query: 157 KDT--------KVDSAEPP-------------------PPPVRETAKRV-------CSGC 182
K++ +VD A PP P E K+ C+ C
Sbjct: 64 KESLLNSLYRFQVDEACPPLVHKPRFTAQATPSGLFPDPMAADELLKQEGPAVEYHCNSC 123
Query: 183 KALCTIACFVCDKY-DLTLCARCYVRGNYRVGVSSSDFKRVEISEET---KTEWTEKETL 238
A C+ + C K D LC C+ G + +SSSDF +E +E +WT++ETL
Sbjct: 124 SADCSRKRYHCPKQADFDLCTECFNSGKFSSDMSSSDFILMEPAEAPGVGSGKWTDQETL 183
Query: 239 NLVEAISHYGDDWKRVSHQVVGRTEKECVAHFLKLPFGEQFLGS--------------AV 284
L+EA+ + ++W ++ V +T+ +C+ HFL++P + FL AV
Sbjct: 184 LLLEALEIFKENWNEIAEHVATKTKAQCMLHFLQMPIEDAFLDQIDYKDPISKDTTDLAV 243
Query: 285 S-DDGCELKQHADESETVASAESNKRMRLTPLADASN 320
S DD LK +E+E + ++ M+ P + N
Sbjct: 244 SKDDNSVLKDAPEEAENKKRVDEDETMKEVPEPEDGN 280
Score = 38.5 bits (88), Expect = 0.007
Identities = 31/105 (29%), Positives = 50/105 (47%), Gaps = 7/105 (6%)
Query: 285 SDDGCELKQHADESETVASAESNKRMRLTPLADASNPIMAQAAFLSALAGPEVAQAAAQA 344
+D+ LK + E V + + + AD NP+M AAFL LAG +VA A+A+A
Sbjct: 322 ADENIALKALTEAFEDVGHSSTPEAS--FSFADLGNPVMGLAAFLVRLAGSDVATASARA 379
Query: 345 ALTTL---SDVYKSTRINY--RSLPKNTLQQDAGVASNGGNNSDS 384
++ +L S + +TR Y P N +++ N D+
Sbjct: 380 SIKSLHSNSGMLLATRHCYILEDPPDNKKDPTKSKSADAEGNDDN 424
>At4g13340 extensin-like protein
Length = 760
Score = 49.3 bits (116), Expect = 4e-06
Identities = 21/46 (45%), Positives = 23/46 (49%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P PPP S PPPP PPPP++ P P P PP P S P
Sbjct: 439 PPPPPPPPPVYSPPPPPPPPPPPPVYSPPPPPPPPPPPPPVYSPPP 484
Score = 46.2 bits (108), Expect = 4e-05
Identities = 20/42 (47%), Positives = 23/42 (54%), Gaps = 4/42 (9%)
Query: 5 SPTTATEPPPSSSA----GTPPPPLPPPPLHQPQPAKPDAPP 42
+P T T PPP S PPPP PPPP++ P P P PP
Sbjct: 419 TPPTLTSPPPPSPPPPVYSPPPPPPPPPPVYSPPPPPPPPPP 460
Score = 44.3 bits (103), Expect = 1e-04
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 1/47 (2%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKP-DAPPSEPKLSATP 51
P+ PPP S PPPP PPPP++ P P +PP P + TP
Sbjct: 485 PSPPPPPPPVYSPPPPPPPPPPPPVYSPPPPPVYSSPPPPPSPAPTP 531
Score = 42.7 bits (99), Expect = 4e-04
Identities = 23/68 (33%), Positives = 28/68 (40%), Gaps = 11/68 (16%)
Query: 6 PTTATEPPPSSSAGTPPPPLP-----------PPPLHQPQPAKPDAPPSEPKLSATPDAN 54
P + PPP + PPPP P PPP H P P + PP EP ++P
Sbjct: 507 PPVYSPPPPPVYSSPPPPPSPAPTPVYCTRPPPPPPHSPPPPQFSPPPPEPYYYSSPPPP 566
Query: 55 VILVPSHS 62
P HS
Sbjct: 567 HSSPPPHS 574
Score = 42.4 bits (98), Expect = 5e-04
Identities = 21/54 (38%), Positives = 25/54 (45%), Gaps = 2/54 (3%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P PPP S P PP PPPP++ P P P PP P + + P V P
Sbjct: 470 PPPPPPPPPVYSPPPPSPPPPPPPVYSPPP--PPPPPPPPPVYSPPPPPVYSSP 521
Score = 42.4 bits (98), Expect = 5e-04
Identities = 20/49 (40%), Positives = 23/49 (46%), Gaps = 2/49 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQP--QPAKPDAPPSEPKLSATP 51
SP PPP +PPPP PPPP P P P PP P + + P
Sbjct: 450 SPPPPPPPPPPPPVYSPPPPPPPPPPPPPVYSPPPPSPPPPPPPVYSPP 498
Score = 42.4 bits (98), Expect = 5e-04
Identities = 18/43 (41%), Positives = 23/43 (52%), Gaps = 6/43 (13%)
Query: 6 PTTATEPPPSSSAGT------PPPPLPPPPLHQPQPAKPDAPP 42
P+ + PPP+ T PPPP PPPP++ P P P PP
Sbjct: 405 PSLPSPPPPAPIFSTPPTLTSPPPPSPPPPVYSPPPPPPPPPP 447
Score = 39.3 bits (90), Expect = 0.004
Identities = 19/54 (35%), Positives = 22/54 (40%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P + PP +PPPP PP H P P PP P LS P + P
Sbjct: 548 PQFSPPPPEPYYYSSPPPPHSSPPPHSPPPPHSPPPPIYPYLSPPPPPTPVSSP 601
Score = 37.7 bits (86), Expect = 0.012
Identities = 18/46 (39%), Positives = 21/46 (45%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPPS PP PPPP P P +PP P S+ P
Sbjct: 477 PPVYSPPPPSPPPPPPPVYSPPPPPPPPPPPPVYSPPPPPVYSSPP 522
Score = 37.0 bits (84), Expect = 0.021
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 10/50 (20%)
Query: 12 PPPSSSAGTPPPPL------PPPPLHQPQPAKPD----APPSEPKLSATP 51
PPP + PPPP+ PPPP+H P P+ +PP P ++P
Sbjct: 643 PPPVYYSSPPPPPVYYSSPPPPPPVHYSSPPPPEVHYHSPPPSPVHYSSP 692
Score = 35.8 bits (81), Expect = 0.047
Identities = 20/55 (36%), Positives = 26/55 (46%), Gaps = 3/55 (5%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPD-APPSEPKLSATPDANVILVP 59
P + PPP S +PPP PPPP P P P +PP P ++P + P
Sbjct: 557 PYYYSSPPPPHS--SPPPHSPPPPHSPPPPIYPYLSPPPPPTPVSSPPPTPVYSP 609
Score = 35.8 bits (81), Expect = 0.047
Identities = 30/97 (30%), Positives = 38/97 (38%), Gaps = 13/97 (13%)
Query: 9 ATEPPPSSSAG--TPPPP-LPPPPLH-----QPQPAKPDAPPSEPKLSATPDANVILVPS 60
++ PPP SS +PPPP PPPP++ P P +PP P S P I P
Sbjct: 561 SSPPPPHSSPPPHSPPPPHSPPPPIYPYLSPPPPPTPVSSPPPTPVYSPPPPPPCIEPPP 620
Query: 61 HSRWFSWDSIHQCEVRHL-----PEFFDSSSKSPRVY 92
+ V H P + SS P VY
Sbjct: 621 PPPCIEYSPPPPPPVVHYSSPPPPPVYYSSPPPPPVY 657
Score = 35.4 bits (80), Expect = 0.062
Identities = 20/57 (35%), Positives = 26/57 (45%), Gaps = 11/57 (19%)
Query: 6 PTTATEPPPSSSAGTPPP-----PLPPPPL--HQPQPAKP----DAPPSEPKLSATP 51
PT + PPP+ PPP P PPPP + P P P +PP P ++P
Sbjct: 595 PTPVSSPPPTPVYSPPPPPPCIEPPPPPPCIEYSPPPPPPVVHYSSPPPPPVYYSSP 651
Score = 34.7 bits (78), Expect = 0.11
Identities = 21/63 (33%), Positives = 25/63 (39%), Gaps = 6/63 (9%)
Query: 1 MATKSPTTATEPPPSSSAGTPPPPLP----PPPLHQPQPAKPDAPPSEPKLSATPDANVI 56
++ + P PPPS + PPPP P PP L P P P P P P V
Sbjct: 392 VSPRPPVVTPLPPPSLPS--PPPPAPIFSTPPTLTSPPPPSPPPPVYSPPPPPPPPPPVY 449
Query: 57 LVP 59
P
Sbjct: 450 SPP 452
Score = 34.3 bits (77), Expect = 0.14
Identities = 17/48 (35%), Positives = 21/48 (43%), Gaps = 14/48 (29%)
Query: 6 PTTATEPPPSSSAGTPPPPL-----------PPPPLHQPQPAKPDAPP 42
P + PPP PPPP+ PPPP++ P P P PP
Sbjct: 447 PVYSPPPPPPPP---PPPPVYSPPPPPPPPPPPPPVYSPPPPSPPPPP 491
Score = 33.9 bits (76), Expect = 0.18
Identities = 17/50 (34%), Positives = 21/50 (42%), Gaps = 4/50 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKP----DAPPSEPKLSATP 51
P EPPP PP PPP +H P P +PP P ++P
Sbjct: 612 PPPCIEPPPPPPCIEYSPPPPPPVVHYSSPPPPPVYYSSPPPPPVYYSSP 661
Score = 33.5 bits (75), Expect = 0.24
Identities = 17/47 (36%), Positives = 23/47 (48%), Gaps = 7/47 (14%)
Query: 6 PTTATEPPPSSSAGTPPPPL----PPPPLHQPQPAKP---DAPPSEP 45
P + PPP+ + PP P+ PPPP +P P P +PP P
Sbjct: 587 PYLSPPPPPTPVSSPPPTPVYSPPPPPPCIEPPPPPPCIEYSPPPPP 633
Score = 32.7 bits (73), Expect = 0.40
Identities = 22/66 (33%), Positives = 27/66 (40%), Gaps = 15/66 (22%)
Query: 6 PTTATEPPPSSSAGTPPPPLPP-----PP-----LHQPQP-----AKPDAPPSEPKLSAT 50
P + PPP + PPP PP PP H P P + P PPS P +
Sbjct: 645 PVYYSSPPPPPVYYSSPPPPPPVHYSSPPPPEVHYHSPPPSPVHYSSPPPPPSAPCEESP 704
Query: 51 PDANVI 56
P A V+
Sbjct: 705 PPAPVV 710
Score = 30.4 bits (67), Expect = 2.0
Identities = 20/54 (37%), Positives = 24/54 (44%), Gaps = 12/54 (22%)
Query: 5 SPTTATEPPPSSSA----GTPPPPL----PPPPL--HQPQP--AKPDAPPSEPK 46
SP + PPP SA PP P+ PPPP+ H P P PP P+
Sbjct: 685 SPVHYSSPPPPPSAPCEESPPPAPVVHHSPPPPMVHHSPPPPVIHQSPPPPSPE 738
Score = 28.5 bits (62), Expect = 7.6
Identities = 16/47 (34%), Positives = 20/47 (42%), Gaps = 2/47 (4%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKP--DAPPSEPKLSATPDANVI 56
PPPS + PPP P P + P P P P + +P VI
Sbjct: 682 PPPSPVHYSSPPPPPSAPCEESPPPAPVVHHSPPPPMVHHSPPPPVI 728
Score = 28.1 bits (61), Expect = 9.9
Identities = 14/37 (37%), Positives = 15/37 (39%)
Query: 15 SSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
S S G P PP P P+ P PP P S P
Sbjct: 385 SFSCGRSVSPRPPVVTPLPPPSLPSPPPPAPIFSTPP 421
>At2g14890 arabinogalactan-protein AGP9
Length = 191
Score = 46.2 bits (108), Expect = 4e-05
Identities = 24/54 (44%), Positives = 29/54 (53%), Gaps = 4/54 (7%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P T PPP ++A PPP PPPP+ P PA P PP+ P A+P V P
Sbjct: 52 PPVTTSPPPVTTA--PPPANPPPPVSSPPPASP--PPATPPPVASPPPPVASPP 101
Score = 44.3 bits (103), Expect = 1e-04
Identities = 22/50 (44%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Query: 3 TKSPTTATEPPP-SSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
T +P A PPP SS PPP PPP+ P P PP+ P ATP
