

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0403.8
(117 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
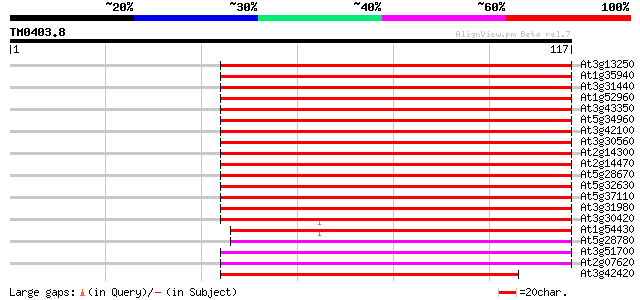
At3g13250 hypothetical protein 75 5e-15
At1g35940 hypothetical protein 75 5e-15
At3g31440 hypothetical protein 75 7e-15
At1g52960 hypothetical protein 74 1e-14
At3g43350 putative protein 74 2e-14
At5g34960 putative protein 73 2e-14
At3g42100 putative protein 71 8e-14
At3g30560 hypothetical protein 71 8e-14
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 71 8e-14
At2g14470 pseudogene 70 2e-13
At5g28670 putative protein 68 8e-13
At5g32630 putative protein 67 1e-12
At5g37110 putative helicase 66 2e-12
At3g31980 hypothetical protein 63 2e-11
At3g30420 hypothetical protein 63 2e-11
At1g54430 hypothetical protein 63 3e-11
At5g28780 putative protein 57 1e-09
At3g51700 unknown protein 57 2e-09
At2g07620 putative helicase 56 3e-09
At3g42420 putative protein 54 1e-08
>At3g13250 hypothetical protein
Length = 1419
Score = 75.1 bits (183), Expect = 5e-15
Identities = 35/73 (47%), Positives = 51/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G+ YIP I++T SD+ LPFK RRQF +++ F MTIN
Sbjct: 1297 LQITQLCSHIVEAKVITGDRIGQIVYIPLINITPSDTKLPFKMRRRQFPLSVAFVMTINK 1356
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V LYL
Sbjct: 1357 SQGQSLEQVGLYL 1369
>At1g35940 hypothetical protein
Length = 1678
Score = 75.1 bits (183), Expect = 5e-15
Identities = 34/73 (46%), Positives = 51/73 (69%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G IP ++LT +D+ LPFK RRQF +++ FAMTIN
Sbjct: 1555 LQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINK 1614
Query: 105 NQGQSLSHVSLYL 117
+QGQSL H+ LYL
Sbjct: 1615 SQGQSLEHIGLYL 1627
>At3g31440 hypothetical protein
Length = 536
Score = 74.7 bits (182), Expect = 7e-15
Identities = 35/73 (47%), Positives = 51/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G IP ++LT +D+ LPFK RRQF +++ FAMTIN
Sbjct: 413 LQITQLFTEIVEAKVITGDRIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINK 472
Query: 105 NQGQSLSHVSLYL 117
+QGQSL HV LYL
Sbjct: 473 SQGQSLEHVGLYL 485
>At1g52960 hypothetical protein
Length = 924
Score = 73.9 bits (180), Expect = 1e-14
Identities = 36/73 (49%), Positives = 50/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + +I A GN++GK IPR+ +T SD+ LPFK RRQF +++ FAMTIN
Sbjct: 826 LQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLPFKMRRRQFPLSVAFAMTINK 885
Query: 105 NQGQSLSHVSLYL 117
+QGQ+L V LYL
Sbjct: 886 SQGQTLESVGLYL 898
>At3g43350 putative protein
Length = 830
Score = 73.6 bits (179), Expect = 2e-14
Identities = 35/73 (47%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + +I A GN++GK IPR+ +T SD+ LPFK R+QF +++ FAMTIN
Sbjct: 616 LQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLPFKMRRKQFALSVAFAMTINK 675
Query: 105 NQGQSLSHVSLYL 117
+QGQ+L V LYL
Sbjct: 676 SQGQTLESVGLYL 688
Score = 71.2 bits (173), Expect = 8e-14
Identities = 34/73 (46%), Positives = 49/73 (66%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + +I A GN++GK IPR+ +T D+ LPFK R+QF +++ FAMTIN
Sbjct: 732 LQVTQMADTVIQARFITGNRVGKIVLIPRMLITPLDTRLPFKMRRKQFALSVAFAMTINK 791
Query: 105 NQGQSLSHVSLYL 117
+QGQ+L V LYL
Sbjct: 792 SQGQTLESVGLYL 804
Score = 31.6 bits (70), Expect = 0.066
Identities = 14/20 (70%), Positives = 16/20 (80%)
