

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0404.10
(989 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
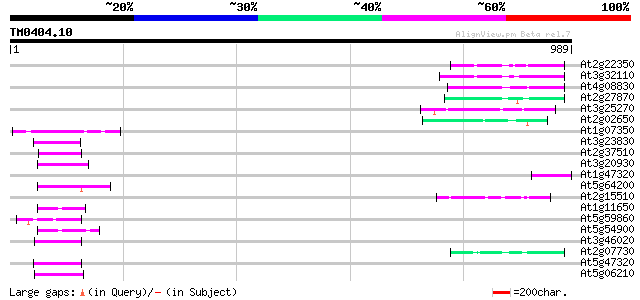
Score E
Sequences producing significant alignments: (bits) Value
At2g22350 putative non-LTR retroelement reverse transcriptase 66 8e-11
At3g32110 non-LTR reverse transcriptase, putative 66 1e-10
At4g08830 putative protein 60 5e-09
At2g27870 putative non-LTR retroelement reverse transcriptase 59 2e-08
At3g25270 hypothetical protein 58 3e-08
At2g02650 putative reverse transcriptase 56 1e-07
At1g07350 transformer-SR ribonucleoprotein like protein 56 1e-07
At3g23830 glycine-rich RNA binding protein, putative 54 3e-07
At2g37510 putative RNA-binding protein 53 7e-07
At3g20930 unknown protein 52 1e-06
At1g47320 hypothetical protein 52 1e-06
At5g64200 unknown protein 52 2e-06
At2g15510 putative non-LTR retroelement reverse transcriptase 52 2e-06
At1g11650 putative DNA binding protein 52 2e-06
At5g59860 RNA-binding protein - like 51 3e-06
At5g54900 unknown protein 51 3e-06
At3g46020 RNA binding protein -like 51 3e-06
At2g07730 putative non-LTR retroelement reverse transcriptase 51 4e-06
At5g47320 40S ribosomal protein S19 50 5e-06
At5g06210 unknown protein 50 5e-06
>At2g22350 putative non-LTR retroelement reverse transcriptase
Length = 321
Score = 66.2 bits (160), Expect = 8e-11
Identities = 52/202 (25%), Positives = 89/202 (43%), Gaps = 19/202 (9%)
Query: 777 LCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPI 836
+C IC E+ H++ CP + +W+ ++ R+++ F +S LLEW + + +
Sbjct: 34 VCQICQGGEETILHVLRDCPAMAGIWSRLVPRDQIRQFFTAS----LLEWI-YKNLRERG 88
Query: 837 MWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQ 896
W + W W+ R +F D + +IK +E + +
Sbjct: 89 SWPTVFVMAVWWGWKWRCGNIFGGNGKCRDRVK---------FIKDLAEEV--AIANAFV 137
Query: 897 HFEKIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFS 956
++R+ V W P G +K N DG+SRGN G + GGVL+D N +G F+
Sbjct: 138 KGNEVRVSRVERLV---SWVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGGFA 194
Query: 957 CAVGSTWAYVAERKALLEGLLL 978
+G A +AE + GL +
Sbjct: 195 VNIGVCSAPLAELWGVYYGLFI 216
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 65.9 bits (159), Expect = 1e-10
Identities = 57/226 (25%), Positives = 99/226 (43%), Gaps = 24/226 (10%)
Query: 758 NIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPS 817
N A+ + R+ + + +C +C ES H++ CP + +W I+ R F
Sbjct: 1600 NQAIMTNSERKRRHLCDSDVCQVCRGGIESILHVLRDCPAMSGIWDRIVPRRLQQSFFTM 1659
Query: 818 SIQGLLLEW--SSLRA--ISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHL 873
S LLEW S+LR +++ W + W W+ R +F + D +
Sbjct: 1660 S----LLEWLYSNLRQGLMTEGSDWSTMFAMAVWWGWKWRCSNIFGENKTCRDRVR---- 1711