Sbjct: 62 TTAPPPANPPPPVSSPPPASPPPATPPPVASPPPPVASPPPATPPPVATP 111
Score = 43.1 bits (100), Expect = 3e-04
Identities = 25/60 (41%), Positives = 29/60 (47%), Gaps = 11/60 (18%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLH--------QPQPAKPDAPPSEPKLSATPDA 53
AT +P T T PPP A TPPP PPP+ P PA P P S P ++ P A
Sbjct: 29 ATPAPPTPTTPPP---AATPPPVSAPPPVTTSPPPVTTAPPPANPPPPVSSPPPASPPPA 85
Score = 41.6 bits (96), Expect = 9e-04
Identities = 23/53 (43%), Positives = 27/53 (50%), Gaps = 5/53 (9%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDAN 54
A SP TAT PP+ + TPPP PPP+ P P P P +A P AN
Sbjct: 22 APTSPPTATPAPPTPT--TPPPAATPPPVSAPPPVTTSPP---PVTTAPPPAN 69
Score = 40.0 bits (92), Expect = 0.003
Identities = 24/58 (41%), Positives = 28/58 (47%), Gaps = 6/58 (10%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPL-PPPPLHQPQPAKPD---APPSEPKLSATPDANV 55
A P ++ PP S TPPP PPPP+ P PA P PP P A+P A V
Sbjct: 68 ANPPPPVSSPPPASPPPATPPPVASPPPPVASPPPATPPPVATPPPAP--LASPPAQV 123
Score = 38.5 bits (88), Expect = 0.007
Identities = 22/53 (41%), Positives = 26/53 (48%), Gaps = 5/53 (9%)
Query: 1 MATKSPTTATEPP--PSSSAGTPPPPLPPPPLHQPQPA---KPDAPPSEPKLS 48
+A+ P A+ PP P A PP PL PP P PA KPD+P P S
Sbjct: 90 VASPPPPVASPPPATPPPVATPPPAPLASPPAQVPAPAPTTKPDSPSPSPSSS 142
Score = 35.4 bits (80), Expect = 0.062
Identities = 21/55 (38%), Positives = 28/55 (50%), Gaps = 7/55 (12%)
Query: 5 SPTTATEPP---PSSSAGTPPPPLPPPPLHQPQPAKPDAPPSE---PKLSATPDA 53
SP AT PP P +PPP PPP+ P PA +PP++ P + PD+
Sbjct: 81 SPPPATPPPVASPPPPVASPPP-ATPPPVATPPPAPLASPPAQVPAPAPTTKPDS 134
Score = 31.2 bits (69), Expect = 1.2
Identities = 17/45 (37%), Positives = 20/45 (43%), Gaps = 4/45 (8%)
Query: 11 EPPPSSSAGTPPPPLP--PPPLHQPQP--AKPDAPPSEPKLSATP 51
+ P S TP PP P PPP P P A P S P ++ P
Sbjct: 21 QAPTSPPTATPAPPTPTTPPPAATPPPVSAPPPVTTSPPPVTTAP 65
>At4g18560 pherophorin - like protein
Length = 637
Score = 45.8 bits (107), Expect = 5e-05
Identities = 25/62 (40%), Positives = 27/62 (43%), Gaps = 7/62 (11%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQP-------AKPDAPPSEPKLSATPDANVIL 57
S +PPP S PPPP PPP L QP P P PP PK + A V
Sbjct: 279 STENRADPPPQKSIPPPPPPPPPPLLQQPPPPPSVSKAPPPPPPPPPPKSLSIASAKVRR 338
Query: 58 VP 59
VP
Sbjct: 339 VP 340
Score = 30.0 bits (66), Expect = 2.6
Identities = 19/44 (43%), Positives = 22/44 (49%), Gaps = 2/44 (4%)
Query: 4 KSPTTA-TEPPPSSSAGTPPPPLP-PPPLHQPQPAKPDAPPSEP 45
KSP+T T+ PS PPPP P PP KP PS+P
Sbjct: 14 KSPSTKKTKDMPSPLPLPPPPPPPLKPPSSGSATTKPPINPSKP 57
>At3g24400 protein kinase, putative
Length = 694
Score = 45.4 bits (106), Expect = 6e-05
Identities = 26/55 (47%), Positives = 29/55 (52%), Gaps = 8/55 (14%)
Query: 5 SPTTATEPPPSSSAGTPP-PPLPPP--PLH-----QPQPAKPDAPPSEPKLSATP 51
SPTT + PPPS S +PP P PPP PL P PA P PP P +TP
Sbjct: 127 SPTTPSPPPPSPSIPSPPLTPSPPPSSPLRPSSPPPPSPATPSTPPRSPPPPSTP 181
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/59 (38%), Positives = 26/59 (43%), Gaps = 5/59 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAP----PSEPKLSATPDANVILVPS 60
P T PP S +PPPPL P PL P P P P P+ P TP + PS
Sbjct: 59 PPPTTVPPIPPSTPSPPPPLTPSPL-PPSPTTPSPPLTPSPTTPSPPLTPSPPPAITPS 116
Score = 40.8 bits (94), Expect = 0.001
Identities = 16/35 (45%), Positives = 20/35 (56%)
Query: 9 ATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPS 43
++ PPP + PP PLP PP QP P P PP+
Sbjct: 2 SSAPPPGGTPSPPPQPLPIPPPPQPLPVTPPPPPT 36
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/56 (41%), Positives = 26/56 (46%), Gaps = 6/56 (10%)
Query: 5 SPTTATEPPPSSSA--GTPPPPLPP----PPLHQPQPAKPDAPPSEPKLSATPDAN 54
SP PPPSS +PPPP P PP P P+ P PP LS P A+
Sbjct: 142 SPPLTPSPPPSSPLRPSSPPPPSPATPSTPPRSPPPPSTPTPPPRVGSLSPPPPAS 197
Score = 37.4 bits (85), Expect = 0.016
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 3/55 (5%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
P T + PPS + +PPPP P P P P PPS P ++P PS
Sbjct: 118 PLTPSPLPPSPTTPSPPPPSPSIP---SPPLTPSPPPSSPLRPSSPPPPSPATPS 169
Score = 36.6 bits (83), Expect = 0.028
Identities = 23/60 (38%), Positives = 27/60 (44%), Gaps = 14/60 (23%)
Query: 5 SPTTATEPPPSSSAGTPPPPL---------PPPPLH----QPQPAKPDAPPSEPKLSATP 51
SPTT + PP + S TP PPL P PPL P P P PP P + + P
Sbjct: 86 SPTTPS-PPLTPSPTTPSPPLTPSPPPAITPSPPLTPSPLPPSPTTPSPPPPSPSIPSPP 144
Score = 34.7 bits (78), Expect = 0.11
Identities = 20/51 (39%), Positives = 23/51 (44%), Gaps = 2/51 (3%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPS--EPKLSATPDANVILVPS 60
PPP +A P P PPPP P P PP+ P +TP L PS
Sbjct: 31 PPPPPTALPPALPPPPPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTPS 81
Score = 34.7 bits (78), Expect = 0.11
Identities = 20/56 (35%), Positives = 22/56 (38%), Gaps = 5/56 (8%)
Query: 1 MATKSPTTATEPPPSSSAGTPPPP-----LPPPPLHQPQPAKPDAPPSEPKLSATP 51
M++ P T PP PPPP PPPP PA P PP A P
Sbjct: 1 MSSAPPPGGTPSPPPQPLPIPPPPQPLPVTPPPPPTALPPALPPPPPPTALPPALP 56
Score = 34.3 bits (77), Expect = 0.14
Identities = 15/40 (37%), Positives = 17/40 (42%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP PP PPPP P P P P L+ +P
Sbjct: 43 PPPPPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTPSP 82
Score = 34.3 bits (77), Expect = 0.14
Identities = 19/45 (42%), Positives = 20/45 (44%), Gaps = 5/45 (11%)
Query: 6 PTTATEPPPSSSAGTPPPPLP---PPPLHQPQPAK--PDAPPSEP 45
P T PP + PPPP P PP L P P P PPS P
Sbjct: 28 PVTPPPPPTALPPALPPPPPPTALPPALPPPPPPTTVPPIPPSTP 72
Score = 33.5 bits (75), Expect = 0.24
Identities = 18/50 (36%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Query: 5 SPTTATEPPPSSSAGTPPP---PLPPPPLHQPQPAKPDAPPSEPKLSATP 51
+P+T PP S TPPP L PPP P + + PS S+ P
Sbjct: 167 TPSTPPRSPPPPSTPTPPPRVGSLSPPPPASPSGGRSPSTPSTTPGSSPP 216
Score = 33.1 bits (74), Expect = 0.31
Identities = 23/62 (37%), Positives = 27/62 (43%), Gaps = 12/62 (19%)
Query: 5 SPTTATEPPPSSSAG------TPPPP---LPPP---PLHQPQPAKPDAPPSEPKLSATPD 52
SP + PPP S A +PPPP PPP L P PA P S S TP
Sbjct: 153 SPLRPSSPPPPSPATPSTPPRSPPPPSTPTPPPRVGSLSPPPPASPSGGRSPSTPSTTPG 212
Query: 53 AN 54
++
Sbjct: 213 SS 214
Score = 32.3 bits (72), Expect = 0.52
Identities = 20/55 (36%), Positives = 23/55 (41%), Gaps = 6/55 (10%)
Query: 3 TKSPTTATEPPPSSS---AGTPPPP---LPPPPLHQPQPAKPDAPPSEPKLSATP 51
T P PPP ++ A PPPP +PP P P P P P P TP
Sbjct: 36 TALPPALPPPPPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTPSPLPPSPTTP 90
Score = 31.6 bits (70), Expect = 0.89
Identities = 19/52 (36%), Positives = 26/52 (49%), Gaps = 9/52 (17%)
Query: 6 PTTATEPPPSS------SAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P+T + PPP + S TP PPL P P P P +PP P ++ +P
Sbjct: 69 PSTPSPPPPLTPSPLPPSPTTPSPPLTPSPT-TPSPPLTPSPP--PAITPSP 117
Score = 30.8 bits (68), Expect = 1.5
Identities = 17/53 (32%), Positives = 20/53 (37%), Gaps = 3/53 (5%)
Query: 2 ATKSPTTATEPPPSSSAGTPP---PPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A P T PP+ PP PP+PP P P P P P + P
Sbjct: 41 ALPPPPPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTPSPLPPSPTTPSPP 93
Score = 28.9 bits (63), Expect = 5.8
Identities = 16/47 (34%), Positives = 22/47 (46%), Gaps = 1/47 (2%)
Query: 6 PTTATEPPPSSSAGTPP-PPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P T + PPS + +PP P P P P+ P A P L+ +P
Sbjct: 77 PLTPSPLPPSPTTPSPPLTPSPTTPSPPLTPSPPPAITPSPPLTPSP 123
>At1g10620 putative serine/threonine protein kinase emb|CAA18823.1
Length = 718
Score = 45.4 bits (106), Expect = 6e-05
Identities = 21/54 (38%), Positives = 28/54 (50%), Gaps = 2/54 (3%)
Query: 6 PTTATEPPPSSSAGTPPPPLPP--PPLHQPQPAKPDAPPSEPKLSATPDANVIL 57
P TA +PPP+ T PPP PP PP P P P P+ + T D+ V++
Sbjct: 62 PATAAQPPPNQPPNTTPPPTPPSSPPPSITPPPSPPQPQPPPQSTPTGDSPVVI 115
Score = 44.3 bits (103), Expect = 1e-04
Identities = 22/52 (42%), Positives = 24/52 (45%), Gaps = 3/52 (5%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLP---PPPLHQPQPAKPDAPPSEPKLSATP 51
T+ P T+ PPS T PPP PPP P P PPS P S TP
Sbjct: 41 TQPPATSPPSPPSPDTQTSPPPATAAQPPPNQPPNTTPPPTPPSSPPPSITP 92