Query: 98 FAMTINNNQGQSLSHVSLYL 117
FAMTIN +QGQ+L V LYL
Sbjct: 553 FAMTINKSQGQTLESVGLYL 572
>At5g34960 putative protein
Length = 1033
Score = 73.2 bits (178), Expect = 2e-14
Identities = 35/73 (47%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+ +G IP ++LT +D+ LPFK RRQF +++ FAMTIN
Sbjct: 910 LQITQLCTQIVEAKVITGDIIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINT 969
Query: 105 NQGQSLSHVSLYL 117
+QGQSL HV LYL
Sbjct: 970 SQGQSLEHVGLYL 982
>At3g42100 putative protein
Length = 1752
Score = 71.2 bits (173), Expect = 8e-14
Identities = 35/73 (47%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L + ++ A V +++G IP I+LT SD+ LPFK RRQF +++ FAMTIN
Sbjct: 1629 LQITQLAKQVVQAKVITRDRIGDIVLIPLINLTPSDTKLPFKMRRRQFPLSVAFAMTINK 1688
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V LYL
Sbjct: 1689 SQGQSLEQVGLYL 1701
>At3g30560 hypothetical protein
Length = 1473
Score = 71.2 bits (173), Expect = 8e-14
Identities = 33/73 (45%), Positives = 48/73 (65%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++ L ++ + G ++GK IPR+ LT SD LPFK RRQF +++ FAMTIN
Sbjct: 1351 LQIVRLGDKLVQGRILTGTRVGKLVIIPRMPLTPSDRRLPFKMKRRQFPLSVAFAMTINK 1410
Query: 105 NQGQSLSHVSLYL 117
+QGQSL +V +YL
Sbjct: 1411 SQGQSLGNVGIYL 1423
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 71.2 bits (173), Expect = 8e-14
Identities = 33/73 (45%), Positives = 48/73 (65%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++ L ++ + G ++GK IPR+SLT SD LPFK RR F +++ FAMTIN
Sbjct: 1150 LQIVRLGDKLVQGRILTGTRVGKLVIIPRMSLTPSDRRLPFKMKRRHFPLSVAFAMTINK 1209
Query: 105 NQGQSLSHVSLYL 117
+QGQSL +V +YL
Sbjct: 1210 SQGQSLGNVGMYL 1222
>At2g14470 pseudogene
Length = 1265
Score = 70.1 bits (170), Expect = 2e-13
Identities = 32/73 (43%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G IP ++LT +++ LPFK RRQF +++ F MTIN
Sbjct: 1159 LQITQLCTQIVEAKVITGDRIGHIILIPTVNLTPTNTKLPFKMRRRQFPLSVAFVMTINK 1218
Query: 105 NQGQSLSHVSLYL 117
++GQSL HV LYL
Sbjct: 1219 SEGQSLEHVGLYL 1231
>At5g28670 putative protein
Length = 647
Score = 67.8 bits (164), Expect = 8e-13
Identities = 32/73 (43%), Positives = 47/73 (63%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++ L ++ + G ++GK IPR+ LT SDS LPF RRQF +++ FAM IN
Sbjct: 566 LQIVRLGDKLVQGRLLTGTRVGKLVLIPRMPLTPSDSRLPFTMQRRQFSLSVAFAMIINK 625
Query: 105 NQGQSLSHVSLYL 117
+QGQSL +V +YL
Sbjct: 626 SQGQSLPNVGIYL 638
>At5g32630 putative protein
Length = 856
Score = 67.4 bits (163), Expect = 1e-12
Identities = 34/73 (46%), Positives = 47/73 (63%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+ +T + + ++ A V G+ +G IP I+LT SD+ LPFK RRQF ++ FAMTIN
Sbjct: 733 LHITQIAKQVVQAKVITGDIIGDIILIPLINLTPSDTKLPFKMRRRQFPLSDAFAMTINK 792
Query: 105 NQGQSLSHVSLYL 117
+QGQSL LYL
Sbjct: 793 SQGQSLEQAGLYL 805
>At5g37110 putative helicase
Length = 1307
Score = 66.2 bits (160), Expect = 2e-12
Identities = 30/73 (41%), Positives = 46/73 (62%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+R+T + ++ A + G++ G IPR+ L SD+ LPF+ R Q + +CFAMTIN
Sbjct: 1195 LRITQMGPFILQAMILTGDRAGHLVLIPRLKLAPSDTKLPFRMRRTQLPLAVCFAMTINK 1254
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V ++L
Sbjct: 1255 SQGQSLKRVGIFL 1267
>At3g31980 hypothetical protein
Length = 1099
Score = 63.2 bits (152), Expect = 2e-11