Query: 874 IRVMWWIKSWWKECPFTVLDFSQHFEKIRIKVAAPKVHSA-DWCPPPPGCLKFNVDGSSR 932
+IK E + ++ ++++ +V+ W PP G K N DG+SR
Sbjct: 1712 -----FIKDLAVEVSIA------YSREVELRLSGLRVNKPIRWTPPMEGWYKINTDGASR 1760
Query: 933 GNSGVSGIGGVLKDANSTMLGRFSCAVGSTWAYVAERKALLEGLLL 978
GN G++ GGVL+++ G F+ +G A +AE + GL +
Sbjct: 1761 GNPGLASAGGVLRNSAGAWCGGFAVNIGRCSAPLAELWGVYYGLYM 1806
>At4g08830 putative protein
Length = 947
Score = 60.5 bits (145), Expect = 5e-09
Identities = 48/208 (23%), Positives = 87/208 (41%), Gaps = 16/208 (7%)
Query: 772 IENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRA 831
I + +C +C E+ H++ CP I +W ++ + Y F S+ G L ++
Sbjct: 712 IGDTSVCQVCKGGDETILHVLKDCPSIAGIWRRLVQVQRSYDFFNGSLFGWLYVNLGMKN 771
Query: 832 ISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTV 891
W + + W W+ R VF + D + ++++ V
Sbjct: 772 AETGYAWATLFAIVVWWSWKWRCGYVFGEVGKCRDRV-------------KFFRDLAAEV 818
Query: 892 LD-FSQHFEKIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANST 950
+ H + ++ ++ + W PP +K N DG+SRGN G++ GGVL+D
Sbjct: 819 SHAHAIHSQNGGLRTRVERLVA--WKPPDGEWVKLNTDGASRGNLGLATTGGVLRDGIGH 876
Query: 951 MLGRFSCAVGSTWAYVAERKALLEGLLL 978
G F+ +G A +AE + GL +
Sbjct: 877 WCGGFALDIGVCSAPLAELWGVYYGLYM 904
>At2g27870 putative non-LTR retroelement reverse transcriptase
Length = 314
Score = 58.5 bits (140), Expect = 2e-08
Identities = 50/216 (23%), Positives = 85/216 (39%), Gaps = 21/216 (9%)
Query: 767 RRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEW 826
RR + + +C IC ++ H++ CP + +W ++ + F S+ L
Sbjct: 11 RRRRHLSDSDICQICKGAEKTIIHILRDCPAMEGIWIRLVPAGKRREFFTQSLLEWLFAN 70
Query: 827 SSLRAISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKE 886
R + W + W W+ R +F +D D + ++K +E
Sbjct: 71 LGDRRKTCESTWSTLFALSIWWAWKWRCGNIFGVQDKCRDRV---------RFLKDLARE 121
Query: 887 CPFTVLDF----SQHFEKIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGG 942
+ H E++ +A W P G K N DG+SRGN G++ GG
Sbjct: 122 TSMAHVIVRTLSGGHGERVERLIA--------WSKPEEGWWKLNTDGASRGNPGLASAGG 173
Query: 943 VLKDANSTMLGRFSCAVGSTWAYVAERKALLEGLLL 978
VL+D G F+ +G A +AE + GL +
Sbjct: 174 VLRDEEGAWRGGFALNIGVCSAPLAELWGVYYGLYI 209
>At3g25270 hypothetical protein
Length = 343
Score = 57.8 bits (138), Expect = 3e-08
Identities = 61/243 (25%), Positives = 99/243 (40%), Gaps = 16/243 (6%)
Query: 725 MGLEIVPAVGSAGGLVTLWKTS----LFQVSQQLMECNIAVKASLIRRGIPIENGGLCDI 780
M EI P G A +WK + +L+ +A +L RR I N C
Sbjct: 1 MDPEIQPPPGKAEIKAKIWKLKTAPKIKHFLWKLLSGALATGDNLKRRHI--RNHPQCHR 58
Query: 781 CGLIGESESHLMLHCPKIWLLWTTI-LAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWE 839
C E+ HL C +W + +E+ + + L SS A P ++