Score = 32.3 bits (72), Expect = 0.52
Identities = 23/58 (39%), Positives = 27/58 (45%), Gaps = 12/58 (20%)
Query: 3 TKSPTTATEPPPS-SSAGTPPPPLPPP--------PLHQPQPAKPDAPPSEPKLSATP 51
T PT + PPPS + +PP P PPP P+ P P KP PP P L P
Sbjct: 77 TPPPTPPSSPPPSITPPPSPPQPQPPPQSTPTGDSPVVIPFP-KPQLPP--PSLFPPP 131
Score = 29.3 bits (64), Expect = 4.4
Identities = 16/45 (35%), Positives = 22/45 (48%), Gaps = 8/45 (17%)
Query: 10 TEPPPSSSAGTPPPP---LPPPPLHQPQPAKPDAPPSEPKLSATP 51
T PP++S +PP P PPP QP PP++P + P
Sbjct: 40 TTQPPATSPPSPPSPDTQTSPPPATAAQP-----PPNQPPNTTPP 79
Score = 28.5 bits (62), Expect = 7.6
Identities = 22/66 (33%), Positives = 27/66 (40%), Gaps = 11/66 (16%)
Query: 5 SPTTATEPPPSSSAGTPP-------PPLPPPPLHQPQP---AKPDAPPSEPKLSATPDAN 54
SP PP S+ G P P LPPP L P PD P++ + P N
Sbjct: 95 SPPQPQPPPQSTPTGDSPVVIPFPKPQLPPPSLFPPPSLVNQLPDPRPNDNNI-LEPINN 153
Query: 55 VILVPS 60
I +PS
Sbjct: 154 PISLPS 159
>At1g49490 hypothetical protein
Length = 847
Score = 45.1 bits (105), Expect = 8e-05
Identities = 26/59 (44%), Positives = 31/59 (52%), Gaps = 6/59 (10%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVIL--VPSHS 62
P + PPP +S PPPP PPPP+H P P +PP P S P + V PSHS
Sbjct: 570 PHVYSPPPPVAS---PPPPSPPPPVHSPPPPPVFSPP-PPVFSPPPPSPVYSPPPPSHS 624
Score = 41.2 bits (95), Expect = 0.001
Identities = 23/70 (32%), Positives = 30/70 (42%), Gaps = 13/70 (18%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSE-----------PKLSATPD 52
P PPP S +PPPP PPPP++ P P PP+ P + TP
Sbjct: 603 PPPVFSPPPPSPVYSPPPPSHSPPPPVYSPPPPTFSPPPTHNTNQPPMGAPTPTQAPTPS 662
Query: 53 ANVILVPSHS 62
+ VP+ S
Sbjct: 663 SETTQVPTPS 672
Score = 40.0 bits (92), Expect = 0.003
Identities = 21/51 (41%), Positives = 24/51 (46%), Gaps = 5/51 (9%)
Query: 6 PTTATEPPPSSSA---GTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATP 51
P A+ PPPS PPPP+ PPPP+ P P P P P S P
Sbjct: 577 PPVASPPPPSPPPPVHSPPPPPVFSPPPPVFSPPPPSPVYSPPPPSHSPPP 627
Score = 37.4 bits (85), Expect = 0.016
Identities = 20/54 (37%), Positives = 23/54 (42%), Gaps = 7/54 (12%)
Query: 5 SPTTATEPPPSSSAGTPPPP---LPPPPLHQPQPAKP----DAPPSEPKLSATP 51
SP PP +PPPP PPPP+ P P P +PP P S P
Sbjct: 551 SPPPPVHSPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPPPVFSPPP 604
Score = 37.0 bits (84), Expect = 0.021
Identities = 20/51 (39%), Positives = 22/51 (42%), Gaps = 4/51 (7%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDAN 54
P PPP + PP P+ PPPP H P P PP P S P N
Sbjct: 596 PPPVFSPPPPVFSPPPPSPVYSPPPPSHSPPPPVYSPPP--PTFSPPPTHN 644
Score = 36.6 bits (83), Expect = 0.028
Identities = 22/63 (34%), Positives = 28/63 (43%), Gaps = 7/63 (11%)
Query: 7 TTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPP-----SEPKLSATPDANVILVP 59
T P P + +PP P+ PPPP+H P P +PP S P A+P P
Sbjct: 531 TRVPPPQPPMPSPSPPSPIYSPPPPVHSPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPP 590
Query: 60 SHS 62
HS
Sbjct: 591 VHS 593
Score = 34.7 bits (78), Expect = 0.11
Identities = 22/67 (32%), Positives = 26/67 (37%), Gaps = 12/67 (17%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQ--------PQPAKPDAPPSE----PKLSATPD 52
SP + PP PPP PPP H P P + P SE P S+ D
Sbjct: 617 SPPPPSHSPPPPVYSPPPPTFSPPPTHNTNQPPMGAPTPTQAPTPSSETTQVPTPSSESD 676
Query: 53 ANVILVP 59
+ IL P
Sbjct: 677 QSQILSP 683
Score = 34.7 bits (78), Expect = 0.11
Identities = 21/63 (33%), Positives = 30/63 (47%), Gaps = 7/63 (11%)
Query: 4 KSPTTATE----PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
++PTT++E P PSS + P P P L +P P S+P S TP + + P
Sbjct: 726 QAPTTSSETSQVPTPSSESNQSPSQAPTPIL---EPVHAPTPNSKPVQSPTPSSEPVSSP 782
Query: 60 SHS 62
S
Sbjct: 783 EQS 785
Score = 33.5 bits (75), Expect = 0.24
Identities = 20/60 (33%), Positives = 27/60 (44%), Gaps = 8/60 (13%)
Query: 6 PTTATEPPPSSSAGTPPPPL----PPPPLHQPQP--AKPDAPPSEPKLSATPDANVILVP 59
P+ PPP +PPPP+ PPP ++ P P A P P P + + P V P
Sbjct: 546 PSPIYSPPPP--VHSPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPPPVFSPP 603
Score = 33.5 bits (75), Expect = 0.24
Identities = 18/49 (36%), Positives = 19/49 (38%), Gaps = 2/49 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPP--LPPPPLHQPQPAKPDAPPSEPKLSATP 51
SP PP +PPPP PPPP P P P P S P
Sbjct: 586 SPPPPVHSPPPPPVFSPPPPVFSPPPPSPVYSPPPPSHSPPPPVYSPPP 634
Score = 32.7 bits (73), Expect = 0.40
Identities = 19/60 (31%), Positives = 25/60 (41%), Gaps = 12/60 (20%)
Query: 12 PPPSSSAGTPPPPL--PPPPLHQ----------PQPAKPDAPPSEPKLSATPDANVILVP 59
P P S +PPPP+ PPPP++ P P PPS P +P + P
Sbjct: 543 PSPPSPIYSPPPPVHSPPPPVYSSPPPPHVYSPPPPVASPPPPSPPPPVHSPPPPPVFSP 602
Score = 32.0 bits (71), Expect = 0.68
Identities = 18/50 (36%), Positives = 19/50 (38%), Gaps = 1/50 (2%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
PPP T PP P PP+ P P P P P S P P H
Sbjct: 523 PPPPKVEDTRVPP-PQPPMPSPSPPSPIYSPPPPVHSPPPPVYSSPPPPH 571
Score = 31.6 bits (70), Expect = 0.89
Identities = 15/36 (41%), Positives = 20/36 (54%), Gaps = 1/36 (2%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAP-PSEPK 46
P PS S+ P P P +PQP K ++P P EP+
Sbjct: 414 PKPSDSSKPETPKTPEQPSPKPQPPKHESPKPEEPE 449
Score = 30.8 bits (68), Expect = 1.5
Identities = 18/55 (32%), Positives = 25/55 (44%), Gaps = 6/55 (10%)
Query: 3 TKSPTTATEPPPSSSAGTP-----PPPLPPPPLHQ-PQPAKPDAPPSEPKLSATP 51
+K +P SS TP P P P PP H+ P+P +P+ PK +P
Sbjct: 407 SKPEPVMPKPSDSSKPETPKTPEQPSPKPQPPKHESPKPEEPENKHELPKQKESP 461
Score = 30.0 bits (66), Expect = 2.6
Identities = 21/73 (28%), Positives = 25/73 (33%), Gaps = 16/73 (21%)
Query: 6 PTTATEPPPSSSAGTPPPPL----------PPPP------LHQPQPAKPDAPPSEPKLSA 49
PT PP + TP P PPPP + PQP P P P S
Sbjct: 493 PTKPVSPPNEAQGPTPDDPYDASPVKNRRSPPPPKVEDTRVPPPQPPMPSPSPPSPIYSP 552
Query: 50 TPDANVILVPSHS 62
P + P +S
Sbjct: 553 PPPVHSPPPPVYS 565
Score = 30.0 bits (66), Expect = 2.6
Identities = 17/57 (29%), Positives = 22/57 (37%), Gaps = 7/57 (12%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPP-------PLHQPQPAKPDAPPSEPKLSATPDAN 54
+PT P PSS P P P+ P P + P SEP TP ++
Sbjct: 652 APTPTQAPTPSSETTQVPTPSSESDQSQILSPVQAPTPVQSSTPSSEPTQVPTPSSS 708
Score = 28.9 bits (63), Expect = 5.8
Identities = 13/37 (35%), Positives = 22/37 (59%), Gaps = 1/37 (2%)
Query: 16 SSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPD 52
S+ G+ P P PP +P+P+KP+ +P S+ P+
Sbjct: 388 SNGGSSPSP-NPPRTSEPKPSKPEPVMPKPSDSSKPE 423
>At5g07750 putative protein
Length = 1289
Score = 44.7 bits (104), Expect = 1e-04
Identities = 22/51 (43%), Positives = 27/51 (52%), Gaps = 3/51 (5%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP---SEPKLSATPDANVILVP 59
PPP S G+PPPP PPPP + P P PP S P P ++V +P
Sbjct: 960 PPPPPSYGSPPPPPPPPPGYGSPPPPPPPPPSYGSPPPPPPPPFSHVSSIP 1010
Score = 43.5 bits (101), Expect = 2e-04
Identities = 17/31 (54%), Positives = 19/31 (60%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
PPP S G+PPPP PPPP + P P PP
Sbjct: 947 PPPPPSYGSPPPPPPPPPSYGSPPPPPPPPP 977
Score = 42.4 bits (98), Expect = 5e-04
Identities = 28/77 (36%), Positives = 35/77 (45%), Gaps = 13/77 (16%)
Query: 6 PTTATEPPPSSS----AGT--PPPPLPPPPLHQPQP-----AKPDAPPSEPKLSATPDAN 54
P PPP SS +GT PPPP PPPP +P P PP P S P++
Sbjct: 668 PPPPPPPPPFSSERPNSGTVLPPPPPPPPPFSSERPNSGTVLPPPPPPPLPFSSERPNSG 727
Query: 55 VILVPSHSRWFSWDSIH 71
+L P S W S++
Sbjct: 728 TVLPPPPSP--PWKSVY 742
Score = 42.4 bits (98), Expect = 5e-04
Identities = 17/31 (54%), Positives = 19/31 (60%)
Query: 13 PPSSSAGTPPPPLPPPPLHQPQPAKPDAPPS 43
PP S G+PPPP PPPP + P P PPS
Sbjct: 935 PPPPSYGSPPPPPPPPPSYGSPPPPPPPPPS 965
Score = 42.0 bits (97), Expect = 7e-04
Identities = 17/31 (54%), Positives = 18/31 (57%), Gaps = 1/31 (3%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
PPP G PPPP PPPP+H P P PP
Sbjct: 1014 PPPPMHGGAPPPP-PPPPMHGGAPPPPPPPP 1043
Score = 42.0 bits (97), Expect = 7e-04
Identities = 17/31 (54%), Positives = 18/31 (57%), Gaps = 1/31 (3%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
PPP G PPPP PPPP+H P P PP
Sbjct: 1027 PPPPMHGGAPPPP-PPPPMHGGAPPPPPPPP 1056