Identities = 30/73 (41%), Positives = 47/73 (64%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + ++ A V G+++G I + ++ SD+ LPF+ RRQF + + FAMTIN
Sbjct: 975 LQVTQMGNHILEARVITGDRVGDKVIIIKSQISPSDTKLPFRMRRRQFPIAVAFAMTINK 1034
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V +YL
Sbjct: 1035 SQGQSLKEVGIYL 1047
>At3g30420 hypothetical protein
Length = 837
Score = 63.2 bits (152), Expect = 2e-11
Identities = 34/74 (45%), Positives = 46/74 (61%), Gaps = 1/74 (1%)
Query: 45 MRVTHLTQ*MIVATVFPGN-KLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTIN 103
+ VTHL ++ A + K K IPRI L+ DS PF RRQF V +C+AMT+N
Sbjct: 707 LTVTHLGDKVLKAEILSDTTKKRKKVLIPRIILSPQDSKHPFTLRRRQFPVRMCYAMTVN 766
Query: 104 NNQGQSLSHVSLYL 117
+QGQ+L+ V+LYL
Sbjct: 767 KSQGQTLNRVALYL 780
>At1g54430 hypothetical protein
Length = 1639
Score = 62.8 bits (151), Expect = 3e-11
Identities = 35/72 (48%), Positives = 45/72 (61%), Gaps = 1/72 (1%)
Query: 47 VTHLTQ*MIVATVFPGN-KLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNN 105
VTHL ++ A + K K IPRI L+ DS PF RRQF V +C+AMTIN +
Sbjct: 1511 VTHLGDKVLKAEILSDTTKERKKVLIPRIILSPQDSKHPFTLRRRQFPVRMCYAMTINKS 1570
Query: 106 QGQSLSHVSLYL 117
QGQ+L+ V+LYL
Sbjct: 1571 QGQTLNRVALYL 1582
>At5g28780 putative protein
Length = 337
Score = 57.4 bits (137), Expect = 1e-09
Identities = 32/71 (45%), Positives = 41/71 (57%)
Query: 47 VTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
VT+L + +I A + G GK IPR L+ S PF R+QF + +C+AMTI NQ
Sbjct: 212 VTNLGEQVIEAQIVTGTHAGKMVSIPRFILSPPQSEHPFTLRRQQFPMRVCYAMTIIKNQ 271
Query: 107 GQSLSHVSLYL 117
GQSL LYL
Sbjct: 272 GQSLKSDVLYL 282
>At3g51700 unknown protein
Length = 344
Score = 56.6 bits (135), Expect = 2e-09
Identities = 29/73 (39%), Positives = 43/73 (58%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T + ++ A + GN G+ IPRI ++ P K RRQF V L FAMTI+
Sbjct: 212 LQITRVETFVLEAMIITGNNHGEKVLIPRIPSDLREAKFPIKMRRRQFPVKLAFAMTIDE 271
Query: 105 NQGQSLSHVSLYL 117
+Q Q+LS V +YL
Sbjct: 272 SQRQTLSKVGIYL 284
>At2g07620 putative helicase
Length = 1241
Score = 55.8 bits (133), Expect = 3e-09
Identities = 27/73 (36%), Positives = 44/73 (59%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + ++ A V G+++ I + ++ SD+ LPF+ RRQF + + FAM I
Sbjct: 1117 LQVTQMGNHILEARVITGDRVRDKVIIIKAQISPSDTKLPFRMRRRQFPIAVAFAMRIKK 1176
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V +YL
Sbjct: 1177 SQGQSLKEVEIYL 1189
>At3g42420 putative protein
Length = 1018
Score = 54.3 bits (129), Expect = 1e-08
Identities = 27/62 (43%), Positives = 41/62 (65%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T +T ++ A V G++ G IP I++T S+ LPF+ RRQF V+L FAMTIN
Sbjct: 957 LQITQMTIQVLQAKVIIGDRSGDIVLIPLINITPSNMKLPFRMRRRQFPVSLAFAMTINK 1016
Query: 105 NQ 106
+Q
Sbjct: 1017 SQ 1018
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.353 0.153 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,022,115
Number of Sequences: 26719
Number of extensions: 61284
Number of successful extensions: 301
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 274
Number of HSP's gapped (non-prelim): 28
length of query: 117
length of database: 11,318,596
effective HSP length: 93
effective length of query: 24
effective length of database: 8,833,729
effective search space: 212009496
effective search space used: 212009496
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0403.8