Sbjct: 59 CCQEDETSQHLFFDCFYAQQVWRASGIPHQELRTTGITMETKMELLLSSCLANRQPQLFN 118
Query: 840 IIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFE 899
+ + + W +W++RN LVF QK S W L R ++ W++ V +Q
Sbjct: 119 LAIWIL-WRLWKSRNQLVFQQKSIS----WQNTLQRARNDVQE-WEDTNTYVQSLNQQVH 172
Query: 900 KIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAV 959
R + P + W PP +K+N DG+ + + G +++D N +G A+
Sbjct: 173 SSRHQ--QPTMARTKWQRPPSTWIKYNYDGAFNHQTRNAKAGWLMRDENGVYMGS-GQAI 229
Query: 960 GST 962
GST
Sbjct: 230 GST 232
>At2g02650 putative reverse transcriptase
Length = 365
Score = 55.8 bits (133), Expect = 1e-07
Identities = 53/230 (23%), Positives = 91/230 (39%), Gaps = 24/230 (10%)
Query: 729 IVPAVGSAGGLVTLWKTSLF-QVSQQLMECNIAVKASLIR-RGIPIENGGLCDICGLIGE 786
I P GS +WK + ++ L C A+ R R I+ +C C + E
Sbjct: 24 IQPPPGSTEVKQAIWKLHVAPKIKHFLWRCVTGALATNTRLRSRNIDADPICQRCCIEEE 83
Query: 787 SESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQ---GLLLEWSSLRAISDPIMWEIIPY 843
+ H+M +CP +W + PSS + L++ S + + + +P+
Sbjct: 84 TIHHIMFNCPYTQSVWRSANIIIGNQWGPPSSFEDNLNRLIQLSKTQTTNS--LDRFLPF 141
Query: 844 AMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRI 903
+ W +W++RN +F QK S DY + W L+ ++ E +
Sbjct: 142 WIMWRLWKSRNVFLFQQKCQSPDYEARKGIQDATEW------------LNANETTENTNV 189
Query: 904 KVAAPKVH-----SADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDAN 948
VA + S+ W PPP G +K N D S + G +++ N
Sbjct: 190 HVATNPIQTSRRDSSQWNPPPEGWVKCNFDSGYTQGSPYTRSGWTIRECN 239
>At1g07350 transformer-SR ribonucleoprotein like protein
Length = 382
Score = 55.8 bits (133), Expect = 1e-07
Identities = 47/191 (24%), Positives = 80/191 (41%), Gaps = 17/191 (8%)
Query: 6 YSPATIAWNKRHNTSAQSTKSRANKWSGLGNNRRADGIHRRIGKSLFVDGLTDQTTHLQV 65
YSP+ ++KR S + SR+ R G SL+V GL+ + T +
Sbjct: 39 YSPSLSPYDKRRGRSVSRSLSRSP-------TRSVSSDAENPGNSLYVTGLSHRVTERDL 91
Query: 66 KNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMNGSILNGATINVSLA 125
++ FA G++ V + + FGF+ DA +R ++ S+L G I V A
Sbjct: 92 EDHFAKEGKVTDVHLVLDPWTRESRGFGFISMKSVGDANRCIRSLDHSVLQGRVITVEKA 151
Query: 126 RLPQDQHTRPRPTDNQWKGV-SEWGEHSKPNKGNRRSARKEWIPLKRNSQSDYPLNYDTL 184
R + + PT ++ G+ + G H P+ RRS + R+ Y +
Sbjct: 152 RRRRGR----TPTPGKYLGLRTARGRHKSPSYSPRRS-----VSCSRSRSRSYSSDRGRS 202
Query: 185 FLPSIEEQDRN 195
+ PS + R+
Sbjct: 203 YSPSYGRRGRS 213
>At3g23830 glycine-rich RNA binding protein, putative
Length = 136
Score = 54.3 bits (129), Expect = 3e-07
Identities = 33/84 (39%), Positives = 43/84 (50%)
Query: 42 GIHRRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDS 101
G R + LFV GL+ T +K AF SFG + V + R FGFV FS +
Sbjct: 28 GSLRYMSSKLFVGGLSWGTDDSSLKQAFTSFGEVTEATVIADRETGRSRGFGFVSFSCED 87
Query: 102 DAEVAMRRMNGSILNGATINVSLA 125