Score = 41.2 bits (95), Expect = 0.001
Identities = 16/31 (51%), Positives = 18/31 (57%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
PPP S + PPP PPPP+H P P PP
Sbjct: 1000 PPPFSHVSSIPPPPPPPPMHGGAPPPPPPPP 1030
Score = 39.3 bits (90), Expect = 0.004
Identities = 16/30 (53%), Positives = 17/30 (56%), Gaps = 1/30 (3%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAP 41
PPP G PPPP PPPP+H P P P
Sbjct: 1040 PPPPMHGGAPPPP-PPPPMHGGAPPPPPPP 1068
Score = 38.1 bits (87), Expect = 0.010
Identities = 16/31 (51%), Positives = 17/31 (54%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
PPP S G+PPPP PPP H P PP
Sbjct: 986 PPPPPSYGSPPPPPPPPFSHVSSIPPPPPPP 1016
Score = 38.1 bits (87), Expect = 0.010
Identities = 21/55 (38%), Positives = 24/55 (43%), Gaps = 8/55 (14%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPP----PPLHQPQP----AKPDAPPSEPKLSATP 51
SP PPPS + PPPP PP PP P P + P PP P + P
Sbjct: 942 SPPPPPPPPPSYGSPPPPPPPPPSYGSPPPPPPPPPGYGSPPPPPPPPPSYGSPP 996
Score = 38.1 bits (87), Expect = 0.010
Identities = 19/45 (42%), Positives = 21/45 (46%), Gaps = 4/45 (8%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPP----PPLHQPQPAKPDAPPSEP 45
SP PPPS + PPPP PP PP P P +PP P
Sbjct: 955 SPPPPPPPPPSYGSPPPPPPPPPGYGSPPPPPPPPPSYGSPPPPP 999
Score = 38.1 bits (87), Expect = 0.010
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 5/55 (9%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
S A+ PPP PPPPLP +Q P PP P S P++ +L P
Sbjct: 641 SQAVASPPPPP-----PPPPLPTYSHYQTSQLPPPPPPPPPFSSERPNSGTVLPP 690
Score = 37.7 bits (86), Expect = 0.012
Identities = 18/40 (45%), Positives = 20/40 (50%), Gaps = 8/40 (20%)
Query: 12 PPPSSSAGTPPPPLPPPPLH------QPQPAKPDAPPSEP 45
PPP G PPPP PPP+H P P + APP P
Sbjct: 1101 PPPPMRGGAPPPP--PPPMHGGAPPPPPPPMRGGAPPPPP 1138
Score = 37.7 bits (86), Expect = 0.012
Identities = 18/40 (45%), Positives = 19/40 (47%), Gaps = 1/40 (2%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP G+PPPP PPPP P P PP S P
Sbjct: 973 PPPPPGYGSPPPP-PPPPPSYGSPPPPPPPPFSHVSSIPP 1011
Score = 37.0 bits (84), Expect = 0.021
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 4/50 (8%)
Query: 16 SSAGTPPPPLPPPPLHQPQPAKPD----APPSEPKLSATPDANVILVPSH 61
S++ +PPPP PPPP K + PP P S P +V ++P H
Sbjct: 855 STSSSPPPPPPPPPFSPLNTTKANDYILPPPPLPYTSIAPSPSVKILPLH 904
Score = 36.6 bits (83), Expect = 0.028
Identities = 18/40 (45%), Positives = 18/40 (45%), Gaps = 6/40 (15%)
Query: 12 PPPSSSAGTPPPPLP------PPPLHQPQPAKPDAPPSEP 45
PPP G PPPP P PPP P P APP P
Sbjct: 1113 PPPPMHGGAPPPPPPPMRGGAPPPPPPPGGRGPGAPPPPP 1152
Score = 36.6 bits (83), Expect = 0.028
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 10/49 (20%)
Query: 5 SPTTATEPPPSSSAGTPPPP--------LPPPPLHQPQPAKPDAPPSEP 45
SP PPPS + PPPP +PPPP P P APP P
Sbjct: 981 SPPPPPPPPPSYGSPPPPPPPPFSHVSSIPPPP--PPPPMHGGAPPPPP 1027
Score = 36.2 bits (82), Expect = 0.036
Identities = 17/38 (44%), Positives = 18/38 (46%), Gaps = 4/38 (10%)
Query: 12 PPPSSSAGTPPPPLPP----PPLHQPQPAKPDAPPSEP 45
PPP G PPPP PP P P P + APP P
Sbjct: 1077 PPPPMRGGAPPPPPPPMRGGAPPPPPPPMRGGAPPPPP 1114
Score = 35.4 bits (80), Expect = 0.062
Identities = 21/59 (35%), Positives = 24/59 (40%), Gaps = 12/59 (20%)
Query: 5 SPTTATEPPP--------SSSAGTPPPPL----PPPPLHQPQPAKPDAPPSEPKLSATP 51
SP T PPP + S +PPPP PPPP P P PP P +P
Sbjct: 911 SPPVKTAPPPPPPPPFSNAHSVLSPPPPSYGSPPPPPPPPPSYGSPPPPPPPPPSYGSP 969
Score = 34.7 bits (78), Expect = 0.11
Identities = 28/104 (26%), Positives = 36/104 (33%), Gaps = 17/104 (16%)
Query: 3 TKSPTTATEPPPSSSAG------------TPPPPLPPPPL-----HQPQPAKPDAPPSEP 45
T SPT PP S G +PPPP PPPP + P PP P
Sbjct: 759 TSSPTPPPPPPAYYSVGQKSSDLQTSQLPSPPPPPPPPPFASVRRNSETLLPPPPPPPPP 818
Query: 46 KLSATPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKSP 89
++ + L+P W S++ SSS P
Sbjct: 819 PFASVRRNSETLLPPPPPPPPWKSLYASTFETHEACSTSSSPPP 862
Score = 34.7 bits (78), Expect = 0.11
Identities = 23/71 (32%), Positives = 28/71 (39%), Gaps = 15/71 (21%)
Query: 2 ATKSPTTATEPPP--------SSSAGTPPPPLPPPPLHQPQP-----AKPDAPPSEPKLS 48
A SP PPP +S PPP PPPP +P P PP P S
Sbjct: 643 AVASPPPPPPPPPLPTYSHYQTSQLPPPPP--PPPPFSSERPNSGTVLPPPPPPPPPFSS 700
Query: 49 ATPDANVILVP 59
P++ +L P
Sbjct: 701 ERPNSGTVLPP 711
Score = 34.7 bits (78), Expect = 0.11
Identities = 19/40 (47%), Positives = 20/40 (49%), Gaps = 8/40 (20%)
Query: 12 PPPSSSAGTPPPPLPPPPLH---QPQPAKP---DAPPSEP 45
PPP G PPPP PPP+ QP P P APP P
Sbjct: 1053 PPPPMHGGAPPPP--PPPMFGGAQPPPPPPMRGGAPPPPP 1090
Score = 34.3 bits (77), Expect = 0.14
Identities = 20/56 (35%), Positives = 22/56 (38%), Gaps = 9/56 (16%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPP----PPLHQPQP-----AKPDAPPSEPKLSATP 51
SP PPP + PPPP PP PP P P + P PP P P
Sbjct: 968 SPPPPPPPPPGYGSPPPPPPPPPSYGSPPPPPPPPFSHVSSIPPPPPPPPMHGGAP 1023
Score = 33.5 bits (75), Expect = 0.24
Identities = 17/40 (42%), Positives = 18/40 (44%), Gaps = 8/40 (20%)
Query: 12 PPPSSSAGTPPPPLPPPPLH------QPQPAKPDAPPSEP 45
PPP G PPPP PPP+ P P APP P
Sbjct: 1089 PPPPMRGGAPPPP--PPPMRGGAPPPPPPPMHGGAPPPPP 1126
Score = 33.1 bits (74), Expect = 0.31
Identities = 15/34 (44%), Positives = 15/34 (44%)
Query: 9 ATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
A PPP G PPP PPP P P PP
Sbjct: 1121 APPPPPPPMRGGAPPPPPPPGGRGPGAPPPPPPP 1154
Score = 32.7 bits (73), Expect = 0.40
Identities = 16/40 (40%), Positives = 18/40 (45%), Gaps = 2/40 (5%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP G PPPP PPP + A P PP + P
Sbjct: 1125 PPPPMRGGAPPPP--PPPGGRGPGAPPPPPPPGGRAPGPP 1162
Score = 32.7 bits (73), Expect = 0.40
Identities = 18/39 (46%), Positives = 19/39 (48%), Gaps = 5/39 (12%)
Query: 12 PPPS-SSAGTPPPPLP----PPPLHQPQPAKPDAPPSEP 45
PPPS S PPPP P PPP P P+ PP P
Sbjct: 936 PPPSYGSPPPPPPPPPSYGSPPPPPPPPPSYGSPPPPPP 974
Score = 32.7 bits (73), Expect = 0.40
Identities = 16/38 (42%), Positives = 17/38 (44%), Gaps = 4/38 (10%)
Query: 12 PPPSSSAGTPPPPLPP----PPLHQPQPAKPDAPPSEP 45
PPP G PPP PP P P P + APP P
Sbjct: 1065 PPPPMFGGAQPPPPPPMRGGAPPPPPPPMRGGAPPPPP 1102
Score = 32.0 bits (71), Expect = 0.68
Identities = 19/56 (33%), Positives = 21/56 (36%), Gaps = 10/56 (17%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPP-------LHQPQP---AKPDAPPSEPKLSATP 51
P PS T PPP PPPP L P P + P PP P + P
Sbjct: 902 PLHGISSAPSPPVKTAPPPPPPPPFSNAHSVLSPPPPSYGSPPPPPPPPPSYGSPP 957
Score = 32.0 bits (71), Expect = 0.68
Identities = 17/45 (37%), Positives = 17/45 (37%), Gaps = 7/45 (15%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKP-------DAPPSEPKLSA 49
PPP PPP PPP P P P PP P L A
Sbjct: 1138 PPPGGRGPGAPPPPPPPGGRAPGPPPPPGPRPPGGGPPPPPMLGA 1182
Score = 31.2 bits (69), Expect = 1.2
Identities = 20/51 (39%), Positives = 22/51 (42%), Gaps = 8/51 (15%)
Query: 3 TKSPTTATEPPPSSS----AGTPPPPLPPPPL----HQPQPAKPDAPPSEP 45
T P PPP SS +GT PP PPPPL +P PP P
Sbjct: 686 TVLPPPPPPPPPFSSERPNSGTVLPPPPPPPLPFSSERPNSGTVLPPPPSP 736
Score = 30.8 bits (68), Expect = 1.5
Identities = 16/46 (34%), Positives = 18/46 (38%), Gaps = 6/46 (13%)
Query: 12 PPPSSSAGTPPPP------LPPPPLHQPQPAKPDAPPSEPKLSATP 51
PPP PPPP PPPP + P PP + A P
Sbjct: 1066 PPPMFGGAQPPPPPPMRGGAPPPPPPPMRGGAPPPPPPPMRGGAPP 1111
Score = 30.8 bits (68), Expect = 1.5
Identities = 17/52 (32%), Positives = 18/52 (33%), Gaps = 5/52 (9%)
Query: 5 SPTTATEPPPSSSAGTPPPPLP-----PPPLHQPQPAKPDAPPSEPKLSATP 51
+P PP A PPPP P PPP P PP P P
Sbjct: 1035 APPPPPPPPMHGGAPPPPPPPPMHGGAPPPPPPPMFGGAQPPPPPPMRGGAP 1086
Score = 28.9 bits (63), Expect = 5.8
Identities = 14/34 (41%), Positives = 15/34 (43%), Gaps = 2/34 (5%)
Query: 9 ATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP 42
A PPP P PP PP P +P P PP
Sbjct: 1147 APPPPPPPGGRAPGPPPPPGP--RPPGGGPPPPP 1178
>At4g32710 putative protein kinase
Length = 731
Score = 44.7 bits (104), Expect = 1e-04
Identities = 29/79 (36%), Positives = 30/79 (37%), Gaps = 22/79 (27%)
Query: 6 PTTATEPPPSSSAGTPPPP-L---PPPPLHQPQ------------------PAKPDAPPS 43
P E PPS +PPPP L PPPPL P PAKP PPS
Sbjct: 64 PPLPVESPPSPPIESPPPPLLESPPPPPLESPSPPSPHVSAPSGSPPLPFLPAKPSPPPS 123
Query: 44 EPKLSATPDANVILVPSHS 62
P P N I P S
Sbjct: 124 SPPSETVPPGNTISPPPRS 142
Score = 43.9 bits (102), Expect = 2e-04
Identities = 23/63 (36%), Positives = 30/63 (47%), Gaps = 5/63 (7%)
Query: 5 SPTTATEPPPSSSAGTPPPPL-----PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
SP PPP S+ PPPL P PP+ P P ++PP P S +P + + P
Sbjct: 46 SPPPLPSPPPLSAPTASPPPLPVESPPSPPIESPPPPLLESPPPPPLESPSPPSPHVSAP 105
Query: 60 SHS 62
S S
Sbjct: 106 SGS 108
Score = 37.4 bits (85), Expect = 0.016
Identities = 23/52 (44%), Positives = 28/52 (53%), Gaps = 9/52 (17%)
Query: 5 SPTTATEPPPSSSAGTPPPPLP-PPPLHQPQPAKP----DAPPSEPKLSATP 51
+P + PPP SS PPPLP PPPL P + P ++PPS P S P
Sbjct: 34 APPLSPLPPPLSS----PPPLPSPPPLSAPTASPPPLPVESPPSPPIESPPP 81
Score = 35.4 bits (80), Expect = 0.062
Identities = 20/52 (38%), Positives = 21/52 (39%), Gaps = 5/52 (9%)
Query: 5 SPTTATEPPPSSSAGTPPPPLP-----PPPLHQPQPAKPDAPPSEPKLSATP 51
SP + PP S T PP P PPPL P P P S P S P
Sbjct: 14 SPPAMSLPPADSVPDTSSPPAPPLSPLPPPLSSPPPLPSPPPLSAPTASPPP 65
Score = 34.7 bits (78), Expect = 0.11
Identities = 22/59 (37%), Positives = 29/59 (48%), Gaps = 9/59 (15%)
Query: 2 ATKSPTTATEPPPSSSAG-TPPPPLPPPPLH----QPQPAKPD----APPSEPKLSATP 51
+T TA+ PPPS + P P PPP++ P P P +PP +P LS TP
Sbjct: 147 STPPVNTASPPPPSPPRRRSGPKPSFPPPINSSPPNPSPNTPSLPETSPPPKPPLSTTP 205
Score = 32.0 bits (71), Expect = 0.68
Identities = 22/66 (33%), Positives = 24/66 (36%), Gaps = 6/66 (9%)
Query: 3 TKSPTTATEPPPSS--SAGTPP----PPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVI 56
T P PPP S S TPP P PP P + KP PP P N
Sbjct: 129 TVPPGNTISPPPRSLPSESTPPVNTASPPPPSPPRRRSGPKPSFPPPINSSPPNPSPNTP 188
Query: 57 LVPSHS 62
+P S
Sbjct: 189 SLPETS 194
Score = 29.3 bits (64), Expect = 4.4
Identities = 17/49 (34%), Positives = 21/49 (42%), Gaps = 9/49 (18%)
Query: 12 PPPSSSAGTPPPP---LPPPPLHQPQPAKPDA------PPSEPKLSATP 51
PPPSS PP + PPP P + P PPS P+ + P
Sbjct: 120 PPPSSPPSETVPPGNTISPPPRSLPSESTPPVNTASPPPPSPPRRRSGP 168
>At3g50580 proline-rich protein
Length = 265
Score = 44.7 bits (104), Expect = 1e-04
Identities = 23/72 (31%), Positives = 31/72 (42%), Gaps = 4/72 (5%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP----DANVILVPSHSRWFSW 67
P P S PP P PPP + P+ P PP PK S P ++ P H + W
Sbjct: 120 PTPKKSPSPPPTPSLPPPAPKKSPSTPSLPPPTPKKSPPPPPSHHSSSPSNPPHHQQNPW 179
Query: 68 DSIHQCEVRHLP 79
+ I +C + P
Sbjct: 180 EHIERCMINMGP 191
Score = 34.3 bits (77), Expect = 0.14
Identities = 24/66 (36%), Positives = 30/66 (45%), Gaps = 14/66 (21%)
Query: 5 SPTTATEPPPSSSAGTPPPPLP---PPPLHQPQPAKPDAPPS--------EPKLSATPDA 53
S + +T P + S +PPPP P PPP P P K +PPS PK S +P
Sbjct: 74 SISPSTPIPSTPSTPSPPPPAPKKSPPP---PTPKKSPSPPSLTPFVPHPTPKKSPSPPP 130
Query: 54 NVILVP 59
L P
Sbjct: 131 TPSLPP 136
Score = 33.5 bits (75), Expect = 0.24
Identities = 18/53 (33%), Positives = 22/53 (40%)
Query: 7 TTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
+ + P S S TP P P P P K PP PK S +P + VP
Sbjct: 66 SNGSAPAISISPSTPIPSTPSTPSPPPPAPKKSPPPPTPKKSPSPPSLTPFVP 118
Score = 33.1 bits (74), Expect = 0.31
Identities = 16/38 (42%), Positives = 19/38 (49%), Gaps = 6/38 (15%)
Query: 4 KSPTTATEPPPSSSAGTPPPPL------PPPPLHQPQP 35
KSP+T + PPP+ PPPP PP HQ P
Sbjct: 141 KSPSTPSLPPPTPKKSPPPPPSHHSSSPSNPPHHQQNP 178
>At1g31810 hypothetical protein
Length = 1201
Score = 44.7 bits (104), Expect = 1e-04
Identities = 23/47 (48%), Positives = 24/47 (50%), Gaps = 8/47 (17%)
Query: 6 PTTATEPPPSSSAGTPPPPLP----PPPLHQPQPAKPDAPPSEPKLS 48
P T PPP PPPPLP PPPL QP P +P PP P S
Sbjct: 567 PINKTPPPPPP----PPPPLPSRSIPPPLAQPPPPRPPPPPPPPPSS 609
Score = 41.6 bits (96), Expect = 9e-04
Identities = 25/50 (50%), Positives = 27/50 (54%), Gaps = 7/50 (14%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
A K P PP S+ G PPPP PPPPL +K APP P LS TP
Sbjct: 695 APKPPAPPPLPPSSTRLGAPPPP-PPPPL-----SKTPAPP-PPPLSKTP 737
Score = 39.7 bits (91), Expect = 0.003
Identities = 21/50 (42%), Positives = 25/50 (50%), Gaps = 2/50 (4%)
Query: 4 KSPTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATP 51
K+P PPP + + PPPL PPPP P P P + S P SA P
Sbjct: 570 KTPPPPPPPPPPLPSRSIPPPLAQPPPPRPPPPPPPPPSSRSIPSPSAPP 619
Score = 39.3 bits (90), Expect = 0.004
Identities = 23/63 (36%), Positives = 28/63 (43%), Gaps = 13/63 (20%)
Query: 2 ATKSPTTATEPPPSSSAGT---------PPPPLPPPPLH----QPQPAKPDAPPSEPKLS 48
A K PPP+S +G+ PPPP PPP + PA P PPS +L
Sbjct: 653 AAKCAPPPPPPPPTSHSGSIRVGPPSTPPPPPPPPPKANISNAPKPPAPPPLPPSSTRLG 712
Query: 49 ATP 51
A P
Sbjct: 713 APP 715
Score = 38.9 bits (89), Expect = 0.006
Identities = 22/55 (40%), Positives = 24/55 (43%), Gaps = 11/55 (20%)
Query: 6 PTTATEPPPSS----SAGTPPPPLPPPP-------LHQPQPAKPDAPPSEPKLSA 49
P PPPSS S PPPP PPPP Q QP P PP ++ A
Sbjct: 599 PPPPPPPPPSSRSIPSPSAPPPPPPPPPSFGSTGNKRQAQPPPPPPPPPPTRIPA 653
Score = 37.7 bits (86), Expect = 0.012
Identities = 21/52 (40%), Positives = 23/52 (43%), Gaps = 10/52 (19%)
Query: 6 PTTATEPPP------SSSAGTPPPPLPPPPLHQP----QPAKPDAPPSEPKL 47
P PPP S S PPPP PPPPL P++P PP P L
Sbjct: 483 PPPPPPPPPLFTSTTSFSPSQPPPPPPPPPLFMSTTSFSPSQPPPPPPPPPL 534
Score = 35.4 bits (80), Expect = 0.062
Identities = 17/40 (42%), Positives = 18/40 (44%), Gaps = 3/40 (7%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEP 45
P A PPP PPPP PP P P+ P PP P
Sbjct: 589 PPLAQPPPPRPP---PPPPPPPSSRSIPSPSAPPPPPPPP 625
Score = 34.7 bits (78), Expect = 0.11
Identities = 19/51 (37%), Positives = 20/51 (38%), Gaps = 7/51 (13%)
Query: 5 SPTTATEPPPS-------SSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLS 48
+P PPPS A PPPP PPPP P PP P S
Sbjct: 617 APPPPPPPPPSFGSTGNKRQAQPPPPPPPPPPTRIPAAKCAPPPPPPPPTS 667
Score = 34.3 bits (77), Expect = 0.14
Identities = 19/49 (38%), Positives = 23/49 (46%), Gaps = 3/49 (6%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDA--PPSEPKLSATP 51
+P PPP ++ P P PPPL P + A PP P LS TP
Sbjct: 679 TPPPPPPPPPKANISNAPKPPAPPPL-PPSSTRLGAPPPPPPPPLSKTP 726
Score = 34.3 bits (77), Expect = 0.14
Identities = 16/35 (45%), Positives = 17/35 (47%), Gaps = 1/35 (2%)
Query: 12 PPPSSSAGTP-PPPLPPPPLHQPQPAKPDAPPSEP 45
PPP + P PPP PPPP P APP P
Sbjct: 588 PPPLAQPPPPRPPPPPPPPPSSRSIPSPSAPPPPP 622
Score = 33.9 bits (76), Expect = 0.18
Identities = 24/71 (33%), Positives = 29/71 (40%), Gaps = 13/71 (18%)
Query: 6 PTTATEPPPSS--SAGTPPPPLPPPPLHQ-------PQPAKPDAPPSEPKLSAT----PD 52
P PPP+ +A PPP PPPP P P PP PK + + P
Sbjct: 640 PPPPPPPPPTRIPAAKCAPPPPPPPPTSHSGSIRVGPPSTPPPPPPPPPKANISNAPKPP 699
Query: 53 ANVILVPSHSR 63
A L PS +R
Sbjct: 700 APPPLPPSSTR 710
Score = 33.5 bits (75), Expect = 0.24
Identities = 20/52 (38%), Positives = 22/52 (41%), Gaps = 11/52 (21%)
Query: 12 PPPSSSAGTPPPPLPPPPLHQPQP----------AKPDAPPSEPKLSATPDA 53
PPP SS P P PPPP P P A+P PP P + P A
Sbjct: 604 PPPPSSRSIPSPSAPPPP-PPPPPSFGSTGNKRQAQPPPPPPPPPPTRIPAA 654
Score = 33.1 bits (74), Expect = 0.31
Identities = 19/51 (37%), Positives = 22/51 (42%), Gaps = 8/51 (15%)
Query: 5 SPTTATEPPPSSSAGT---PPPPLPPPP-----LHQPQPAKPDAPPSEPKL 47
+P PPSS PPPP PPPP P++P PP P L
Sbjct: 463 NPLNLPSDPPSSGDHVTLLPPPPPPPPPPLFTSTTSFSPSQPPPPPPPPPL 513
Score = 32.7 bits (73), Expect = 0.40
Identities = 21/58 (36%), Positives = 25/58 (42%), Gaps = 15/58 (25%)
Query: 5 SPTTATEPPPS----------SSAGTPPPPLPPPPLHQP----QPAKPDAPPSEPKLS 48
SP+ PPP S + PPPP PPPPL P++P PP P S
Sbjct: 500 SPSQPPPPPPPPPLFMSTTSFSPSQPPPPP-PPPPLFTSTTSFSPSQPPPPPPLPSFS 556
Score = 32.7 bits (73), Expect = 0.40
Identities = 23/69 (33%), Positives = 28/69 (40%), Gaps = 22/69 (31%)
Query: 5 SPTTATEPPP------SSSAGTPPPPLPPPPL------------HQP----QPAKPDAPP 42
SP+ PPP S+++ +P P PPPPL HQP P P PP
Sbjct: 521 SPSQPPPPPPPPPLFTSTTSFSPSQPPPPPPLPSFSNRDPLTTLHQPINKTPPPPPPPPP 580
Query: 43 SEPKLSATP 51
P S P
Sbjct: 581 PLPSRSIPP 589
Score = 30.4 bits (67), Expect = 2.0
Identities = 18/52 (34%), Positives = 21/52 (39%), Gaps = 5/52 (9%)