A A++ M+G LNG I V+LA
Sbjct: 88 SANNAIKEMDGKELNGRQIRVNLA 111
>At2g37510 putative RNA-binding protein
Length = 142
Score = 53.1 bits (126), Expect = 7e-07
Identities = 31/76 (40%), Positives = 44/76 (57%)
Query: 51 LFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRM 110
LFV GL+ TT+ ++++AFASFG+++ V + R FGFV ++ DAE A M
Sbjct: 36 LFVSGLSRLTTNEKLQDAFASFGQLVDARVITDRDSGRSKGFGFVTYATIEDAEKAKAEM 95
Query: 111 NGSILNGATINVSLAR 126
N L+G I V AR
Sbjct: 96 NAKFLDGWVIFVDPAR 111
>At3g20930 unknown protein
Length = 374
Score = 52.4 bits (124), Expect = 1e-06
Identities = 27/90 (30%), Positives = 49/90 (54%)
Query: 49 KSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
K LF+ GL+ T+ ++ AF FG ++ V + K +R + F+ ++ + A A++
Sbjct: 282 KKLFITGLSFYTSEKTLRAAFEGFGELVEVKIIMDKISKRSKGYAFLEYTTEEAAGTALK 341
Query: 109 RMNGSILNGATINVSLARLPQDQHTRPRPT 138
MNG I+NG I V +A+ + R +P+
Sbjct: 342 EMNGKIINGWMIVVDVAKTKPFRQNRSQPS 371
>At1g47320 hypothetical protein
Length = 259
Score = 52.4 bits (124), Expect = 1e-06
Identities = 29/69 (42%), Positives = 38/69 (55%)
Query: 921 GCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVGSTWAYVAERKALLEGLLLCG 980
GCLK N DG+SRGN G++ GGVL+D G FS +G + A +AE GL L
Sbjct: 99 GCLKINTDGASRGNPGLATAGGVLQDNEGRWCGGFSLNIGRSSAPMAELWGAYYGLYLAW 158
Query: 981 KIYSCKLRL 989
+ S + L
Sbjct: 159 ERKSSHIEL 167
>At5g64200 unknown protein
Length = 303
Score = 52.0 bits (123), Expect = 2e-06
Identities = 37/138 (26%), Positives = 65/138 (46%), Gaps = 11/138 (7%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL V +T +TT + FA +G+++ VF+ +R F FVR+ + +A A+ R
Sbjct: 17 SLLVLNITFRTTADDLYPLFAKYGKVVDVFIPRDRRTGDSRGFAFVRYKYKDEAHKAVER 76
Query: 110 MNGSILNGATINVSLA-------RLPQDQHTRPRPTDNQWKGVSEWGEHS---KPNKGNR 159
++G +++G I V A ++ + + P P + + S S + R
Sbjct: 77 LDGRVVDGREITVQFAKYGPNAEKISKGRVVEPPPKSRRSRSRSPRRSRSPRRSRSPPRR 136
Query: 160 RSARKEWIPLKRNSQSDY 177
RS R+ P +R S+ DY
Sbjct: 137 RSPRRSRSP-RRRSRDDY 153
>At2g15510 putative non-LTR retroelement reverse transcriptase
Length = 1138
Score = 51.6 bits (122), Expect = 2e-06
Identities = 48/205 (23%), Positives = 88/205 (42%), Gaps = 22/205 (10%)
Query: 753 QLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILAREEVY 812
+++ + V +IRRG+ I+ C ICG GES +H++ C +W +
Sbjct: 837 RILSAALPVADQIIRRGMSIDPR--CQICGDEGESTNHVLFTCSMARQVWALSGVPTPEF 894
Query: 813 CCFPSSIQG---LLLEWSSLRAISDPIM--WEIIPYAMFWSIWRARNDLVFNQKDFSADY 867
+SI L E + + D + W P+ + W +W+ RN L F+ F
Sbjct: 895 GFQNASIFANIQFLFELKKMILVPDLVKRSW---PWVL-WRLWKNRNKLFFDGITFCPLN 950
Query: 868 IWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRIKVAAPKVHSADWCPPPPGCLKFNV 927
++++ W++ + S+ +++ P HS W PPP G +K N+