Query: 5 SPTTATEPPPSSSAGTPPPPL-----PPPPLHQPQPAKPDAPPSEPKLSATP 51
+P PP S + PPPPL PPPP + PP K S P
Sbjct: 713 APPPPPPPPLSKTPAPPPPPLSKTPVPPPPPGLGRGTSSGPPPLGAKGSNAP 764
Score = 29.6 bits (65), Expect = 3.4
Identities = 18/59 (30%), Positives = 23/59 (38%), Gaps = 19/59 (32%)
Query: 6 PTTATEPPPSSSAGTPPPP------------LPPPPLHQPQPAK-------PDAPPSEP 45
P++ + P PS+ PPPP PPPP P P + P PP P
Sbjct: 607 PSSRSIPSPSAPPPPPPPPPSFGSTGNKRQAQPPPPPPPPPPTRIPAAKCAPPPPPPPP 665
Score = 29.6 bits (65), Expect = 3.4
Identities = 20/65 (30%), Positives = 23/65 (34%), Gaps = 12/65 (18%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPP-------LHQPQPAKPDAPPSEPKLSATPDAN 54
A P PP A PP PPPP + P+ P PP P P AN
Sbjct: 637 AQPPPPPPPPPPTRIPAAKCAPPPPPPPPTSHSGSIRVGPPSTPPPPPPPP-----PKAN 691
Query: 55 VILVP 59
+ P
Sbjct: 692 ISNAP 696
>At2g15880 unknown protein
Length = 727
Score = 43.9 bits (102), Expect = 2e-04
Identities = 24/63 (38%), Positives = 29/63 (45%), Gaps = 6/63 (9%)
Query: 6 PTTATEPPPSSSAGTPPPPL----PPPPLHQPQPAKP--DAPPSEPKLSATPDANVILVP 59
P + PPPS PPPP+ PPPP++ P P P PP P S P + P
Sbjct: 496 PPVHSPPPPSPIHSPPPPPVYSPPPPPPVYSPPPPPPVYSPPPPPPVHSPPPPVHSPPPP 555
Query: 60 SHS 62
HS
Sbjct: 556 VHS 558
Score = 43.1 bits (100), Expect = 3e-04
Identities = 23/60 (38%), Positives = 26/60 (43%), Gaps = 2/60 (3%)
Query: 5 SPTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
SP PP +PPPP+ PPPP+H P P PP P S P P HS
Sbjct: 579 SPPPPVYSPPPPPVHSPPPPVHSPPPPVHSPPPPVYSPPPPPPVHSPPPPVFSPPPPVHS 638
Score = 42.0 bits (97), Expect = 7e-04
Identities = 23/59 (38%), Positives = 28/59 (46%), Gaps = 4/59 (6%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P + PPP + PPPP+ PPPP+H P P PP P S P + P HS
Sbjct: 523 PVYSPPPPPPVYSPPPPPPVHSPPPPVHSPPPPVHSPPP--PVHSPPPPVHSPPPPVHS 579
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/61 (37%), Positives = 27/61 (43%), Gaps = 3/61 (4%)
Query: 5 SPTTATEPPPSSSAGTPPPPL---PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
SP + PPP S PPP PPPP++ P P P P P S P + P H
Sbjct: 505 SPIHSPPPPPVYSPPPPPPVYSPPPPPPVYSPPPPPPVHSPPPPVHSPPPPVHSPPPPVH 564
Query: 62 S 62
S
Sbjct: 565 S 565
Score = 40.4 bits (93), Expect = 0.002
Identities = 17/39 (43%), Positives = 20/39 (50%), Gaps = 2/39 (5%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPP 42
P PPP +PPPP+ PPPP+H P P PP
Sbjct: 610 PPPVYSPPPPPPVHSPPPPVFSPPPPVHSPPPPVYSPPP 648
Score = 39.7 bits (91), Expect = 0.003
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 5/59 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P + PPP S PPPP+ PPPP++ P P P P P S P + P +S
Sbjct: 590 PPVHSPPPPVHS---PPPPVHSPPPPVYSPPPPPPVHSPPPPVFSPPPPVHSPPPPVYS 645
Score = 38.5 bits (88), Expect = 0.007
Identities = 17/40 (42%), Positives = 19/40 (47%), Gaps = 2/40 (5%)
Query: 5 SPTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPP 42
SP PP PPPP+ PPPP+H P P PP
Sbjct: 572 SPPPPVHSPPPPVYSPPPPPVHSPPPPVHSPPPPVHSPPP 611
Score = 38.5 bits (88), Expect = 0.007
Identities = 17/35 (48%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Query: 12 PPPSSSAGTPPPP-LPPPPLHQPQPAKPDAPPSEP 45
PPP+SS T PP P P+H+PQP K P++P
Sbjct: 447 PPPASSPPTSPPVHSTPSPVHKPQPPKESPQPNDP 481
Score = 38.5 bits (88), Expect = 0.007
Identities = 19/45 (42%), Positives = 24/45 (53%), Gaps = 5/45 (11%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLS 48
P + PPP S PPPP+ PPPP++ P P +PP P S
Sbjct: 627 PPVFSPPPPVHS---PPPPVYSPPPPVYSPPPPPVKSPPPPPVYS 668
Score = 38.1 bits (87), Expect = 0.010
Identities = 24/65 (36%), Positives = 29/65 (43%), Gaps = 11/65 (16%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPD------APPSEPKLSATPDANVIL 57
P + PPP S PPPP+ PPPP+H P P +PP P S P +
Sbjct: 547 PPVHSPPPPVHS---PPPPVHSPPPPVHSPPPPVHSPPPPVYSPPPPPVHSPPPPVHSPP 603
Query: 58 VPSHS 62
P HS
Sbjct: 604 PPVHS 608
Score = 38.1 bits (87), Expect = 0.010
Identities = 20/55 (36%), Positives = 24/55 (43%), Gaps = 7/55 (12%)
Query: 4 KSPTTATEPPPSSSAGTPPPPLP-----PPPLHQPQPAKP--DAPPSEPKLSATP 51
+SP PP +PPPP P PPP++ P P P PP P S P
Sbjct: 484 QSPVKFRRSPPPPPVHSPPPPSPIHSPPPPPVYSPPPPPPVYSPPPPPPVYSPPP 538
Score = 37.7 bits (86), Expect = 0.012
Identities = 22/58 (37%), Positives = 29/58 (49%), Gaps = 7/58 (12%)
Query: 6 PTTATEPPPSSSAGTPPPPL---PPPPLHQPQPAKPDAPP-SEPKLSATPDANVILVP 59
P + PPP S PPPP+ PPPP+ P P +PP PK+S+ P + P
Sbjct: 634 PPVHSPPPPVYS---PPPPVYSPPPPPVKSPPPPPVYSPPLLPPKMSSPPTQTPVNSP 688
Score = 35.8 bits (81), Expect = 0.047
Identities = 19/48 (39%), Positives = 23/48 (47%), Gaps = 5/48 (10%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP S PPPP+ PPPP++ P P PP K P
Sbjct: 620 PPVHSPPPPVFS---PPPPVHSPPPPVYSPPPPVYSPPPPPVKSPPPP 664
Score = 35.8 bits (81), Expect = 0.047
Identities = 23/61 (37%), Positives = 27/61 (43%), Gaps = 6/61 (9%)
Query: 6 PTTATEPPPSSSAGTPPPP--L--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
P + PPP + PPPP PPPP+H P P PP P S P + P H
Sbjct: 514 PVYSPPPPPPVYSPPPPPPVYSPPPPPPVHSPPPPVHSPPP--PVHSPPPPVHSPPPPVH 571
Query: 62 S 62
S
Sbjct: 572 S 572
Score = 35.4 bits (80), Expect = 0.062
Identities = 23/68 (33%), Positives = 28/68 (40%), Gaps = 10/68 (14%)
Query: 2 ATKSPTTATEPPPSSSAGTPP---PPL------PPPPLHQPQPAKP-DAPPSEPKLSATP 51
+T SP +PP S P P+ PPPP+H P P P +PP P S P
Sbjct: 461 STPSPVHKPQPPKESPQPNDPYDQSPVKFRRSPPPPPVHSPPPPSPIHSPPPPPVYSPPP 520
Query: 52 DANVILVP 59
V P
Sbjct: 521 PPPVYSPP 528
Score = 34.3 bits (77), Expect = 0.14
Identities = 15/33 (45%), Positives = 20/33 (60%), Gaps = 3/33 (9%)
Query: 16 SSAGTP---PPPLPPPPLHQPQPAKPDAPPSEP 45
SS TP P P+P P+H+PQP K P++P
Sbjct: 389 SSQATPSKSPSPVPTRPVHKPQPPKESPQPNDP 421
Score = 34.3 bits (77), Expect = 0.14
Identities = 16/40 (40%), Positives = 17/40 (42%), Gaps = 2/40 (5%)
Query: 5 SPTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPP 42
SP PP PPP PPPP+H P P PP
Sbjct: 565 SPPPPVHSPPPPVHSPPPPVYSPPPPPVHSPPPPVHSPPP 604
Score = 32.7 bits (73), Expect = 0.40
Identities = 16/54 (29%), Positives = 22/54 (40%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P + PPP S PP + PP P + P PS+ + P I+ P
Sbjct: 657 PVKSPPPPPVYSPPLLPPKMSSPPTQTPVNSPPPRTPSQTVEAPPPSEEFIIPP 710
Score = 32.7 bits (73), Expect = 0.40
Identities = 15/39 (38%), Positives = 18/39 (45%), Gaps = 1/39 (2%)
Query: 5 SPTTATEPPPSSSAGTPPPP-LPPPPLHQPQPAKPDAPP 42
SP PP PPP PPPP++ P P +PP
Sbjct: 558 SPPPPVHSPPPPVHSPPPPVHSPPPPVYSPPPPPVHSPP 596
Score = 32.3 bits (72), Expect = 0.52
Identities = 20/62 (32%), Positives = 26/62 (41%), Gaps = 11/62 (17%)
Query: 5 SPTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKP---------DAPPSEPKLSATPDA 53
SP A+ PP S + P P+ P PP PQP P +PP P S P +
Sbjct: 446 SPPPASSPPTSPPVHSTPSPVHKPQPPKESPQPNDPYDQSPVKFRRSPPPPPVHSPPPPS 505
Query: 54 NV 55
+
Sbjct: 506 PI 507
Score = 32.0 bits (71), Expect = 0.68
Identities = 15/46 (32%), Positives = 23/46 (49%), Gaps = 4/46 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP + PPP PP L P K +PP++ +++ P
Sbjct: 648 PPVYSPPPPPVKSPPPPPVYSPPLL----PPKMSSPPTQTPVNSPP 689
Score = 29.6 bits (65), Expect = 3.4
Identities = 16/48 (33%), Positives = 22/48 (45%), Gaps = 4/48 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPP--LHQPQPAKPDAPPSEPKLSATP 51
P P +PPPP P +H P PA +PP+ P + +TP
Sbjct: 418 PNDPYNQSPVKFRRSPPPPQQPHHHVVHSPPPAS--SPPTSPPVHSTP 463
>At1g35230 arabinogalactan-protein AGP5
Length = 133
Score = 43.9 bits (102), Expect = 2e-04
Identities = 25/58 (43%), Positives = 29/58 (49%), Gaps = 4/58 (6%)
Query: 2 ATKSPTT---ATEPPPSSSAGTPPP-PLPPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
AT +P+ AT P PS SA PP P PP+ QP P APP+ S P NV
Sbjct: 35 ATPTPSQSPRATAPAPSPSANPPPSAPTTAPPVSQPPTESPPAPPTSTSPSGAPGTNV 92
Score = 31.6 bits (70), Expect = 0.89
Identities = 20/60 (33%), Positives = 27/60 (44%), Gaps = 5/60 (8%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPP---PPLHQPQPAKP--DAPPSEPKLSATPDANVI 56
A P+ T PP S T PP PP P P P +A P++ LS +P+A +
Sbjct: 54 ANPPPSAPTTAPPVSQPPTESPPAPPTSTSPSGAPGTNVPSGEAGPAQSPLSGSPNAAAV 113
Score = 29.6 bits (65), Expect = 3.4
Identities = 21/59 (35%), Positives = 24/59 (40%), Gaps = 3/59 (5%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP-SEPKLSATPDANVILVPS 60
T SP AT P S T P P P + P A APP S+P + P PS
Sbjct: 29 TISPLPATPTPSQSPRATAPAP--SPSANPPPSAPTTAPPVSQPPTESPPAPPTSTSPS 85
>At1g23040 predicted GPI-anchored protein
Length = 144
Score = 43.9 bits (102), Expect = 2e-04
Identities = 20/40 (50%), Positives = 22/40 (55%), Gaps = 4/40 (10%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSE 44
SP+ PPP S PPPP PPPP P PA P PP +
Sbjct: 52 SPSCIQNPPPPS----PPPPSPPPPACPPPPALPPPPPKK 87
Score = 35.0 bits (79), Expect = 0.081
Identities = 19/54 (35%), Positives = 25/54 (46%), Gaps = 2/54 (3%)
Query: 4 KSPTTATEPPPSSS--AGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
++P + PPPS A PPP LPPPP + P PP+ P N+
Sbjct: 57 QNPPPPSPPPPSPPPPACPPPPALPPPPPKKVSSYCPPPPPANFLYITGPPGNL 110
>At4g33970 extensin-like protein
Length = 699
Score = 43.5 bits (101), Expect = 2e-04
Identities = 21/53 (39%), Positives = 27/53 (50%), Gaps = 6/53 (11%)
Query: 4 KSPTTATEPPPSSSAGTPPPPL-----PPPPLHQPQPAKPDAPPSEPKLSATP 51
+SP T PP + +PPPP+ PPPP+H P P +PP P S P
Sbjct: 509 QSPVTKRRSPPPAPVNSPPPPVYSPPPPPPPVHSP-PPPVHSPPPPPVYSPPP 560
Score = 43.1 bits (100), Expect = 3e-04
Identities = 24/61 (39%), Positives = 30/61 (48%), Gaps = 5/61 (8%)
Query: 4 KSPTTA---TEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILV 58
+SP A + PPP S PPPP+ PPPP+H P P +PP P +P V
Sbjct: 516 RSPPPAPVNSPPPPVYSPPPPPPPVHSPPPPVHSPPPPPVYSPPPPPPPVHSPPPPVFSP 575
Query: 59 P 59
P
Sbjct: 576 P 576
Score = 42.0 bits (97), Expect = 7e-04
Identities = 20/55 (36%), Positives = 24/55 (43%), Gaps = 9/55 (16%)
Query: 6 PTTATEPPPSSSAGTPPPPL---------PPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP +PPPP+ PPPP+H P P PP P S P
Sbjct: 553 PPVYSPPPPPPPVHSPPPPVFSPPPPVYSPPPPVHSPPPPVHSPPPPAPVHSPPP 607
Score = 41.2 bits (95), Expect = 0.001
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 2/57 (3%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
P PPP + PP P+ PPPP+H P P P P P S P + +V S
Sbjct: 583 PPPVHSPPPPVHSPPPPAPVHSPPPPVHSPPPPPPVYSPPPPVFSPPPSQSPPVVYS 639
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/56 (41%), Positives = 26/56 (46%), Gaps = 5/56 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P + PPP S PPPP+ PPPP+H P P P P P S P V P
Sbjct: 570 PPVFSPPPPVYS---PPPPVHSPPPPVHSPPPPAPVHSPPPPVHSPPPPPPVYSPP 622
Score = 40.8 bits (94), Expect = 0.001
Identities = 19/53 (35%), Positives = 27/53 (50%), Gaps = 7/53 (13%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKP-----DAPPSEPKLSATP 51
P PPP + PPPP+ PPPP+ P P++ PP PK+++ P
Sbjct: 599 PAPVHSPPPPVHSPPPPPPVYSPPPPVFSPPPSQSPPVVYSPPPRPPKINSPP 651
Score = 38.1 bits (87), Expect = 0.010
Identities = 20/50 (40%), Positives = 23/50 (46%), Gaps = 7/50 (14%)
Query: 6 PTTATEPPPSSSAGTPPPPL----PPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP S PPPP+ PP P+H P P PP P S P
Sbjct: 577 PPVYSPPPPVHS---PPPPVHSPPPPAPVHSPPPPVHSPPPPPPVYSPPP 623
Score = 38.1 bits (87), Expect = 0.010
Identities = 22/58 (37%), Positives = 25/58 (42%), Gaps = 4/58 (6%)
Query: 5 SPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
SP PP +PPP PPPP+H P P P P P S P + P HS
Sbjct: 542 SPPPPVHSPPPPPVYSPPP--PPPPVHSPPP--PVFSPPPPVYSPPPPVHSPPPPVHS 595
Score = 37.4 bits (85), Expect = 0.016
Identities = 18/45 (40%), Positives = 24/45 (53%), Gaps = 5/45 (11%)
Query: 5 SPTTAT-EPPPSSSAGTPPPPL----PPPPLHQPQPAKPDAPPSE 44
SP PPP + +PPPP+ PPPP++ P P PPS+
Sbjct: 588 SPPPPVHSPPPPAPVHSPPPPVHSPPPPPPVYSPPPPVFSPPPSQ 632
Score = 35.4 bits (80), Expect = 0.062
Identities = 19/60 (31%), Positives = 26/60 (42%), Gaps = 6/60 (10%)
Query: 6 PTTATEPPPSSSAGT--PPPPLPP----PPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P PPPS S PPP PP PP+ P PA + + P + P + ++P
Sbjct: 622 PPPVFSPPPSQSPPVVYSPPPRPPKINSPPVQSPPPAPVEKKETPPAHAPAPSDDEFIIP 681
Score = 32.3 bits (72), Expect = 0.52
Identities = 22/66 (33%), Positives = 28/66 (42%), Gaps = 17/66 (25%)
Query: 3 TKSPTTATEPPPSSSAGTP---PPPLPPPPLHQPQP-------------AKPDAPPSEPK 46
T PTT + P S TP P P+P P+H+P P ++P P PK
Sbjct: 440 TPVPTTPVQKP-SPVPTTPVQKPSPVPTTPVHEPSPVLATPVDKPSPVPSRPVQKPQPPK 498
Query: 47 LSATPD 52
S PD
Sbjct: 499 ESPQPD 504
Score = 32.0 bits (71), Expect = 0.68
Identities = 17/43 (39%), Positives = 21/43 (48%), Gaps = 4/43 (9%)
Query: 6 PTTATEPPPSSSAGTP---PPPLPPPPLHQPQPAKPDAPPSEP 45
PTT P S TP P P+P P+ +PQP K P +P
Sbjct: 465 PTTPVHEP-SPVLATPVDKPSPVPSRPVQKPQPPKESPQPDDP 506
Score = 28.9 bits (63), Expect = 5.8
Identities = 23/62 (37%), Positives = 27/62 (43%), Gaps = 6/62 (9%)
Query: 3 TKSPTTATEPPPSSSAGTP---PPPLPPPPLHQPQPAKPDAPPSEPK-LSATPDANVILV 58
T PTT P TP P P+P P+ +P P P P EP + ATP V
Sbjct: 429 TPVPTTPVHKPTPVPT-TPVQKPSPVPTTPVQKPSPV-PTTPVHEPSPVLATPVDKPSPV 486
Query: 59 PS 60
PS
Sbjct: 487 PS 488
>At3g19020 hypothetical protein
Length = 951
Score = 43.5 bits (101), Expect = 2e-04
Identities = 24/59 (40%), Positives = 28/59 (46%), Gaps = 5/59 (8%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P + PPP S PPPP+ PPPP+H P P P P P S P L P+ S
Sbjct: 771 PPVHSPPPPVHS---PPPPVHSPPPPVHSPPPPSPIYSPPPPVFSPPPKPVTPLPPATS 826
Score = 42.0 bits (97), Expect = 7e-04
Identities = 21/48 (43%), Positives = 24/48 (49%), Gaps = 5/48 (10%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP S PPPP+ PPPP+H P P PP P S P
Sbjct: 764 PPVHSPPPPVHS---PPPPVHSPPPPVHSPPPPVHSPPPPSPIYSPPP 808
Score = 41.6 bits (96), Expect = 9e-04
Identities = 23/56 (41%), Positives = 28/56 (49%), Gaps = 6/56 (10%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVP 59
P + PPP S PPPP+ PPPP+H P P +PP P S P A + P
Sbjct: 696 PPVHSPPPPVHS---PPPPVHSPPPPVHSP-PPPVQSPPPPPVFSPPPPAPIYSPP 747
Score = 41.2 bits (95), Expect = 0.001
Identities = 23/62 (37%), Positives = 27/62 (43%), Gaps = 2/62 (3%)
Query: 1 MATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
M + P + PPP S PP PPPP+H P P PP P S P + P
Sbjct: 662 MHSPPPPVYSPPPPVHSPPPPPVHSPPPPVHSPPPPVHSPPP--PVHSPPPPVHSPPPPV 719
Query: 61 HS 62
HS
Sbjct: 720 HS 721
Score = 40.8 bits (94), Expect = 0.001
Identities = 24/59 (40%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P + PPP S PPPP+ PPPP+H P P +PP P S P + P HS
Sbjct: 653 PPVFSPPPPMHS---PPPPVYSPPPPVHSPPPPPVHSPP-PPVHSPPPPVHSPPPPVHS 707
Score = 40.4 bits (93), Expect = 0.002
Identities = 22/59 (37%), Positives = 27/59 (45%), Gaps = 8/59 (13%)
Query: 12 PPPSSSAGTPPPPL--PPPPLHQP------QPAKPDAPPSEPKLSATPDANVILVPSHS 62
PPP +PPPP+ PPPP+H P P +PP P S P + P HS
Sbjct: 642 PPPPPPVHSPPPPVFSPPPPMHSPPPPVYSPPPPVHSPPPPPVHSPPPPVHSPPPPVHS 700
Score = 39.7 bits (91), Expect = 0.003
Identities = 23/67 (34%), Positives = 28/67 (41%), Gaps = 12/67 (17%)
Query: 6 PTTATEPPPSSSAGTPPPPL----------PPPPLHQPQPAKPDAPPSEPKLSATPDANV 55
P + PPP+ PPPP+ PPPP+H P P PP P S P +
Sbjct: 732 PPVFSPPPPAPIYSPPPPPVHSPPPPVHSPPPPPVHSPPPPVHSPPP--PVHSPPPPVHS 789
Query: 56 ILVPSHS 62
P HS
Sbjct: 790 PPPPVHS 796
Score = 38.9 bits (89), Expect = 0.006
Identities = 20/58 (34%), Positives = 25/58 (42%), Gaps = 4/58 (6%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPP----SEPKLSATPDANVILVP 59
P + PPP S PP PPPP+H P P PP P + + P + I P
Sbjct: 749 PPVHSPPPPVHSPPPPPVHSPPPPVHSPPPPVHSPPPPVHSPPPPVHSPPPPSPIYSP 806
Score = 37.7 bits (86), Expect = 0.012
Identities = 23/70 (32%), Positives = 27/70 (37%), Gaps = 13/70 (18%)
Query: 5 SPTTATEPPPSSSAGTPPPPL------------PPPPLHQPQPAKPDAPPSEPKLSATPD 52
SP PP PPPP+ PPPP+H P P +PP P S P
Sbjct: 714 SPPPPVHSPPPPVQSPPPPPVFSPPPPAPIYSPPPPPVHSP-PPPVHSPPPPPVHSPPPP 772
Query: 53 ANVILVPSHS 62
+ P HS
Sbjct: 773 VHSPPPPVHS 782
Score = 37.7 bits (86), Expect = 0.012
Identities = 30/102 (29%), Positives = 37/102 (35%), Gaps = 34/102 (33%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSA---------------- 49