Sbjct: 951 ----SIVKIQEDTLEWFQAQSQIRVSESEEGQRV-----IPFTHS--WEPPPEGWVKCNI 999
Query: 928 DGSSRGNSGVSGIGGVLKDANSTML 952
+ G V G VL+D + +++
Sbjct: 1000 GSAWSGKKKVCGGAWVLRDEHGSVI 1024
>At1g11650 putative DNA binding protein
Length = 405
Score = 51.6 bits (122), Expect = 2e-06
Identities = 31/84 (36%), Positives = 46/84 (53%), Gaps = 6/84 (7%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
++FV GL T +KN F+ +G I+ V + KR GFV+FS S AE A+R
Sbjct: 262 TVFVGGLDASVTDDHLKNVFSQYGEIVHVKIPAGKR------CGFVQFSEKSCAEEALRM 315
Query: 110 MNGSILNGATINVSLARLPQDQHT 133
+NG L G T+ +S R P ++ +
Sbjct: 316 LNGVQLGGTTVRLSWGRSPSNKQS 339
Score = 32.7 bits (73), Expect = 1.0
Identities = 22/64 (34%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 50 SLFVDGLT-DQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
++FV L D T ++ ++ AS+ + G V + R +GFVRFS +S+ AM
Sbjct: 156 TIFVGDLAADVTDYILLETFRASYPSVKGAKVVIDRVTGRTKGYGFVRFSDESEQIRAMT 215
Query: 109 RMNG 112
MNG
Sbjct: 216 EMNG 219
>At5g59860 RNA-binding protein - like
Length = 157
Score = 51.2 bits (121), Expect = 3e-06
Identities = 37/127 (29%), Positives = 59/127 (46%), Gaps = 23/127 (18%)
Query: 13 WNKRHNTSAQSTKSRANKW----------SGLGNNRRADGIHRRIGKSLFV---DGLTDQ 59
W N+ A+ +++ NK S + RR G GK+L GL+
Sbjct: 27 WQPHKNSGAKDEEAKFNKRTKYKMRIRPRSSISKRRRLIG-----GKTLTCVSNSGLSAY 81
Query: 60 TTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMNGSILNGAT 119
TT ++ FA F R++ ++ QR FGF+ F + DA+ A++ +NG I+NG
Sbjct: 82 TTDQSLRQLFAPFARLI-----KDQQTQRPKGFGFITFESEDDAQKALKALNGKIVNGRL 136
Query: 120 INVSLAR 126
I V A+
Sbjct: 137 IFVETAK 143
>At5g54900 unknown protein
Length = 387
Score = 51.2 bits (121), Expect = 3e-06
Identities = 34/108 (31%), Positives = 53/108 (48%), Gaps = 11/108 (10%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
++FV GL T ++K+ F FG +L V + KR GFV+++ + AE A+
Sbjct: 261 TIFVGGLDANVTDDELKSIFGQFGELLHVKIPPGKR------CGFVQYANKASAEHALSV 314
Query: 110 MNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWGEHSKPNKG 157
+NG+ L G +I +S R P Q + QW G +G +P G
Sbjct: 315 LNGTQLGGQSIRLSWGRSPNKQSDQA-----QWNGGGYYGYPPQPQGG 357
Score = 33.5 bits (75), Expect = 0.61
Identities = 19/64 (29%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Query: 50 SLFVDGLTDQTTHLQVKNAFAS-FGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
++FV L + T + + F + +G + G V + R +GFVRF+ +++ AM
Sbjct: 155 TIFVGDLAPEVTDYMLSDTFKNVYGSVKGAKVVLDRTTGRSKGYGFVRFADENEQMRAMT 214
Query: 109 RMNG 112
MNG
Sbjct: 215 EMNG 218
>At3g46020 RNA binding protein -like
Length = 102
Score = 51.2 bits (121), Expect = 3e-06
Identities = 27/82 (32%), Positives = 44/82 (52%)
Query: 45 RRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAE 104
+RI LFV L+ TT ++ F+ FG+I + QR FGF+ F + DA
Sbjct: 3 KRISAQLFVSRLSAYTTDQSLRQLFSPFGQIKEARLIRDSETQRPKGFGFITFDSEDDAR 62
Query: 105 VAMRRMNGSILNGATINVSLAR 126
A++ ++G I++G I V +A+
Sbjct: 63 KALKSLDGKIVDGRLIFVEVAK 84
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 50.8 bits (120), Expect = 4e-06
Identities = 49/204 (24%), Positives = 79/204 (38%), Gaps = 26/204 (12%)
Query: 778 CDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIM 837
C +C ES H++ CP + +W +L + F EW L DP
Sbjct: 685 CSVCNGADESILHVLRDCPAMTPIWQRLLPQRRQNEFFSQ------FEW--LFTNLDPAK 736
Query: 838 --WEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFT-VLDF 894
W + W W+ R VF ++ D + +IK +E V
Sbjct: 737 GDWPTLFSMGIWWAWKWRCGDVFGERKLCRDRLK---------FIKDIAEEVRKAHVGTL 787
Query: 895 SQHFEKIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGR 954
+ H ++ R++ W P +K DG+SRG+ G++ G + + LG
Sbjct: 788 NNHVKRARVE------RMIRWKAPSDRWVKLTTDGASRGHQGLAAASGAILNLQGEWLGG 841
Query: 955 FSCAVGSTWAYVAERKALLEGLLL 978
F+ +GS A +AE GLL+
Sbjct: 842 FALNIGSCDAPLAELWGAYYGLLI 865
>At5g47320 40S ribosomal protein S19
Length = 212
Score = 50.4 bits (119), Expect = 5e-06
Identities = 29/85 (34%), Positives = 44/85 (51%)
Query: 42 GIHRRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDS 101
G R + L++ GL+ T +K+AF+SF + V K R +GFV F +
Sbjct: 24 GSLRYMSTKLYIGGLSPGTDEHSLKDAFSSFNGVTEARVMTNKVTGRSRGYGFVNFISED 83
Query: 102 DAEVAMRRMNGSILNGATINVSLAR 126
A A+ MNG LNG I+V++A+
Sbjct: 84 SANSAISAMNGQELNGFNISVNVAK 108
>At5g06210 unknown protein
Length = 146
Score = 50.4 bits (119), Expect = 5e-06
Identities = 28/85 (32%), Positives = 43/85 (49%)
Query: 45 RRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAE 104
R + LF+ GL+ TT + AF+ G+++ + + R FGFV F+ +A+
Sbjct: 30 RGVASKLFIGGLSFCTTEQGLSEAFSKCGQVVEAQIVMDRVSDRSKGFGFVTFASADEAQ 89
Query: 105 VAMRRMNGSILNGATINVSLARLPQ 129
A+ NG LNG TI V A+ Q
Sbjct: 90 KALMEFNGQQLNGRTIFVDYAKAKQ 114
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.134 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,424,010
Number of Sequences: 26719
Number of extensions: 1052768
Number of successful extensions: 3528
Number of sequences better than 10.0: 189
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 64
Number of HSP's that attempted gapping in prelim test: 3235
Number of HSP's gapped (non-prelim): 307
length of query: 989
length of database: 11,318,596
effective HSP length: 109
effective length of query: 880
effective length of database: 8,406,225
effective search space: 7397478000
effective search space used: 7397478000
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0404.10