P PPP S +PPPP+ PP P+P P P + P +A
Sbjct: 791 PPPVHSPPPPSPIYSPPPPVFSPP---PKPVTPLPPATSPMANAPTPSSSESGEISTPVQ 847
Query: 50 --TPDANVILVPSHSRWFSWDSIHQCEVRHLPEFFDSSSKSP 89
TPD+ I PS S H P F S + SP
Sbjct: 848 APTPDSEDIEAPSDS-------------NHSPVFKSSPAPSP 876
Score = 37.4 bits (85), Expect = 0.016
Identities = 19/49 (38%), Positives = 24/49 (48%), Gaps = 6/49 (12%)
Query: 6 PTTATEPPPSSSAGTPPPPL---PPPPLHQPQPAKPDAPPSEPKLSATP 51
P + PPP S PPPP+ PPPP+ P P P P P + + P
Sbjct: 710 PPVHSPPPPVHS---PPPPVQSPPPPPVFSPPPPAPIYSPPPPPVHSPP 755
Score = 36.2 bits (82), Expect = 0.036
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 4/59 (6%)
Query: 6 PTTATEPPPSSSAGTPP--PPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
P+ +TE ++S +PP P PPPP+H P P P P P S P P HS
Sbjct: 622 PSPSTEETKTTSPQSPPVHSPPPPPPVHSPPP--PVFSPPPPMHSPPPPVYSPPPPVHS 678
Score = 35.0 bits (79), Expect = 0.081
Identities = 22/60 (36%), Positives = 24/60 (39%), Gaps = 4/60 (6%)
Query: 5 SPTTATEPPPSSSAGTPPPP--LPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSHS 62
SP PP PPP PPPP+H P P PP P S P + P HS
Sbjct: 657 SPPPPMHSPPPPVYSPPPPVHSPPPPPVHSPPPPVHSPPP--PVHSPPPPVHSPPPPVHS 714
Score = 32.0 bits (71), Expect = 0.68
Identities = 16/51 (31%), Positives = 23/51 (44%), Gaps = 5/51 (9%)
Query: 11 EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPSH 61
+P P+ A P + PP L +P KP+ P + S P+ PSH
Sbjct: 405 KPTPTPKAPEPKKEINPPNLEEPSKPKPEESPKPQQPSPKPE-----TPSH 450
Score = 31.6 bits (70), Expect = 0.89
Identities = 21/61 (34%), Positives = 26/61 (42%), Gaps = 7/61 (11%)
Query: 6 PTTATEPPPSSSAGTPPPPL--PPPPLHQPQPAK--PDAPPSEPKLSATPDANVILVPSH 61
P + PPP S PPPP+ PPPP+ P P PP+ P + P H
Sbjct: 703 PPVHSPPPPVHS---PPPPVHSPPPPVQSPPPPPVFSPPPPAPIYSPPPPPVHSPPPPVH 759
Query: 62 S 62
S
Sbjct: 760 S 760
Score = 31.2 bits (69), Expect = 1.2
Identities = 16/53 (30%), Positives = 22/53 (41%), Gaps = 5/53 (9%)
Query: 4 KSPTTATEPPPSSSAGTPPPPLP-----PPPLHQPQPAKPDAPPSEPKLSATP 51
K ++ EPP + P PP P P P Q P ++P +P TP
Sbjct: 503 KQESSKQEPPKPEESPKPEPPKPEESPKPQPPKQETPKPEESPKPQPPKQETP 555
Score = 30.4 bits (67), Expect = 2.0
Identities = 16/42 (38%), Positives = 21/42 (49%), Gaps = 4/42 (9%)
Query: 4 KSPTTATEPPPSSSAGTPPPPL---PPPPLHQPQPAKPDAPP 42
++ TT+ + PP S PPPP+ PPP P P PP
Sbjct: 628 ETKTTSPQSPPVHSP-PPPPPVHSPPPPVFSPPPPMHSPPPP 668
Score = 30.0 bits (66), Expect = 2.6
Identities = 14/41 (34%), Positives = 17/41 (41%)
Query: 11 EPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
+PP S+ T PP+H P P P P P S P
Sbjct: 620 QPPSPSTEETKTTSPQSPPVHSPPPPPPVHSPPPPVFSPPP 660
Score = 28.5 bits (62), Expect = 7.6
Identities = 16/55 (29%), Positives = 21/55 (38%), Gaps = 7/55 (12%)
Query: 4 KSPTTATEPPPSSSAGTPPP-------PLPPPPLHQPQPAKPDAPPSEPKLSATP 51
+ P E P S+ PP P PP P P+P P +P+ S P
Sbjct: 493 EQPKPKPESPKQESSKQEPPKPEESPKPEPPKPEESPKPQPPKQETPKPEESPKP 547
Score = 28.5 bits (62), Expect = 7.6
Identities = 20/63 (31%), Positives = 26/63 (40%), Gaps = 15/63 (23%)
Query: 3 TKSPTTATEPPPSSSAGTPPP---PLPPPPLHQ---------PQPAKPDAPP--SEPKLS 48
T P + +P P TP P P P PP + PQP K + PP PK+
Sbjct: 538 TPKPEESPKPQPPKQE-TPKPEESPKPQPPKQETPKPEESPKPQPPKQEQPPKTEAPKMG 596
Query: 49 ATP 51
+ P
Sbjct: 597 SPP 599
>At1g68690 protein kinase, putative
Length = 708
Score = 43.5 bits (101), Expect = 2e-04
Identities = 20/46 (43%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P T PPP ++ G PPPPLP PP P +P P P S P
Sbjct: 54 PPVTTSPPPVAN-GAPPPPLPKPPESSSPPPQPVIPSPPPSTSPPP 98
Score = 41.6 bits (96), Expect = 9e-04
Identities = 20/54 (37%), Positives = 27/54 (49%), Gaps = 1/54 (1%)
Query: 5 SPTTATEPPPSSSAGTPPPPL-PPPPLHQPQPAKPDAPPSEPKLSATPDANVIL 57
SP +T PPP +PPP PPP L P P+ P P S P +P +++
Sbjct: 89 SPPPSTSPPPQPVIPSPPPSASPPPALVPPLPSSPPPPASVPPPRPSPSPPILV 142
Score = 38.9 bits (89), Expect = 0.006
Identities = 19/51 (37%), Positives = 24/51 (46%), Gaps = 2/51 (3%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPP--PLHQPQPAKPDAPPSEPKLSATP 51
T+SP + P P S T PP PP P P P P +PPS+ + P
Sbjct: 164 TQSPPPPSPPSPPSERPTQSPPSPPSERPTQSPPPPSPPSPPSDRPSQSPP 214
Score = 38.5 bits (88), Expect = 0.007
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 1/56 (1%)
Query: 6 PTTATEPPPSSS-AGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
P + PPPS +PPPP PP P + P +PPSE + P + PS
Sbjct: 151 PIQSPPPPPSDRPTQSPPPPSPPSPPSERPTQSPPSPPSERPTQSPPPPSPPSPPS 206
Score = 37.4 bits (85), Expect = 0.016
Identities = 18/47 (38%), Positives = 21/47 (44%), Gaps = 1/47 (2%)
Query: 6 PTTATEPPPS-SSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATP 51
P PPPS +PPPP P P P P +PPSE + P
Sbjct: 139 PILVRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPSPPSERPTQSPP 185
Score = 37.4 bits (85), Expect = 0.016
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 3/55 (5%)
Query: 3 TKSPTTATEPPP---SSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDAN 54
+ SP + PPP ++S TPPP P P P P + PP S P AN
Sbjct: 11 SNSPPVTSPPPPLNNATSPATPPPVTSPLPPSAPPPNRAPPPPPPVTTSPPPVAN 65
Score = 37.4 bits (85), Expect = 0.016
Identities = 24/65 (36%), Positives = 28/65 (42%), Gaps = 5/65 (7%)
Query: 2 ATKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAP----PSEPKLSATPDANVIL 57
AT P T+ PP + PPP PPP P P AP P P+ S+ P VI
Sbjct: 30 ATPPPVTSPLPPSAPPPNRAPPP-PPPVTTSPPPVANGAPPPPLPKPPESSSPPPQPVIP 88
Query: 58 VPSHS 62
P S
Sbjct: 89 SPPPS 93
Score = 36.6 bits (83), Expect = 0.028
Identities = 21/55 (38%), Positives = 27/55 (48%), Gaps = 8/55 (14%)
Query: 6 PTTATEPPPSSSAGTPPPPLPPPPLHQPQPAKPDAPPSEPKLSATPDANVILVPS 60
P ++ PPP +PPP PP PQP P PPS ++ P A V +PS
Sbjct: 75 PPESSSPPPQPVIPSPPPSTSPP----PQPVIPSPPPS----ASPPPALVPPLPS 121
Score = 35.4 bits (80), Expect = 0.062
Identities = 22/57 (38%), Positives = 27/57 (46%), Gaps = 9/57 (15%)
Query: 4 KSPTTATEPP----PSSSAGTPPPPLP----PPPLHQPQPA-KPDAPPSEPKLSATP 51
K P +++ PP PS T PPP P PPP P PA P P S P ++ P
Sbjct: 74 KPPESSSPPPQPVIPSPPPSTSPPPQPVIPSPPPSASPPPALVPPLPSSPPPPASVP 130
Score = 35.0 bits (79), Expect = 0.081
Identities = 24/60 (40%), Positives = 32/60 (53%), Gaps = 6/60 (10%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPPPLHQPQPAK--PDAPPSEPKLSATP-DANVILVP 59
T+SP + P P S + PP PPP +PQP + P++PP P S+ P ILVP
Sbjct: 193 TQSPPPPSPPSPPSDRPSQSPP-PPPEDTKPQPPRRSPNSPP--PTFSSPPRSPPEILVP 249
Score = 35.0 bits (79), Expect = 0.081
Identities = 20/51 (39%), Positives = 24/51 (46%), Gaps = 9/51 (17%)
Query: 7 TTATEPPPSSSAGTPPPPLPPPPLHQ------PQPAKPDAPPSEPKLSATP 51
TT +PP S+S PP PPPPL+ P P PPS P + P
Sbjct: 3 TTPVQPPVSNS---PPVTSPPPPLNNATSPATPPPVTSPLPPSAPPPNRAP 50
Score = 33.9 bits (76), Expect = 0.18
Identities = 16/45 (35%), Positives = 21/45 (46%), Gaps = 1/45 (2%)
Query: 3 TKSPTTATEPPPSSS-AGTPPPPLPPPPLHQPQPAKPDAPPSEPK 46
++ PT + PPS +PPPP PP P P PP + K
Sbjct: 177 SERPTQSPPSPPSERPTQSPPPPSPPSPPSDRPSQSPPPPPEDTK 221
Score = 33.5 bits (75), Expect = 0.24
Identities = 17/48 (35%), Positives = 21/48 (43%), Gaps = 2/48 (4%)
Query: 3 TKSPTTATEPPPSSSAGTPPPPLPPP--PLHQPQPAKPDAPPSEPKLS 48
T+SP + P+ S P PP PP P P P D P P+ S
Sbjct: 181 TQSPPSPPSERPTQSPPPPSPPSPPSDRPSQSPPPPPEDTKPQPPRRS 228
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.131 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,890,716
Number of Sequences: 26719
Number of extensions: 530302
Number of successful extensions: 13474
Number of sequences better than 10.0: 432
Number of HSP's better than 10.0 without gapping: 296
Number of HSP's successfully gapped in prelim test: 149
Number of HSP's that attempted gapping in prelim test: 3782
Number of HSP's gapped (non-prelim): 4228
length of query: 434
length of database: 11,318,596
effective HSP length: 102
effective length of query: 332
effective length of database: 8,593,258
effective search space: 2852961656
effective search space used: 2852961656
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0438.5


